Analysis of the Polymorphisms and Expression Levels of the BCL2, BAX and c-MYC Genes in Patients with Ovarian Cancer
Abstract
:1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Patients
4.2. Polymorphism Analysis of BCL2, BAX and c-MYC Genes
4.3. Expression Analysis of BCL2, BAX and c-MYC Genes
4.4. ELISA
4.5. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Nowacka, M.; Ginter-Matuszewska, B.; Świerczewska, M.; Nowicki, M.; Zabel, M.; Sterzyńska, K.; Januchowski, R. The significance of HERC5, IFIH1, SAMD4, SEMA3A and MCTP1 genes expression in resistance to cytotoxic drugs in ovarian cancer cell lines. Med. J. Cell Biol. 2021, 9, 138–147. [Google Scholar] [CrossRef]
- Koshiyama, M.; Matsumura, N.; Konishi, I. Subtypes of Ovarian Cancer and Ovarian Cancer Screening. Diagnostics 2017, 7, 12. [Google Scholar] [CrossRef] [PubMed]
- Gaona-Luviano, P.; Medina-Gaona, L.A.; Magaña-Pérez, K. Epidemiology of ovarian cancer. Chin. Clin. Oncol. 2020, 9, 47. [Google Scholar] [CrossRef] [PubMed]
- Sarivalasis, A.; Morotti, M.; Mulvey, A.; Imbimbo, M.; Coukos, G. Cell therapies in ovarian cancer. Ther. Adv. Med. Oncol. 2021, 13, 17588359211008399. [Google Scholar] [CrossRef]
- Trabert, B.; Ness, R.B.; Lo-Ciganic, W.-H.; Murphy, M.A.; Goode, E.L.; Poole, E.M.; Brinton, L.A.; Webb, P.M.; Nagle, C.M.; Jordan, S.J.; et al. Aspirin, nonaspirin nonsteroidal anti-inflammatory drug, and acetaminophen use and risk of invasive epithelial ovarian cancer: A pooled analysis in the Ovarian Cancer Association Consortium. J. Natl. Cancer Inst. 2014, 106, djt431. [Google Scholar] [CrossRef]
- Bakir, M.S.; Birge, O.; Karadag, C.; Ilhan, Y.; Tuncer, H.A.; Göksu, S.S.; Simsek, T. Bevacizumab in recurrent ovarian cancer. JBUON 2021, 26, 1271–1278. [Google Scholar]
- García-Martínez, E.; Pérez-Fidalgo, J.A. Immunotherapies in ovarian cancer. EJC Suppl. 2020, 15, 87–95. [Google Scholar] [CrossRef]
- Rastogi, M.; Gupta, S.; Sachan, M. Biomarkers towards Ovarian Cancer Diagnostics: Present and Future Prospects. Braz. Arch. Biol. Technol. 2016, 59, e16160070. [Google Scholar] [CrossRef]
- Xu, Y.; Yu, Q.; Wang, P.; Wu, Z.; Zhang, L.; Wu, S.; Li, M.; Wu, B.; Li, H.; Zhuang, H.; et al. Selective Small-Molecule c-Myc Degrader Potently Regresses Lethal c-Myc Overexpressing Tumors. Adv. Sci. 2022, 9, 2104344. [Google Scholar] [CrossRef]
- Shi, Z.E.; Zhang, M.Y.; Liu, J.Y.; Zhang, W.D.; Hu, D.M.; Wang, Q.X.; Li, X.L.; Jiang, Y.Y.; Qu, Y.Q. Autophagy Induced by BCL2-Related ceRNA Network Participates in the Occurrence of COPD. Int. J. Chron. Obstruct Pulmon Dis. 2022, 17, 791–808. [Google Scholar] [CrossRef]
- Wang, C.; Qiao, X.; Wang, J.; Yang, J.; Yang, C.; Qiao, Y.; Guan, Y.; Wen, A.; Jiang, L. Amelioration of DMH-induced colon cancer by eupafolin through the reprogramming of apoptosis-associated p53/Bcl2/Bax signaling in rats. Eur. J. Inflamm. 2022, 20, 1–15. [Google Scholar] [CrossRef]
- Ozoran, E.; Can Trabulus, F.D.; Erhan, D.; Batar, B.; Guven, M. Association of XRCC3, XRCC4, BAX, and BCL-2 Polymorphisms with the Risk of Breast Cancer. Int. J. Breast Cancer 2022, 2022, 5817841. [Google Scholar] [CrossRef]
- Yuan, J.; Lan, H.; Jiang, X.; Zeng, D.; Xiao, S. Bcl-2 family: Novel insight into individualized therapy for ovarian cancer (Review). Int. J. Mol. Med. 2020, 46, 1255–1265. [Google Scholar] [CrossRef] [PubMed]
- Beale, P.J.; Rogers, P.; Boxall, F.; Sharp, S.Y.; Kelland, L.R. BCL-2 family protein expression and platinum drug resistance in ovarian carcinoma. Br. J. Cancer 2000, 82, 436–440. [Google Scholar] [CrossRef] [PubMed]
- Liang, M.; Zhao, J. Protein expressions of AIB1, p53 and Bcl-2 in epithelial ovarian cancer and their correlations with the clinical pathological features and prognosis. Eur. Rev. Med. Pharmacol. Sci. 2008, 22, 5134–5139. [Google Scholar]
- Faderl, S.; Keating, M.; Do, K.-A.; Liang, S.-Y.; Kantarjian, H.; O’Brien, S.; Garcia-Manero, G.; Manshouri, T.; Albitar, M. Expression profile of 11 proteins and their prognostic significance in patients with chronic lymphocytic leukemia (CLL). Leukemia 2002, 16, 1045–1052. [Google Scholar] [CrossRef]
- Callagy, G.M.; Webber, M.J.; Pharoah, P.D.; Caldas, C. Meta-analysis confirms BCL2 is an independent prognostic marker in breast cancer. BMC Cancer 2008, 8, 153. [Google Scholar] [CrossRef]
- Buglioni, S.; D’Agnano, I.; Cosimelli, M.; Vasselli, S.; D’Angelo, C.; Tedesco, M.; Zupi, G.; Mottolese, M. Evaluation of multiple bio-pathological factors in colorectal adenocarcinomas: Independent prognostic role of p53 and bcl-2. Int. J. Cancer 1999, 84, 545–552. [Google Scholar] [CrossRef]
- Dawson, S.J.; Makretsov, N.; Blows, F.M.; Driver, K.E.; Provenzano, E.; Quesne, J.L.; Baglietto, L.; Severi, G.; Giles, G.G.; Mclean, C.A.; et al. BCL2 in breast cancer: A favourable prognostic marker across molecular subtypes and independent of adjuvant therapy received. Br. J. Cancer 2010, 103, 668–675. [Google Scholar] [CrossRef]
- Ali, H.; Dawson, S.-J.; Blows, F.M.; Provenzano, E.; Leung, S.; Nielsen, T.; Pharoah, P.D.; Caldas, C. A Ki67/BCL2 index based on immunohistochemistry is highly prognostic in ER-positive breast cancer. J. Pathol. 2012, 226, 97–107. [Google Scholar] [CrossRef]
- Nuckel, H.; Frey, U.H.; Bau, M.; Sellmann, L.; Stanelle, J.; Düring, J.; Jöckel, K.-H.; Dührsen, U.; Siffert, W. Association of a novel regulatory polymorphism (-938C>A) in the BCL2 gene promotor with disease progression and survival in chronic lymphocytic leukemia. Blood 2007, 109, 290–297. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, H.S.; Bachmann, H.S.; Otterbach, F.; Callies, R.; Nückel, H.; Bau, M.; Schmid, K.W.; Siffert, W.; Kimmig, R. The AA genotype of the regulatory BCL2 promoter polymorphism (938C>A) is associated with a favorable outcome in lymph node negative invasive breast cancer patients. Clin. Cancer Res. 2007, 13, 5790–5797. [Google Scholar] [CrossRef] [PubMed]
- Lehnerdt, G.F.; Franz, P.; Bankfalvi, A.; Grehl, S.; Kelava, A.; Nückel, H.; Lang, S.; Schmid, K.W.; Siffert, W.; Bachmann, H.S. The regulatory BCL2 promoter polymorphism (-938C>A) is associated with relapse and survival of patients with oropharyngeal squamous cell carcinoma. Ann. Oncol. 2009, 20, 1094–1099. [Google Scholar] [CrossRef] [PubMed]
- Hirata, H.; Hinoda, Y.; Nakajima, K.; Kikuno, N.; Suehiro, Y.; Tabatabai, Z.L.; Ishii, N.; Dahiya, R. The bcl2 -938CC genotype has poor prognosis and lower survival in renal cancer. J. Urol. 2009, 182, 721–727. [Google Scholar] [CrossRef]
- Searle, C.J.; Brock, I.W.; Cross, S.S.; Balasubramanian, S.P.; Reed, M.W.R.; Cox, A. A BCL2 promoter polymorphism rs2279115 is not associated with BCL2 protein expression or patient survival in breast cancer patients. Springerplus 2012, 1, 38. [Google Scholar] [CrossRef]
- Zhang, N.; Li, X.; Tao, K.; Jiang, L.; Ma, T.; Yan, S.; Yuan, C.; Moran, M.S.; Liang, F.; Haffty, B.G.; et al. BCL-2 (-938C>a) polymorphism is associated with breast cancer susceptibility. BMC Med. Genet. 2011, 12, 48. [Google Scholar] [CrossRef]
- Pirocanac, E.C.; Nassirpour, R.; Yang, M.; Wang, J.; Nardin, S.R.; Gu, J.; Fang, B.; Moossa, A.; Hoffman, R.M.; Bouvet, M. Bax-Induction Gene Therapy of Pancreatic Cancer. J. Surg. Res. 2002, 106, 346–351. [Google Scholar] [CrossRef]
- Miquel, C.; Borrini, F.; Grandjouan, S.; Aupérin, A.; Viguier, J.; Velasco, V.; Duvillard, P.; Praz, F.; Sabourin, J.C. Role of bax Mutations in Apoptosis in Colorectal Cancers With Microsatellite Instability. Am. J. Clin. Pathol. 2005, 123, 562–570. [Google Scholar] [CrossRef]
- Yigit, S.; Demir, L.; Tarhan, M.O.; Cabuk, F.K.; Ellidokuz, H.; Erten, C.; Somali, I.; Dirican, A.; Cakalagaoglu, F.; Çakalaogaoglu, F. The clinicopathological significance of Bax and Bcl-2 protein expression with tumor infiltrating lymphocytes in ovarian carcinoma. Neoplasma 2012, 59, 475–485. [Google Scholar] [CrossRef]
- Jeong, S.H.; Lee, H.W.; Han, J.H.; Kang, S.Y.; Choi, J.-H.; Jung, Y.M.; Choi, H.; Oh, Y.T.; Park, K.J.; Hwang, S.C.; et al. Low Expression of Bax Predicts Poor Prognosis in Resected Non-small Cell Lung Cancer Patients with Non-squamous Histology. Jpn. J. Clin. Oncol. 2008, 38, 661–669. [Google Scholar] [CrossRef]
- Lopes-Costa, P.V.; dos Santos, A.R.; da Silva, B.B. The effect of raloxifene on Bax protein expression in breast carcinomas of postmenopausal women. Diagn. Cytopathol. 2012, 40, 570–574. [Google Scholar] [CrossRef] [PubMed]
- Tai, Y.T.; Lee, S.; Niloff, E.; Weisman, C.; Strobel, T.; Cannistra, S.A. BAX protein expression and clinical outcome in epithelial ovarian cancer. J. Clin. Oncol. 1998, 16, 2583–2590. [Google Scholar] [CrossRef] [PubMed]
- Moawadh, M.S.; Mir, R.; Tayeb, F.J.; Asim, O.; Ullah, M.F. Molecular Evaluation of the Impact of Polymorphic Variants in Apoptotic (Bcl-2/Bax) and Proinflammatory Cytokine (TNF-α/IL-8) Genes on the Susceptibility and Progression of Myeloproliferative Neoplasms: A Case-Control Biomarker Study. Curr. Issues Mol. Biol. 2023, 45, 3933–3952. [Google Scholar] [CrossRef] [PubMed]
- Knudson, C.M.; Tung, K.S.K.; Tourtellotte, W.G.; Brown, G.A.J.; Korsmeyer, S.J. Bax-Deficient Mice with Lymphoid Hyperplasia and Male Germ Cell Death. Science 1995, 270, 96–99. [Google Scholar] [CrossRef]
- Alam, M.; Kashyap, T.; Mishra, P.; Panda, A.K.; Nagini, S.; Mishra, R. Role and regulation of pro-apoptotic Bax in oral squamous cell carcinoma and drug resistance. Head Neck 2019, 41, 185–197. [Google Scholar] [CrossRef]
- Feng, Y.; Chen, X.L.; Zheng, Y.; Liu, Q.; Chen, H.W.; Cai, Y.H.; Cao, L.X.; Lai, X.L.; Pan, L.L.; Li, Y.; et al. Prognostic value and susceptibility of Bax rs4645878 polymorphism in cancer A systematic review and meta-analysis. Medicine 2018, 97, e11591. [Google Scholar] [CrossRef]
- Kholoussi, N.M.; el-Nabi, S.E.H.; Esmaiel, N.N.; Abd el-Bary, N.M.; el-Kased, A.F. Evaluation of Bax and Bak gene mutations and expression in breast cancer. BioMed Res. Int. 2014, 2014, 249372. [Google Scholar] [CrossRef]
- Wang, X.; Lin, Y.; Lan, F.; Yu, Y.; Ouyang, X.; Liu, W.; Xie, F.; Wang, X.; Huang, Q. BAX and CDKN1A polymorphisms correlated with clinical outcomes of gastric cancer patients treated with postoperative chemotherapy. Med. Oncol. 2014, 31, 249. [Google Scholar] [CrossRef]
- Gu, S.; Wu, Q.; Zhao, X.; Wu, W.; Gao, Z.; Tan, X.; Qian, J.; Chen, H.; Xie, Y.; Jin, L.; et al. Association of CASP3 polymorphism with hematologic toxicity in patients with advanced non-small-cell lung carcinoma treated with platinum-based chemotherapy. Cancer Sci. 2012, 103, 1451–1459. [Google Scholar] [CrossRef]
- Saxena, A.; Moshynska, O.; Sankaran, K.; Viswanathan, S.; Sheridan, D.P. Association of a novel single nucleotide polymorphism, G(-248)A, in the 5′-UTR of BAX gene in chronic lymphocytic leukemia with disease progression and treatment resistance. Cancer Lett. 2002, 187, 199–205. [Google Scholar] [CrossRef]
- Starczynski, J.; Pepper, C.; Pratt, G.; Hooper, L.; Thomas, A.; Milligan, D.; Bentley, P.; Fegan, C. Common polymorphism G(-248)A in the promoter region of the bax gene results in significantly shorter survival in patients with chronic lymphocytic leukemia once treatment is initiated. J. Clin. Oncol. 2005, 23, 1514–1521. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.-K.; Guo, Y.-L.; Tan, W.; Lin, D.-X. Functional Bax polymorphism associated with lung cancer susceptibility. Zhonghua Zhong Liu Za Zhi. 2010, 32, 324–327. [Google Scholar] [PubMed]
- Skogsberg, S.; Tobin, G.; Kröber, A.; Kienle, D.; Thunberg, U.; Åleskog, A.; Karlsson, K.; Laurell, A.; Merup, M.; Vilpo, J.; et al. The G(-248)A polymorphism in the promoter region of the Bax gene does not correlate with prognostic markers or overall survival in chronic lymphocytic leukemia. Leukemia 2006, 20, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Sohn, S.K.; Jung, J.T.; Kim, D.H.; Kim, J.G.; Kwak, E.K.; Park, T.I.; Shin, D.G.; Sohn, K.R.; Lee, K.B. Prognostic significance of bcl-2, bax, and p53 expression in diffuse large B-cell lymphoma. Am. J. Hematol. 2003, 73, 101–107. [Google Scholar] [CrossRef]
- Soucek, L.; Whitfield, J.; Martins, C.P.; Finch, A.J.; Murphy, D.J.; Sodir, N.M.; Karnezis, A.N.; Swigart, L.B.; Nasi, S.; Evan, G.I. Modelling Myc inhibition as a cancer therapy. Nature 2008, 455, 679–683. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.H.; van Riggelen, J.; Yetil, A.; Fan, A.C.; Bachireddy, P.; Felsher, D.W. Cellular senescence is an important mechanism of tumor regression upon c-Myc inactivation. Proc. Natl. Acad. Sci. USA 2007, 104, 13028–13033. [Google Scholar] [CrossRef]
- Haiman, C.A.; Patterson, N.; Freedman, M.L.; Myers, S.R.; Pike, M.C.; Waliszewska, A.; Neubauer, J.; Tandon, A.; Schirmer, C.; McDonald, G.J.; et al. Multiple regions within 8q24 independently affect risk for prostate cancer. Nat. Genet. 2007, 39, 638–644. [Google Scholar] [CrossRef]
- Liu, J.; Hua, R.-X.; Fu, W.; Zhu, J.; Jia, W.; Zhang, J.; Zhou, H.; Cheng, J.; Xia, H.; Liu, G.; et al. MYC gene associated polymorphisms and Wilms tumor risk in Chinese children: A four-center case-control study. Ann. Transl. Med. 2019, 7, 475. [Google Scholar] [CrossRef]
- Pan, J.; Zhu, J.; Wang, M.; Yang, T.; Hu, C.; Yang, J.; Zhang, J.; Cheng, J.; Zhou, H.; Xia, H.; et al. Association of MYC gene polymorphisms with neuroblastoma risk in Chinese children: A four-center case–control study. J. Gene Med. 2020, 22, e3190. [Google Scholar] [CrossRef]
- Yang, T.; Wen, Y.; Li, J.; Tan, T.; Yang, J.; Pan, J.; Hu, C.; Yao, Y.; Zhang, J.; Li, S.; et al. Association of CMYC polymorphisms with hepatoblastoma risk. Transl. Cancer Res. 2020, 9, 849–855. [Google Scholar] [CrossRef]
- Amatori, S.; Persico, G.; Fanelli, M. Real-time quantitative PCR array to study drug-induced changes of gene expression in tumor cell lines. J. Cancer Metastasis Treat. 2017, 3, 90–99. [Google Scholar] [CrossRef]
- Sarkan, S.A.; Sharna, R.P. Modulation of c-myc, max, and mad gene expression during neural differentiation of embryonic stem cells by all-trans-retinoic acid. Gene Expr. 2002, 10, 125–135. [Google Scholar]
- Santos, M.M.; Tannuri, A.C.A.; Coelho, M.C.M.; Goncalves, J.O.; Serafini, S.; Ferraz da Silva, L.F.; Tannuri, U. Immediate expression of c-fos and c-jun mRNA in a model of intestinal autotransplantation and ischemia-reperfusion in situ. Clinics 2015, 70, 373–379. [Google Scholar] [CrossRef] [PubMed]
| Parameters | Patients (N = 98) | Controls (N = 100) |
|---|---|---|
| Age (years) | N (%) | N (%) |
| <40 | 9 (9.18%) | 12 (12.00%) |
| >40 | 89 (90.12%) | 88 (88.00%) |
| Education | N (%) | N (%) |
| Primary | 0 (0.00%) | 0 (0.00%) |
| Lower secondary | 0 (0.00%) | 0 (0.00%) |
| Vocational | 32 (32.65%) | 32 (32.00%) |
| Secondary | 36 (36.73%) | 38 (38.00%) |
| Higher | 30 (30.62%) | 30 (30.00%) |
| Occupation | N (%) | N (%) |
| Farmers | 16 (16.33%) | 14 (14.00%) |
| Other manual workers | 52 (53.06%) | 42 (42.00%) |
| Office workers or other specialists | 12 (12.25%) | 21 (21.00%) |
| Health professionals | 6 (6.12%) | 7 (7.00%) |
| Teachers/educators | 8 (8.16%) | 10 (10.00%) |
| Unemployed | 4 (4.08) | 6 (6.00%) |
| Reported economic status | N (%) | N (%) |
| Bad | 0 (0.00%) | 0 (0.00%) |
| Average | 32 (32.65%) | 38 (38.00%) |
| Good | 58 (59.18%) | 56 (56.00%) |
| Very good | 8 (8.17%) | 6 (6.00%) |
| Parameter | Group | Mean ± SD | Median | 95% CI | p |
|---|---|---|---|---|---|
| Leukocytes 109/L | OC | 8.44 ± 4.48 | 7.62 | 7.63–8.99 | 0.006 |
| Control | 6.98 ± 3.61 | 6.87 | 6.51–7.85 | ||
| Erythrocytes 1012/L | OC | 4.58 ± 0.68 | 4.42 | 4.36–4.50 | 0.150 |
| Control | 4.39 ± 0.53 | 4.41 | 4.28–4.49 | ||
| Platelets 109/L | OC | 330.87 ± 155.42 | 288.00 | 295.75–370.94 | 0.016 |
| Control | 266.48 ± 64.87 | 262.50 | 253.47–279.48 | ||
| Hemoglobin g/dL | OC | 7.56 ± 1.01 | 7.54 | 7.36–7.68 | 0.036 |
| Control | 7.61 ± 0.98 | 7.84 | 7.42–7.92 | ||
| Hematocrit | OC | 0.38 ± 0.37 | 0.37 | 0.36–0.38 | 0.061 |
| Control | 0.47 ± 0.88 | 0.39 | 0.32–0.65 | ||
| Glucose mg/dL | OC | 98.14 ± 18.48 | 94.00 | 93.46–99.88 | <0.001 |
| Control | 88.91 ± 15.96 | 85.86 | 72.32–91.91 | ||
| Sodium mmol/L | OC | 139.55 ± 2.93 | 138.98 | 138.98–140.11 | 0.037 |
| Control | 139.48 ± 2.65 | 139.20 | 138.85–139.96 | ||
| Potassium mmol/L | OC | 4.36 ± 0.42 | 4.36 | 4.30–4.50 | 0.419 |
| Control | 4.28 ± 0.35 | 4.25 | 4.21–4.34 | ||
| Creatinine mg/dL | OC | 0.85 ± 0.49 | 0.74 | 0.74–0.95 | 0.644 |
| Control | 0.83 ± 0.26 | 0.77 | 0.72–0.94 | ||
| eGFR mL/min/1.73 m2 | OC | 85.63 ± 32.53 | 86.21 | 77.42–94.54 | 0.292 |
| Control | 95.97 ± 24.65 | 96.97 | 82.96–107.34 | ||
| Total protein g/dL | OC | 6.98 ± 0.84 | 6.98 | 6.78–7.23 | 0.321 |
| Control | 7.09 ± 0.42 | 7.15 | 6.98–7.38 | ||
| Uric acid mg/dL | OC | 5.28 ± 1.78 | 4.86 | 4.69–5.59 | 0.728 |
| Control | 5.24 ± 1.56 | 5.14 | 4.37–5.98 | ||
| Urea mg/dL | OC | 32.30 ± 18.99 | 27.70 | 27.22–36.65 | 0.286 |
| Control | 33.42 ± 10.48 | 31.00 | 28.09–37.87 | ||
| D-dimer ng/mL | OC | 3349.653 ± 2024.42 | 1931.00 | 2559.53–4299.54 | <0.001 |
| Control | 790.69 ± 424.54 | 466.50 | 398.20–1282.28 | ||
| Fibrinogen g/L | OC | 7.65 ± 5.34 | 4.62 | 0.27–13.94 | <0.001 |
| Control | 2.98 ± 0.78 | 2.99 | 2.81–3.22 | ||
| INR | OC | 1.18 ± 0.24 | 1.13 | 1.13–1.24 | 0.075 |
| Control | 1.19 ± 0.07 | 1.12 | 1.09–1.14 | ||
| PTT | OC | 12.99 ± 2.62 | 12.50 | 12.46–13.57 | 0.068 |
| Control | 12.28 ± 0.67 | 12.22 | 11.84–12.37 | ||
| APTT | OC | 30.10 ± 3.72 | 30.20 | 29.27–30.94 | 0.965 |
| Control | 30.41 ± 3.22 | 30.55 | 28.84–31.98 | ||
| Systolic pressure mmHg | OC | 125.27 ± 13.27 | 124.00 | 121.39–126.76 | 0.175 |
| Control | 121.81 ± 14.88 | 120.00 | 118.81–124.82 | ||
| Diastolic pressure mmHg | OC | 78.99 ± 14.69 | 80.00 | 76.98–82.89 | 0.741 |
| Control | 79.32 ± 8.45 | 80.00 | 77.62–81.01 | ||
| CA-125 U/mL | OC | 778.51 ± 444.25 | 295.00 | 505.27–1055.74 | <0.001 |
| Control | 128.29 ± 100.43 | 20.81 | 8.49–266.64 | ||
| HE4 pmol/L | OC | 1709.42 ± 1208.42 | 364.45 | 128.20–3987.76 | 0.008 |
| Control | 84.47 ± 31.30 | 74.26 | 8.16–141.83 |
| Women with Ovarian Cancer | Healthy Group | p a | OR | |||
|---|---|---|---|---|---|---|
| Observed Values n (%) | Expected Values (%) | Observed Values n (%) | Expected Values (%) | |||
| BCL2 rs2279115 C>A | ||||||
| CC | 23 (23.47%) | 23.99 | 16 (16.00%) | 15.60 | 0.046 | 1.61 |
| CA | 50 (51.02%) | 49.98 | 47 (47.00%) | 47.80 | 1.17 | |
| AA | 25 (25.51%) | 26.03 | 37 (37.00%) | 36.60 | 1.74 | |
| Total | 98 (100%) | 100.0 | 100 (100%) | 100.0 | ||
| Allele | ||||||
| C | 96 (48.98%) | - | 79 (39.50%) | - | 0.039 | 1.24 |
| A | 100 (51.02%) | - | 121 (60.50%) | - | 1.24 | |
| Total | 196 (100.0%) | - | 200 (100.0%) | - | ||
| BAX rs4645878 G>A | ||||||
| GG | 74 (75.51%) | 76.13 | 73 (73.00%) | 73.10 | 0.401 | 1.14 |
| GA | 23 (23.47%) | 22.24 | 25 (25.00%) | 24.80 | 1.09 | |
| AA | 1 (1.02%) | 1.63 | 2 (2.00%) | 2.10 | 1.10 | |
| Total | 98 (100%) | 100.0 | 100 (100%) | 100.0 | ||
| Allele | ||||||
| G | 171 (87.25%) | - | 171 (85.50%) | - | 0.062 | 1.16 |
| A | 25 (12.75%) | - | 29 (14.50%) | - | 1.16 | |
| Total | 196 (100.0%) | - | 200 (100.0%) | - | ||
| c-MYC rs4645943 C>T | ||||||
| AA | 33 (33.67%) | 30.36 | 38 (38.00%) | 34.81 | 0.331 | 0.82 |
| AG | 42 (42.86%) | 49.48 | 42 (42.00%) | 48.38 | 0.96 | |
| GG | 23 (23.47%) | 20.16 | 20 (20.00%) | 16.81 | 0.81 | |
| Total | 98 (100%) | 100.0 | 100 (100%) | 100.0 | ||
| Allele | ||||||
| A | 108 (55.10%) | - | 118 (59.00%) | - | 0.642 | 0.76 |
| G | 88 (44.9%) | - | 62 (41.00%) | - | 0.76 | |
| Total | 196 (100.0%) | - | 200 (100.0%) | - | ||
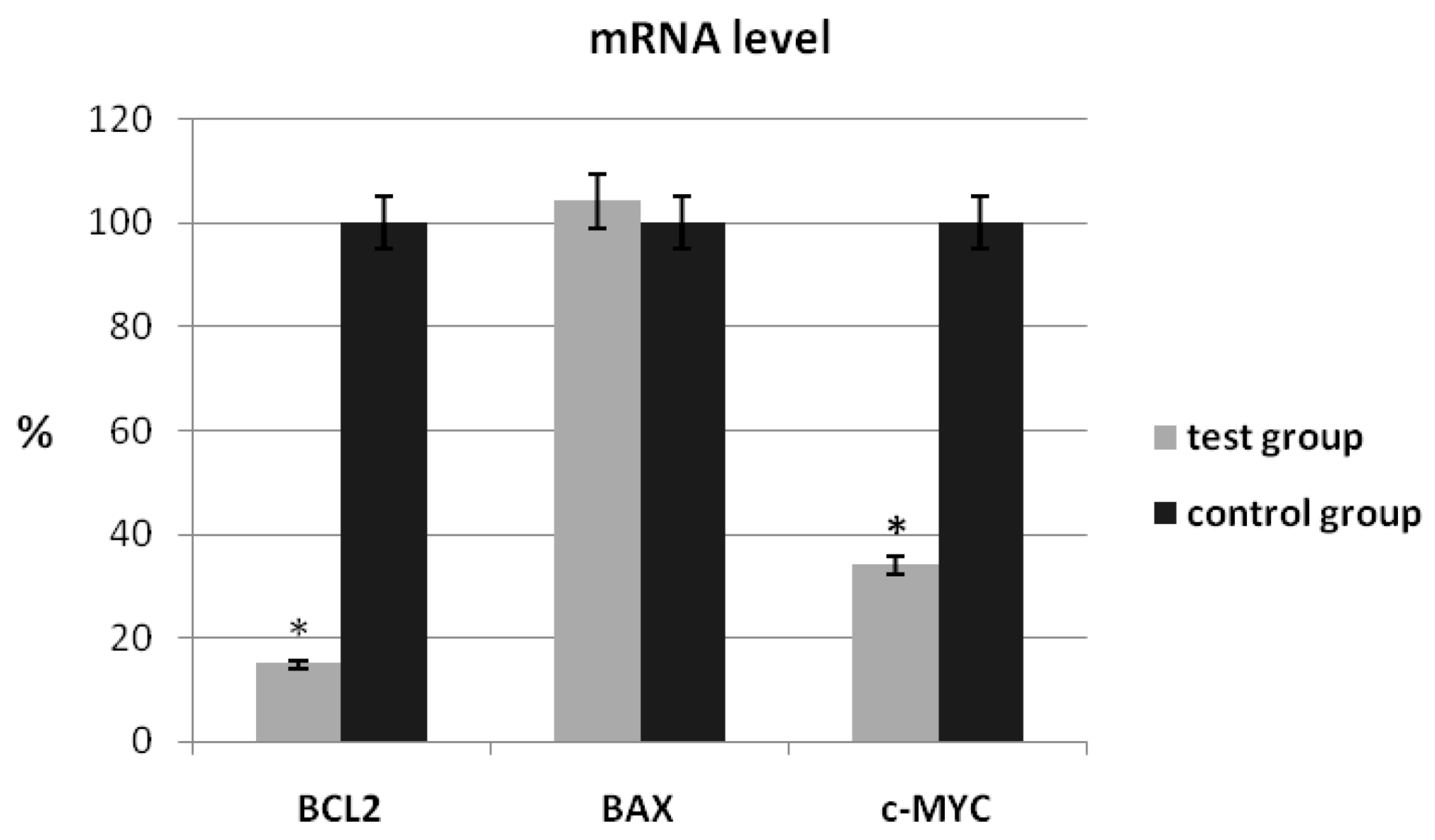
| Gene | Patients with Ovarian Cancer | Control Group | p-Value * |
|---|---|---|---|
| BCL2 | 1.452 ± 0.002 | 2.324 ± 0.002 | 0.046 |
| BAX | 0.982 ± 0.002 | 0.921 ± 0.001 | 0.314 |
| c-MYC | 0.725 ± 0.003 | 1.122 ± 0.003 | 0.058 |
| Gene | Forward 5′-3′ | Reverse 5′-3′ | Reference |
|---|---|---|---|
| BCL2 | CTGGTGGACAACATCGCCCT | TCTTCAGAGACAGCCAGGAGAAAT | [51] |
| BAX | CAAACTGGTGCTCAAGGCCC | GGGCGTCCCAAAGTAGGAGA | [51] |
| c-MYC | ATCTGCGACGAGGAAGAGAA | ATCGCAGATGAAGCTCTGGT | [52] |
| GAPDH | GCAAATTCCATGGCACCGT | TCGCCCCACTTGATTTTGG | [51] |
| β-ACTIN | GCCAGAGCGGGAGTGGTGAA | GGCTTGGGCTCAGGGTCATT | [53] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Olbromski, P.J.; Bogacz, A.; Bukowska, M.; Kamiński, A.; Moszyński, R.; Pawlik, P.; Szeliga, A.; Kotrych, K.; Czerny, B. Analysis of the Polymorphisms and Expression Levels of the BCL2, BAX and c-MYC Genes in Patients with Ovarian Cancer. Int. J. Mol. Sci. 2023, 24, 16309. https://doi.org/10.3390/ijms242216309
Olbromski PJ, Bogacz A, Bukowska M, Kamiński A, Moszyński R, Pawlik P, Szeliga A, Kotrych K, Czerny B. Analysis of the Polymorphisms and Expression Levels of the BCL2, BAX and c-MYC Genes in Patients with Ovarian Cancer. International Journal of Molecular Sciences. 2023; 24(22):16309. https://doi.org/10.3390/ijms242216309
Chicago/Turabian StyleOlbromski, Piotr Józef, Anna Bogacz, Marta Bukowska, Adam Kamiński, Rafał Moszyński, Piotr Pawlik, Anna Szeliga, Katarzyna Kotrych, and Bogusław Czerny. 2023. "Analysis of the Polymorphisms and Expression Levels of the BCL2, BAX and c-MYC Genes in Patients with Ovarian Cancer" International Journal of Molecular Sciences 24, no. 22: 16309. https://doi.org/10.3390/ijms242216309





