Multiomics of GCN4-Dependent Replicative Lifespan Extension Models Reveals Gcn4 as a Regulator of Protein Turnover in Yeast
Abstract
:1. Introduction
2. Results
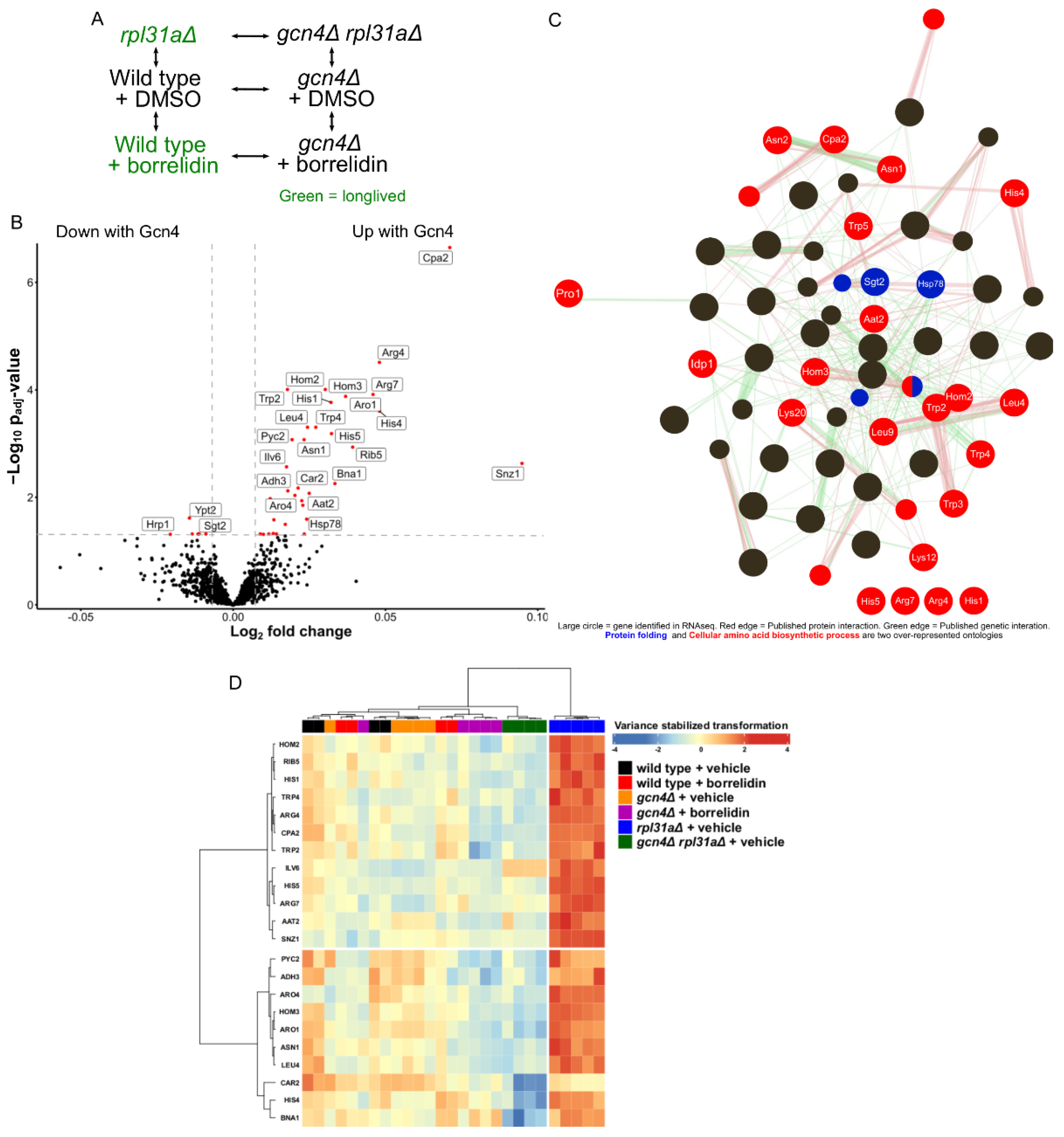
2.1. GCN4 Impacts Proteins Related to Amino Acid Biosynthesis in Yeast
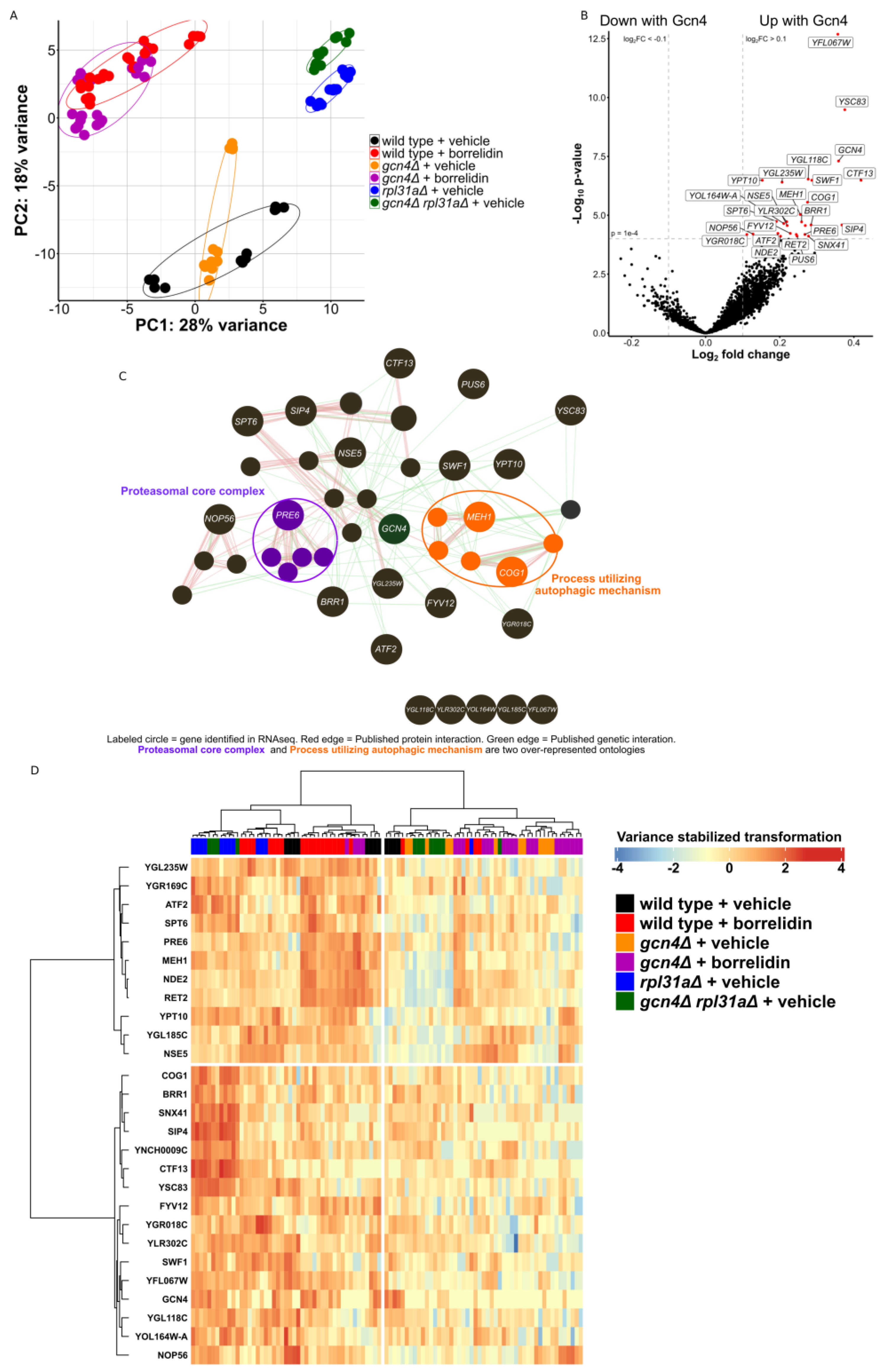
2.2. GCN4 Impacts the Transcription of Protein Degradation Mechanisms in Yeast
2.3. Conserved Genes Involved in Protein Degradation Are Differentially Expressed in This Yeast Dataset
3. Discussion
4. Materials and Methods
4.1. Yeast Culture
4.2. Protein Extraction, Mass Spectrometry, and Proteome Analysis
4.3. RNA Extraction, Sequencing, and Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- López-Otín, C.; Blasco, M.A.; Partridge, L.; Serrano, M.; Kroemer, G. The Hallmarks of Aging. Cell 2013, 153, 1194–1217. [Google Scholar] [CrossRef] [PubMed]
- López-Otín, C.; Blasco, M.A.; Partridge, L.; Serrano, M.; Kroemer, G. Hallmarks of aging: An expanding universe. Cell 2023, 186, 243–278. [Google Scholar] [CrossRef] [PubMed]
- Hoppe, T.; Cohen, E. Organismal Protein Homeostasis Mechanisms. Genetics 2020, 215, 889–901. [Google Scholar] [CrossRef] [PubMed]
- Hansen, M.; Taubert, S.; Crawford, D.; Libina, N.; Lee, S.; Kenyon, C. Lifespan extension by conditions that inhibit translation in Caenorhabditis elegans. Aging Cell 2007, 6, 95–110. [Google Scholar] [CrossRef]
- McCormick, M.A.; Delaney, J.R.; Tsuchiya, M.; Tsuchiyama, S.; Shemorry, A.; Sim, S.; Chou, A.C.-Z.; Ahmed, U.; Carr, D.; Murakami, C.J.; et al. A Comprehensive Analysis of Replicative Lifespan in 4,698 Single-Gene Deletion Strains Uncovers Conserved Mechanisms of Aging. Cell Metab. 2015, 22, 895–906. [Google Scholar] [CrossRef]
- Steffen, K.K.; MacKay, V.L.; Kerr, E.O.; Tsuchiya, M.; Hu, D.; Fox, L.A.; Dang, N.; Johnston, E.D.; Oakes, J.A.; Tchao, B.N.; et al. Yeast Life Span Extension by Depletion of 60S Ribosomal Subunits Is Mediated by Gcn4. Cell 2008, 133, 292–302. [Google Scholar] [CrossRef]
- Chondrogianni, N.; Sakellari, M.; Lefaki, M.; Papaevgeniou, N.; Gonos, E.S. Proteasome activation delays aging in vitro and in vivo. Free Radic. Biol. Med. 2014, 71, 303–320. [Google Scholar] [CrossRef]
- Jana, N.R. Protein homeostasis and aging: Role of ubiquitin protein ligases. Neurochem. Int. 2012, 60, 443–447. [Google Scholar] [CrossRef]
- Rubinsztein, D.C.; Marino, G.; Kroemer, G. Autophagy and Aging. Cell 2011, 146, 682–695. [Google Scholar] [CrossRef]
- Coux, O.; Tanaka, K.; Goldberg, A.L. Structure and functions of the 20S and 26S proteasomes. Annu. Rev. Biochem. 1996, 65, 801–847. [Google Scholar] [CrossRef]
- Nguyen, N.N.; Rana, A.; Goldman, C.; Moore, R.; Tai, J.; Hong, Y.; Shen, J.; Walker, D.W.; Hur, J.H. Proteasome β5 subunit overexpression improves proteostasis during aging and extends lifespan in Drosophila melanogaster. Sci. Rep. 2019, 9, 3170. [Google Scholar] [CrossRef] [PubMed]
- Vilchez, D.; Morantte, I.; Liu, Z.; Douglas, P.M.; Merkwirth, C.; Rodrigues, A.P.C.; Manning, G.; Dillin, A. RPN-6 determines C. elegans longevity under proteotoxic stress conditions. Nature 2012, 489, 263–268. [Google Scholar] [CrossRef] [PubMed]
- Kruegel, U.; Robison, B.; Dange, T.; Kahlert, G.; Delaney, J.R.; Kotireddy, S.; Tsuchiya, M.; Tsuchiyama, S.; Murakami, C.J.; Schleit, J.; et al. Elevated Proteasome Capacity Extends Replicative Lifespan in Saccharomyces cerevisiae. PLoS Genet. 2011, 7, e1002253. [Google Scholar] [CrossRef] [PubMed]
- Ghazi, A.; Henis-Korenblit, S.; Kenyon, C. Regulation of Caenorhabditis elegans lifespan by a proteasomal E3 ligase complex. Proc. Natl. Acad. Sci. USA 2007, 104, 5947–5952. [Google Scholar] [CrossRef]
- Jensen, M.B.; Jasper, H. Mitochondrial Proteostasis in the Control of Aging and Longevity. Cell Metab. 2014, 20, 214–225. [Google Scholar] [CrossRef]
- Nakatogawa, H.; Suzuki, K.; Kamada, Y.; Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: Lessons from yeast. Nat. Rev. Mol. Cell Biol. 2009, 10, 458–467. [Google Scholar] [CrossRef]
- Wong, S.Q.; Kumar, A.V.; Mills, J.; Lapierre, L.R. Autophagy in aging and longevity. Hum. Genet. 2020, 139, 277–290. [Google Scholar] [CrossRef]
- Cebollero, E.; Reggiori, F.; Kraft, C. Reticulophagy and Ribophagy: Regulated Degradation of Protein Production Factories. Int. J. Cell Biol. 2012, 2012, e182834. [Google Scholar] [CrossRef]
- Tolkovsky, A.M. Mitophagy. Biochim. Biophys. Acta-Mol. Cell Res. 2009, 1793, 1508–1515. [Google Scholar] [CrossRef]
- Zaffagnini, G.; Martens, S. Mechanisms of Selective Autophagy. J. Mol. Biol. 2016, 428, 1714–1724. [Google Scholar] [CrossRef]
- Klionsky, D.J.; Abdel-Aziz, A.K.; Abdelfatah, S.; Abdellatif, M.; Abdoli, A.; Abel, S.; Abeliovich, H.; Abildgaard, M.H.; Abudu, Y.P.; Acevedo-Arozena, A.; et al. Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition)1. Autophagy 2021, 17, 1–382. [Google Scholar] [CrossRef] [PubMed]
- Ponpuak, M.; Mandell, M.A.; Kimura, T.; Chauhan, S.; Cleyrat, C.; Deretic, V. Secretory autophagy. Curr. Opin. Cell Biol. 2015, 35, 106–116. [Google Scholar] [CrossRef] [PubMed]
- Pilli, M.; Arko-Mensah, J.; Ponpuak, M.; Roberts, E.; Master, S.; Mandell, M.A.; Dupont, N.; Ornatowski, W.; Jiang, S.; Bradfute, S.B.; et al. TBK-1 promotes autophagy-mediated antimicrobial defense by controlling autophagosome maturation. Immunity 2012, 37, 223–234. [Google Scholar] [CrossRef] [PubMed]
- Mandell, M.A.; Jain, A.; Arko-Mensah, J.; Chauhan, S.; Kimura, T.; Dinkins, C.; Silvestri, G.; Münch, J.; Kirchhoff, F.; Simonsen, A.; et al. TRIM Proteins Regulate Autophagy and Can Target Autophagic Substrates by Direct Recognition. Dev. Cell 2014, 30, 394–409. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, H.; Zhang, S.; Mizushima, N. Autophagy genes in biology and disease. Nat. Rev. Genet. 2023, 24, 382–400. [Google Scholar] [CrossRef]
- Seo, H.; Sonntag, K.-C.; Kim, W.; Cattaneo, E.; Isacson, O. Proteasome Activator Enhances Survival of Huntington’s Disease Neuronal Model Cells. PLoS ONE 2007, 2, e238. [Google Scholar] [CrossRef]
- Yazgili, A.S.; Ebstein, F.; Meiners, S. The Proteasome Activator PA200/PSME4: An Emerging New Player in Health and Disease. Biomolecules 2022, 12, 1150. [Google Scholar] [CrossRef]
- Jung, R.; Lechler, M.C.; Fernandez-Villegas, A.; Chung, C.W.; Jones, H.C.; Choi, Y.H.; Thompson, M.A.; Rödelsperger, C.; Röseler, W.; Schierle, G.S.K.; et al. A safety mechanism enables tissue-specific resistance to protein aggregation during aging in C. elegans. PLoS Biol. 2023, 21, e3002284. [Google Scholar] [CrossRef]
- David, D. Aging and the aggregating proteome. Front. Genet. 2012, 3, 247. [Google Scholar] [CrossRef]
- David, D.C.; Ollikainen, N.; Trinidad, J.C.; Cary, M.P.; Burlingame, A.L.; Kenyon, C. Widespread Protein Aggregation as an Inherent Part of Aging in C. elegans. PLoS Biol. 2010, 8, e1000450. [Google Scholar] [CrossRef]
- Hinnebusch, A.G.; Natarajan, K. Gcn4p, a Master Regulator of Gene Expression, Is Controlled at Multiple Levels by Diverse Signals of Starvation and Stress. Eukaryot. Cell 2002, 1, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, A.G. Mechanisms of gene regulation in the general control of amino acid biosynthesis in Saccharomyces cerevisiae. Microbiol. Rev. 1988, 52, 248–273. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, A.G. Translational regulation of GCN4 and the general amino acid control of yeast. Annu. Rev. Microbiol. 2005, 59, 407–450. [Google Scholar] [CrossRef] [PubMed]
- Robbins, C.E.; Patel, B.; Sawyer, D.L.; Wilkinson, B.; Kennedy, B.K.; McCormick, M.A. Cytosolic and mitochondrial tRNA synthetase inhibitors increase lifespan in a GCN4/atf-4-dependent manner. iScience 2022, 25, 105410. [Google Scholar] [CrossRef]
- Statzer, C.; Meng, J.; Venz, R.; Bland, M.; Robida-Stubbs, S.; Patel, K.; Petrovic, D.; Emsley, R.; Liu, P.; Morantte, I.; et al. ATF-4 and hydrogen sulfide signalling mediate longevity in response to inhibition of translation or mTORC1. Nat. Commun. 2022, 13, 967. [Google Scholar] [CrossRef]
- Molenaars, M.; Janssens, G.E.; Williams, E.G.; Jongejan, A.; Lan, J.; Rabot, S.; Joly, F.; Moerland, P.D.; Schomakers, B.V.; Lezzerini, M.; et al. A Conserved Mito-Cytosolic Translational Balance Links Two Longevity Pathways. Cell Metab. 2020, 31, 549–563.e7. [Google Scholar] [CrossRef]
- Hu, Z.; Xia, B.; Postnikoff, S.D.; Shen, Z.-J.; Tomoiaga, A.S.; Harkness, T.A.; Seol, J.H.; Li, W.; Chen, K.; Tyler, J.K. Ssd1 and Gcn2 suppress global translation efficiency in replicatively aged yeast while their activation extends lifespan. eLife 2018, 7, e35551. [Google Scholar] [CrossRef]
- Li, W.; Li, X.; Miller, R.A. ATF4 activity: A common feature shared by many kinds of slow-aging mice. Aging Cell 2014, 13, 1012–1018. [Google Scholar] [CrossRef]
- Li, W.; Miller, R.A. Elevated ATF4 Function in Fibroblasts and Liver of Slow-Aging Mutant Mice. J. Gerontol. Ser. A 2015, 70, 263–272. [Google Scholar] [CrossRef]
- Mariner, B.L.; Rodriguez, A.S.; Heath, O.C.; McCormick, M.A. Induction of proteasomal activity in mammalian cells by lifespan-extending tRNA synthetase inhibitors. In GeroScience; Springer: Berlin/Heidelberg, Germany, 2023. [Google Scholar] [CrossRef]
- Lu, P.D.; Harding, H.P.; Ron, D. Translation reinitiation at alternative open reading frames regulates gene expression in an integrated stress response. J. Cell Biol. 2004, 167, 27–33. [Google Scholar] [CrossRef]
- Harding, H.P.; Novoa, I.; Zhang, Y.; Zeng, H.; Wek, R.; Schapira, M.; Ron, D. Regulated Translation Initiation Controls Stress-Induced Gene Expression in Mammalian Cells. Mol. Cell 2000, 6, 1099–1108. [Google Scholar] [CrossRef]
- Rolfes, R.J.; Hinnebusch, A.G. Translation of the yeast transcriptional activator GCN4 is stimulated by purine limitation: Implications for activation of the protein kinase GCN2. Mol. Cell. Biol. 1993, 13, 5099–5111. [Google Scholar] [CrossRef]
- Hinnebusch, A.G. The Scanning Mechanism of Eukaryotic Translation Initiation. Annu. Rev. Biochem. 2014, 83, 779–812. [Google Scholar] [CrossRef]
- Hinnebusch, A.G. Molecular mechanism of scanning and start codon selection in eukaryotes. Microbiol. Mol. Biol. Rev. 2011, 75, 434–467. [Google Scholar] [CrossRef]
- Hinnebusch, A.G.; Dever, T.E.; Asano, K. Mechanism of Translation Initiation in the Yeast Saccharomyces cerevisiae. Cold Spring Harb. Monogr. Arch. 2007, 48, 225–268. [Google Scholar]
- Pakos-Zebrucka, K.; Koryga, I.; Mnich, K.; Ljujic, M.; Samali, A.; Gorman, A.M. The integrated stress response. EMBO Rep. 2016, 17, 1374–1395. [Google Scholar] [CrossRef]
- Postnikoff, S.D.L.; Johnson, J.E.; Tyler, J.K. The integrated stress response in budding yeast lifespan extension. Microb. Cell 2017, 4, 368–375. [Google Scholar] [CrossRef]
- Habibi, D.; Ogloff, N.; Jalili, R.B.; Yost, A.; Weng, A.P.; Ghahary, A.; Ong, C.J. Borrelidin, a small molecule nitrile-containing macrolide inhibitor of threonyl-tRNA synthetase, is a potent inducer of apoptosis in acute lymphoblastic leukemia. Investig. New Drugs 2012, 30, 1361–1370. [Google Scholar] [CrossRef]
- Steffen, K.K.; McCormick, M.A.; Pham, K.M.; MacKay, V.L.; Delaney, J.R.; Murakami, C.J.; Kaeberlein, M.; Kennedy, B.K. Ribosome Deficiency Protects Against ER Stress in Saccharomyces cerevisiae. Genetics 2012, 191, 107–118. [Google Scholar] [CrossRef]
- McCormick, M.; Chen, K.; Ramaswamy, P.; Kenyon, C. New genes that extend Caenorhabditis elegans’ lifespan in response to reproductive signals. Aging Cell 2012, 11, 192–202. [Google Scholar] [CrossRef]
- Montojo, J.; Zuberi, K.; Rodriguez, H.; Kazi, F.; Wright, G.; Donaldson, S.L.; Morris, Q.; Bader, G.D. GeneMANIA Cytoscape plugin: Fast gene function predictions on the desktop. Bioinformatics 2010, 26, 2927–2928. [Google Scholar] [CrossRef]
- Mostafavi, S.; Ray, D.; Warde-Farley, D.; Grouios, C.; Morris, Q. GeneMANIA: A real-time multiple association network integration algorithm for predicting gene function. Genome Biol. 2008, 9, S4. [Google Scholar] [CrossRef]
- Pavlova, N.N.; King, B.; Josselsohn, R.H.; Violante, S.; Macera, V.L.; Vardhana, S.A.; Cross, J.R.; Thompson, C.B. Translation in amino-acid-poor environments is limited by tRNAGln charging. eLife 2020, 9, e62307. [Google Scholar] [CrossRef]
- Piérard, A.; Schröter, B. Structure-Function Relationships in the Arginine Pathway Carbamoylphosphate Synthase of Saccharomyces cerevisiae. J. Bacteriol. 1978, 134, 167–176. [Google Scholar] [CrossRef]
- Lusty, C.J.; Widgren, E.E.; Broglie, K.E.; Nyunoya, H. Yeast carbamyl phosphate synthetase. Structure of the yeast gene and homology to Escherichia coli carbamyl phosphate synthetase. J. Biol. Chem. 1983, 258, 14466–14477. [Google Scholar] [CrossRef]
- Penn, M.D.; Galgoci, B.; Greer, H. Identification of AAS genes and their regulatory role in general control of amino acid biosynthesis in yeast. Proc. Natl. Acad. Sci. USA 1983, 80, 2704–2708. [Google Scholar] [CrossRef]
- von Janowsky, B.; Major, T.; Knapp, K.; Voos, W. The disaggregation activity of the mitochondrial ClpB homolog Hsp78 maintains Hsp70 function during heat stress. J. Mol. Biol. 2006, 357, 793–807. [Google Scholar] [CrossRef]
- Moczko, M.; Schönfisch, B.; Voos, W.; Pfanner, N.; Rassow, J. The mitochondrial ClpB homolog Hsp78 cooperates with matrix Hsp70 in maintenance of mitochondrial function. J. Mol. Biol. 1995, 254, 538–543. [Google Scholar] [CrossRef]
- Röttgers, K.; Zufall, N.; Guiard, B.; Voos, W. The ClpB homolog Hsp78 is required for the efficient degradation of proteins in the mitochondrial matrix. J. Biol. Chem. 2002, 277, 45829–45837. [Google Scholar] [CrossRef]
- Ishikawa, Y.; Nishino, S.; Fukuda, S.; Nguyet, V.T.A.; Izawa, S. Severe ethanol stress induces the preferential synthesis of mitochondrial disaggregase Hsp78 and formation of DUMPs in Saccharomyces cerevisiae. Biochimica Et Biophysica Acta. Gen. Subj. 2022, 1866, 130147. [Google Scholar] [CrossRef]
- Kiktev, D.A.; Patterson, J.C.; Müller, S.; Bariar, B.; Pan, T.; Chernoff, Y.O. Regulation of chaperone effects on a yeast prion by cochaperone Sgt2. Mol. Cell. Biol. 2012, 32, 4960–4970. [Google Scholar] [CrossRef]
- Wang, F.; Brown, E.C.; Mak, G.; Zhuang, J.; Denic, V. A chaperone cascade sorts proteins for posttranslational membrane insertion into the endoplasmic reticulum. Mol. Cell 2010, 40, 159–171. [Google Scholar] [CrossRef]
- Huh, W.-K.; Falvo, J.V.; Gerke, L.C.; Carroll, A.S.; Howson, R.W.; Weissman, J.S.; O’Shea, E.K. Global analysis of protein localization in budding yeast. Nature 2003, 425, 686–691. [Google Scholar] [CrossRef]
- Tkach, J.M.; Yimit, A.; Lee, A.Y.; Riffle, M.; Costanzo, M.; Jaschob, D.; Hendry, J.A.; Ou, J.; Moffat, J.; Boone, C.; et al. Dissecting DNA damage response pathways by analysing protein localization and abundance changes during DNA replication stress. Nat. Cell Biol. 2012, 14, 966–976. [Google Scholar] [CrossRef]
- Mittal, N.; Guimaraes, J.C.; Gross, T.; Schmidt, A.; Vina-Vilaseca, A.; Nedialkova, D.D.; Aeschimann, F.; Leidel, S.A.; Spang, A.; Zavolan, M. The Gcn4 transcription factor reduces protein synthesis capacity and extends yeast lifespan. Nat. Commun. 2017, 8, 457. [Google Scholar] [CrossRef]
- Harding, H.P.; Zhang, Y.; Bertolotti, A.; Zeng, H.; Ron, D. Perk Is Essential for Translational Regulation and Cell Survival during the Unfolded Protein Response. Mol. Cell 2000, 5, 897–904. [Google Scholar] [CrossRef]
- Tameire, F.; Verginadis, I.I.; Leli, N.M.; Polte, C.; Conn, C.S.; Ojha, R.; Salinas, C.S.; Chinga, F.; Monroy, A.M.; Fu, W.; et al. ATF4 couples MYC-dependent translational activity to bioenergetic demands during tumour progression. Nat. Cell Biol. 2019, 21, 889–899. [Google Scholar] [CrossRef]
- Yofe, I.; Weill, U.; Meurer, M.; Chuartzman, S.; Zalckvar, E.; Goldman, O.; Ben-Dor, S.; Schütze, C.; Wiedemann, N.; Knop, M.; et al. One library to make them all: Streamlining the creation of yeast libraries via a SWAp-Tag strategy. Nat. Methods 2016, 13, 371–378. [Google Scholar] [CrossRef]
- Rocco, V.; Daly, M.J.; Matre, V.; Lichten, M.; Nicolas, A. Identification of two divergently transcribed genes centromere-proximal to the ARG4 locus on chromosome VIII of Saccharomyces cerevisiae. Yeast 1993, 9, 1111–1120. [Google Scholar] [CrossRef]
- Onodera, J.; Ohsumi, Y. Ald6p Is a Preferred Target for Autophagy in Yeast, Saccharomyces cerevisiae. J. Biol. Chem. 2004, 279, 16071–16076. [Google Scholar] [CrossRef]
- Delorme-Axford, E.; Wen, X.; Klionsky, D.J. The yeast transcription factor Stb5 acts as a negative regulator of autophagy by modulating cellular metabolism. Autophagy 2023, 19, 2719–2732. [Google Scholar] [CrossRef] [PubMed]
- Takeshige, K.; Baba, M.; Tsuboi, S.; Noda, T.; Ohsumi, Y. Autophagy in yeast demonstrated with proteinase-deficient mutants and conditions for its induction. J. Cell Biol. 1992, 119, 301–311. [Google Scholar] [CrossRef]
- Pereira, M.D.; Eleutherio, E.C.; Panek, A.D. Acquisition of tolerance against oxidative damage in Saccharomyces cerevisiae. BMC Microbiol. 2001, 1, 11. [Google Scholar] [CrossRef]
- Kim, B.; Lee, Y.; Choi, H.; Huh, W.-K. The trehalose-6-phosphate phosphatase Tps2 regulates ATG8 transcription and autophagy in Saccharomyces cerevisiae. Autophagy 2021, 17, 1013–1027. [Google Scholar] [CrossRef]
- Michaillat, L.; Mayer, A. Identification of Genes Affecting Vacuole Membrane Fragmentation in Saccharomyces cerevisiae. PLoS ONE 2013, 8, e54160. [Google Scholar] [CrossRef]
- Lang, T.; Reiche, S.; Straub, M.; Bredschneider, M.; Thumm, M. Autophagy and the cvt Pathway Both Depend on AUT9. J. Bacteriol. 2000, 182, 2125–2133. [Google Scholar] [CrossRef]
- Verghese, J.; Abrams, J.; Wang, Y.; Morano, K.A. Biology of the Heat Shock Response and Protein Chaperones: Budding Yeast (Saccharomyces cerevisiae) as a Model System. Microbiol. Mol. Biol. Rev. 2012, 76, 115–158. [Google Scholar] [CrossRef]
- Eising, S.; Esch, B.; Wälte, M.; Duarte, P.V.; Walter, S.; Ungermann, C.; Bohnert, M.; Fröhlich, F. A lysosomal biogenesis map reveals the cargo spectrum of yeast vacuolar protein targeting pathways. J. Cell Biol. 2022, 221, e202107148. [Google Scholar] [CrossRef]
- Whyte, J.R.; Munro, S. The Sec34/35 Golgi transport complex is related to the exocyst, defining a family of complexes involved in multiple steps of membrane traffic. Dev. Cell 2001, 1, 527–537. [Google Scholar] [CrossRef]
- Loh, E.; Hong, W. The binary interacting network of the conserved oligomeric Golgi tethering complex. J. Biol. Chem. 2004, 279, 24640–24648. [Google Scholar] [CrossRef]
- Corbacho, I.; Olivero, I.; Hernández, L.M. Identification of low-dye-binding (ldb) mutants of Saccharomyces cerevisiae. FEMS Yeast Res. 2004, 4, 437–444. [Google Scholar] [CrossRef] [PubMed]
- Heinemeyer, W.; Troendle, N.; Albrecht, G.; Wolf, D.H. PRE5 and PRE6, the last missing genes encoding 20S proteasome subunits from yeast? Indication for a set of 14 different subunits in the eukaryotic proteasome core. Biochemistry 1994, 33, 12229–12237. [Google Scholar] [CrossRef] [PubMed]
- Dooley, H.C.; Razi, M.; Polson, H.E.; Girardin, S.E.; Wilson, M.I.; Tooze, S.A. WIPI2 links LC3 conjugation with PI3P, autophagosome formation, and pathogen clearance by recruiting Atg12-5-16L1. Mol. Cell 2014, 55, 238–252. [Google Scholar] [CrossRef]
- Mercer, T.J.; Gubas, A.; Tooze, S.A. A molecular perspective of mammalian autophagosome biogenesis. J. Biol. Chem. 2018, 293, 5386–5395. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.I.; Dooley, H.C.; Tooze, S.A. WIPI2b and Atg16L1: Setting the stage for autophagosome formation. Biochem. Soc. Trans. 2014, 42, 1327–1334. [Google Scholar] [CrossRef] [PubMed]
- Strømhaug, P.E.; Reggiori, F.; Guan, J.; Wang, C.-W.; Klionsky, D.J. Atg21 is a phosphoinositide binding protein required for efficient lipidation and localization of Atg8 during uptake of aminopeptidase I by selective autophagy. Mol. Biol. Cell 2004, 15, 3553–3566. [Google Scholar] [CrossRef]
- Meiling-Wesse, K.; Barth, H.; Voss, C.; Eskelinen, E.-L.; Epple, U.D.; Thumm, M. Atg21 is required for effective recruitment of Atg8 to the preautophagosomal structure during the Cvt pathway. J. Biol. Chem. 2004, 279, 37741–37750. [Google Scholar] [CrossRef]
- Meijer, W.H.; van der Klei, I.J.; Veenhuis, M.; Kiel, J.A.K.W. ATG genes involved in non-selective autophagy are conserved from yeast to man, but the selective Cvt and pexophagy pathways also require organism-specific genes. Autophagy 2007, 3, 106–116. [Google Scholar] [CrossRef]
- Bhatia, P.K.; Verhage, R.A.; Brouwer, J.; Friedberg, E.C. Molecular cloning and characterization of Saccharomyces cerevisiae RAD28, the yeast homolog of the human Cockayne syndrome A (CSA) gene. J. Bacteriol. 1996, 178, 5977–5988. [Google Scholar] [CrossRef]
- Bilodeau, P.S.; Urbanowski, J.L.; Winistorfer, S.C.; Piper, R.C. The Vps27p Hse1p complex binds ubiquitin and mediates endosomal protein sorting. Nat. Cell Biol. 2002, 4, 534–539. [Google Scholar] [CrossRef]
- Bilodeau, P.S.; Winistorfer, S.C.; Kearney, W.R.; Robertson, A.D.; Piper, R.C. Vps27-Hse1 and ESCRT-I complexes cooperate to increase efficiency of sorting ubiquitinated proteins at the endosome. J. Cell Biol. 2003, 163, 237–243. [Google Scholar] [CrossRef] [PubMed]
- Stolz, A.; Schweizer, R.S.; Schäfer, A.; Wolf, D.H. Dfm1 forms distinct complexes with Cdc48 and the ER ubiquitin ligases and is required for ERAD. Traffic 2010, 11, 1363–1369. [Google Scholar] [CrossRef] [PubMed]
- Kandel, R.; Jung, J.; Syau, D.; Kuo, T.; Songster, L.; Horn, C.; Chapman, C.; Aguayo, A.; Duttke, S.; Benner, C.; et al. Yeast derlin Dfm1 employs a chaperone-like function to resolve misfolded membrane protein stress. PLoS Biol. 2023, 21, e3001950. [Google Scholar] [CrossRef]
- Santt, O.; Pfirrmann, T.; Braun, B.; Juretschke, J.; Kimmig, P.; Scheel, H.; Hofmann, K.; Thumm, M.; Wolf, D.H. The yeast GID complex, a novel ubiquitin ligase (E3) involved in the regulation of carbohydrate metabolism. Mol. Biol. Cell 2008, 19, 3323–3333. [Google Scholar] [CrossRef] [PubMed]
- Menssen, R.; Schweiggert, J.; Schreiner, J.; Kušević, D.; Reuther, J.; Braun, B.; Wolf, D.H. Exploring the topology of the Gid complex, the E3 ubiquitin ligase involved in catabolite-induced degradation of gluconeogenic enzymes. J. Biol. Chem. 2012, 287, 25602–25614. [Google Scholar] [CrossRef]
- Shen, Z.-J.; Postnikoff, S.; Tyler, J.K. Is Gcn4-induced autophagy the ultimate downstream mechanism by which hormesis extends yeast replicative lifespan? Curr. Genet. 2019, 65, 717–720. [Google Scholar] [CrossRef] [PubMed]
- Patil, C.K.; Li, H.; Walter, P. Gcn4p and Novel Upstream Activating Sequences Regulate Targets of the Unfolded Protein Response. PLoS Biol. 2004, 2, e246. [Google Scholar] [CrossRef]
- Yeast Protocols Handbook PT3024–1; CLONTECH: Mountain View, CA, USA, 2009.
- Sepulveda, J.L. Using R and Bioconductor in Clinical Genomics and Transcriptomics. J. Mol. Diagn. 2020, 22, 3–20. [Google Scholar] [CrossRef]
- Small, E.M.; Felker, D.P.; Heath, O.C.; Cantergiani, R.J.; Robbins, C.E.; Osley, M.A.; McCormick, M.A. SPOCK, an R based package for high-throughput analysis of growth rate, survival, and chronological lifespan in yeast. Transl. Med. Aging 2020, 4, 141–148. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Smyth, G.K.; Shi, W. Feature Counts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [PubMed]
- Law, C.W.; Chen, Y.; Shi, W.; Smyth, G.K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014, 15, R29. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Liu, S.; Wang, Z.; Zhu, R.; Wang, F.; Cheng, Y.; Liu, Y. Three Differential Expression Analysis Methods for RNA Sequencing: Limma, EdgeR, DESeq2. JOVE 2021, 175, e62528. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mariner, B.L.; Felker, D.P.; Cantergiani, R.J.; Peterson, J.; McCormick, M.A. Multiomics of GCN4-Dependent Replicative Lifespan Extension Models Reveals Gcn4 as a Regulator of Protein Turnover in Yeast. Int. J. Mol. Sci. 2023, 24, 16163. https://doi.org/10.3390/ijms242216163
Mariner BL, Felker DP, Cantergiani RJ, Peterson J, McCormick MA. Multiomics of GCN4-Dependent Replicative Lifespan Extension Models Reveals Gcn4 as a Regulator of Protein Turnover in Yeast. International Journal of Molecular Sciences. 2023; 24(22):16163. https://doi.org/10.3390/ijms242216163
Chicago/Turabian StyleMariner, Blaise L., Daniel P. Felker, Ryla J. Cantergiani, Jack Peterson, and Mark A. McCormick. 2023. "Multiomics of GCN4-Dependent Replicative Lifespan Extension Models Reveals Gcn4 as a Regulator of Protein Turnover in Yeast" International Journal of Molecular Sciences 24, no. 22: 16163. https://doi.org/10.3390/ijms242216163



