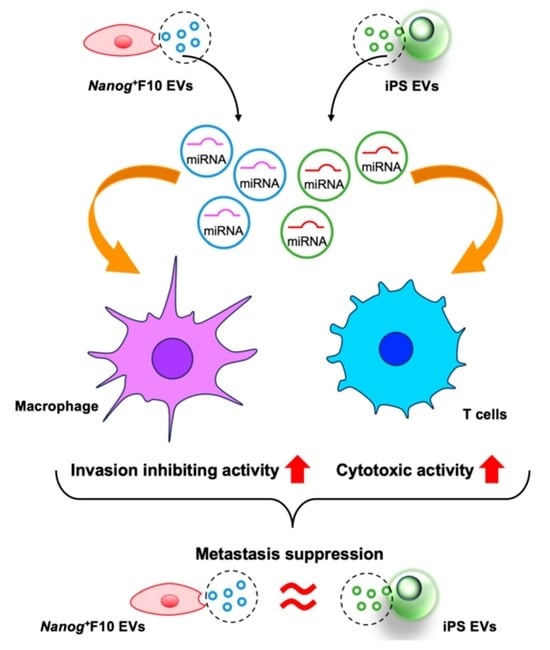
Comparative Study of Metastasis Suppression Effects of Extracellular Vesicles Derived from Anaplastic Cell Lines, Nanog-Overexpressing Melanoma, and Induced Pluripotent Stem Cells
Abstract
:1. Introduction
2. Results
2.1. Transfer of EVs to Various Organs
2.2. Effects of EVs as a Vaccine



2.3. Effects on Immune Cells



2.4. Results of miRNA Analysis




3. Discussion
4. Material and Methods
4.1. Cell Culture
4.2. Preparation of EVs
4.3. Western Analysis of CD81 and Gapdh
4.4. Analysis of the Transfer of EVs to Various Organs in a Mouse
4.5. In Vivo Test of the Effects of EVs on the Metastasis of Melanoma
4.6. Invasion Ability Test
4.7. Phagocytic Activity Test
4.8. Cytotoxic Activity Test
4.9. miRNA Analysis
4.10. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wattanawongdon, W.; Hahnvajanawong, C.; Namwat, N.; Kanchanawat, S.; Boonmars, T.; Earanaikoon, P.; Leelayuwat, C.; Techasen, A.; Seubwai, W. Establishment and characterization of gemcitabine-resistant human cholangiocarcinoma cell lines with multidrug resistance and enhanced invasiveness. Int. J. Oncol. 2015, 47, 398–410. [Google Scholar] [CrossRef] [PubMed]
- Kajiyama, H.; Shibata, K.; Terauchi, M.; Yamashita, M.; Ino, K.; Nawa, A.; Kikkawa, F. Chemoresistance to paclitaxel induces epithelial-mesenchymal transition and enhances metastatic potential for epithelial ovarian carcinoma cells. Int. J. Oncol. 2007, 31, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Speers, C.; Zhao, S.; Liu, M.; Bartelink, H.; Pierce, L.J.; Feng, F.Y. Development and validation of a novel radiosensitivity signature in human breast cancer. Clin. Cancer Res. 2015, 21, 3667–3677. [Google Scholar] [CrossRef] [PubMed]
- Carlos-Reyes, A.; Muñiz-Lino, M.A.; Romero-Garcia, S.; López-Camarillo, C.; Hernández-de la Cruz, O.N. Biological adaptations of tumor cells to radiation therapy. Front. Oncol. 2021, 11, 718636. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Delgado, I.; Calzada-Fraile, D.; Sánchez-Madrid, S. Immune regulation by dendritic cell extracellular vesicles in cancer immunotherapy and vaccines. Cancers 2020, 12, 3558. [Google Scholar] [CrossRef] [PubMed]
- Santos, P.; Almeida, F. Exosome-based vaccines: History, current state, and clinical trials. Front. Immunol. 2021, 12, 711565. [Google Scholar] [CrossRef] [PubMed]
- Lucotti, S.; Kenific, C.M.; Zhang, H.; Lyden, D. Extracellular vesicles and particles impact the systemic landscape of cancer. EMBO J. 2022, 41, e109288. [Google Scholar] [CrossRef]
- Buzas, E.I. The roles of extracellular vesicles in the immune system. Nat. Rev. Immunol. 2023, 23, 236–250. [Google Scholar] [CrossRef]
- Loric, S.; Denis, J.A.; Desbene, C.; Sabbah, M.; Conti, M. Extracellular vesicles in breast cancer: From biology and function to clinical diagnosis and therapeutic management. Int. J. Mol. Sci. 2023, 24, 7208. [Google Scholar] [CrossRef]
- Damo, M.; Wilson, D.S.; Simeoni, E.; Hibbell, J.A. TLR-3 stimulation improves anti-tumor immunity elicited by dendritic cell exosome-based vaccines in a murine model of melanoma. Sci. Rep. 2015, 5, 17622. [Google Scholar] [CrossRef]
- Kitai, Y.; Kawasaki, T.; Sueyoshi, T.; Kobiyama, K.; Ishii, K.J.; Zou, J.; Akira, S.; Matsuda, T.; Kawai, T. DNA-containing exosomes derived from cancer cells treated with topotecan activate a STING-dependent pathway and reinforce antitumor immunity. J. Immunol. 2017, 198, 1649–1659. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Wu, J.; Xu, A.; Ahmeqd, S.; Sami, A.; Chibbar, R.; Freywald, A.; Zheng, C.; Xiang, J. Heterologous human/rat HER2-specific exosome-targeted T cell vaccine stimulates potent humoral and CTL responses leading to enhanced circumvention of HER2 tolerance in double transgenic HLA-A2/HER2 mice. Vaccine 2018, 36, 1414–1422. [Google Scholar] [CrossRef] [PubMed]
- Nakazawa, Y.; Nishiyama, N.; Koizumi, H.; Kanemaru, K.; Nakahashi-Oda, C.; Shibuya, A. Tumor-derived extracellular vesicles regulate tumor-infiltrating regulatory T cells via the inhibitory immunoreceptor CD300a. eLife 2021, 10, e61999. [Google Scholar] [CrossRef] [PubMed]
- Rezaei, R.; Baghaei, K.; Hashemi, S.M.; Zali, M.R.; Ghanbarian, H.; Amani, D. Tumor-derived exosomes enriched by miRNA-124 promote anti-tumor immune response in CT-26 tumor-bearing mice. Front. Med. 2021, 8, 619939. [Google Scholar] [CrossRef] [PubMed]
- Claridge, B.; Lozanto, J.; Poh, Q.H.; Greening, D.W. Development of extracellular vesicle therapeutics: Challenges, considerations, and opportunities. Front. Cell Dev. Biol. 2021, 9, 734720. [Google Scholar] [CrossRef] [PubMed]
- van Neerven, S.M.; Tieken, M.; Vermeulen, L.; Bijlsma, M.F. Bidirectional interconversion of stem and non-stem cancer cell populations: A reassessment of theoretical models for tumor heterogeneity. Mol. Cell. Oncol. 2016, 3, e1098791. [Google Scholar] [CrossRef]
- Saito, M.; Kishi, R.; Sasai, T.; Hatakenaka, T.; Matsuki, N.; Minagawa, S. Effect of Nanog overexpression on the metastatic potential of a mouse melanoma cell line B16-BL. Mol. Cell. Biochem. 2021, 476, 2651–2661. [Google Scholar] [CrossRef]
- Hatakenaka, T.; Matsuki, N.; Minagawa, S.; May, C.K.S.; Saito, M. Anti-metastatic function of extracellular vesicles derived from Nanog-overexpressing melanoma. Curr. Oncol. 2022, 29, 1029–1046. [Google Scholar] [CrossRef]
- Chen, C.L.; Kumar, D.B.U.; Punj, V.; Xu, J.; Sher, L.; Tahara, S.M.; Hess, S.; Machida, K. NANOG metabolically reprograms tumor-initiating stem-like cells through tumorigenic changes in oxidative phosphorylation and fatty acid metabolism. Cell Metab. 2016, 23, 206–219. [Google Scholar] [CrossRef]
- Gawlik-Rzemieniewska, N.; Bednarek, I. The role of NANOG transcriptional factor in the development of malignant phenotype of cancer cells. Cancer Biol. Ther. 2016, 17, 1–10. [Google Scholar] [CrossRef]
- Saito, M. Novel roles of Nanog in cancer cells and their extracellular vesicles. Cells 2022, 11, 3881. [Google Scholar] [CrossRef] [PubMed]
- Adamiak, M.; Cheng, G.; Bobis-Wozowicz, S.; Zhao, L.; Kedracka-Krok, S.; Samanta, A.; Karnas, E.; Xuan, Y.T.; Skupien-Rabian, B.; Chen, X.; et al. Induced pluripotent stem cell (iPSC)–derived extracellular vesicles are safer and more effective for cardiac repair than iPSCs. Circ. Res. 2018, 122, 296–309. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.X.F.; Johansson, H.; Wood, M.J.A.; Andaloussi, S.E. Considerations and implications in the purification of extracellular vesicles—A cautionary tale. Front. Neurosci. 2019, 13, 1067. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Xu, J.; Li, H.; Hu, Y.; Yu, G. Identification of metastasis-asspciated microRNAs in metastatic melanoma by miRNA expression profile and experimental validation. Front. Genet. 2021, 12, 663110. [Google Scholar] [CrossRef]
- Mathavan, S.; Kue, C.S.; Kumar, S. Identification of potential candidate genes for lip and oral cavity cancer using network analysis. Genomics Inform. 2021, 19, e4. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, M.; Hong, C.P.; Oshima, H.; Sakai, E.; Kim, S.-J.; Oshima, M. Loss of wild-type p53 promotes mutant p53-driven metastasis through acquisition of survival and tumor-initiating properties. Nat. Comm. 2020, 11, 2333. [Google Scholar] [CrossRef] [PubMed]
- Maaty, M.A.A.; Terzic, J.; Keime, C.; Rovito, D.; Lutzing, R.; Yanushko, D.; Parisotto, M.; Grelet, E.; Namer, I.J.; Lindner, V.; et al. Hypoxia-mediated stabilization of HIF1A in prostatic intraepithelial neoplasia promotes cell plasticity and malignant progression. Sci. Adv. 2022, 8, eabo2295. [Google Scholar] [CrossRef]


| miRNA | Log2(FC) | Number of Target Genes | |||
|---|---|---|---|---|---|
| p < 0.05CWCS ≤ −0.4 | Targeted by Each miRNA among Top30 Genes | ||||
| Nanog+F10-EVs/F10-EVs | miR-18a-5p | 2.46 | 42 | 301 | 4 |
| miR-3473e | 1.75 | 6 | 0 | ||
| miR-19a-3p | 1.20 | 126 | 11 | ||
| miR-3473b | 1.18 | 127 | 5 | ||
| miR-210-3p | −3.62 | 6 | 104 | 0 | |
| miR-369-3p | −2.52 | 7 | 2 | ||
| miR-122-5p | −1.79 | 29 | 5 | ||
| miR-706 | −1.28 | 62 | 3 | ||
| Sum | 405 | 30 | |||
| iPS-EVs/Nanog+F10-EVs | miR-466h-3p | 7.62 | 58 | 275 | 1 |
| miR-706 | 6.23 | 62 | 3 | ||
| miR-323-3p | 6.23 | 10 | 0 | ||
| miR-466f-3p | 6.07 | 145 | 5 | ||
| miR-30a-3p | −4.99 | 49 | 394 | 0 | |
| miR-342-5p | −4.74 | 335 | 20 | ||
| let-7c-5p | −4.63 | 2 | 0 | ||
| miR-423-3p | −4.36 | 8 | 1 | ||
| Sum | 669 | 30 | |||
| Gene | MCC Score | Degree | Regulation by mi RNA | |
|---|---|---|---|---|
| Nanog+F10-EVs | Trp53 | 1058 | 23 | up-regulated-miR-3473b |
| Hif1a | 990 | 19 | up-regulated-miR-18a-5p | |
| Esr1 | 919 | 11 | up-regulated-miR-18a-5p | |
| Atm | 872 | 9 | up-regulated-miR-18a-5p | |
| Cdkn1b | 847 | 7 | down-regulated-miR-706 | |
| iPS-EVs | Ins1 | 13,801 | 19 | up-regulated-miR-466f-3p |
| Kitl | 13,493 | 15 | up-regulated-miR-466f-3p | |
| Fgf16 | 13,254 | 11 | up-regulated-miR-466f-3p | |
| Grb2 | 11,784 | 15 | up-regulated-miR-466f-3p | |
| Fgf1 | 11,640 | 11 | down-regulated-miR-342-5p |
| Specific Keywords | |||
|---|---|---|---|
| Immune Responses | Immune Cells | Inflammation Signaling | |
| Immune | T cell | Inflammation | JAK/Stat |
| Cytotoxicity | B cell | Cytokine | CD3 |
| Phagocytosis | Dendritic | Interleukin | CD4 |
| Helper | NF-kappaB | CD8 | |
| Macrophage | TGF-beta | ||
| Myeloid | |||
| Gene with 4 Degrees | Regulation by miRNA | Associated Keywords | |
|---|---|---|---|
| Nanog+F10-EVs | Rnf11 | up-regulated-miR-19a-3p | Immune, Cytokine, NF-kappaB, Tgf-beta |
| iPS-EVs | Malt1 | up-regulated-miR-466f-3p | Immune, T cell, B cell, Helper cell |
| Il1f6 | down-regulated-miR-30a-3p | T cell, Dendritic cell, Interleukin, NF-kappaB | |
| Cmklr1 | down-regulated-miR-342-5p | Immune, Macrophage, Inflammation, NF-kappaB | |
| Siglec1 | down-regulated-miR-342-5p | T cell, Macrophage, B cell, CD8 | |
| Cd28 | down-regulated-miR-342-5p | Cytotoxicity, Immune, T cell, CD4 |
| EVs | miRNA | Target Genes | Prediction By | Hub Gene Candidates |
|---|---|---|---|---|
| Nanog+F10-EVs/F10- EVs | up-regulated miRNAs (miR-18a-5p, miR-19a-3p, miR-3473b) | down-regulation of target genes | GD-based analysis | Trp53 |
| Hif1a | ||||
| Esr1 | ||||
| Atm | ||||
| SK-based analysis | Rnf11 | |||
| down-regulated miRNAs (miR-706) | up-regulation of target genes | GD-based analysis | Cdkn1b | |
| SK-based analysis | - | |||
| iPS-EVs/Nanog+F10-EVs | up-regulated miRNAs (miR-466f-3p) | down-regulation of target genes | GD-based analysis | Ins1 |
| Kitl | ||||
| Fgf16 | ||||
| Grb2 | ||||
| SK-based analysis | Malt1 | |||
| down-regulated miRNAs (miR-342-5p) | up-regulation of target genes | GD-based analysis | Fgf1 | |
| SK-based analysis | Il1f6 | |||
| Cmklr1 | ||||
| Siglec1 | ||||
| Cd28 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khoo, C.S.M.; Henmi, T.; Saito, M. Comparative Study of Metastasis Suppression Effects of Extracellular Vesicles Derived from Anaplastic Cell Lines, Nanog-Overexpressing Melanoma, and Induced Pluripotent Stem Cells. Int. J. Mol. Sci. 2023, 24, 17206. https://doi.org/10.3390/ijms242417206
Khoo CSM, Henmi T, Saito M. Comparative Study of Metastasis Suppression Effects of Extracellular Vesicles Derived from Anaplastic Cell Lines, Nanog-Overexpressing Melanoma, and Induced Pluripotent Stem Cells. International Journal of Molecular Sciences. 2023; 24(24):17206. https://doi.org/10.3390/ijms242417206
Chicago/Turabian StyleKhoo, Celine Swee May, Takuya Henmi, and Mikako Saito. 2023. "Comparative Study of Metastasis Suppression Effects of Extracellular Vesicles Derived from Anaplastic Cell Lines, Nanog-Overexpressing Melanoma, and Induced Pluripotent Stem Cells" International Journal of Molecular Sciences 24, no. 24: 17206. https://doi.org/10.3390/ijms242417206
APA StyleKhoo, C. S. M., Henmi, T., & Saito, M. (2023). Comparative Study of Metastasis Suppression Effects of Extracellular Vesicles Derived from Anaplastic Cell Lines, Nanog-Overexpressing Melanoma, and Induced Pluripotent Stem Cells. International Journal of Molecular Sciences, 24(24), 17206. https://doi.org/10.3390/ijms242417206