Comprehensive Analysis of NAC Transcription Factors Reveals Their Evolution in Malvales and Functional Characterization of AsNAC019 and AsNAC098 in Aquilaria sinensis
Abstract
:1. Introduction
2. Results
2.1. Identification of the NAC Gene in A. sinensis and Eleven Other Species
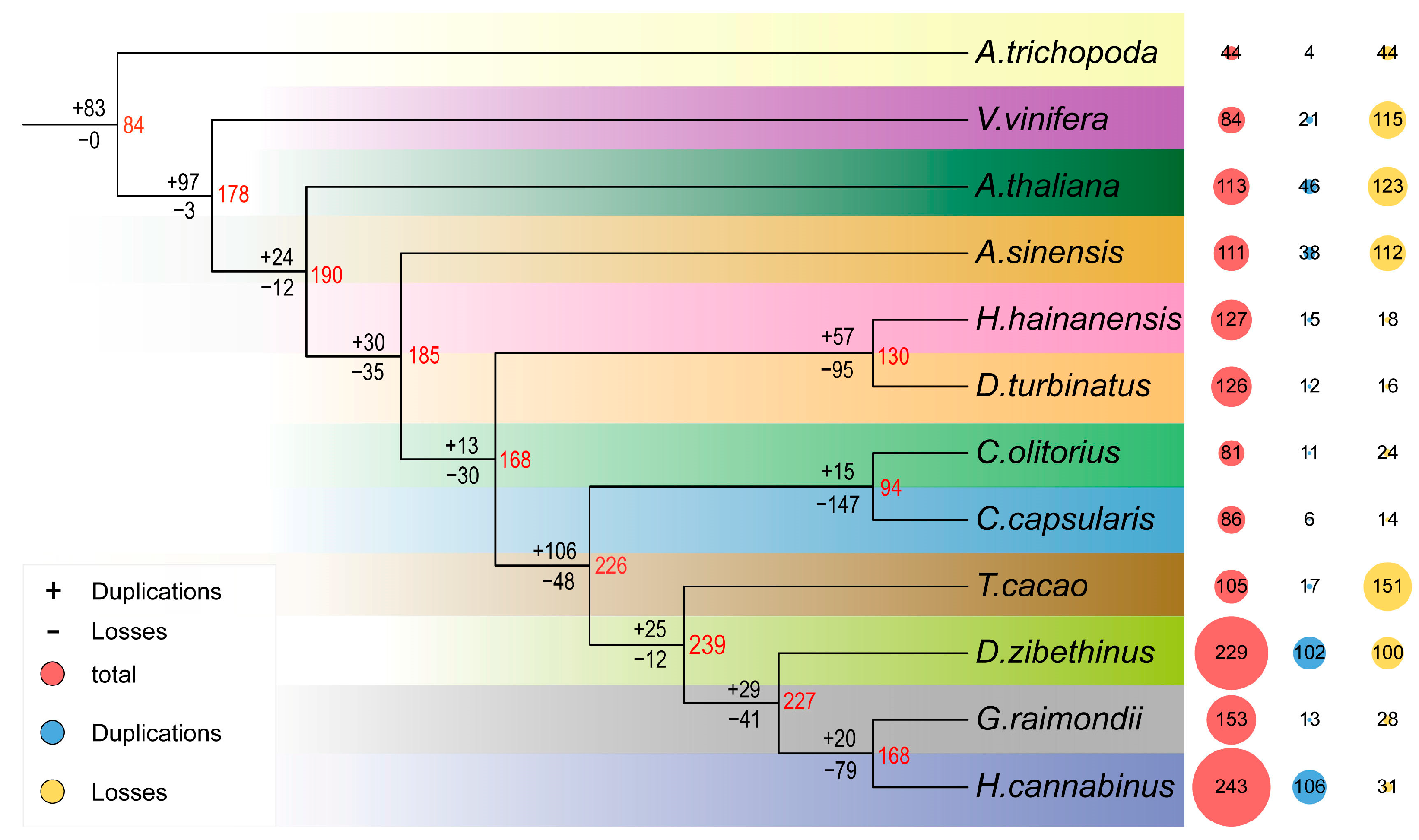
2.2. Comparative Evolutionary Analysis of the NAC Family in Malvales
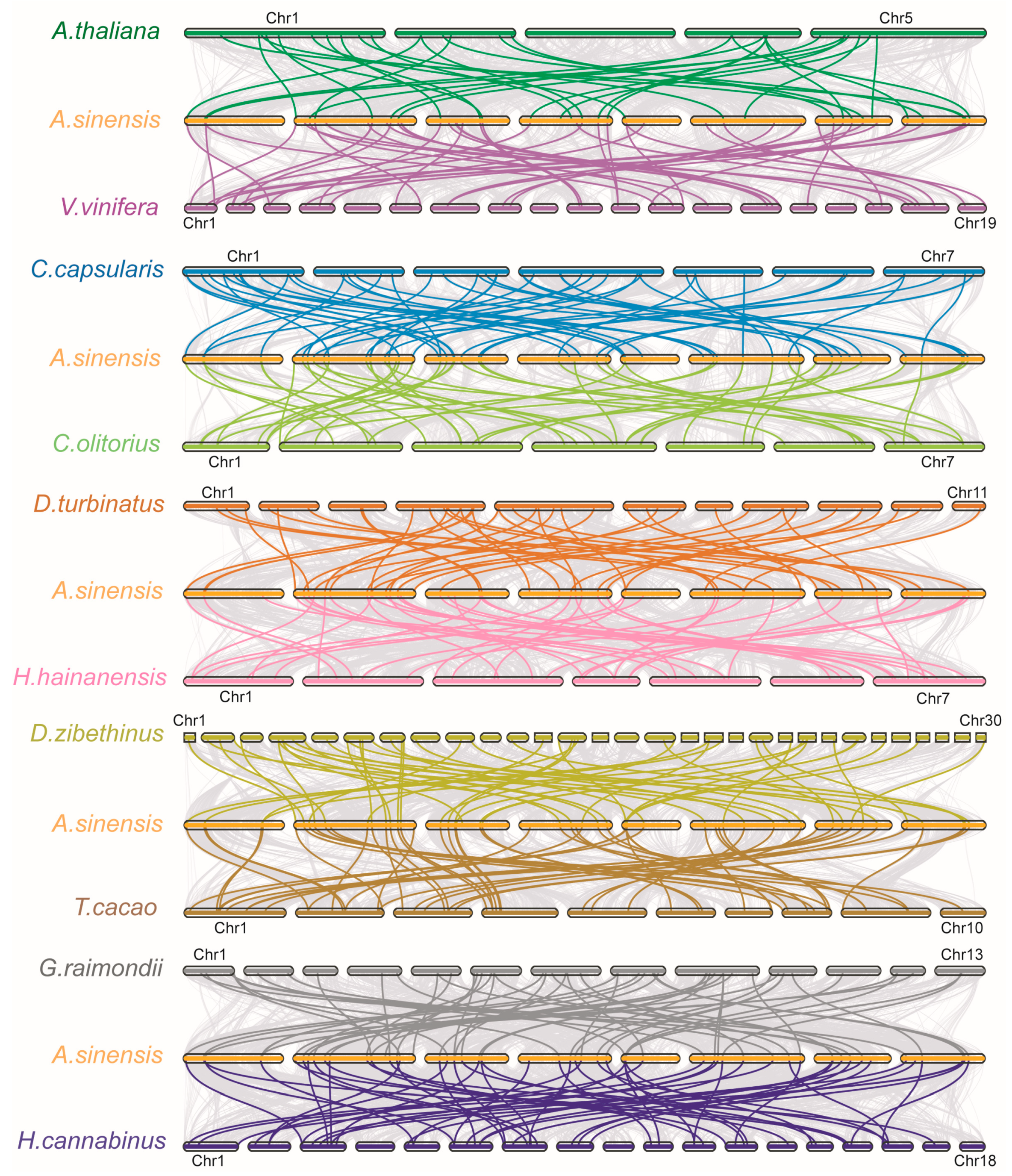
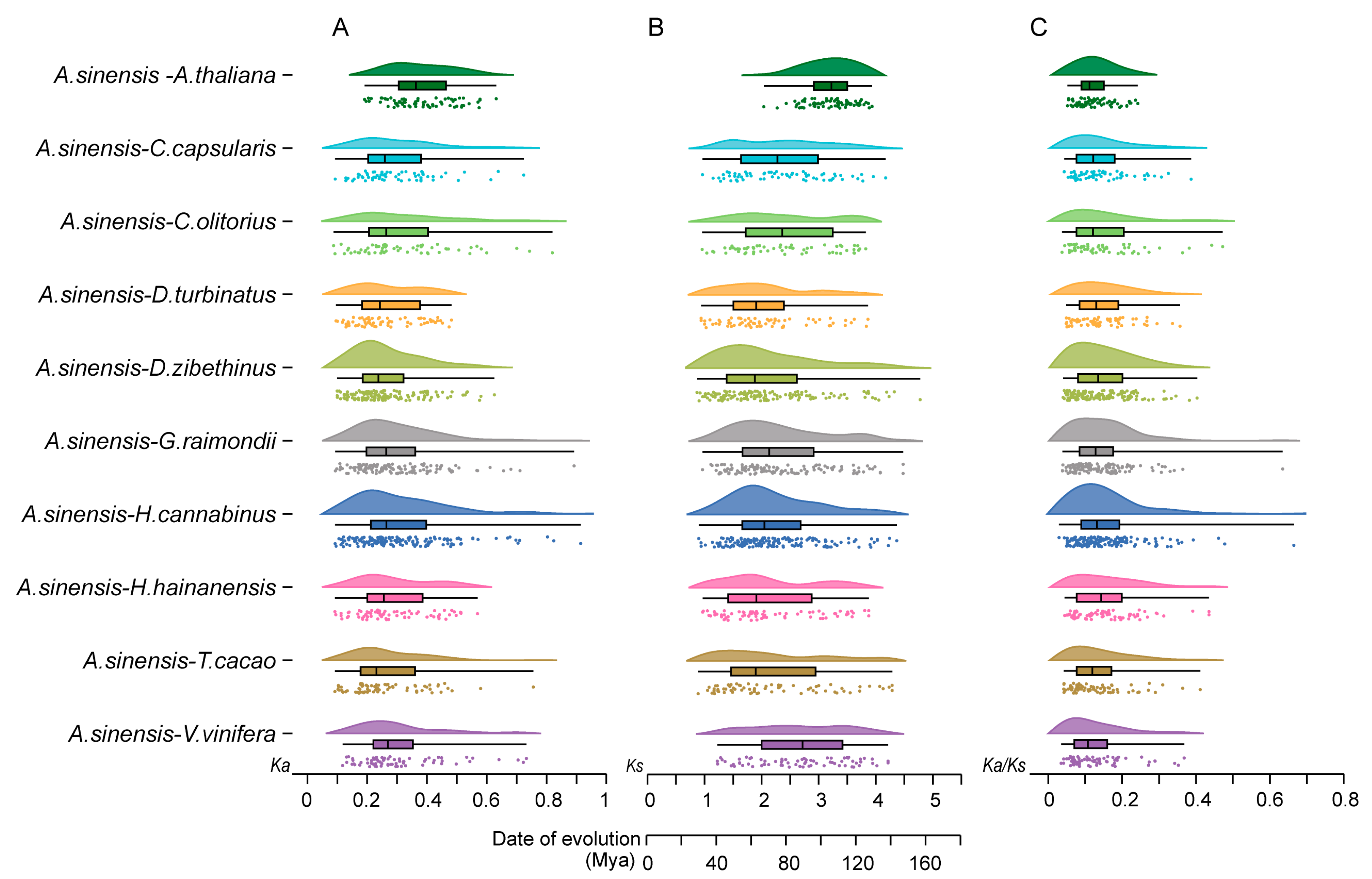
2.3. Interspecific Synteny Analysis of AsNAC and NAC from Ten Other Species
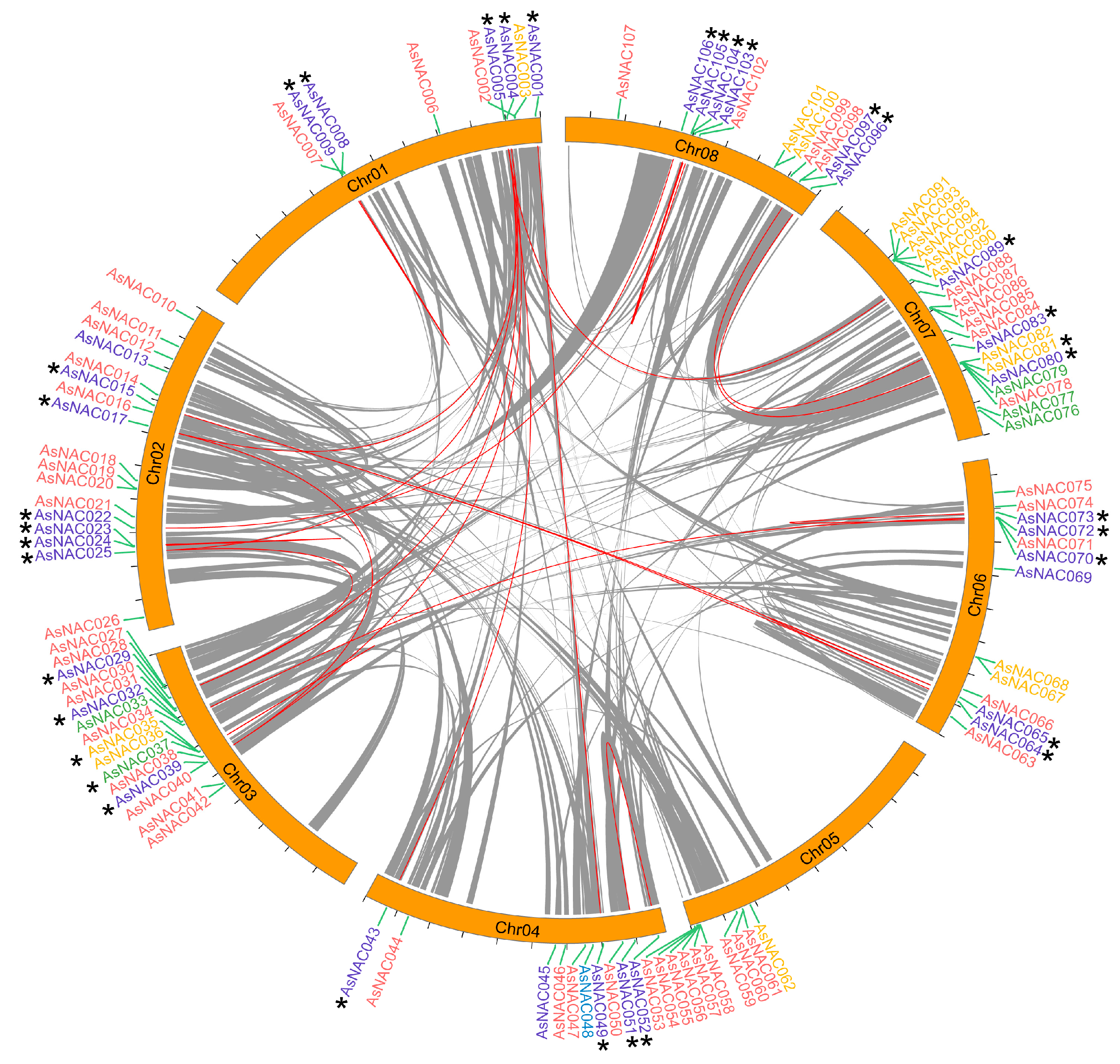
2.4. Characterization, Chromosomal Localization and Intraspecific Syntenic Analysis of the AsNAC Genes
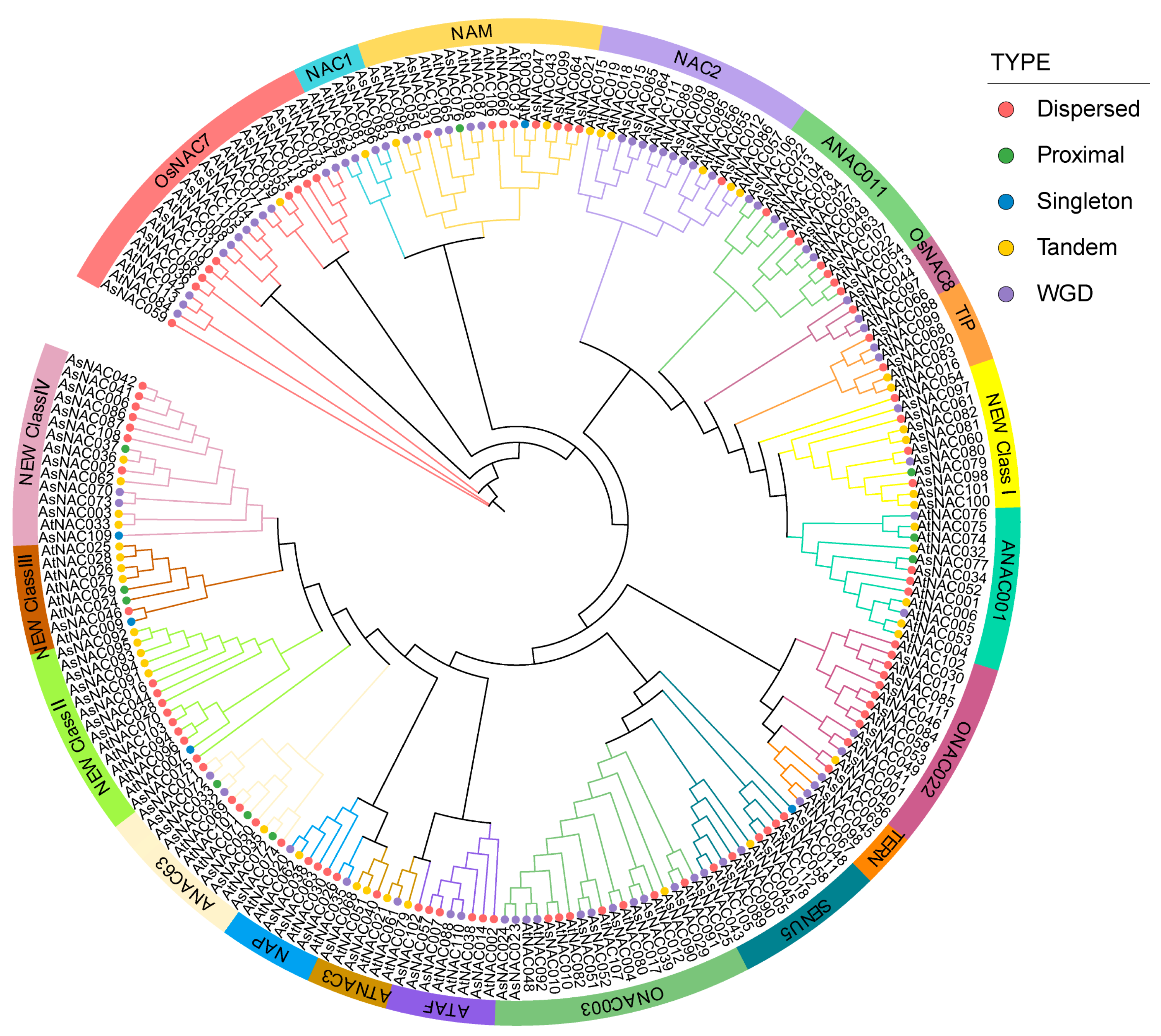
2.5. Phylogenetic Analysis of AsNAC and AtNAC
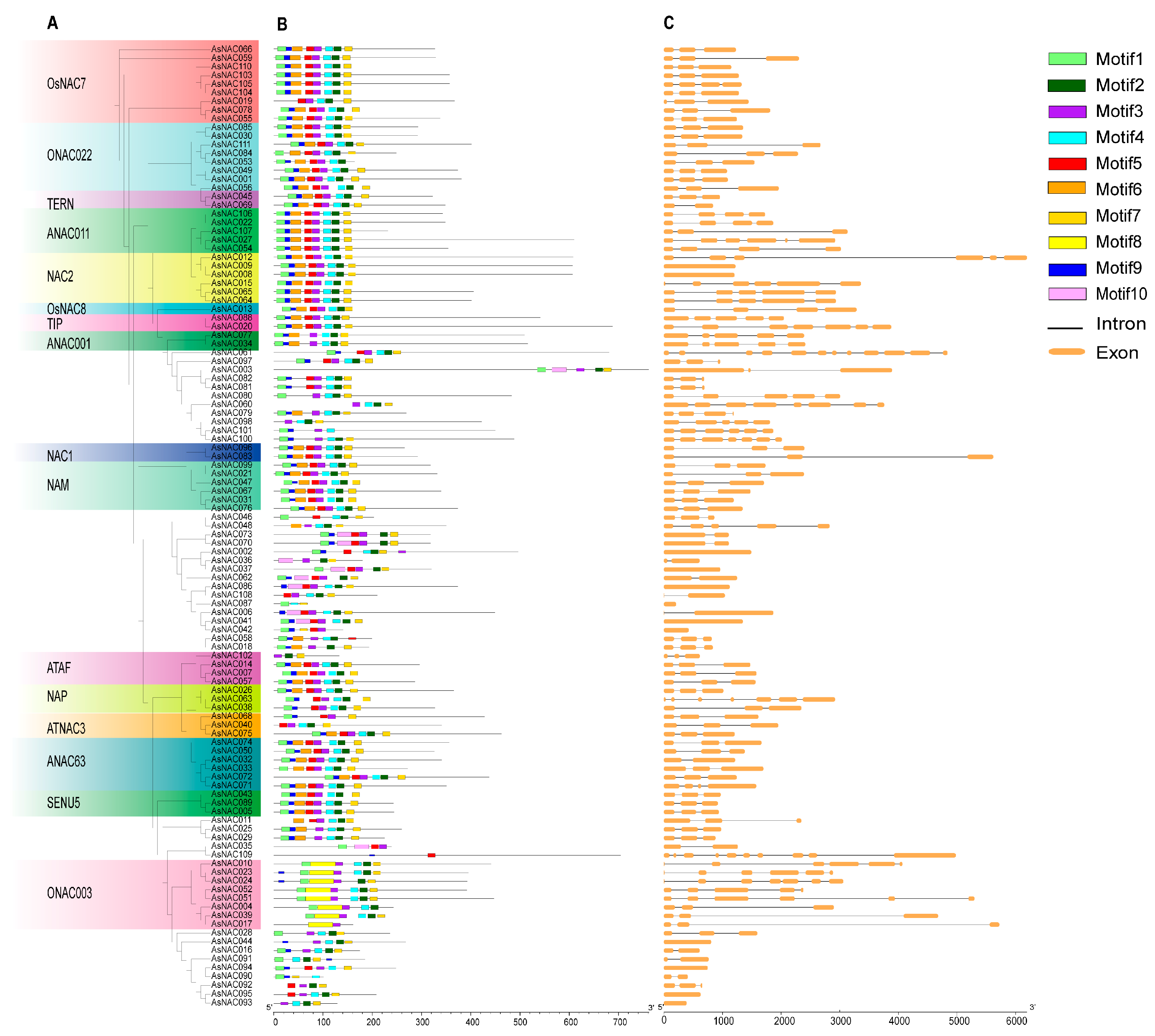
2.6. Gene Structure and Conserved Motif Analysis of the AsNAC Gene Family
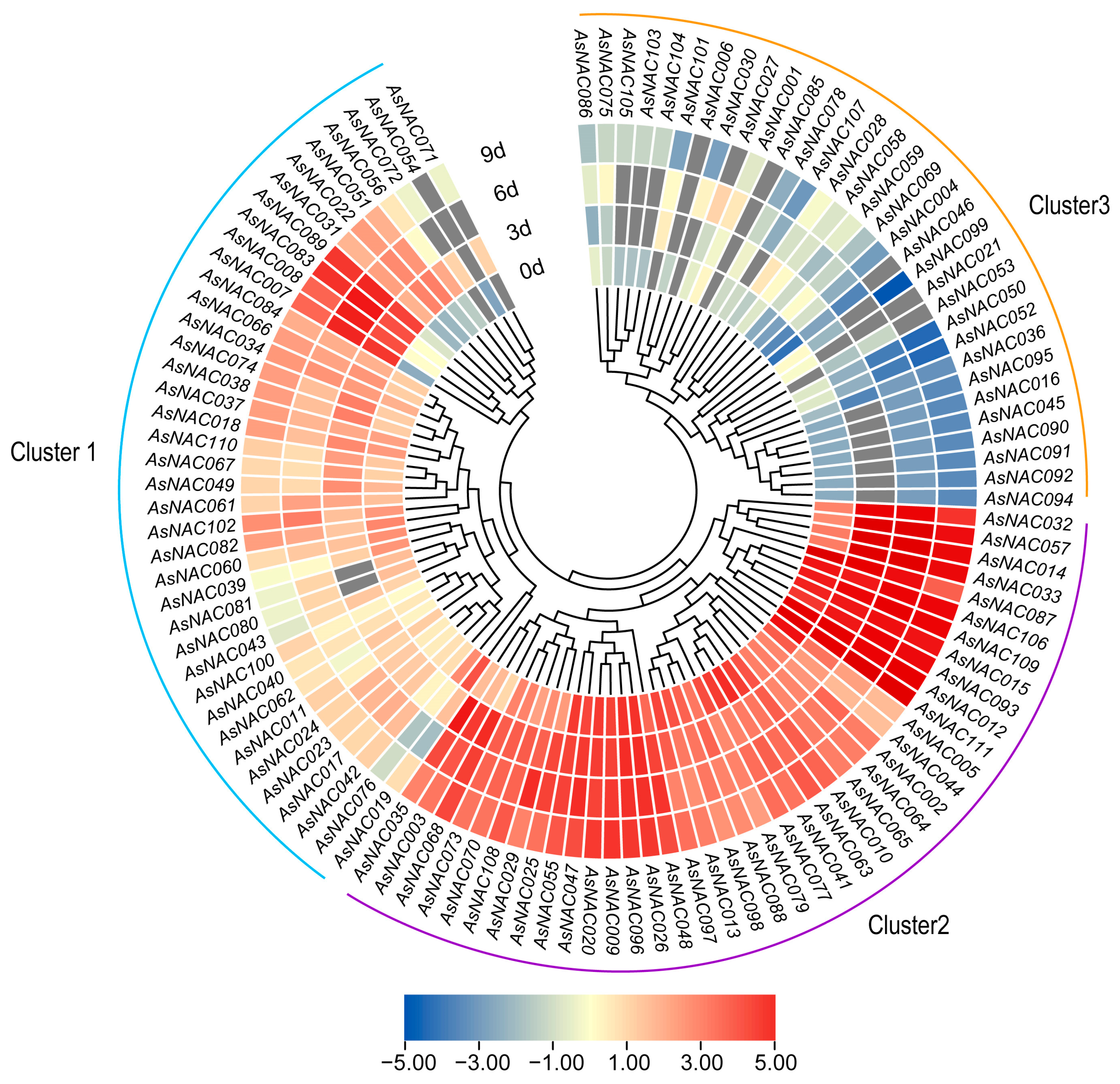
2.7. Expression Pattern of AsNAC Genes under Agarwood Inducer Treatment
2.8. Expression of AsNAC Genes under Injury Stress
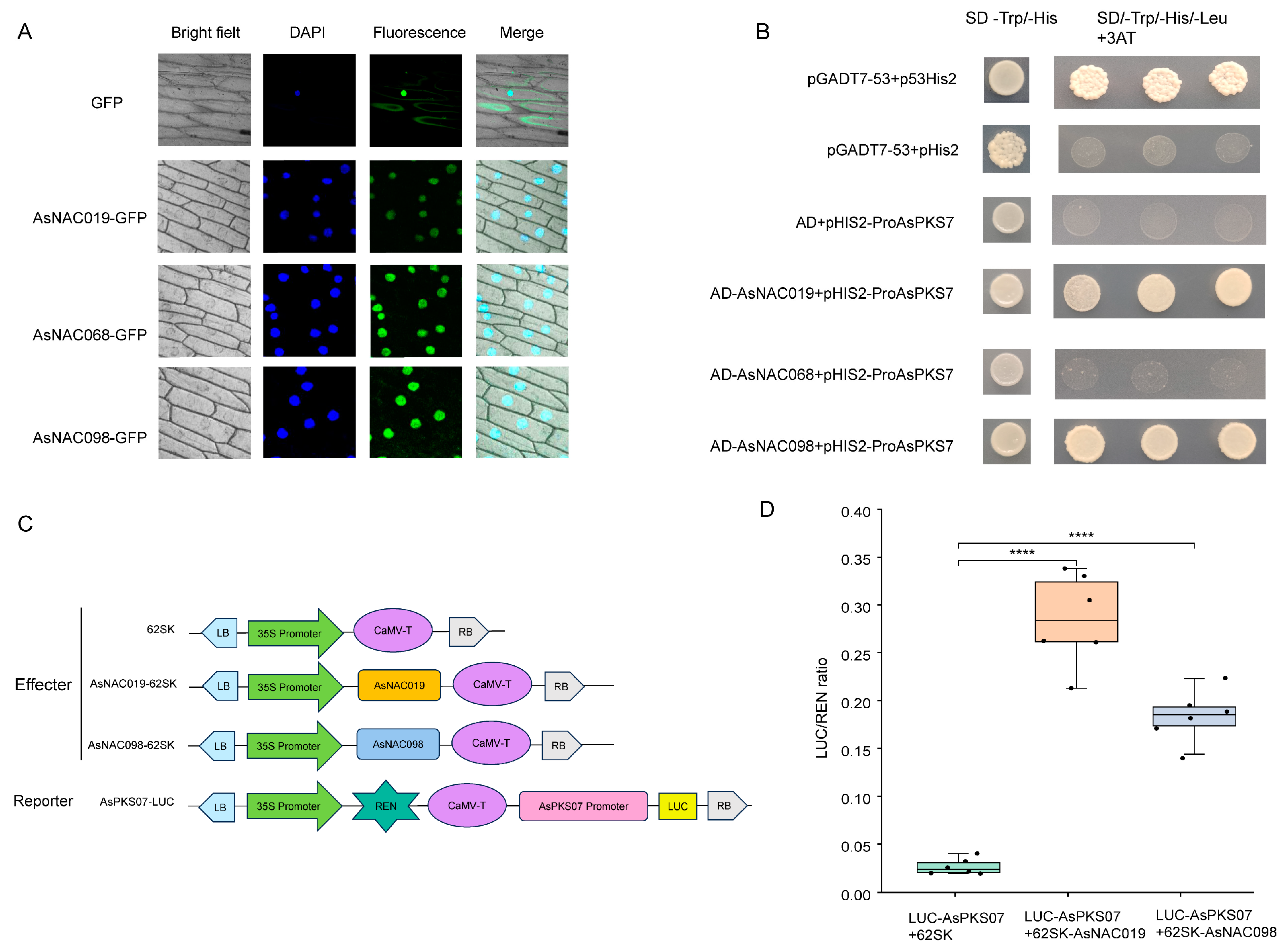
2.9. Subcellular Localization of AsNAC019, AsNAC068 and AsNAC098
2.10. Transcriptional Activation of AsPKS07 by Interaction with AsNAC019, AsNAC068 and AsNAC098
3. Discussion
3.1. Identification and Evolution of NAC Transcription Factors
3.2. Potential Function of NAC Genes in A. sinensis
3.3. Possible Role of NAC Genes in Secondary Metabolite Synthesis in A. sinensis
4. Materials and Methods
4.1. Plant Material and Treatment
4.2. Identification of NAC Gene Family in the Genomes of A. sinensis and 11 Other Plants
4.3. Expansion and Contraction of NAC Genes
4.4. Identification of Syntenic Genes and Calculation of Ka, Ks and Ka/Ks Values
4.5. Phylogenetic Analysis and Chromosomal Localization of the AsNAC Transcription Factors
4.6. Gene Structure and Conserved Motif Analyses
4.7. Real-Time Quantitative PCR (RT-qPCR) Analysis
4.8. Gene Cloning and Subcellular Localization of AsNAC Genes
4.9. Yeast One-Hybrid (Y1H) Assay
4.10. Dual-Luciferase Assay
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Liu, X.; Wang, T.; Bartholomew, E.; Black, K.; Dong, M.; Zhang, Y.; Yang, S.; Cai, Y.; Xue, S.; Weng, Y.; et al. Comprehensive analysis of NAC transcription factors and their expression during fruit spine development in cucumber (Cucumis sativus L.). Hortic. Res. 2018, 5, 31. [Google Scholar] [CrossRef] [PubMed]
- Ooka, H.; Satoh, K.; Doi, K.; Nagata, T.; Otomo, Y.; Murakami, K.; Matsubara, K.; Osato, N.; Kawai, J.; Carninci, P.; et al. Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res. 2003, 10, 239–247. [Google Scholar] [CrossRef]
- Shen, S.Y.; Zhang, Q.R.; Shi, Y.; Sun, Z.M.; Zhang, Q.Q.; Hou, S.J.; Wu, R.L.; Jiang, L.B.; Zhao, X.Y.; Guo, Y.Q. Genome-wide analysis of the NAC domain transcription factor gene family in Theobroma cacao. Genes 2020, 11, 35. [Google Scholar] [CrossRef] [PubMed]
- Yue, Y.Z.; Li, L.; Li, Y.L.; Li, H.Y.; Ding, W.J.; Shi, T.T.; Chen, G.W.; Yang, X.L.; Wang, L.G. Genome-wide analysis of NAC transcription factors and characterization of the cold stress response in Sweet osmanthus. Plant Mol. Biol. Rep. 2020, 38, 314–330. [Google Scholar] [CrossRef]
- Munir, N.; Chen, Y.K.; Chen, X.H.; Nawaz, M.A.; Iftikhar, J.; Rizwan, H.M.; Xu, S.; Lin, Y.L.; Xu, X.H.; Lai, Z.X. Genome-wide identification and comprehensive analyses of NAC transcription factor gene family and expression patterns during somatic embryogenesis in Dimocarpus longan Lour. Plant Physiol. Bioch. 2020, 157, 169–184. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Qiao, Y.; Li, C.; Hou, B. The NAC transcription factors play core roles in flowering and ripening fundamental to fruit yield and quality. Front. Plant Sci. 2023, 14, 1095967. [Google Scholar] [CrossRef]
- Han, K.J.; Zhao, Y.; Sun, Y.H.; Li, Y. NACs, generalist in plant life. Plant Bio. J. 2023, 21, 1–25. [Google Scholar] [CrossRef]
- Moyano, E.; Martinez-Rivas, F.J.; Blanco-Portales, R.; Molina-Hidalgo, F.J.; Ric-Varas, P.; Matas-Arroyo, A.J.; Caballero, J.L.; Munoz-Blanco, J.; Rodriguez-Franco, A. Genome-wide analysis of the NAC transcription factor family and their expression during the development and ripening of the Fragaria x ananassa fruits. PLoS ONE 2018, 13, e0196953. [Google Scholar] [CrossRef]
- Min, X.Y.; Jin, X.Y.; Zhang, Z.S.; Wei, X.Y.; Ndayambaza, B.; Wang, Y.R.; Liu, W.X. Genome-wide identification of NAC transcription factor family and functional analysis of the abiotic stress-responsive genes in Medicago sativa L. J. Plant Growth Regul. 2020, 39, 324–337. [Google Scholar] [CrossRef]
- Hussey, S.G.; Saidi, M.N.; Hefer, C.A.; Myburg, A.A.; Grima-Pettenati, J. Structural, evolutionary and functional analysis of the NAC domain protein family in Eucalyptus. New Phytol. 2015, 206, 1337–1350. [Google Scholar] [CrossRef]
- Yang, X.F.; Kim, M.Y.; Ha, J.; Lee, S.H. Overexpression of the soybean NAC gene GmNAC109 increases lateral root formation and abiotic stress tolerance in transgenic Arabidopsis plants. Front. Plant Sci. 2019, 10, 1036. [Google Scholar] [CrossRef]
- Ma, N.N.; Feng, H.L.; Meng, X.; Li, D.; Yang, D.Y.; Wu, C.G.; Meng, Q.W. Overexpression of tomato SlNAC1 transcription factor alters fruit pigmentation and softening. BMC Plant Biol. 2014, 14, 351. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.F.; Tian, S.W.; Yu, Y.T.; Ren, Y.; Guo, S.G.; Zhang, J.; Li, M.Y.; Zhang, H.Y.; Gong, G.Y.; Wang, M.; et al. Natural variation in the NAC transcription factor NONRIPENING contributes to melon fruit ripening. J. Integr. Plant Biol. 2022, 64, 1448–1461. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Guo, C.; Li, Z.Y.; Sun, J.H.; Deng, Z.C.; Wen, L.C.; Li, X.X.; Guo, Y.F. Potato NAC transcription factor StNAC053 enhances salt and drought tolerance in transgenic Arabidopsis. Int. J. Mol. Sci. 2021, 22, 2568. [Google Scholar] [CrossRef]
- Trishla, V.S.; Kirti, B. Structure-function relationship of Gossypium hirsutum NAC transcription factor, GhNAC4 with regard to ABA and abiotic stress responses. Plant Sci. 2021, 302, 110718. [Google Scholar] [CrossRef] [PubMed]
- Nuruzzaman, M.; Sharoni, A.M.; Kikuchi, S. Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Front. Microbiol. 2013, 4, 248. [Google Scholar] [CrossRef]
- Liu, M.Y.; Ma, Z.T.; Sun, W.J.; Huang, L.; Wu, Q.; Tang, Z.Z.; Bu, T.L.; Li, C.L.; Chen, H. Genome-wide analysis of the NAC transcription factor family in Tartary buckwheat (Fagopyrum tataricum). BMC Genom. 2019, 20, 113. [Google Scholar] [CrossRef]
- Li, W.H.; Zeng, Y.L.; Yin, F.L.; Wei, R.; Mao, X.F. Genome-wide identification and comprehensive analysis of the NAC transcription factor family in sunflower during salt and drought stress. Sci. Rep. 2021, 11, 19865. [Google Scholar] [CrossRef]
- Jensen, M.K.; Hagedorn, P.H.; de Torres-Zabala, M.; Grant, M.R.; Rung, J.H.; Collinge, D.B.; Lyngkjaer, M.F. Transcriptional regulation by an NAC (NAM-ATAF1,2-CUC2) transcription factor attenuates ABA signalling for efficient basal defence towards Blumeria graminis f. sp. hordei in Arabidopsis. Plant J. 2008, 56, 867–880. [Google Scholar] [CrossRef]
- Jung, S.E.; Kim, T.H.; Shim, J.S.; Bang, S.W.; Bin Yoon, H.; Oh, S.H.; Kim, Y.S.; Oh, S.J.; Seo, J.S.; Kim, J.K. Rice NAC17 transcription factor enhances drought tolerance by modulating lignin accumulation. Plant Sci. 2022, 323, 111404. [Google Scholar] [CrossRef]
- Guan, C.J.; Wang, Y.; Zhang, Y.N. Molecular characterization and transcription profiling of NAC genes in Lilium pumilum under abiotic stresses. Int. J. Agric. Biol. 2020, 23, 484–492. [Google Scholar]
- Cheng, Z.H.; Zhang, X.M.; Zhao, K.; Zhou, B.R.; Jiang, T.B. Ectopic expression of a poplar gene NAC13 confers enhanced tolerance to salinity stress in transgenic Nicotiana tabacum. J. Plant Res. 2020, 133, 727–737. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Lin, X.; Zhang, D.; Li, Q.; Zhao, X.; Chen, S. Genome-wide analysis of NAC gene family in Betula pendula. Forests 2019, 10, 741. [Google Scholar] [CrossRef]
- Li, W.; Li, X.X.; Chao, J.T.; Zhang, Z.L.; Wang, W.F.; Guo, Y.F. NAC family transcription factors in tobacco and their potential role in regulating leaf senescence. Front. Plant Sci. 2018, 9, 1900. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Seo, P.J.; Lee, H.J.; Park, C.M. A NAC transcription factor NTL4 promotes reactive oxygen species production during drought-induced leaf senescence in Arabidopsis. Plant J. 2012, 70, 831–844. [Google Scholar] [CrossRef]
- de Zelicourt, A.; Diet, A.; Marion, J.; Laffont, C.; Ariel, F.; Moison, M.L.; Zahaf, O.; Crespi, M.; Gruber, V.; Frugier, F. Dual involvement of a Medicago truncatula NAC transcription factor in root abiotic stress response and symbiotic nodule senescence. Plant J. 2012, 70, 220–230. [Google Scholar] [CrossRef]
- He, H.; Li, Q.Y.; Fang, L.; Yang, W.; Xu, F.C.; Yan, Y.; Mao, R.J. Comprehensive analysis of NAC transcription factors in Scutellaria baicalensis and their response to exogenous ABA and GA3. Int. J. Biol. Macromol. 2023, 244, 125290. [Google Scholar] [CrossRef]
- Liu, C.; Zhao, M.Z.; Ma, H.D.; Zhang, Y.; Liu, Q.; Liu, S.Z.; Wang, Y.F.; Wang, K.Y.; Zhang, M.P.; Wang, Y. The NAC transcription factor PgNAC41-2 gene involved in the regulation of ginsenoside biosynthesis in Panax ginseng. Int. J. Mol. Sci. 2023, 24, 11946. [Google Scholar] [CrossRef]
- Wang, Z.; Zhang, Z.P.; Wang, P.P.; Qin, C.; He, L.Q.; Kong, L.Y.; Ren, W.C.; Liu, X.B.; Ma, W. Genome-wide identification of the NAC transcription factors family and regulation of metabolites under salt stress in Isatis indigotica. Int. J. Biol. Macromol. 2023, 240, 124436. [Google Scholar] [CrossRef]
- Ding, X.P.; Mei, W.L.; Lin, Q.; Wang, H.; Wang, J.; Peng, S.Q.; Li, H.L.; Zhu, J.H.; Li, W.; Wang, P.; et al. Genome sequence of the agarwood tree Aquilaria sinensis (Lour.) Spreng: The first chromosome-level draft genome in the Thymelaeceae family. Gigascience 2020, 9, giaa013. [Google Scholar] [CrossRef]
- Li, W.; Chen, H.Q.; Wang, H.; Mei, W.L.; Dai, H.F. Natural products in agarwood and Aquilaria plants: Chemistry, biological activities and biosynthesis. Nat. Prod. Rep. 2021, 38, 528–565. [Google Scholar] [CrossRef] [PubMed]
- Dafni, A.; Böck, B. Medicinal plants of the Bible—Revisited. J. Ethnobiol. Ethnomed. 2019, 15, 57. [Google Scholar] [CrossRef]
- Yang, H.R.; Wang, P.; Liu, F.Z.; Yuan, J.Z.; Cai, C.H.; Wu, F.; Jiang, B.; Mei, W.L.; Dai, H.F. Dimeric 2-(2-phenethyl)chromones from agarwood of Aquilaria filaria. Fitoterapia 2023, 165, 105422. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Huo, H.; Zhang, H.; Wang, L.; Meng, Y.; Jin, F.; Wang, X.; Zhao, Y.; Zhao, Y.; Tu, P. 2-(2-phenylethyl) chromone-enriched extract of the resinous heartwood of Chinese agarwood (Aquilaria sinensis) protects against taurocholic acid-induced gastric epithelial cells apoptosis through Perk/eIF2α/CHOP pathway. Phytomedicine 2022, 98, 153935. [Google Scholar] [CrossRef]
- Chen, L.Y.; Chen, H.Q.; Cai, C.H.; Yuan, J.Z.; Gai, C.J.; Liu, S.B.; Mei, W.L.; Dai, H.F. Seven new 2-(2-phenethyl)chromone derivatives from agarwood of Aquilaria walla. Fitoterapia 2023, 165, 105421. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.N.; Dong, W.H.; Huang, S.Z.; Li, W.; Kong, F.D.; Wang, H.; Wang, J.; Mei, W.L.; Dai, H.F. Three new sesquiterpenoids from agarwood of Aquilaria crassna. Fitoterapia 2016, 114, 7–11. [Google Scholar] [CrossRef]
- Yu, Z.X.; Wang, C.H.; Zheng, W.; Chen, D.L.; Liu, Y.Y.; Yang, Y.; Wei, J.H. Anti-inflammatory 5,6,7,8-tetrahydro-2-(2-phenylethyl)chromones from agarwood of Aquilaria sinensis. Bioorg. Chem. 2020, 99, 103789. [Google Scholar] [CrossRef]
- Mallika, V.; Sivakumar, K.C.; Soniya, E.V. Evolutionary implications and physicochemical analyses of selected proteins of type III polyketide synthase family. Evol. Bioinform. 2011, 7, 41–53. [Google Scholar] [CrossRef]
- Yang, Y.; Zhu, J.H.; Wang, H.; Guo, D.; Wang, Y.; Mei, W.L.; Peng, S.Q.; Dai, H.F. Systematic investigation of the R2R3-MYB gene family in Aquilaria sinensis reveals a transcriptional repressor AsMYB054 involved in 2-(2-phenylethyl) chromone biosynthesis. Int. J. Biol. Macromol. 2023, 244, 125302. [Google Scholar] [CrossRef]
- Jauhal, A.A.; Newcomb, R.D. Assessing genome assembly quality prior to downstream analysis: N50 versus BUSCO. Mol. Ecol. Resour. 2021, 21, 1416–1421. [Google Scholar] [CrossRef]
- Zhang, H.; Ding, X.P.; Wang, H.; Chen, H.Q.; Dong, W.H.; Zhu, J.H.; Wang, J.; Peng, S.Q.; Dai, H.F.; Mei, W.L. Systematic evolution of bZIP transcription factors in Malvales and functional exploration of AsbZIP1 and AsbZIP41 in Aquilaria sinensis. Front. Plant Sci. 2023, 14, 1243323. [Google Scholar] [CrossRef] [PubMed]
- Crow, K.D.; Wagner, G.P.; Investigators, S.T.-N.Y. What is the role of genome duplication in the evolution of complexity and diversity? Mol. Biol. Evol. 2006, 23, 887–892. [Google Scholar] [CrossRef]
- Tiffin, P.; Hahn, M.W. Coding sequence divergence between two closely related plant species: Arabidopsis thaliana and Brassica rapa ssp. pekinensis. J. Mol. Evol. 2002, 54, 746–753. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Suo, X.M.; Zhao, L.; Ma, X.L.; Cheng, R.H.; Wang, G.P.; Zhang, H.S. Molecular evolution, diversification, and expression assessment of MADS gene family in Setaria italica, Setaria viridis, and Panicum virgatum. Plant Cell Rep. 2023, 42, 1003–1024. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.H.; Gao, B.W.; Nakashima, Y.; Mori, T.; Zhang, Z.X.; Kodama, T.; Lee, Y.E.; Zhang, Z.K.; Wong, C.P.; Liu, Q.Q.; et al. Identification of a diarylpentanoid-producing polyketide synthase revealing an unusual biosynthetic pathway of 2-(2-phenylethyl)chromones in agarwood. Nat. Commun. 2022, 13, 348. [Google Scholar] [CrossRef] [PubMed]
- Su, H.; Zhang, S.; Yuan, X.; Chen, C.; Wang, X.-F.; Hao, Y.-J. Genome-wide analysis and identification of stress-responsive genes of the NAM–ATAF1, 2–CUC2 transcription factor family in apple. Plant Physiol. Bioch. 2013, 71, 11–21. [Google Scholar] [CrossRef]
- Gong, X.; Zhao, L.; Song, X.; Lin, Z.; Gu, B.; Yan, J.; Zhang, S.; Tao, S.; Huang, X. Genome-wide analyses and expression patterns under abiotic stress of NAC transcription factors in white pear (Pyrus bretschneideri). BMC Plant Biol. 2019, 19, 161. [Google Scholar] [CrossRef]
- Ma, J.H.; Yuan, M.; Sun, B.; Zhang, D.J.; Zhang, J.; Li, C.X.; Shao, Y.; Liu, W.; Jiang, L.N. Evolutionary divergence and biased expression of NAC transcription factors in hexaploid bread wheat (Triticum aestivum) L. Plants 2021, 10, 382. [Google Scholar] [CrossRef]
- Fu, C.; Liu, M. Genome-wide identification and molecular evolution of NAC gene family in Dendrobium nobile. Front. Plant Sci. 2023, 14, 1232804. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, T.; Xing, W.; Wang, J.; Yu, W.; Zhou, Y. Comprehensive genomic characterization of the NAC transcription factors and their response to drought stress in Dendrobium catenatum. Agronomy 2022, 12, 2753. [Google Scholar] [CrossRef]
- Olsen, A.N.; Ernst, H.A.; Leggio, L.L.; Skriver, K. NAC transcription factors: Structurally distinct, functionally diverse. Trend. Plant Sci. 2005, 10, 79–87. [Google Scholar] [CrossRef] [PubMed]
- So, H.A.; Lee, J.H. NAC transcription factors from soybean (Glycine max L.) differentially regulated by abiotic stress. J. Plant Biol. 2019, 62, 147–160. [Google Scholar] [CrossRef]
- Le, D.T.; Nishiyama, R.; Watanabe, Y.; Mochida, K.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Tran, L.S.P. Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress. DNA Res. 2011, 18, 263–276. [Google Scholar] [CrossRef] [PubMed]
- Ling, L.; Song, L.L.; Wang, Y.J.; Guo, C.H. Genome-wide analysis and expression patterns of the NAC transcription factor family in Medicago truncatula. Physiol. Mol. Biol. Plants 2017, 23, 343–356. [Google Scholar] [CrossRef]
- Jensen, M.K.; Skriver, K. NAC transcription factor gene regulatory and protein-protein interaction networks in plant stress responses and senescence. IUBMB Life 2014, 66, 156–166. [Google Scholar] [CrossRef] [PubMed]
- Vision, T.J.; Brown, D.G.; Tanksley, S.D. The origins of genomic duplications in Arabidopsis. Science 2000, 290, 2114–2117. [Google Scholar] [CrossRef]
- Blanc, G.; Hokamp, K.; Wolfe, K.H. A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome. Genome Res. 2003, 13, 137–144. [Google Scholar] [CrossRef] [PubMed]
- Mohanta, T.K.; Yadav, D.; Khan, A.; Hashem, A.; Tabassum, B.; Khan, A.L.; Abd_Allah, E.F.; Al-Harrasi, A. Genomics, molecular and evolutionary perspective of NAC transcription factors. PLoS ONE 2020, 15, e0231425. [Google Scholar] [CrossRef]
- Birchler, J.A.; Yang, H. The multiple fates of gene duplications: Deletion, hypofunctionalization, subfunctionalization, neofunctionalization, dosage balance constraints, and neutral variation. Plant Cell 2022, 34, 2466–2474. [Google Scholar] [CrossRef]
- Hu, W.; Wei, Y.X.; Xia, Z.Q.; Yan, Y.; Hou, X.W.; Zou, M.L.; Lu, C.; Wang, W.Q.; Peng, M. Genome-wide identification and expression analysis of the NAC transcription factor family in Cassava. PLoS ONE 2015, 10, e0136993. [Google Scholar] [CrossRef]
- Baranwal, V.K.; Khurana, P. Genome-wide analysis, expression dynamics and varietal comparison of NAC gene family at various developmental stages in Morus notabilis. Mol. Genet. Genom. 2016, 291, 1305–1317. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; White, M.J.; MacRae, T.H. Transcription factors and their genes in higher plants functional domains, evolution and regulation. Eur. j. Biochem. 1999, 262, 247–257. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, S.; Mitsuda, N. Reconstitution of a secondary cell wall in a secondary cell wall-deficient Arabidopsis mutant. Plant. Cell Physiol. 2014, 56, 299–310. [Google Scholar] [CrossRef] [PubMed]
- Adachi, S.; Minamisawa, K.; Okushima, Y.; Inagaki, S.; Yoshiyama, K.; Kondou, Y.; Kaminuma, E.; Kawashima, M.; Toyoda, T.; Matsui, M.; et al. Programmed induction of endoreduplication by DNA double-strand breaks in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 10004–10009. [Google Scholar] [CrossRef] [PubMed]
- Yoshiyama, K.O.; Kimura, S.; Maki, H.; Britt, A.B.; Umeda, M. The role of SOG1, a plant-specific transcriptional regulator, in the DNA damage response. Plant Signal Behav. 2014, 9, e28889. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.R.; Deng, Z.Y.; Lai, J.B.; Zhang, Y.Y.; Yang, C.P.; Yin, B.J.; Zhao, Q.Z.; Zhang, L.; Li, Y.; Yang, C.W.; et al. Dual function of Arabidopsis ATAF1 in abiotic and biotic stress responses. Cell Res. 2009, 19, 1279–1290. [Google Scholar] [CrossRef]
- Klok, E.J.; Wilson, I.W.; Wilson, D.; Chapman, S.C.; Ewing, R.M.; Somerville, S.C.; Peacock, W.J.; Dolferus, R.; Dennis, E.S. Expression profile analysis of the low-oxygen response in Arabidopsis root cultures. Plant Cell 2002, 14, 2481–2494. [Google Scholar] [CrossRef]
- Huh, S.U.; Lee, S.B.; Kim, H.H.; Paek, K.H. ATAF2, a NAC transcription factor, binds to the promoter and regulates NIT2 gene expression involved in auxin biosynthesis. Mol. Cells 2012, 34, 305–313. [Google Scholar] [CrossRef]
- Dalman, K.; Wind, J.J.; Nemesio-Gorriz, M.; Hammerbacher, A.; Lunden, K.; Ezcurra, I.; Elfstrand, M. Overexpression of PaNAC03, a stress induced NAC gene family transcription factor in Norway spruce leads to reduced flavonol biosynthesis and aberrant embryo development. BMC Plant Biol. 2017, 17, 6. [Google Scholar] [CrossRef]
- Zhang, H.H.; Xu, J.F.; Chen, H.M.; Jin, W.B.; Liang, Z.S. Characterization of NAC family genes in Salvia miltiorrhiza and NAC2 potentially involved in the biosynthesis of tanshinones. Phytochemistry 2021, 191, 112932. [Google Scholar] [CrossRef]
- Zhou, H.; Kui, L.W.; Wang, H.L.; Gu, C.; Dare, A.P.; Espley, R.V.; He, H.P.; Allan, A.C.; Han, Y.P. Molecular genetics of blood-fleshed peach reveals activation of anthocyanin biosynthesis by NAC transcription factors. Plant J. 2015, 82, 105–121. [Google Scholar] [CrossRef] [PubMed]
- Zou, S.C.; Zhuo, M.G.; Abbas, F.; Hu, G.B.; Wang, H.C.; Huang, X.M. Transcription factor LcNAC002 coregulates chlorophyll degradation and anthocyanin biosynthesis in litchi. Plant Physiol. 2023, 192, 1913–1927. [Google Scholar] [CrossRef] [PubMed]
- Shan, W.; Kuang, J.-F.; Lu, W.-J.; Chen, J.-Y. Banana fruit NAC transcription factor MaNAC1 is a direct target of MaICE1 and involved in cold stress through interacting with MaCBF1. Plant Cell Environ. 2014, 37, 2116–2127. [Google Scholar] [CrossRef] [PubMed]
- Xiang, P.; Chen, H.Q.; Cai, C.H.; Wang, H.; Zhou, L.M.; Mei, W.L.; Dai, H.F. Six new dimeric 2-(2-phenylethyl)chromones from artificial agarwood of Aquilaria sinensis. Fitoterapia 2020, 142, 104542. [Google Scholar] [CrossRef] [PubMed]
- Bisht, R.; Bhattacharyya, A.; Shrivastava, A.; Saxena, P. An overview of the medicinally important plant type III PKS derived polyketides. Front. Plant Sci. 2021, 12, 746908. [Google Scholar] [CrossRef]
- Wang, J.; Wang, X.H.; Liu, X.; Li, J.; Shi, X.P.; Song, Y.L.; Zeng, K.W.; Zhang, L.; Tu, P.F.; Shi, S.P. Synthesis of unnatural 2-substituted quinolones and 1, 3-diketones by a member of type III polyketide synthases from Huperzia serrata. Org. Lett. 2016, 18, 3550–3553. [Google Scholar] [CrossRef]
- Jin, J.; Tian, F.; Yang, D.C.; Meng, Y.Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2016, 45, D1040–D1045. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Paysan-Lafosse, T.; Blum, M.; Chuguransky, S.; Grego, T.; Pinto, B.L.; Salazar, G.A.; Bileschi, M.L.; Bork, P.; Bridge, A.; Colwell, L.; et al. InterPro in 2022. Nucleic Acids Res. 2022, 51, D418–D427. [Google Scholar] [CrossRef]
- Lu, S.N.; Wang, J.Y.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; Gwadz, M.; Hurwitz, D.I.; Marchler, G.H.; Song, J.S.; et al. CDD/SPARCLE: The conserved domain database in 2020. Nucleic Acids Res. 2020, 48, D265–D268. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef] [PubMed]
- Cantalapiedra, C.P.; Hernandez-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. eggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef] [PubMed]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef]
- Emms, D.M.; Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol. 2019, 20, 238. [Google Scholar] [CrossRef] [PubMed]
- Rozewicki, J.; Li, S.L.; Amada, K.M.; Standley, D.M.; Katoh, K. MAFFT-DASH: Integrated protein sequence and structural alignment. Nucleic Acids Res. 2019, 47, W5–W10. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef]
- Chen, K.; Durand, D.; Farach-Colton, M. NOTUNG: A program for dating gene duplications and optimizing gene family trees. J. Comput. Biol. 2000, 7, 429–447. [Google Scholar] [CrossRef]
- Wang, Y.P.; Li, J.P.; Paterson, A.H. MCScanX-transposed: Detecting transposed gene duplications based on multiple colinearity scans. Bioinformatics 2013, 29, 1458–1460. [Google Scholar] [CrossRef]
- Wang, Y.P.; Tang, H.B.; DeBarry, J.D.; Tan, X.; Li, J.P.; Wang, X.Y.; Lee, T.H.; Jin, H.Z.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Sun, P.; Jiao, B.; Yang, Y.; Shan, L.; Li, T.; Li, X.; Xi, Z.; Wang, X.; Liu, J. WGDI: A user-friendly toolkit for evolutionary analyses of whole-genome duplications and ancestral karyotypes. Mol. Plant 2022, 15, 1841–1851. [Google Scholar] [CrossRef]
- Koch, M.A.; Haubold, B.; Mitchell-Olds, T. Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis, Arabis, and related genera (Brassicaceae). Mol. Biol. Evol. 2000, 17, 1483–1498. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; Von Haeseler, A.; Lanfear, R. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [PubMed]
- Trifinopoulos, J.; Nguyen, L.T.; von Haeseler, A.; Minh, B.Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, W232–W235. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed]
- Goll, J.; Rusch, D.B.; Tanenbaum, D.M.; Thiagarajan, M.; Li, K.; Methé, B.A.; Yooseph, S. METAREP: JCVI metagenomics reports—An open source tool for high-performance comparative metagenomics. Bioinformatics 2010, 26, 2631–2632. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Jin, J.P.; Guo, A.Y.; Zhang, H.; Luo, J.C.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Chen, H.L.; Song, X.M.; Shang, Q.; Feng, S.Y.; Ge, W.N. CFVisual: An interactive desktop platform for drawing gene structure and protein architecture. BMC Bioinform. 2022, 23, 178. [Google Scholar] [CrossRef]
- Imai, T.; Ubi, B.E.; Saito, T.; Moriguchi, T. Evaluation of reference genes for accurate normalization of gene expression for real time-quantitative PCR in Pyrus pyrifolia using different tissue samples and seasonal conditions. PLoS ONE 2014, 9, e86492. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.; Mei, W.; Huang, S.; Wang, H.; Zhu, J.; Hu, W.; Ding, Z.; Tie, W.; Peng, S.; Dai, H. Genome survey sequencing for the characterization of genetic background of Dracaena cambodiana and its defense response during dragon’s blood formation. PLoS ONE 2018, 13, e0209258. [Google Scholar] [CrossRef] [PubMed]
- Yan, P.; Tuo, D.; Shen, W.T.; Deng, H.D.; Zhou, P.; Gao, X.Z. A Nimble Cloning-compatible vector system for high-throughput gene functional analysis in plants. Plant Commun. 2023, 4, 100471. [Google Scholar] [CrossRef] [PubMed]

| Species | Duplication Type | Total | ||||
|---|---|---|---|---|---|---|
| Singleton | Dispersed | Proximal | Tandem | WGD | ||
| A. trichopoda * | 44 | / | / | / | / | 44 |
| V. vinifera | 1 (1%) | 36 (43%) | 6 (7%) | 6 (7%) | 35(42%) | 84 |
| A. thaliana | 3 (3%) | 38 (34%) | 5 (4%) | 27 (24%) | 40 (35%) | 113 |
| A. sinensis | 2 (2%) | 53 (48%) | 5 (5%) | 16 (14%) | 35 (32%) | 111 |
| H. hainanensis | / | 18 (14%) | 11 (9%) | / | 98 (77%) | 127 |
| D. turbinatus | / | 22 (17%) | 8 (6%) | 3 (2%) | 93 (74%) | 126 |
| C. olitorius | / | 47 (58%) | 6 (7%) | 5 (6%) | 23 (28%) | 81 |
| C. capsularis | 1 (1%) | 49 (57%) | 6 (7%) | 5 (6%) | 25 (29%) | 86 |
| T. cacao | 2 (2%) | 40 (38%) | 23 (22%) | 12 (11%) | 28 (27%) | 105 |
| D. zibethinus | 1 (1%) | 18 (8%) | 4 (2%) | 70 (31%) | 136 (59%) | 229 |
| G. raimondii | 1 (1%) | 40 (26%) | 7 (5%) | 10 (7%) | 95 (62%) | 153 |
| H. cannabinus | / | 38 (16%) | 23 (9%) | 21 (9%) | 161 (66%) | 243 |
| Total | 55 | 399 | 104 | 175 | 769 | 1502 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Z.; Mei, W.; Wang, H.; Zeng, J.; Dai, H.; Ding, X. Comprehensive Analysis of NAC Transcription Factors Reveals Their Evolution in Malvales and Functional Characterization of AsNAC019 and AsNAC098 in Aquilaria sinensis. Int. J. Mol. Sci. 2023, 24, 17384. https://doi.org/10.3390/ijms242417384
Yang Z, Mei W, Wang H, Zeng J, Dai H, Ding X. Comprehensive Analysis of NAC Transcription Factors Reveals Their Evolution in Malvales and Functional Characterization of AsNAC019 and AsNAC098 in Aquilaria sinensis. International Journal of Molecular Sciences. 2023; 24(24):17384. https://doi.org/10.3390/ijms242417384
Chicago/Turabian StyleYang, Zhuo, Wenli Mei, Hao Wang, Jun Zeng, Haofu Dai, and Xupo Ding. 2023. "Comprehensive Analysis of NAC Transcription Factors Reveals Their Evolution in Malvales and Functional Characterization of AsNAC019 and AsNAC098 in Aquilaria sinensis" International Journal of Molecular Sciences 24, no. 24: 17384. https://doi.org/10.3390/ijms242417384





