DNA Copy Number Alterations and Copy Neutral Loss of Heterozygosity in Adult Ph-Negative Acute B-Lymphoblastic Leukemia: Focus on the Genes Involved
Abstract
:1. Introduction
2. Results
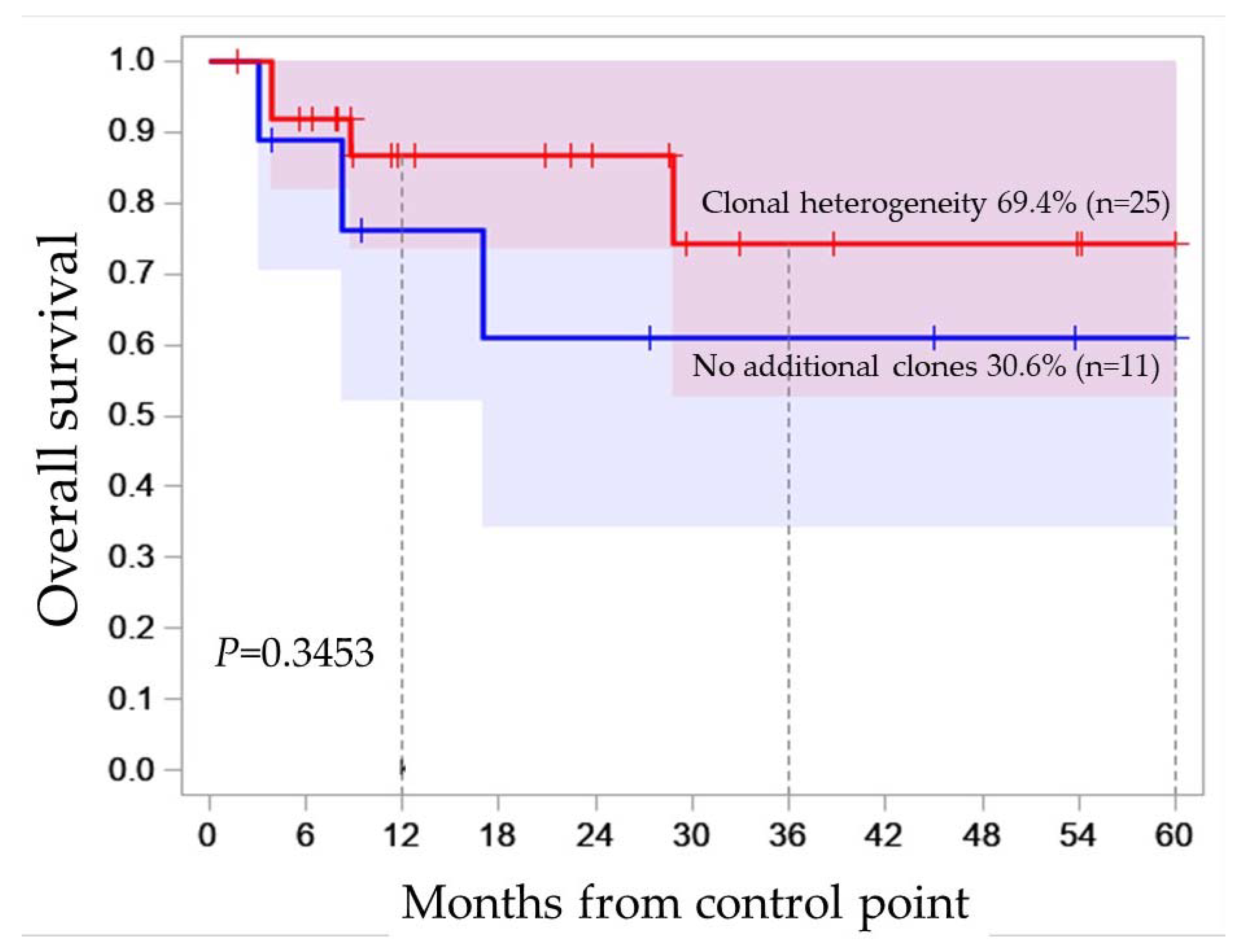
2.1. Clonal Heterogeneity in B-ALL
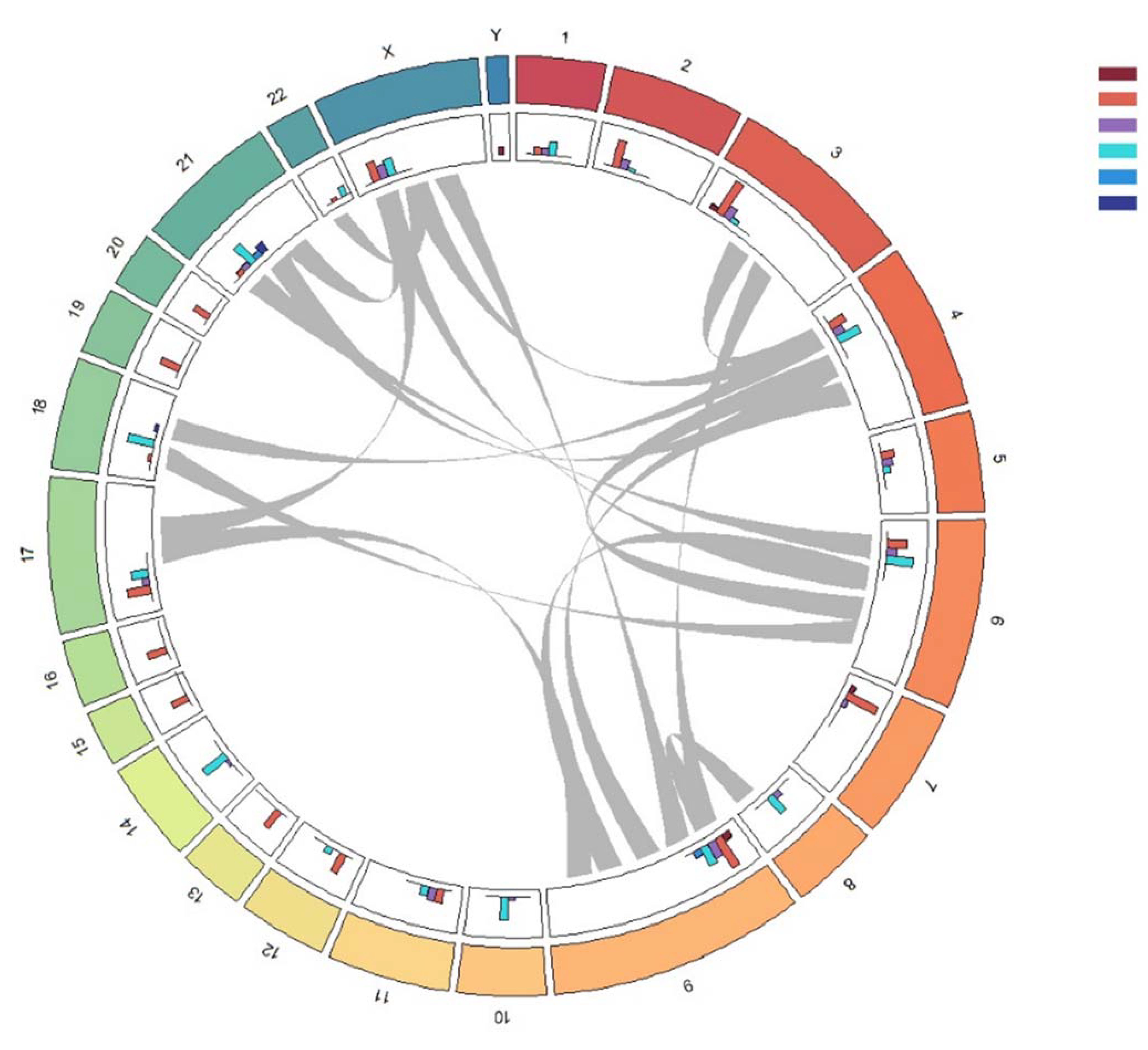
2.2. Molecular Karyotype of B-ALL
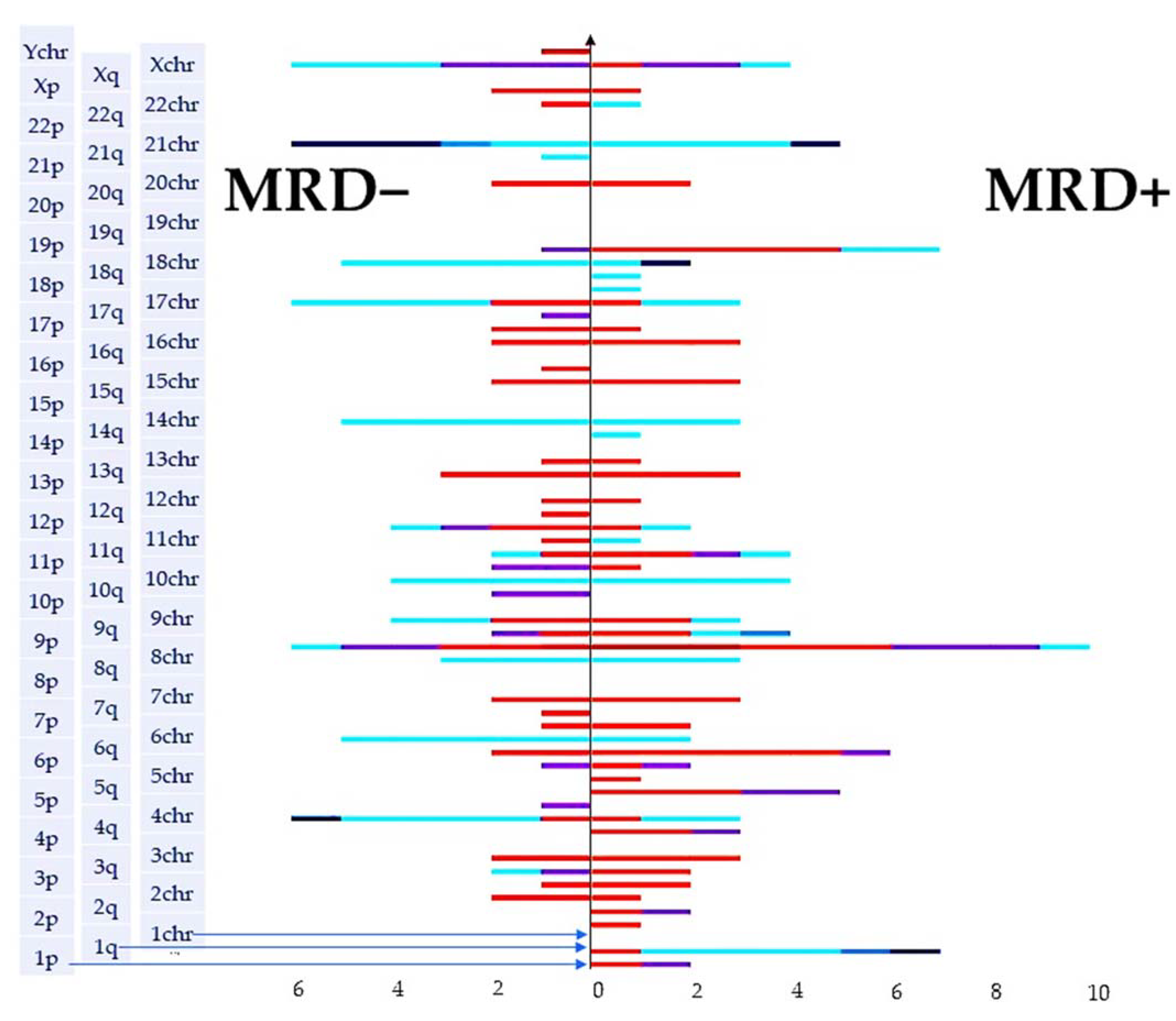
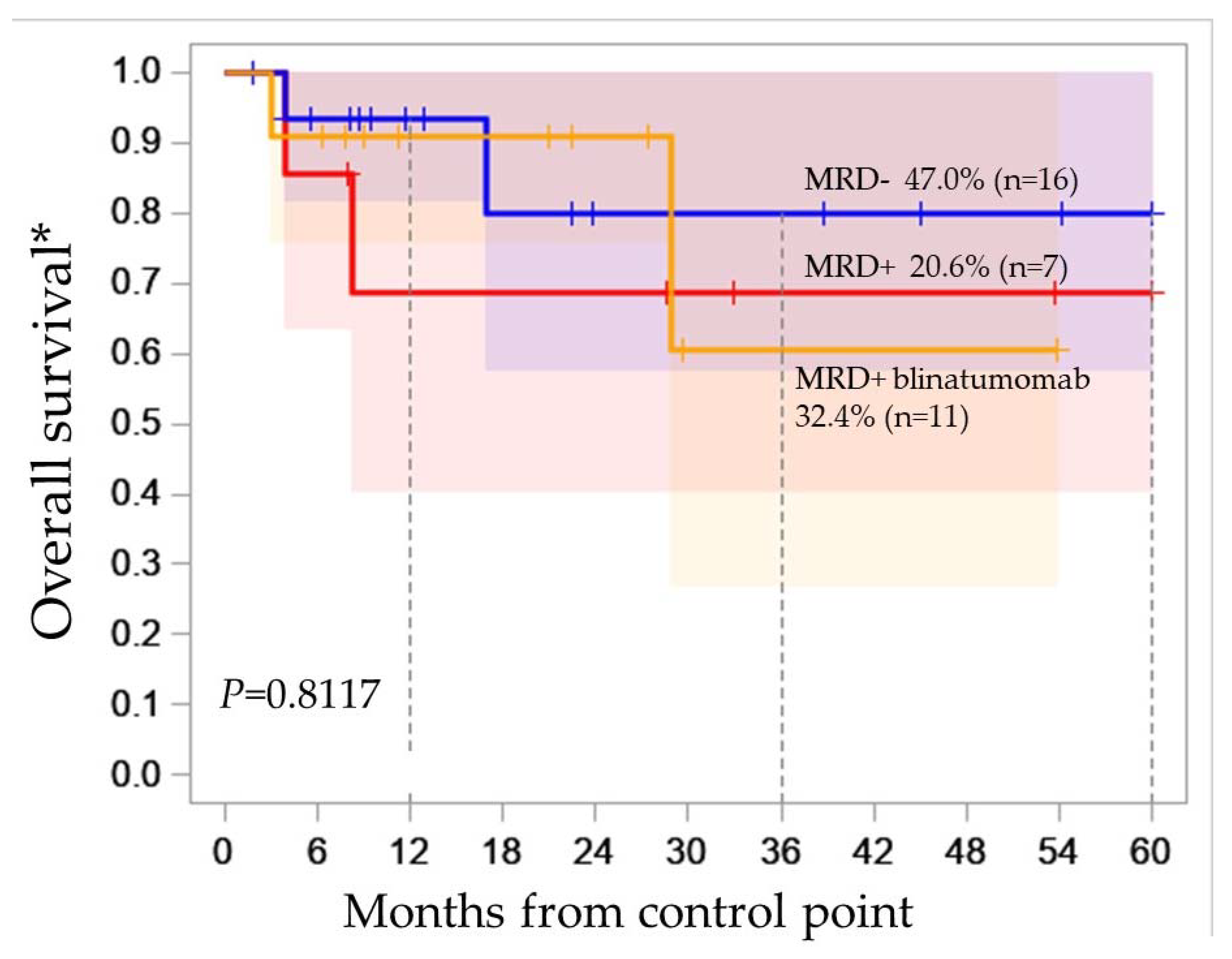
2.3. Molecular Karyotype Is Not Associated with MRD
2.4. Loss of Both Alleles—Biallelic or Homozygous Deletion
2.5. HLA-haplotype Loss
2.6. Loss of 3p21.3 Tumor Suppressor Genes Cluster
2.7. Copy Number Alterations and cnLOH in Genes Involved in the Pathogenesis of B-ALL
2.8. The Prognostic Significance of Various Genetic Abnormalities
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Parovichnikova, E.N.; Aleshina, O.A.; Troitskaya, V.V.; Chabaeva, Y.A.; Sokolov, A.N.; Isinova, G.A.; Kotova, E.S.; Akhmerzaeva, Z.H.; Klyasova, G.A.; Galtseva, I.V.; et al. Comparison of the treatment results in adult patients with acute Ph-negative lymphoblastic leukemia on protocols of the Russian multicenter studies ALL-2009 and ALL-2016. Russ. J. Hematol. Transfusiology 2022, 67, 460–477. (In Russian) [Google Scholar] [CrossRef]
- Thiagalingam, S.; Foy, R.L.; Cheng, K.; Lee, H.J.; Thiagalingam, A.; Ponte, J.F. Loss of heterozygosity as a predictor to map tumor suppressor genes in cancer: Molecular basis of its occurrence. Curr. Opin. Oncol. 2002, 14, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Makishima, H.; Maciejewski, J.P. Pathogenesis and consequences of uniparental disomy in cancer. Clin. Cancer Res. 2011, 17, 3913–3923. [Google Scholar] [CrossRef] [PubMed]
- Schoumans, J.; Suela, J.; Hastings, R.; Muehlematter, D.; Rack, K.; van den Berg, E.; Berna Beverloo, H.; Stevens-Kroef, M. Guidelines for genomic array analysis in acquired haematological neoplastic disorders. Genes Chromosomes Cancer 2016, 55, 480–491. [Google Scholar] [CrossRef] [PubMed]
- Bandaru, S.; Zhou, A.X.; Zhang, Y.; Bergo, M.O.; Cao, Y.; Akyürek, L.M. Targeting filamin B induces tumor growth and metastasis via enhanced activity of matrix metalloproteinase-9 and secretion of VEGF-A. Oncogenesis 2014, 3, e119. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.R.; Cao, L.; Wang, F.; Cheng, C.; Jiang, R.; Zhou, H.; Xie, Z.; Wuermanbieke, S.; Qian, Z. Filamin B extensively regulates transcription and alternative splicing, and is associated with apoptosis in HeLa cells. Oncol Rep. 2020, 43, 1536–1546. [Google Scholar] [CrossRef] [PubMed]
- Mitrakos, A.; Kattamis, A.; Katsibardi, K.; Papadhimitriou, S.; Kitsiou-Tzeli, S.; Kanavakis, E.; Tzetis, M. High resolution Chromosomal Microarray Analysis (CMA) enhances the genetic profile of pediatric B-cell Acute Lymphoblastic Leukemia patients. Leuk Res. 2019, 83, 106177. [Google Scholar] [CrossRef]
- Pös, O.; Radvanszky, J.; Buglyó, G.; Pös, Z.; Rusnakova, D.; Nagy, B.; Szemes, T. DNA copy number variation: Main characteristics, evolutionary significance, and pathological aspects. Biomed J. 2021, 44, 548–559. [Google Scholar] [CrossRef]
- Shao, X.; Lv, N.; Liao, J.; Long, J.; Xue, R.; Ai, N.; Xu, D.; Fan, X. Copy number variation is highly correlated with differential gene expression: A pan-cancer study. BMC Med. Genet. 2019, 20, 175. [Google Scholar] [CrossRef]
- Leary, R.J.; Lin, J.C.; Cummins, J.; Boca, S.; Wood, L.D.; Parsons, D.W.; Jones, S.; Sjöblom, T.; Park, B.H.; Parsons, R.; et al. Integrated analysis of homozygous deletions, focal amplifications, and sequence alterations in breast and colorectal cancers. Proc. Natl. Acad. Sci. USA 2008, 105, 16224–16229. [Google Scholar] [CrossRef]
- Brosens, R.P.M.; Haan, J.C.; Carvalho, B.; Rustenburg, F.; Grabsch, H.; Quirke, P.; Engel, A.F.; Cuesta, M.A.; Maughan, N.; Flens, M.; et al. Candidate driver genes in focal chromosomal aberrations of stage II colon cancer. J. Pathol. 2010, 221, 411–424. [Google Scholar] [CrossRef]
- Krijgsman, O.; Carvalho, B.; Meijer, G.A.; Steenbergen, R.D.M.; Ylstra, B. Focal chromosomal copy number aberrations in cancer-Needles in a genome haystack. Biochim. Biophys. Acta 2014, 1843, 2698–2704. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Yuan, Y.; Lu, K.H.; Zhang, L. Identification of recurrent focal copy number variations and their putative targeted driver genes in ovarian cancer. BMC Bioinf. 2016, 17, 222. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Lu, L.; Zhou, Z.; Liu, J.; Zhang, D.; Nan, K.; Zhao, X.; Li, F.; Tian, L.; Dong, H.; et al. CNV detection from circulating tumor DNA in late stage non-small cell lung cancer patients. Genes 2019, 10, 926. [Google Scholar] [CrossRef]
- Murakami, F.; Tsuboi, Y.; Takahashi, Y.; Horimoto, Y.; Mogushi, K.; Ito, T.; Emi, M.; Matsubara, D.; Shibata, T.; Saito, M.; et al. Short somatic alterations at the site of copy number variation in breast cancer. Cancer Sci. 2021, 112, 444–453. [Google Scholar] [CrossRef] [PubMed]
- Lyu, X.D.; Guo, Z.; Li, Y.W.; Hu, J.Y.; Fan, R.H.; Song, Y.P. Clonal heterogeneity and its prognostic significance in acute lymphoblastic leukemia. Zhonghua Nei Ke Za Zhi 2020, 59, 629–633. [Google Scholar] [CrossRef] [PubMed]
- Garrido, M.A.; Perea, F.; Vilchez, J.R.; Rodríguez, T.; Anderson, P.; Garrido, F.; Ruiz-Cabello, F.; Aptsiauri, N. Copy Neutral LOH Affecting the Entire Chromosome 6 Is a Frequent Mechanism of HLA Class I Alterations in Cancer. Cancers 2021, 13, 5046. [Google Scholar] [CrossRef]
- Ji, L.; Minna, J.D.; Roth, J.A. 3p21.3 tumor suppressor cluster: Prospects for translational applications. Future Oncol. 2005, 1, 79–92. [Google Scholar] [CrossRef]
- Oh, J.J.; West, A.R.; Fishbein, M.C.; Slamon, D.J. A candidate tumor suppressor gene, H37, from the human lung cancer tumor suppressor locus 3p21.3. Cancer Res. 2002, 62, 3207–3213. [Google Scholar]
- Zhao, L.; Li, R.; Shao, C.; Li, P.; Liu, J.; Wang, K. 3p21.3 tumor suppressor gene RBM5 inhibits growth of human prostate cancer PC-3 cells through apoptosis. World J. Surg. Oncol. 2012, 10, 247. [Google Scholar] [CrossRef]
- Oh, J.J.; Taschereau, E.O.; Koegel, A.K.; Ginther, C.L.; Rotow, J.K.; Isfahani, K.Z.; Slamon, D.J. RBM5/H37 tumor suppressor, located at the lung cancer hot spot 3p21.3, alters expression of genes involved in metastasis. Lung Cancer 2010, 70, 253–262. [Google Scholar] [CrossRef] [PubMed]
- Sinclair, P.B.; Blair, H.H.; Ryan, S.L.; Buechler, L.; Cheng, J.; Clayton, J.; Hanna, R.; Hollern, S.; Hawking, Z.; Bashton, M.; et al. Dynamic clonal progression in xenografts of acute lymphoblastic leukemia with intrachromosomal amplification of chromosome 21. Haematologica 2017, 103, 634–644. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Kuang, P.; Liu, T. Prognostic significance of CDKN2A/B deletions in acute lymphoblastic leukaemia: A meta-analysis. Ann Med. 2019, 51, 28–40. [Google Scholar] [CrossRef] [PubMed]
- Piskunova, I.S.; Obukhova, T.N.; Parovichnikova, E.N.; Kulikov, S.M.; Gavrilina, O.A.; Lukyanova, I.A.; Savchenko, V.G. CDKN2A/p16INK4a deletion is not a poor prognostic factor in adult acute lymphoblastic leukemia patiets treated according to protocol RALL-2009. Oncohematology 2017, 12, 17–24. [Google Scholar] [CrossRef]
- Zhang, L.; Cao, Y.; Dai, X.; Zhang, X. Deciphering the role of DOCK8 in tumorigenesis by regulating immunity and the application of nanotechnology in DOCK8 deficiency therapy. Front. Pharmacol. 2022, 13, 1065029. [Google Scholar] [CrossRef] [PubMed]
- Quijada-Álamo, M.; Hernández-Sánchez, M.; Rodríguez-Vicente, A.E.; Pérez-Carretero, C.; Rodríguez-Sánchez, A.; Martín-Izquierdo, M.; Alonso-Pérez, V.; García-Tuñón, I.; Bastida, J.M.; Vidal-Manceñido, M.J.; et al. Biological significance of monoallelic and biallelic BIRC3 loss in del(11q) chronic lymphocytic leukemia progression. Blood Cancer J. 2021, 11, 127. [Google Scholar] [CrossRef] [PubMed]
- Alhourani, E.; Othman, M.A.; Melo, J.B.; Carreira, I.M.; Grygalewicz, B.; Vujić, D.; Zecević, Z.; Joksić, G.; Glaser, A.; Pohle, B.; et al. BIRC3 alterations in chronic and B-cell acute lymphocytic leukemia patients. Oncol. Lett. 2016, 11, 3240–3246. [Google Scholar] [CrossRef]
- Wu, Q.; Berglund, A.E.; MacAulay, R.J.; Etame, A.B. A Novel Role of BIRC3 in Stemness Reprogramming of Glioblastoma. Int. J. Mol. Sci. 2021, 23, 297. [Google Scholar] [CrossRef]
- Dupuy, A.; Simon, R.M. Critical review of published microarray studies for cancer outcome and guidelines on statistical analysis and reporting. J. Natl. Cancer Inst. 2007, 99, 147–157. [Google Scholar] [CrossRef]
- McGowan-Jordan, J.; Ros, J. ISCN 2020: An International System for Human Cytogenomic Nomenclature, 1st ed.; S. Karger AG: Basel, Switzerland, 2020. [Google Scholar] [CrossRef]
- Borowitz, M.J.; Wood, B.L.; Keeney, M.; Hedley, B.D. Measurable Residual Disease Detection in B-Acute Lymphoblastic Leukemia: The Children’s Oncology Group (COG) Method. Curr. Protoc. 2022, 2, e383. [Google Scholar] [CrossRef]
- Pickett, K.L.; Suresh, K.; Campbell, K.R.; Davis, S.; Juarez-Colunga, E. Random survival forests for dynamic predictions of a time-to-event outcome using a longitudinal biomarker. BMC Med. Res. Methodol. 2021, 21, 216. [Google Scholar] [CrossRef] [PubMed]
- Bohannan, Z.S.; Coffman, F.; Mitrofanova, A. Random survival forest model identifies novel biomarkers of event-free survival in high-risk pediatric acute lymphoblastic leukemia. Comput. Struct Biotechnol. J. 2022, 20, 583–597. [Google Scholar] [CrossRef] [PubMed]
- Ishwaran, H.; Kogalur, U. Fast Unified Random Forests for Survival, Regression, and Classification (RF-SRC). R Package Version 3.2.2. 2023. Available online: https://cran.r-project.org/package=randomForestSRC (accessed on 1 November 2023).


| Parameter | B-ALL (n = 36) |
|---|---|
| Male:Female | 21:15 |
| Age, median | 34.1 (19–53) years |
| Leukocytes, 10 × 9/L | 7.66 (1.09–466.53) |
| LDH | 624 (148–7348.8) |
| Blast cells in peripheral blood, % | 40 (0–95) |
| Blast cells in the bone marrow, % | 88.8 (29–98) |
| Immunophenotype, EGIL, WHO | B-I 5 (13.8%) |
| B-II 30 (83.3%) | |
| B-III 1 (2.7%) | |
| B-IV 0 (0%) | |
| Standard cytogenetics | 36 |
| + mitosis | 35 (97.2%) |
| − mitosis | 1 (2.8%) |
| Karyotype | 35 |
| Normal | 7 (20.0%) |
| Abnormal: | 28 (80.0%) |
| CNS leukemia | 3 (8.3%) |
| Extramedullary disease | 14 (38.8%) |
| MRD-status (+70 day) | 34 |
| MRD+ | 18 |
| MRD- | 16 |
| CR: | |
| After 2nd induction (+70 day) | 16 |
| Refractory disease | 1 |
| Early Death | 1 |
| N° | Sex | Age | Diagnosis | CCA Data [mitoses count] | CMA Data | FISH |
|---|---|---|---|---|---|---|
| 1 | f | 31 | B-II | no mitosis | (X,1-22)×2 | n/rvl t(9;22)(q34;q11); t(11q23)/MLL |
| 2 | f | 26 | B-II | 46,XX,t(4;11)(q21;q23)[7]/46,XX [13] | (X,1-22)×2 | 21% t(11q23)/MLL |
| n/rvl t(9;22)(q34;q11) | ||||||
| 3 | f | 54 | B-II | 50,XX,+X,+8,+9,i(9)(q10),+22[10]/46,XX [4] | (X)×3, 2p16.3×1, 5q35.2q35.3×2 hmz, 7p12.3p12.1×1, (8)×3, 9p24.3p21.3×2 hmz, 9p21.3×0, 9p21.3p13.1×2 hmz, (9q)×4, 17q11.2q11.2×2 hmz, (22) ×3 | 94%++ABL/9q34 ++BCR/22q11 |
| n/rvlt(9;22)(q34;q11); t(11q23)/MLL | ||||||
| 4 | f | 34 | B-I | 47,XX,+X,t(4;11)(q21;q23)[17]/46,XX [3] | (X)x3[0.6],1p21.2p21.1x2 hmz,3q25.31q26.1x2 hmz, 3p21.1p14.2×2 hmz,3p22.1p21.31×2 hmz, 4q34.3q34.3×2 hmz,5p13.1q11.2×2 hmz, 6p22.3p22.3×2 hmz,8p23.1p22)×2 hmz, 9q22.31q31.1×2 hmz,10q25.3q26.13×2 hmz, 10q25.1q25.2×2 hmz,11q22.3q23.2×2 hmz | 90% t(11q23)/MLL |
| n/rvl t(9;22)(q34;q11) | ||||||
| 5 | f | 34 | B-II | 46,XX [20] | (X)×1 [0.2], 2q24.1q24.1×2 hmz, (3)×1 [0.19], 4q31.3q32.1×1 [0.2], (7)×1 [0.19],7q31.2q31.31×2 hmz, (8)×3 [0.31], (9)×1 [0.19], (10)×3 [0.29], | 12% + MLL/11q23, +BCR/22q11 |
| (14)×3 [0.3], (15)×1 [0.2],(16)x1 [0.19],(20)×1 [0.17],(21)x3 [0.31] | n/rvl t(9;22)(q34;q11); t(11q23)/MLL | |||||
| 6 | m | 32 | B-II | 46,XY,der(6),-11,+mar or der(11?)[6]/46,XY [14] | 3p21.31×1,6q16.1q22.1×1, 9p24.3×3,11p14.3p12×1, 11p11.2q12.1×1,11q14.2q14.3x1, 11q22.1q23.3×1,11q23.3q24.1×1, 11q24.1×1,12p13.31×3, 19p13.3×1 | n/rvl t(9;22)(q34;q11); t(11q23)/MLL |
| 7 | f | 24 | B-II | 81-85,XX, ?-X, del (6)(q22)*2, add(11)(q2?5),+3-4 mar, inc [cp2]/ | Xp22.33p11.3×1 [0.2],Xp11.3q28×1 [0.5], 3p26.3p12.2×1 [0.3], 3p12.1p11.1×2 hmz, | n/rvl t(9;22)(q34;q11); t(11q23)/MLL |
| 46,XX [18] | 3q25.33q29×3 [0.3], 6q16.3q24.3×1 [0.6], (9)×1 [0.21], (11)×1 [0.3], 12p13.33p13.2×1 [0.22], 12p13.2p12.1×2 hmz [0.5], 12p11.23q22×3 [0.3], 12q22q24.33×1 [0.2], (15)×1 [0.72], 17p13.3q21.31×1 [0.2], 17q21.31q25.3×2 hmz [0.5], 21q22.12q22.3×3 [0.3] | |||||
| 8 | m | 41 | B-I | 46,XY,t(4;11)(q21;q23)[3] | 1p31.1p31.1×2hmz, 5q23.1q23.1x2 hmz | 98% t(4;11)(q21;q23) |
| n/rvl t(9;22)(q34;q11) | ||||||
| 9 | f | 29 | B-II | 56,XX,+X,+4,+4,+6,+10,+14,+17,+18,+21, | (X)×3, (4)×5, (6)×3, (10)×3, | n/rvl t(9;22)(q34;q11); t(11q23)/MLL |
| +21[18]/46,XX [2] | 11p11.2p11.12×2 hmz, (14) ×3, (17) ×3, (18) ×3, (21) ×5 | |||||
| 10 | m | 53 | B-I | 46,XY,t(4;11)(q21;q23) [20] | 2q36.3q37.1×2 hmz, 4q12q13.2×2 hmz, | 88% t(11q23)/MLL |
| 9p24.3p13.3×2 hmz, 9p21.3×1 | ||||||
| 12 | f | 36 | B-II | 46,XX,add(14)(q32) or t(14;?)(q32;?) | 14q31.2q32.33×3 | 92% t/del14q32/IGH(telomeric part); |
| n/rvl t(9;22)(q34;q11); t(11q23)/MLL t(12p13)/ETV6 | ||||||
| 13 | m | 20 | BIII/BIV | 52~54,XY,+X,+4,+8,+14,+14,+18,+21, | (X) ×2 [0.83], (4) ×3, 6q15q23.1×1[0.36], (8) ×3, 9p21.3×0, 9p21.3p21.2×1, 9q22.33q31.1×1, 11q22.3×1, 13q14.2q31.1×1, (14) ×3, (18)x3, (21) ×3 | 94% +cMYC/8q24 |
| +mar[cp16]/46,XY [4] | n/rvl t(8q24)/cMYC; t(11q23)/MLL; t(9;22)(q34;q11) | |||||
| 14 | m | 22 | B-II | 56,XY,+X,+Y,+4,+6,+8,+10,+14,+17,+18, | (X,Y)x1,(1-22)x2 | n/rvl t(9;22)(q34;q11); t(11q23)/MLL |
| +21[9]/46,XY [11] | ||||||
| 15 | m | 19 | B-II | 54~56<2n>,XY,+X,+4,+6,+9,+10,+14,+15,+17,+21,+21[cp13]/ | (X)×2 [0.81], 3p21.31p21.1×2 hmz, (4) ×3 [0.72], (6)×3 [0.69], (9)×3 [0.42], (10)×3, (14)×3, (17)×3, (18)×3, 19p13.3p13.3×1 [0.4], | 30% +ABL/9q34 |
| 54~56,idem,add(19)(p?q?)[5]/46,XY [2] | 19p13.2p13.12×3 [0.3], (21) ×5 | n/rvl t(9;22)(q34;q11) t(11q23)/MLL | ||||
| 16 | m | 20 | B-II | 52~54,XY,+X,+4,+6,+?10,+17,+18,+20,+21,+mar[cp5]/ 52~54, XY, +X, dup(1)(q3?1q44),+4,+6,+?10, +17, +18, +20, +21[cp3]/46,XY [5 ] | (X)×2 [0.78], (4)×3, (6)×3, (10)×3, (17)×3, (18)×5,(21)×3 | n/rvl t(9;22)(q34;q11); t(11q23)/MLL |
| 17 | m | 22 | B-II | 46,XY [20] | (2)×1, (3)×1, (4)×1, (7)×1, (9)×1, (12)×1, (13)×1, (16)×1, (17)×1, (20)×1 | n/d |
| 18 | f | 28 | B-II | 46,XX,der(19)t(1;19)(q23;p13), inc[cp15]/ | 1q21.1q23.3×3 [0.7], 1q23.3q43×5 [0.5], 1q43q44×4 [0.6], | 28% t/del (19p13)/E2A (telomeric part) |
| 46,XX [5] | 9p24.3p12x1 [0.31], 19p13.3x1 | |||||
| 19 | m | 44 | B-II | 36,X,-Y,-2,-3,der(4),-7,der(12),-13,-15,-16,-17, -20x2,-22, +mar[cp6]/46,XY [14] | (Y) ×0 [0.8], (2) ×1, (3) ×1, 4q26q26×2 hmz, 5p13.2p13.1×1, (7)x1, 11q13.3q25×3, 12p13.33p11.22×1, 13q12.11q22.3×1, 13q22.3q34×1 [0.2], | 96% del (22q11)/BCR |
| (15) ×1,(16) ×1,(17) ×1,18q11.2×3,(20) ×1,(22) ×1 | n/rvl t(9;22)(q34;q11) | |||||
| 21 | f | 46 | B-II | 46,X,-X, | (X) ×1 [0.1], 1p36.13p34.31×1 [0.2],1q23.3q44×3, 3q13.12q13.2×1 [0.5],4q21.22q28.1×1 [0.2], 4q35.1q35.2×1 [0.2],5q11.2q12.1×1 [0.2], 6q11.1q21×1 [0.5],6q22.32q22.33×2 hmz, 6q24.3q25.1×1 [0.5],6q25.3q27×1 [0.5], 8q11.1q11.21×2 hmz,9p24.3p24.2×0 [0.3],9p24.1p21.3×1, 9p21.3p21.3×0 [0.3], 9p21.3p11.2×1,9q21.11q34.3×3, 13q12.11q12.12×1 [0.5],13q13.3q31.1×1 [0.5], 18p11.32p11.31×1 [0.5],19p13.3×1,(21) ×3 | n/d |
| add(1)(p3?3),-4, | ||||||
| i(9)(q10), | ||||||
| der(19)t(1;19)(q23;p13),+?21,+mar [15]/ | ||||||
| 46,XX [5] | ||||||
| 22 | f | 44 | B-II | 46,XX[29]/ | 7q35q36.2×1 [0.3],9p21.3p21.2×1 [0.4] | n/rvl t(9;22)(q34;q11) |
| 47,XX,+12[1] | ||||||
| 23 | f | 39 | B-II | 52~54,?XX,+?6,+?11,+?17,+?18,+?20,+?21, | (X) ×3,(4) ×3,(6) ×3,(8) ×3, 9p24.3p13.1×2 hmz,9p21.3p21.3×0 [0.5],(10) ×3,(14) ×3, | n/rvl t(9;22)(q34;q11) |
| inc[cp3]/46,XX [12] | 14q23.1q23.2×2 hmz,(17) ×3,(18) ×3,(21) ×5 | |||||
| 24 | f | 36 | B-II | 44-45, X, -X,der(2),der(3),der(5),der(7),der(17),add(p11),+mar[cp5]/ | Xp22.33p22.32×1 [0.37], 2p13.1p12×1 [0.31], 3p24.1p22.3×1 [0.27], 3q13.12q13.13×1 [0.42], 5q11.2q12.1×1 [0.26], 5q13.3q14.1×1 [0.31], 7p12.2p12.1×1 [0.44], 13q14.2q14.3×1 [0.2], 17p13.3p11.1×1 [0.35] | n/rvl t(11q23)/MLL, t(9;22)(q34;q11 |
| 46,XX [15] | ||||||
| 25 | m | 26 | B-II | 45,XY,-7,der(14) t(7;14)(q11;q?32), del(17)(p10)[6]/ | 2p11.2p11.2×1, 7p14.1p11.2×1 [0.16], 7q34×0, 9p24.3p13.2×2 hmz, 9p21.3×0,16p13.3×1, 17p13.3p11.2×1 | 90% del (17p13)/TP53; |
| 46,XY [14] | n/rvl t(9;22)(q34;q11); t(14q32)/IgH; t(11q23)/MLL | |||||
| 26 | f | 38 | B-II | 46,XY [20] | 5 q14.3q14.3×2 hmz | n/rvl t(11q23)/MLL; t(9;22)(q34;q11) |
| 27 | m | 23 | B-I | 46,XY,der(19)[20] | (X,Y) ×1,(1-22) ×2 | n/rvl t(9;22)(q34;q11); t(11q23)/MLL; t(19p13)/ E2A |
| 28 | f | 23 | B-II | 46,XX,add(1)(q44),der(9), inc [9]/46,XX [11] | 1q21.1q32.1×3, 6q16.2q16.3×2 hmz, 9q21.13q31.1×1, 11q22.1q22.3×2 hmz | n/rvl t(9;22)(q34;q11) |
| 29 | m | 42 | B-II | 46,XY [13] | 11p11.2p11.12×2 hmz, 21q21.1q21.3×2 hmz | n/rvl t(11q23)/MLL; t(9;22)(q34;q11) |
| 30 | m | 21 | B-II | 57,XY,+X,+4,+6,+8,+10,+14,+17,+18,+21x2,+mar[14] | (X)×2 [0.81],(Y)×0 [0.13],(4)×3, (6)×3,(8)×3,(9)×3 [0.19],(10)×3,(14)×3,(17) ×3,(18) ×3,(21)×4 | 20% +ABL/9q34 |
| n/rvl t(9;22)(q34;q11) t(11q23)/MLL | ||||||
| 31 | f | 47 | B-II | 46,XX,del(9)(p?21)[5]/46,XX[15] | 2q36.1×1,3q26.32q26.32×0, 3q26.32q26.33×1 | 11% del(9p21)/ CDKN2A |
| n/rvl t(9;22)(q34;q11) | ||||||
| 32 | f | 24 | B-II | 46, XX [20] | (1q)×3 [0.19],(2)×1 [0.16],(3)×1 [0.15],(7)×1 [0.15],(8)×3 [0.15], 9p24.3q22.32×3 [0.12],9q22.32q34.3×1 [0.17],(10)×3 [0.43] (11)×3 [0.15],(12) ×1 [0.18],(13)×1 [0.15],(14)×3 [0.2],(15)×1 [0.18],(16)×1 [0.15], (18p)×3 [0.46],(18q)×3 [0.25],19p13.3p13.11×3 [0.15], 19p13.11q13.43×1 [0.2],(21)×3 [0.19] | 5% +MLL/11q23 |
| n/rvl t(9;22)(q34;q11) t(11q23)/MLL | ||||||
| 34 | f | 44 | B-II | 46,XY,der(9),add(13)(p11),add(15)(p11), | 2q32.3q33.1×2 hmz,9p24.3×3,(21)×3 [0.15] | n/rvl t(9;22)(q34;q11); t(11q23)/MLL |
| der(17) or mar,der(22) or mar,inc[9]/46,XY[2] | ||||||
| 35 | f | 40 | B-II | 46,XX [20] | (X,1-22)×2 | n/rvl t(9;22)(q34;q11); t(11q23)/MLL; |
| 36 | f | 30 | B-II | ?54-55,XX,?+4,+8,?+8 or 10,+9,+11,+?14,+21,+21,+mar[cp6]/46,XX[19]. | (X,1-22)×2 | 90%+ MLL (11q23), +ABL (9q34) |
| n/rvl t(11q23)/MLL; t(9;22)(q34;q11) | ||||||
| 37 | m | 27 | B-II | 46,der(X),Y,del(6)(q22), ?der(16)[20] | 6p21.1×1, 6q14.3q22.31×1,6q25.2q25.3×1, 12p13.2p13.1×1,19q12×1, 19q13.11×1 | n/rvl t(9;22)(q34;q11); t(11q23)/MLL |
| 38 | f | 38 | B-II | 46,XX[20] | 2p11.2×1, 3p14.3×0, 12q13.12×1, 21q21.1×1 | n/rvl t(9;22)(q34;q11) |
| 39 | m | 41 | B-I | 34~38,XY, | (3) ×1 [0.5], (4) ×1 [0.5], (5) ×1 [0.5],6p22.2p21.32×2 hmz,(7) ×1 [0.5],(9) ×1 [0.5], 11q14.1q25×3 [0.6],(15) ×1 [0.5],(16) ×1 [0.5],(17) ×1 [0.5],(20) ×1 [0.5] | n/d |
| +2-3mar[12]/46,XY[8] |
| N° Pat | Locus | Genes Affected by Biallelic/Homozygous Deletion | Nature of Loss |
|---|---|---|---|
| 38 | 3p14.3 | FLNB | biallelic |
| 31 | 3q26.32 | TBL1XR1, KCNMB2 | biallelic |
| 25 | 7q34 | TCAF2, PRSS1, PRSS2 | biallelic |
| 25 | 9p21.3 | MTAP, CDKN2A, CDKN2B-AS1, CDKN2B, DMRTA1 | homozygous |
| 13 | 9p21.3 | IFNB1, IFNW1, IFNA21, IFNA4, IFNA7, IFNA10, IFNA16, IFNA17, IFNA14, IFNA5, KLHL9, IFNA6, IFNA13, IFNA2, IFNA8, IFNA1, MIR31HG, IFNE, MIR31, MTAP, CDKN2A, CDKN2B-AS1, CDKN2B, DMRTA1, ELAVL2, IZUMO3 | biallelic |
| 23 | 9p21.3 | MTAP, CDKN2A, CDKN2B-AS1, CDKN2B, DMRTA1 | homozygous |
| 3 | 9p21.3 | CDKN2A, CDKN2B-AS1, CDKN2B, DMRTA1, ELAVL2 | homozygous |
| 21 | 9p21.3 | DMRTA1 | biallelic |
| 21 | 9p24.3 | DOCK8, KANK1, DMRT1, DMRT3, DMRT2, SMARCA2, VLDLR, KCNV2, PUM3, RFX3, GLIS3, SLC1A1 | biallelic |
| Gene 1 | Chr. Location | Count of Event | Type of Events |
|---|---|---|---|
| CDKN2A/B | 9p21. 3 | 15 | 4loss0/8loss/3gain |
| DMRTA | 9p21.3 | 15 | 4loss0/6loss/2cnLOH/3gain |
| DOCK8 | 9p24.3 | 15 | 1loss0/5loss/4cnLOH/5gain |
| TP53 | 17p13.1 | 13 | 7loss/6gain |
| SMARCA2 | 9p24.3 | 13 | 6loss/4cnLOH/3gain |
| PAX5 | 9p13 | 12 | 6loss/ 3cnLOH/ 3gain |
| XPA; FOXE1; HEMGN | 9q22.33 | 12 | 7loss/1cnLOH/4gain |
| USP45 | 6q16.2 | 12 | 4loss/1cnLOH/7gain |
| RUNX1 | 21q22.2 | 11 | 7 gain/1gainX4/3gainX5 |
| NF1 | 17q11.2 | 11 | 5loss/ 1cnLOH/ 5gain |
| IGF2BP1 | 17q21.32 | 10 | 4loss/1cnLOH/5gain |
| ERG; TMPRSS2 | 21q22.2 | 9 | 5gain/1gain4x/3gain5x |
| CRLF2 | Xp22.23 | 9 | 4loss/ 5gain |
| FGFR3 | 4p16.3 | 9 | 2loss/7gain |
| FLNB | 3p14.3 | 8 | 7 loss/1 cnLOH |
| IKZF1 | 7p12.2 | 8 | 8 loss |
| RUNX2 | 6p21.1 | 8 | 1loss/7gain |
| ARID1B | 6q25.3 | 8 | 1loss/7gain |
| CIP2A | 3q13.13 | 7 | 7 loss |
| PIK3CA | 3q26.32 | 7 | 6loss/1gain |
| ATM | 11q22.3 | 6 | 3loss/3gain |
| RB1 | 13q14.2 | 6 | 6 loss |
| BIRC3 | 11q22.3 | 6 | 2loss/1cnLOH/3gain |
| MYC | 8q24.21 | 6 | 6gain |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Risinskaya, N.; Gladysheva, M.; Abdulpatakhov, A.; Chabaeva, Y.; Surimova, V.; Aleshina, O.; Yushkova, A.; Dubova, O.; Kapranov, N.; Galtseva, I.; et al. DNA Copy Number Alterations and Copy Neutral Loss of Heterozygosity in Adult Ph-Negative Acute B-Lymphoblastic Leukemia: Focus on the Genes Involved. Int. J. Mol. Sci. 2023, 24, 17602. https://doi.org/10.3390/ijms242417602
Risinskaya N, Gladysheva M, Abdulpatakhov A, Chabaeva Y, Surimova V, Aleshina O, Yushkova A, Dubova O, Kapranov N, Galtseva I, et al. DNA Copy Number Alterations and Copy Neutral Loss of Heterozygosity in Adult Ph-Negative Acute B-Lymphoblastic Leukemia: Focus on the Genes Involved. International Journal of Molecular Sciences. 2023; 24(24):17602. https://doi.org/10.3390/ijms242417602
Chicago/Turabian StyleRisinskaya, Natalya, Maria Gladysheva, Abdulpatakh Abdulpatakhov, Yulia Chabaeva, Valeriya Surimova, Olga Aleshina, Anna Yushkova, Olga Dubova, Nikolay Kapranov, Irina Galtseva, and et al. 2023. "DNA Copy Number Alterations and Copy Neutral Loss of Heterozygosity in Adult Ph-Negative Acute B-Lymphoblastic Leukemia: Focus on the Genes Involved" International Journal of Molecular Sciences 24, no. 24: 17602. https://doi.org/10.3390/ijms242417602





