Proteomic Analysis of Proteins Related to Defense Responses in Arabidopsis Plants Transformed with the rolB Oncogene
Abstract
1. Introduction
2. Results
2.1. AtB-1 Line
2.2. General Description of Proteomic Analysis
2.3. Defense Reactions and Plant Immunity
2.4. Primary Metabolism
2.5. Chaperones
2.6. Photosynthesis
2.7. RACK1-Associated Proteins
2.8. CERK1-Associated Proteins
2.9. Gene Expression
3. Discussion
3.1. Defense Response
3.2. ROS Metabolism and Abiotic Stress Response
4. Materials and Methods
4.1. Plant Material
4.2. 2D-Gel Electrophoresis
4.3. Quantification of Protein Expression
4.4. Experimental Design and Statistical Rationale
4.5. MALDI-TOF Mass Spectrometry and Protein Identification
4.6. RNA Isolation, cDNA Synthesis, and Real-Time PCR
4.6.1. RNA Isolation and cDNA Synthesis
4.6.2. Quantitative Real-Time PCR
4.6.3. Absolute Quantification
4.6.4. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Otten, L. The Agrobacterium Phenotypic Plasticity (Plast) Genes. Curr. Top Microbiol. Immunol. 2018, 418, 375–419. [Google Scholar] [CrossRef]
- Mauro, M.L.; Bettini, P.P. Agrobacterium rhizogenes rolB oncogene: An intriguing player for many roles. Plant Physiol. Biochem. 2021, 165, 10–18. [Google Scholar] [CrossRef]
- Delbarre, A.; Muller, P.; Imhoff, V.; Barbier-Brygoo, H.; Maurel, C.; Leblanc, N.; Perrot-Rechenmann, C.; Guern, J. The rolB gene of Agrobacterium rhizogenes does not increase the auxin sensitivity of tobacco protoplasts by modifying the intracellular auxin concentration. Plant Physiol. 1994, 105, 563–569. [Google Scholar] [CrossRef]
- Cecchetti, V.; Pomponi, M.; Altamura, M.M.; Pezzotti, M.; Marsilio, S.; D’Angeli, S.; Tornielli, G.B.; Costantino, P.; Cardarelli, M. Expression of rolB in tobacco flowers affects the coordinated processes of anther dehiscence and style elongation. Plant J. 2004, 38, 512–525. [Google Scholar] [CrossRef] [PubMed]
- Altamura, M.M.; Capitani, F.; Gazza, L.; Capone, I.; Costantino, P. The plant oncogene rolB stimulates the formation of flowers and root meristemoids in tobacco thin cell layers. New Phytol. 1994, 126, 283–293. [Google Scholar] [CrossRef]
- Koltunow, A.M.; Johnson, S.D.; Lynch, M.; Yoshihara, T.; Costantino, P. Expression of rolB in apomictic Hieracium piloselloides Vill. causes ectopic meristems in planta and changes in ovule formation, where apomixis initiates at higher frequency. Planta 2001, 214, 196–205. [Google Scholar] [CrossRef]
- Bulgakov, V.P.; Gorpenchenko, T.Y.; Veremeichik, G.N.; Shkryl, Y.N.; Tchernoded, G.K.; Bulgakov, D.V.; Aminin, D.L.; Zhuravlev, Y.N. The rolB gene suppresses reactive oxygen species in transformed plant cells through the sustained activation of antioxidant defense. Plant Physiol. 2012, 158, 1371–1381. [Google Scholar] [CrossRef] [PubMed]
- Gorpenchenko, T.Y.; Aminin, D.L.; Vereshchagina, Y.V.; Shkryl, Y.N.; Veremeichik, G.N.; Tchernoded, G.K.; Bulgakov, V.P. Can plant oncogenes inhibit programmed cell death? The rolB oncogene reduces apoptosis-like symptoms in transformed plant cells. Plant Signal. Behav. 2012, 7, 1058–1061. [Google Scholar] [CrossRef]
- Veremeichik, G.N.; Shkryl, Y.N.; Rusapetova, T.V.; Silantieva, S.A.; Grigorchuk, V.P.; Velansky, P.V.; Brodovskaya, E.V.; Konnova, Y.A.; Khopta, A.A.; Bulgakov, D.V.; et al. Overexpression of the A4-rolB gene from the pRiA4 of Rhizobium rhizogenes modulates hormones homeostasis and leads to an increase of flavonoid accumulation and drought tolerance in Arabidopsis thaliana transgenic plants. Planta 2022, 256, 8. [Google Scholar] [CrossRef]
- Bettini, P.P.; Marvasi, M.; Fani, F.; Lazzara, L.; Cosi, E.; Melani, L.; Mauro, M.L. Agrobacterium rhizogenes rolB gene affects photosynthesis and chlorophyll content in transgenic tomato (Solanum lycopersicum L.) plants. J. Plant Physiol. 2016, 204, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Bettini, P.P.; Lazzara, L.; Massi, L.; Fani, F.; Mauro, M.L. Effect of far-red light exposure on photosynthesis and photoprotection in tomato plants transgenic for the Agrobacterium rhizogenes rolB gene. J. Plant Physiol. 2020, 245, 153095. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.H.; Lin, H.H.; Liu, C.T.; Lin, T.C.; Liu, L.Y.; Lee, K.T. Transcriptomic analysis reveals that reactive oxygen species and genes encoding lipid transfer protein are associated with tobacco hairy root growth and branch development. Mol. Plant Microbe Interact. 2014, 27, 678–687. [Google Scholar] [CrossRef] [PubMed]
- Favero, B.T.; Tan, Y.; Chen, X.; Müller, R.; Lütken, H. Kalanchoë blossfeldiana naturally transformed with Rhizobium rhizogenes exhibits superior root phenotype. Plant Sci. 2022, 321, 111323. [Google Scholar] [CrossRef] [PubMed]
- Favero, B.T.; Tan, Y.; Lin, Y.; Hansen, H.B.; Shadmani, N.; Xu, J.; He, J.; Müller, R.; Almeida, A.; Lütken, H. Transgenic Kalanchoë blossfeldiana, containing individual rol genes and open reading frames under 35S promoter, exhibit compact habit, reduced plant growth, and altered ethylene tolerance in flowers. Front. Plant Sci. 2021, 12, 672023. [Google Scholar] [CrossRef]
- Shkryl, Y.; Yugay, Y.; Vasyutkina, E.; Chukhlomina, E.; Rusapetova, T.; Bulgakov, V. The RolB/RolC homolog from sweet potato promotes early flowering and triggers premature leaf senescence in transgenic Arabidopsis thaliana plants. Plant Physiol. Biochem. 2022, 193, 50–60. [Google Scholar] [CrossRef]
- Arshad, W.; Haq, I.U.; Waheed, M.T.; Mysore, K.S.; Mirza, B. Agrobacterium-mediated transformation of tomato with rolB gene results in enhancement of fruit quality and foliar resistance against fungal pathogens. PLoS ONE 2014, 9, e96979. [Google Scholar] [CrossRef]
- Clément, B.; Perot, J.; Geoffroy, P.; Legrand, M.; Zon, J.; Otten, L. Abnormal accumulation of sugars and phenolics in tobacco roots expressing the Agrobacterium T-6b oncogene and the role of these compounds in 6b-induced growth. Mol. Plant Microbe Interact. 2007, 20, 53–62. [Google Scholar] [CrossRef]
- Bulgakov, V.P.; Shkryl, Y.N.; Veremeichik, G.N.; Gorpenchenko, T.Y.; Vereshchagina, Y.V. Recent advances in the understanding of Agrobacterium rhizogenes-derived genes and their effects on stress resistance and plant metabolism. In Biotechnology of Hairy Root Systems; Advances in Biochemical Engineering/Biotechnology Series; Springer: Berlin/Heidelberg, Germany, 2013; Volume 134, pp. 1–22. [Google Scholar] [CrossRef]
- Bulgakov, V.P.; Vereshchagina, Y.V.; Bulgakov, D.V.; Veremeichik, G.N.; Shkryl, Y.N. The rolB plant oncogene affects multiple signaling protein modules related to hormone signaling and plant defense. Sci. Rep. 2018, 8, 2285. [Google Scholar] [CrossRef]
- Adams, D.R.; Ron, D.; Kiely, P.A. RACK1, A multifaceted scaffolding protein: Structure and function. Cell Comm. Signal. 2011, 9, 22. [Google Scholar] [CrossRef]
- Islas-Flores, T.; Rahman, A.; Ullah, H.; Villanueva, M.A. The Receptor for Activated C Kinase in plant signaling: Tale of a promiscuous little molecule. Front. Plant Sci. 2015, 6, 1090. [Google Scholar] [CrossRef]
- Alshammari, S.O.; Dakshanamurthy, S.; Ullah, H. Small compounds targeting tyrosine phosphorylation of scaffold protein Receptor for Activated C Kinase1A (RACK1A) regulate auxin mediated lateral root development in Arabidopsis. Plant Signal. Behav. 2021, 16, 1899488. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Z.; Li, J.F.; Niu, Y.; Zhang, X.C.; Woody, O.Z.; Xiong, Y.; Djonović, S.; Millet, Y.; Bush, J.; McConkey, B.J.; et al. Pathogen-secreted proteases activate a novel plant immune pathway. Nature 2015, 521, 213–216. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Shan, L.; He, P. Stack Heterotrimeric G Proteins and MAPK Cascades on a RACK. Mol. Plant 2015, 8, 1691–1693. [Google Scholar] [CrossRef]
- Nakashima, A.; Chen, L.; Thao, N.P.; Fujiwara, M.; Wong, H.L.; Kuwano, M.; Umemura, K.; Shirasu, K.; Kawasaki, T.; Shimamoto, K. RACK1 functions in rice innate immunity by interacting with the Rac1 Immune Complex. Plant Cell 2008, 20, 2265–2279. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Yu, J.; Zhu, D.; Chang, Y.; Zhao, Q. Maize ZmRACK1 is involved in the plant response to fungal phytopathogens. Int. J. Mol. Sci. 2014, 15, 9343–9359. [Google Scholar] [CrossRef]
- Hyodo, K.; Suzuki, N.; Okuno, T. Hijacking a host scaffold protein, RACK1, for replication of a plant RNA virus. New Phytol. 2019, 221, 935–945. [Google Scholar] [CrossRef]
- Majzoub, K.; Hafirassou, M.L.; Meignin, C.; Goto, A.; Marzi, S.; Fedorova, A.; Verdier, Y.; Vinh, J.; Hoffmann, J.A.; Martin, F.; et al. RACK1 controls IRES-mediated translation of viruses. Cell 2014, 159, 1086–1095. [Google Scholar] [CrossRef]
- Ullah, H.; Hou, W.; Dakshanamurthy, S.; Tang, Q. Host targeted antiviral (HTA): Functional inhibitor compounds of scaffold protein RACK1 inhibit herpes simplex virus proliferation. Oncotarget 2019, 10, 3209–3226. [Google Scholar] [CrossRef]
- Chen, J.G.; Ullah, H.; Temple, B.; Liang, J.; Guo, J.; Alonso, J.M.; Ecker, J.R.; Jones, A.M. RACK1 mediates multiple hormone responsiveness and developmental processes in Arabidopsis. J. Exp. Bot. 2006, 57, 2697–2708. [Google Scholar] [CrossRef]
- Guo, J.; Hu, Y.; Zhou, Y.; Zhu, Z.; Sun, Y.; Li, J.; Wu, R.; Miao, Y.; Sun, X. Profiling of the Receptor for Activated C Kinase 1a (RACK1a) interaction network in Arabidopsis thaliana. Biochem. Biophys. Res. Commun. 2019, 520, 366–372. [Google Scholar] [CrossRef]
- Kodahl, N.; Müller, R.; Lütken, H. The Agrobacterium rhizogenes oncogenes rolB and ORF13 increase formation of generative shoots and induce dwarfism in Arabidopsis thaliana (L.) Heynh. Plant Sci. 2016, 252, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Yamada, K.; Hara-Nishimura, I.; Nishimura, M. Unique defense strategy by the endoplasmic reticulum body in plants. Plant Cell Physiol. 2011, 52, 2039–2049. [Google Scholar] [CrossRef] [PubMed]
- Nakano, R.T.; Yamada, K.; Bednarek, P.; Nishimura, M.; Hara-Nishimura, I. ER bodies in plants of the Brassicales order: Biogenesis and association with innate immunity. Front. Plant Sci. 2014, 5, 73. [Google Scholar] [CrossRef] [PubMed]
- Nakano, R.T.; Piślewska-Bednarek, M.; Yamada, K.; Edger, P.P.; Miyahara, M.; Kondo, M.; Böttcher, C.; Mori, M.; Nishimura, M.; Schulze-Lefert, P.; et al. PYK10 myrosinase reveals a functional coordination between endoplasmic reticulum bodies and glucosinolates in Arabidopsis thaliana. Plant J. 2017, 89, 204–220. [Google Scholar] [CrossRef]
- Bertini, L.; Proietti, S.; Aleandri, M.P.; Mondello, F.; Sandini, S.; Caporale, C.; Caruso, C. Modular structure of HEL protein from Arabidopsis reveals new potential functions for PR-4 proteins. Biol. Chem. 2012, 393, 1533–1546. [Google Scholar] [CrossRef] [PubMed]
- Ibeas, J.I.; Yun, D.J.; Damsz, B.; Narasimhan, M.L.; Uesono, Y.; Ribas, J.C.; Lee, H.; Hasegawa, P.M.; Bressan, R.A.; Pardo, J.M. Resistance to the plant PR-5 protein osmotin in the model fungus Saccharomyces cerevisiae is mediated by the regulatory effects of SSD1 on cell wall composition. Plant J. 2001, 25, 271–280. [Google Scholar] [CrossRef]
- Hao, J.; Wu, W.; Wang, Y.; Yang, Z.; Liu, Y.; Lv, Y.; Zhai, Y.; Yang, J.; Liang, Z.; Huang, K.; et al. Arabidopsis thaliana defense response to the ochratoxin A-producing strain (Aspergillus ochraceus 3.4412). Plant Cell Rep. 2015, 34, 705–719. [Google Scholar] [CrossRef]
- Park, E.J.; Kim, T.H. Arabidopsis OSMOTIN 34 functions in the ABA signaling pathway and is regulated by proteolysis. Int. J. Mol. Sci. 2021, 22, 7915. [Google Scholar] [CrossRef]
- Berger, S.; Mitchell-Olds, T.; Stotz, H.U. Local and differential control of vegetative storage protein expression in response to herbivore damage in Arabidopsis thaliana. Physiol. Plant. 2002, 114, 85–91. [Google Scholar] [CrossRef]
- Shah, S.J.; Anjam, M.S.; Mendy, B.; Anwer, M.A.; Habash, S.S.; Lozano-Torres, J.L.; Grundler, F.M.W.; Siddique, S. Damage-associated responses of the host contribute to defence against cyst nematodes but not root-knot nematodes. J. Exp. Bot. 2017, 68, 5949–5960. [Google Scholar] [CrossRef]
- Liu, N.; Sun, Y.; Wang, P.; Duan, H.; Ge, X.; Li, X.; Pei, Y.; Li, F.; Hou, Y. Mutation of key amino acids in the polygalacturonase-inhibiting proteins CkPGIP1 and GhPGIP1 improves resistance to Verticillium wilt in cotton. Plant J. 2018, 96, 546–561. [Google Scholar] [CrossRef] [PubMed]
- Nicaise, V.; Joe, A.; Jeong, B.R.; Korneli, C.; Boutrot, F.; Westedt, I.; Staiger, D.; Alfano, J.R.; Zipfel, C. Pseudomonas HopU1 modulates plant immune receptor levels by blocking the interaction of their mRNAs with GRP7. EMBO J. 2013, 32, 701–712. [Google Scholar] [CrossRef] [PubMed]
- Makowski, W.; Królicka, A.; Tokarz, B.; Miernicka, K.; Kołton, A.; Pięta, Ł.; Malek, K.; Ekiert, H.; Szopa, A.; Tokarz, K.M. Response of physiological parameters in Dionaea muscipula J. Ellis teratomas transformed with rolB oncogene. BMC Plant Biol. 2021, 21, 564. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Zhang, Y.; Zhu, J.; Sun, Y.; Guo, B.; Liu, F.; Huang, J.; Wang, H.; Dong, S.; Wang, Y.; et al. Phytophthora sojae apoplastic effector AEP1 mediates sugar uptake by mutarotation of extracellular aldose and is recognized as a MAMP. Plant Physiol. 2021, 187, 321–335. [Google Scholar] [CrossRef]
- Mohajjel-Shoja, H.; Clément, B.; Perot, J.; Alioua, M.; Otten, L. Biological activity of the Agrobacterium rhizogenes-derived trolC gene of Nicotiana tabacum and its functional relation to other plast genes. Mol. Plant Microbe Interact. 2011, 24, 44–53. [Google Scholar] [CrossRef]
- Wiszniewski, A.A.; Bussell, J.D.; Long, R.L.; Smith, S.M. Knockout of the two evolutionarily conserved peroxisomal 3-ketoacyl-CoA thiolases in Arabidopsis recapitulates the abnormal inflorescence meristem 1 phenotype. J. Exp. Bot. 2014, 65, 6723–6733. [Google Scholar] [CrossRef]
- Bussell, J.D.; Reichelt, M.; Wiszniewski, A.A.; Gershenzon, J.; Smith, S.M. Peroxisomal ATP-binding cassette transporter COMATOSE and the multifunctional protein abnormal INFLORESCENCE MERISTEM are required for the production of benzoylated metabolites in Arabidopsis seeds. Plant Physiol. 2014, 164, 48–54. [Google Scholar] [CrossRef]
- Ray, S.K.; Macoy, D.M.; Kim, W.Y.; Lee, S.Y.; Kim, M.G. Role of RIN4 in Regulating PAMP-Triggered Immunity and Effector-Triggered Immunity: Current Status and Future Perspectives. Mol. Cells 2019, 42, 503–511. [Google Scholar] [CrossRef]
- Weng, Y.; Ge, L.; Jia, S.; Mao, P.; Ma, X. Cyclophilin AtROC1S58F confers Arabidopsis cold tolerance by modulating jasmonic acid signaling and antioxidant metabolism. Plant Physiol. Biochem. 2020, 152, 81–89. [Google Scholar] [CrossRef]
- Jung, H.; Jo, S.H.; Park, H.J.; Lee, A.; Kim, H.S.; Lee, H.J.; Cho, H.S. Golgi-localized cyclophilin 21 proteins negatively regulate ABA signalling via the peptidyl prolyl isomerase activity during early seedling development. Plant Mol. Biol. 2020, 102, 19–38. [Google Scholar] [CrossRef]
- Abdeeva, I.A.; Pogorelko, G.V.; Maloshenok, L.G.; Mokrykova, M.V.; Fursova, O.V.; Bruskin, S.A. Search for partner proteins of A. thaliana immunophilins involved in the control of plant immunity. Molecules 2018, 23, 953. [Google Scholar] [CrossRef] [PubMed]
- Toribio, R.; Mangano, S.; Fernández-Bautista, N.; Muñoz, A.; Castellano, M.M. HOP, a co-chaperone involved in response to stress in plants. Front. Plant Sci. 2020, 11, 591940. [Google Scholar] [CrossRef] [PubMed]
- Kovacs, D.; Kalmar, E.; Torok, Z.; Tompa, P. Chaperone activity of ERD10 and ERD14, two disordered stress-related plant proteins. Plant Physiol. 2008, 147, 381–390. [Google Scholar] [CrossRef]
- Flannery, S.E.; Hepworth, C.; Wood, W.H.J.; Pastorelli, F.; Hunter, C.N.; Dickman, M.J.; Jackson, P.J.; Johnson, M.P. Developmental acclimation of the thylakoid proteome to light intensity in Arabidopsis. Plant J. 2021, 105, 223–244. [Google Scholar] [CrossRef] [PubMed]
- Singh, H.; Kaur, K.; Singh, M.; Kaur, G.; Singh, P. Plant cyclophilins: Multifaceted proteins with versatile roles. Front. Plant Sci. 2020, 11, 585212. [Google Scholar] [CrossRef] [PubMed]
- Dixon, D.P.; Edwards, R. Selective binding of glutathione conjugates of fatty acid derivatives by plant glutathione transferases. J. Biol. Chem. 2009, 284, 21249–21256. [Google Scholar] [CrossRef]
- Zhao, L.; Wang, C.; Zhu, F.; Li, Y. Mild osmotic stress promotes 4-methoxy indolyl-3-methyl glucosinolate biosynthesis mediated by the MKK9-MPK3/MPK6 cascade in Arabidopsis. Plant Cell Rep. 2017, 36, 543–555. [Google Scholar] [CrossRef]
- Lyou, S.H.; Park, H.J.; Jung, C.; Sohn, H.B.; Lee, G.; Kim, C.H.; Kim, M.; Choi, Y.D.; Cheong, J.J. The Arabidopsis AtLEC gene encoding a lectin-like protein is up-regulated by multiple stimuli including developmental signal, wounding, jasmonate, ethylene, and chitin elicitor. Mol. Cells 2009, 27, 75–81. [Google Scholar] [CrossRef]
- Wang, R.; Zeng, Z.; Guo, H.; Tan, H.; Liu, A.; Zhao, Y.; Chen, L. Over-expression of the Arabidopsis formate dehydrogenase in chloroplasts enhances formaldehyde uptake and metabolism in transgenic tobacco leaves. Planta 2018, 247, 339–354. [Google Scholar] [CrossRef]
- Myouga, F.; Hosoda, C.; Umezawa, T.; Iizumi, H.; Kuromori, T.; Motohashi, R.; Shono, Y.; Nagata, N.; Ikeuchi, M.; Shinozaki, K. A heterocomplex of iron superoxide dismutases defends chloroplast nucleoids against oxidative stress and is essential for chloroplast development in Arabidopsis. Plant Cell 2008, 20, 3148–3162. [Google Scholar] [CrossRef]
- Barros, J.; Escamilla-Trevino, L.; Song, L.; Rao, X.; Serrani-Yarce, J.C.; Palacios, M.D.; Engle, N.; Choudhury, F.K.; Tschaplinski, T.J.; Venables, B.J.; et al. 4-Coumarate 3-hydroxylase in the lignin biosynthesis pathway is a cytosolic ascorbate peroxidase. Nat. Commun. 2019, 10, 1994. [Google Scholar] [CrossRef] [PubMed]
- Jamet, E.; Roujol, D.; San-Clemente, H.; Irshad, M.; Soubigou-Taconnat, L.; Renou, J.P.; Pont-Lezica, R. Cell wall biogenesis of Arabidopsis thaliana elongating cells: Transcriptomics complements proteomics. BMC Genom. 2009, 10, 505. [Google Scholar] [CrossRef]
- DiMario, R.J.; Clayton, H.; Mukherjee, A.; Ludwig, M.; Moroney, J.V. Plant carbonic anhydrases: Structures, locations, evolution, and physiological roles. Mol. Plant 2017, 10, 30–46. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Sakamoto, W. Possible function of VIPP1 in maintaining chloroplast membranes. Biochim. Biophys. Acta 2015, 1847, 831–837. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Ding, P.; Sun, T.; Nitta, Y.; Dong, O.; Huang, X.; Yang, W.; Li, X.; Botella, J.R.; Zhang, Y. Heterotrimeric G proteins serve as a converging point in plant defense signaling activated by multiple receptor-like kinases. Plant Physiol. 2013, 161, 2146–2158. [Google Scholar] [CrossRef] [PubMed]
- Yamada, K.; Yamaguchi, K.; Shirakawa, T.; Nakagami, H.; Mine, A.; Ishikawa, K.; Fujiwara, M.; Narusaka, M.; Narusaka, Y.; Ichimura, K.; et al. The Arabidopsis CERK1-associated kinase PBL27 connects chitin perception to MAPK activation. EMBO J. 2016, 35, 2468–2483. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, B.; Xu, Y.; Li, H.; Li, S.; Zhang, D.; Mao, Z.; Guo, S.; Yang, C.; Weng, Y.; et al. The cyclophilin CYP20-2 modulates the conformation of BRASSINAZOLE-RESISTANT1, which binds the promoter of FLOWERING LOCUS D to regulate flowering in Arabidopsis. Plant Cell 2013, 25, 2504–2521. [Google Scholar] [CrossRef]
- Fu, Z.Q.; Guo, M.; Jeong, B.R.; Tian, F.; Elthon, T.E.; Cerny, R.L.; Staiger, D.; Alfano, J.R. A type III effector ADP-ribosylates RNA-binding proteins and quells plant immunity. Nature 2007, 447, 284–288. [Google Scholar] [CrossRef]
- Truman, W.; de Zabala, M.T.; Grant, M. Type III effectors orchestrate a complex interplay between transcriptional networks to modify basal defence responses during pathogenesis and resistance. Plant J. 2006, 46, 14–33. [Google Scholar] [CrossRef]
- Filippini, F.; Rossi, V.; Marin, O.; Trovato, M.; Costantino, P.; Downey, P.M.; Schiavo, F.L.; Terzi, M. A plant oncogene as a phosphatase. Nature 1996, 379, 499–500. [Google Scholar] [CrossRef]
- Espinosa, A.; Guo, M.; Tam, V.C.; Fu, Z.Q.; Alfano, J.R. The Pseudomonas syringae type III-secreted protein HopPtoD2 possesses protein tyrosine phosphatase activity and suppresses programmed cell death in plants. Mol. Microbiol. 2003, 49, 377–387. [Google Scholar] [CrossRef] [PubMed]
- Bretz, J.R.; Mock, N.M.; Charity, J.C.; Zeyad, S.; Baker, C.J.; Hutcheson, S.W. A translocated protein tyrosine phosphatase of Pseudomonas syringae pv. tomato DC3000 modulates plant defence response to infection. Mol. Microbiol. 2003, 49, 389–400. [Google Scholar] [CrossRef]
- Wang, M.; Soyano, T.; Machida, S.; Yang, J.; Jung, C.; Chua, N.; Yuan, Y.A. Molecular insights into plant cell proliferation disturbance by Agrobacterium protein 6b. Genes Dev. 2011, 25, 64–76. [Google Scholar] [CrossRef]
- Su, J.; Xu, J.; Zhang, S. RACK1, scaffolding a heterotrimeric G protein and a MAPK cascade. Trends Plant Sci. 2015, 20, 405–407. [Google Scholar] [CrossRef] [PubMed]
- Bulgakov, V.P.; Veremeichik, G.N.; Grigorchuk, V.P.; Rybin, V.G.; Shkryl, Y.N. The rolB gene activates secondary metabolism in Arabidopsis calli via selective activation of genes encoding MYB and bHLH transcription factors. Plant Physiol. Biochem. 2016, 102, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ahn, J.E.; Datta, S.; Salzman, R.A.; Moon, J.; Huyghues-Despointes, B.; Pittendrigh, B.; Murdock, L.L.; Koiwa, H.; Zhu-Salzman, K. Arabidopsis vegetative storage protein is an anti-insect acid phosphatase. Plant Physiol. 2005, 139, 1545–1556. [Google Scholar] [CrossRef]
- Jia, X.; Zeng, H.; Wang, W.; Zhang, F.; Yin, H. Chitosan oligosaccharide induces resistance to Pseudomonas syringae pv. tomato DC3000 in Arabidopsis thaliana by activating both salicylic acid- and jasmonic acid-mediated pathways. Mol. Plant Microbe Interact. 2018, 31, 1271–1279. [Google Scholar] [CrossRef]
- Howlader, P.; Bose, S.K.; Jia, X.; Zhang, C.; Wang, W.; Yin, H. Oligogalacturonides induce resistance in Arabidopsis thaliana by triggering salicylic acid and jasmonic acid pathways against Pst DC3000. Int. J. Biol. Macromol. 2020, 164, 4054–4064. [Google Scholar] [CrossRef]
- Mukherjee, A.K.; Carp, M.J.; Zuchman, R.; Ziv, T.; Horwitz, B.A.; Gepstein, S. Proteomics of the response of Arabidopsis thaliana to infection with Alternaria brassicicola. J. Proteom. 2010, 73, 709–720. [Google Scholar] [CrossRef]
- Moreno, J.E.; Tao, Y.; Chory, J.; Ballaré, C.L. Ecological modulation of plant defense via phytochrome control of jasmonate sensitivity. Proc. Natl. Acad. Sci. USA 2009, 106, 4935–4940. [Google Scholar] [CrossRef]
- Giovannoni, M.; Marti, L.; Ferrari, S.; Tanaka-Takada, N.; Maeshima, M.; Ott, T.; De Lorenzo, G.; Mattei, B. The plasma membrane-associated Ca2+-binding protein, PCaP1, is required for oligogalacturonide and flagellin-induced priming and immunity. Plant Cell Environ. 2021, 44, 3078–3093. [Google Scholar] [CrossRef]
- Huang, Y.P.; Huang, Y.W.; Chen, I.H.; Shenkwen, L.L.; Hsu, Y.H.; Tsai, C.H. Plasma membrane-associated cation-binding protein 1-like protein negatively regulates intercellular movement of BaMV. J. Exp. Bot. 2017, 68, 4765–4774. [Google Scholar] [CrossRef] [PubMed]
- Choi du, S.; Lim, C.W.; Hwang, B.K. Proteomics and functional analyses of Arabidopsis nitrilases involved in the defense response to microbial pathogens. Planta 2016, 244, 449–465. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Liu, J.; Wang, J.; Chen, S.; Chen, L.; Wang, J.; Wang, H.B.; Liu, B. The juxtamembrane domains of Arabidopsis CERK1, BAK1, and FLS2 play a conserved role in chitin-induced signaling. J. Integr. Plant Biol. 2020, 62, 556–562. [Google Scholar] [CrossRef] [PubMed]
- Gong, B.Q.; Wang, F.Z.; Li, J.F. Hide-and-seek: Chitin-triggered plant immunity and fungal counterstrategies. Trends Plant Sci. 2020, 25, 805–816. [Google Scholar] [CrossRef] [PubMed]
- Romano, P.G.; Horton, P.; Gray, J.E. The Arabidopsis cyclophilin gene family. Plant Physiol. 2004, 134, 1268–1282. [Google Scholar] [CrossRef]
- Speth, C.; Willing, E.M.; Rausch, S.; Schneeberger, K.; Laubinger, S. RACK1 scaffold proteins influence miRNA abundance in Arabidopsis. Plant J. 2013, 76, 433–445. [Google Scholar] [CrossRef]
- Bulgakov, V.P.; Veremeichik, G.N.; Shkryl, Y.N. The rolB gene activates the expression of genes encoding microRNA processing machinery. Biotechnol. Lett. 2015, 37, 921–925. [Google Scholar] [CrossRef]
- Veremeichik, G.; Bulgakov, V.; Shkryl, Y. Modulation of NADPH-oxidase gene expression in rolB-transformed calli of Arabidopsis thaliana and Rubia cordifolia. Plant Physiol. Biochem. 2016, 105, 282–289. [Google Scholar] [CrossRef]
- Shkryl, Y.N.; Veremeichik, G.N.; Bulgakov, V.P.; Avramenko, T.V.; Günter, E.A.; Ovodov, Y.S.; Muzarok, T.I.; Zhuravlev, Y.N. The production of class III plant peroxidases in transgenic callus cultures transformed with the rolB gene of Agrobacterium rhizogenes. J. Biotechnol. 2013, 168, 64–70. [Google Scholar] [CrossRef]
- Daudi, A.; Cheng, Z.; O’Brien, J.A.; Mammarella, N.; Khan, S.; Ausubel, F.M.; Bolwell, G.P. The apoplastic oxidative burst peroxidase in Arabidopsis is a major component of pattern-triggered immunity. Plant Cell 2012, 24, 275–287. [Google Scholar] [CrossRef] [PubMed]
- Kámán-Tót, E.; Dankó, T.; Gullner, G.; Bozsó, Z.; Palkovics, L.; Pogány, M. Contribution of cell wall peroxidase- and NADPH oxidase-derived reactive oxygen species to Alternaria brassicicola-induced oxidative burst in Arabidopsis. Mol. Plant Pathol. 2019, 20, 485–499. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, F.K.; Rivero, R.M.; Blumwald, E.; Mittler, R. Reactive oxygen species, abiotic stress and stress combination. Plant J. 2017, 90, 856–867. [Google Scholar] [CrossRef] [PubMed]
- Shkryl, Y.; Yugay, Y.; Avramenko, T.; Grigorchuk, V.; Gorpenchenko, T.; Grischenko, O.; Bulgakov, V. CRISPR/Cas9-mediated knockout of HOS1 reveals its role in the regulation of secondary metabolism in Arabidopsis thaliana. Plants 2021, 10, 104. [Google Scholar] [CrossRef] [PubMed]
- Spena, A.; Schmülling, T.; Koncz, C.; Schell, J.S. Independent and synergistic activity of rolA, B and C loci in stimulating abnormal growth in plants. EMBO J. 1987, 6, 3891–3899. [Google Scholar] [CrossRef] [PubMed]
- Perez-Riverol, Y.; Bai, J.; Bandla, C.; García-Seisdedos, D.; Hewapathirana, S.; Kamatchinathan, S.; Kundu, D.J.; Prakash, A.; Frericks-Zipper, A.; Eisenacher, M.; et al. The PRIDE database resources in 2022: A hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 2022, 50, D543–D552. [Google Scholar] [CrossRef]
- Czechowski, T.; Stitt, M.; Altmann, T.; Udvardi, M.K.; Scheible, W.R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 2005, 139, 5–17. [Google Scholar] [CrossRef]
- Brzezinka, K.; Altmann, S.; Czesnick, H.; Nicolas, P.; Gorka, M.; Benke, E.; Kabelitz, T.; Jähne, F.; Graf, A.; Kappel, C.; et al. Arabidopsis FORGETTER1 mediates stress-induced chromatin memory through nucleosome remodeling. Elife 2016, 5, e17061. [Google Scholar] [CrossRef]
- Dhanasekaran, S.; Doherty, T.M.; Kenneth, J.; TB Trials Study Group. Comparison of different standards for real-time PCR-based absolute quantification. J. Immunol. Methods 2010, 354, 34–39. [Google Scholar] [CrossRef]
- Meyer, S.A.; Jirtle, R.L. Old dance with a new partner: EGF receptor as the phenobarbital receptor mediating Cyp2B expression. Sci. Signal. 2013, 6, pe16. [Google Scholar] [CrossRef]
- Sabila, M.; Kundu, N.; Smalls, D.; Ullah, H. Tyrosine phosphorylation based homo-dimerization of Arabidopsis RACK1A proteins regulates oxidative stress signaling pathways in yeast. Front. Plant Sci. 2016, 7, 176. [Google Scholar] [CrossRef] [PubMed]
- Grishchenko, O.V.; Kiselev, K.V.; Tchernoded, G.K.; Fedoreyev, S.A.; Veselova, M.V.; Bulgakov, V.P.; Zhuravlev, Y.N. RolB gene-induced production of isoflavonoids in transformed Maackia amurensis cells. Appl. Microbiol. Biotechnol. 2016, 100, 7479–7489. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Li, L.; Wang, L.; Wang, S.; Li, S.; Du, J.; Zhang, S.; Shou, H. MPK3/MPK6 are involved in iron deficiency-induced ethylene production in Arabidopsis. Front. Plant Sci. 2015, 6, 953. [Google Scholar] [CrossRef] [PubMed]
- Wingenter, K.; Trentmann, O.; Winschuh, I.; Hörmiller, I.I.; Heyer, A.G.; Reinders, J.; Schulz, A.; Geiger, D.; Hedrich, R.; Neuhaus, H.E. A member of the mitogen-activated protein 3-kinase family is involved in the regulation of plant vacuolar glucose uptake. Plant J. 2011, 68, 890–900. [Google Scholar] [CrossRef] [PubMed]
- Arraño-Salinas, P.; Domínguez-Figueroa, J.; Herrera-Vásquez, A.; Zavala, D.; Medina, J.; Vicente-Carbajosa, J.; Meneses, C.; Canessa, P.; Moreno, A.A.; Blanco-Herrera, F. WRKY7, -11 and -17 transcription factors are modulators of the bZIP28 branch of the unfolded protein response during PAMP-triggered immunity in Arabidopsis thaliana. Plant Sci. 2018, 277, 242–250. [Google Scholar] [CrossRef] [PubMed]

| UniProtKB Code | Name of the Protein/Short Name | Protein Function 1 | Activation, Folds 2 | |
|---|---|---|---|---|
| Chaperones | ||||
| 1 | Q38867 | Peptidyl-prolyl cis-trans isomerase CYP19-3/ROC2 | Protein peptidyl-prolyl isomerization | 3.8 |
| 2 | P34790 | Peptidyl-prolyl cis-trans isomerase CYP18-3/ROC1 | Protein peptidyl-prolyl isomerization | 2.0 |
| 3 | Q42406 | Peptidyl-prolyl cis-trans isomerase CYP18-4/ROC5 | Protein peptidyl-prolyl isomerization | 1.7 |
| 4 | Q38900 | Peptidyl-prolyl cis-trans isomerase CYP19-1/ROC3 | Protein peptidyl-prolyl isomerization | 1.5 |
| 5 | Q9LKR3 Q39043 | Heat shock 70 kDa protein BIP1/AtHsp70-11 Heat shock 70 kDa protein BIP2/AtHsp70-12 | Protein refolding; required for pollen development and pollen tube growth | 2.8 |
| Plant defense | ||||
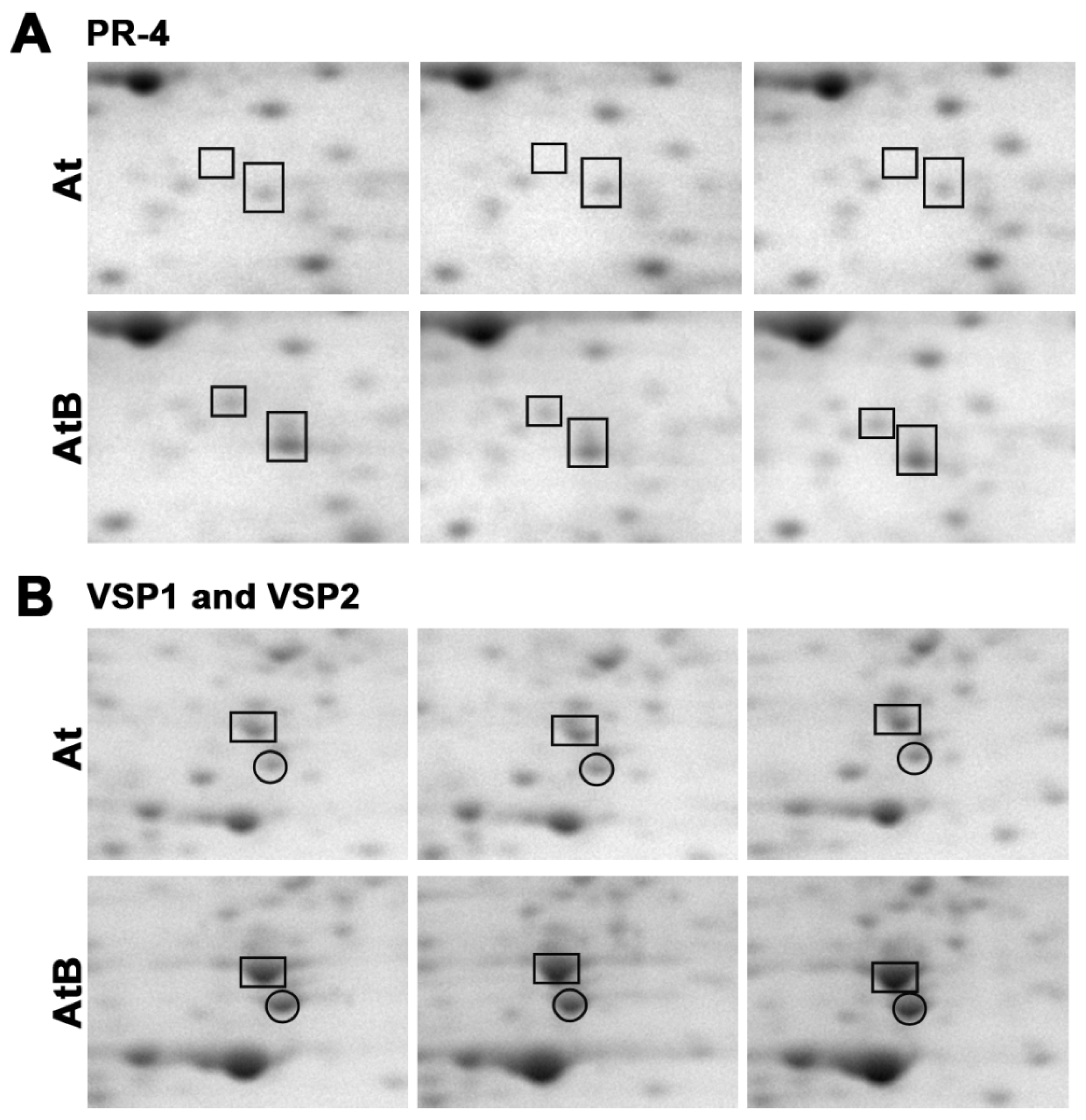
| 1 | P43082 | Hevein-like preproprotein/PR-4/ HEL | Defense response to bacterium, fungus, and virus; systemic acquired resistance | 10 |
| 2 | P50700 | Osmotin-like protein OSM34 | Defense response to fungus | 10 |
| 3 | Q9LJR2 | Lectin-like protein LEC | Defense response to fungus, triggered by jasmonate, ethylene and chitin | 10 |
| 4 | O49195 | Vegetative storage protein 1 | Defense response, response to jasmonic acid | 3.4 |
| 5 | O82122 | Vegetative storage protein 2 | Defense response to insects; response to wounding, oxidative stress, and jasmonic acid | 6.7 |
| 6 | Q9SR37 | Beta-glucosidase 23/ BGLU23/PYK10 | Major component of the ER body; glucosinolate catabolic process | 10 |
| 7 | Q9C5C2 | Myrosinase 2/Beta-glucosidase 37/ BGLU37 | Defense response to insect; glucosinolate catabolic process; role in abscisic acid and methyl jasmonate signaling | 2.0 |
| 8 | Q9SE50 | Beta-D-glucopyranosyl abscisate beta-glucosidase/BGLU18 | Defense response to fungus and insect; response to salt stress and water deprivation | 2.0 |
| 9 | Q9LJG3 | GDSL esterase/lipase ESM1 | Response to insect; glucosinolate catabolic process | 1.7 |
| 10 | O04314 | PYK10-binding protein 1 (Jacalin-related lectin 30)/PBP1 | Regulates the correct polymerization of BGLU23/PYK10 upon tissue damage | 10 |
| 12 | O80948 | Jacalin-related lectin 23/JAL23 | Polymerization and activation of BGLU23/PYK10 upon tissue damage | 3.3 |
| 13 | O04309 | Jacalin-related lectin 35/JAL35 | Component of the PYK10 complex; JA-responsive protein | 3.0 |
| 14 | Q9LK72 | Lectin-like protein At3g16530 | Response to oomycetes | 3.2 |
| 15 | Q9LNN2 | Lectin-like protein At1g53070 | May be involved in response to insect | 2.8 |
| 16 | Q93XW5 | Nitrile-specifier protein 5/NSP5 | Glucosinolate catabolic process, nitrile biosynthetic process | 4.5 |
| 17 | Q9M5J9 | Polygalacturonase inhibitor 1/PGIP-1 | Inhibitor of fungal polygalacturonase. An important factor for plant resistance to phytopathogenic fungi | 5.0 |
| 18 | O81862 | Class V chitinase | Hydrolyzes glycol chitin and chitin oligosaccharides; response to abscisic acid and jasmonic acid; response to salt stress | 5.1 |
| 19 | Q9MBH2 | Protein AIG2 B / Avirulence-induced gene 2 protein B | Response to bacterium | 4.7 |
| 20 | Q96262 | Plasma membrane-associated cation-binding protein 1/PCaP1 | Response to oligogalacturonides and flagellin; restricts viral cell-to-cell movement; response to cold, salt, and water deprivation | 1.7 |
| 21 | P32962 | Nitrilase 2/NIT2 | Involved in plant defense and R gene-mediated resistant responses to microbial pathogens | 2.0 |
| Response to abiotic stress | ||||
| 1 | Q9SYT0 | Annexin D1/ANN1 | Response to multiple stresses, including water deprivation | 3.8 |
| 2 | Q0PGJ6 | NADPH-dependent aldo-keto reductase, chloroplastic | Response to cold, salt, and water deprivation | 10 |
| 3 | O80944 | Aldo-keto reductase family 4 member C8 | Response to cold, salt, and water deprivation | 1.5 |
| 4 | O49629 | Probable plastid-lipid-associated protein 2, chloroplastic/AtPap2 | Probably involved in light/cold stress-related jasmonate biosynthesis | 3.8 |
| 5 | P46422 | Glutathione S-transferase F2 | Regulation of binding and transport of defense-related compounds during plant stress; binds auxin, flavonoids and camalexin | 2.8 |
| 6 | O80852 | Glutathione S-transferase F9 | Stress response; detoxification | 2.3 |
| 7 | P42761 | Glutathione S-transferase F10 | Response to water deprivation | 2.0 |
| 8 | Q9S7E4 | Formate dehydrogenase, chloroplastic/mitochondrial/FDH | Oxidoreductase; cell stress response | 1.7 |
| 9 | Q9LZ66 | Assimilatory sulfite reductase (ferredoxin), chloroplastic/AtSiR | Sulfate assimilation; response to cold | 2.5 |
| Photosynthesis | ||||
| 1 | Q9M0V6 | Ferredoxin--NADP reductase, root isozyme 1, chloroplastic /RFNR 1 | Photosynthesis; ferredoxin-NADP+ reductase activity | 4.9 |
| 2 | Q9LHH7 | Bifunctional protein FolD 2 | Photorespiration; one-carbon metabolic process | 2.3 |
| Oxidative stress and redox homeostasis | ||||
| 1 | Q9SMU8 | Peroxidase 34 | Hydrogen peroxide catabolic process; response to oxidative stress; defense response to bacterium and fungus | 7.2 |
| 2 | Q9XEX2 | Peroxiredoxin-2B | Cell redox homeostasis, hydrogen peroxide catabolic process | 2.3 |
| 3 | Q9M7T0 | Peroxiredoxin-2F, mitochondrial | Cell redox homeostasis; hydrogen peroxide catabolism | 1.6 |
| 4 | Q949U7 | Peroxiredoxin-2E, chloroplastic | Cell redox homeostasis, hydrogen peroxide catabolic process | 1.6 |
| 5 | Q9FWR4 | Glutathione S-transferase DHAR1, mitochondrial | Key component of the ascorbate recycling system; involved in the redox homeostasis, especially in ROS scavenging under oxidative stress | 1.8 |
| 6 | Q05431 | L-ascorbate peroxidase 1, cytosolic | Key role in hydrogen peroxide removal; lignin biosynthetic process; phenylpropanoid biosynthetic process | 2.0 |
| 7 | P42760 | Glutathione S-transferase F6 | Response to oxidative stress; defense response to bacterium; involved in camalexin biosynthesis; response to water deprivation | 5.0 |
| 8 | Q9ZRW8 | Glutathione S-transferase U19 | Response to oxidative stress Response to water deprivation | 2.0 |
| 9 | O78310 | Superoxide dismutase [Cu-Zn] 2, chloroplastic/CSD2 | Removal of superoxide radicals; response to light intensity, UV-B; response to salt stress | 2.2 |
| 10 | P21276 | Superoxide dismutase [Fe] 1, chloroplastic | Removal of superoxide radicals. Response to light intensity | 1.8 |
| 11 | P52577 | Isoflavone reductase homolog P3 | Response to oxidative stress | 3.4 |
| 12 | Q9LSQ5 | NAD(P)H dehydrogenase (quinone) FQR1 | Response to oxidative stress | 2.4 |
| 13 | Q9SV68 | Chloroplast envelope quinone oxidoreductase homolog/ceQORH | Reduces toxic stress-related oxidized lipids produced under oxidative stress conditions | 1.8 |
| Development | ||||
| 1 | Q9FGY1 | Beta-D-xylosidase 1 | Seed coat development | 4.4 |
| 2 | O65351 | Subtilisin-like protease SBT1.7 | Seed coat development | 4.7 |
| 3 | O64530 | Thiosulfate/3-mercaptopyruvate sulfurtransferase 1, mitochondrial/MST1 | Embryo development ending in seed dormancy | 3.9 |
| 4 | P24806 | Xyloglucan endotransglucosylase/hydrolase protein 24 | Cell wall biogenesis | 4.6 |
| 5 | P31265 | Translationally-controlled tumor protein 1/TCTP1 | General regulator required for the development of the entire plant | 1.5 |
| Metabolism | ||||
| 1 | Q9STT3 | Aldose 1-epimerase | Carbohydrate metabolism; glucose metabolic process | 10 |
| 2 | A0A097PMS4 | Galactose mutarotase-like superfamily protein | Carbohydrate metabolism | 5.0 |
| 3 | Q9SPK5 | Formate--tetrahydrofolate ligase | One-carbon metabolism | 4.0 |
| 4 | Q9LYG3 | NADP-dependent malic enzyme 2 | Malate and pyruvate metabolic processes; pentose-phosphate shunt, oxidative branch | 3.0 |
| 5 | Q9SK66 | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 9, mitochondrial | Ubiquinone-6 biosynthetic process; electron transport | 2.9 |
| 6 | Q9SMN0 | Probable carboxylesterase 12 | Carboxylic ester hydrolase activity | 3.1 |
| 7 | Q56WD9 | 3-ketoacyl-CoA thiolase 2, peroxisomal/ PED1/KAT2 | Fatty acid biosynthesis; links fatty acid beta-oxidation with redox regulation; required for the accumulation of benzoylated glucosinolates | 3.4 |
| 8 | O48917 | UDP-sulfoquinovose synthase, chloroplastic | Glycolipid and sulfolipid biosynthetic processes | 3.0 |
| 9 | Q8VY84 | Probable UMP-CMP kinase1/UMK1 | Pyrimidine biosynthesis | 2.7 |
| 10 | P34066 | Proteasome subunit alpha type-1-A/PAF1 | Ubiquitin-dependent protein catabolic process | 2.5 |
| 11 | Q9XI05 | Proteasome subunit beta type-3-A/PBC1 | Protein catabolic process | 1.7 |
| 12 | Q56WN1 | Glutamine synthetase cytosolic isozyme 1-1 | Glutamine biosynthetic process | 2.6 |
| 13 | P47999 | Cysteine synthase, chloroplastic/chromoplastic/OASB | A major cysteine synthase | 2.2 |
| 14 | Q9LK23 | Glucose-6-phosphate 1-dehydrogenase 5, cytoplasmic | Pentose-phosphate shunt, oxidative branch | 2.1 |
| 15 | Q9FIE8 | UDP-glucuronic acid decarboxylase 3 | UDP-D-xylose biosynthetic process | 1.9 |
| 16 | Q9S9W2 | Short-chain dehydrogenase/reductase SDRA | Fatty acid metabolic process | 1.6 |
| 17 | Q9T034 | Phenylalanine--tRNA ligase alpha subunit, cytoplasmic | Protein biosynthesis | 1.5 |
| 18 | Q8L934 | Nucleoid DNA-binding-like protein | Proteolysis | 10 |
| Other proteins | ||||
| 1 | Q9ZSK4 | Actin-depolymerizing factor 3/ADF-3 | Actin filament depolymerization | 4.7 |
| 2 | Q29Q34 | NAD(P)-binding Rossmann-fold superfamily protein / At5g19440 | Oxidoreductase activity; response to Agrobacterium tumefaciens | 1.5 |
| 3 | O23016 | Probable voltage-gated potassium channel subunit beta | Potassium ion transport | 1.9 |
| 4 | Q9XI10 | DPP6 N-terminal domain-like protein | Unknown function | 4.7 |
| 5 | Q8GYZ3 | RmlC-like jelly roll fold protein | Unknown function | 3.1 |
| 6 | Q9SSK5 | MLP-like protein 43 | Unknown function | 3.0 |
| UniProtKB Code | Name of the Protein | Protein Function 1 | Inhibition, Folds 2 | |
|---|---|---|---|---|
| 1 | O24456 | Receptor for activated C kinase 1A/ RACK1A | Major component of the RACK1 regulatory proteins that play a role in multiple signal transduction pathways | 1.5 |
| 2 | Q9C4Z6 | Receptor for activated C kinase 1B/RACK1B | Minor component of the RACK1 regulatory proteins | 3.8 |
| 3 | Q9LV28 | Receptor for activated C kinase 1C /RACK1C | Minor component of the RACK1 regulatory proteins | 4.2 |
| Chaperones | ||||
| 1 | Q9LF37 | Chaperone protein ClpB3, chloroplastic | Molecular chaperone essential for chloroplast development and seedling viability; response to heat | 3.0 |
| 2 | P42730 | Chaperone protein ClpB1/Heat shock protein 101 | Molecular chaperone that plays an important role in thermotolerance | 3.0 |
| 3 | Q9FI56 | Chaperone protein ClpC1, chloroplastic | Regulation of chlorophyll biosynthetic process; chloroplast organization | 3.1 |
| 4 | Q9SAR5 | Ankyrin repeat domain-containing protein 2A | Chaperone; protein targeting to chloroplast | 2.2 |
| 5 | Q03250 | Glycine-rich RNA-binding protein 7/AtGR-RBP7/ GRP7 | Chaperone; innate immunity; plant defense; target of the Pseudomonas syringae type III effector HopU1 | 2.0 |
| 6 | P42763 | Dehydrin ERD14 | Chaperone, protein stabilization; response to cold and water deprivation | 1.9 |
| 7 | Q9LNB6 | Hsp70-Hsp90 organizing protein 1/AtHop1 | Mediates the association of the molecular chaperones HSP70 and HSP90; stress response | 1.5 |
| Plant defense | ||||
| 1 | Q9FN05 | Probable glucan 1,3-alpha-glucosidase | Defense response to bacterium; required for sustained activation of EFR-mediated signaling | 3.1 |
| 2 | Q9FXA2 | Polyadenylate-binding protein 8/PABP-8 | Host-virus interaction | 1.7 |
| 3 | Q9ZUU4 | RNA-binding protein CP29B, chloroplastic | Innate immune response; potential target of HopU1 | 1.7 |
| 4 | Q9FGS0 | RNA-binding protein CP31B, chloroplastic | Innate immune response; potential target of HopU1 | 1.7 |
| Response to stress | ||||
| 1 | P42759 | Dehydrin ERD10 | Protein stabilization; cold acclimation; response to abscisic acid and water deprivation | 3.9 |
| 2 | P54887 | Delta-1-pyrroline-5-carboxylate synthase A | Plays a key role in proline biosynthesis. Response to abscisic acid, cold, desiccation, salt stress, and oxidative stress | 2.4 |
| 3 | Q9FLT0 | Ribonuclease TUDOR 2 | Cytoprotective ribonuclease, which is essential for resistance to abiotic stress | 2.3 |
| 4 | O23523 | RGG repeats nuclear RNA binding protein A | Promotes stomata closure in drought conditions; involved in resistance to salt and drought | 1.5 |
| Photosynthesis | ||||
| 1 | Q9SW18 | Magnesium protoporphyrin IX methyltransferase, chloroplastic | Chlorophyll biosynthesis | 3.0 |
| 2 | Q9LR75 | Coproporphyrinogen-III oxidase 1, chloroplastic | Chlorophyll biosynthesis | 5.0 |
| 3 | O22886 | Uroporphyrinogen decarboxylase 2, chloroplastic | Chlorophyll biosynthesis | 2.8 |
| 4 | P21218 | Protochlorophyllide reductase B, chloroplastic | Chlorophyll biosynthesis | 2.4 |
| 5 | O48741 | Protochlorophyllide reductase C, chloroplastic | Chlorophyll biosynthesis | 2.4 |
| 6 | Q9FMD5 | Protein TIC 40, chloroplastic | Chloroplast organization, protein import into chloroplast stroma | 2.3 |
| 7 | Q9FKW6 | Ferredoxin--NADP reductase, leaf isozyme 1, chloroplastic/FNR 1 | Photosynthetic electron transport chain | 2.0 |
| 8 | Q8W493 | Ferredoxin--NADP reductase, leaf isozyme 2, chloroplastic /FNR 2 | Regulates photosynthetic electron flow during the transition from dark to light | 1.7 |
| 9 | Q01908 | ATP synthase gamma chain 1, chloroplastic/ATPC1 | ATP biosynthetic process; photosynthetic electron transport in photosystem II | 1.8 |
| 10 | P42699 | Plastocyanin major isoform, chloroplastic | Electron transfer between P700 and the cytochrome b6-f complex in photosystem I | 3.0 |
| 11 | Q9ZR03 | Cytochrome b6-f complex iron-sulfur subunit, chloroplastic | Electron transfer between photosystem II and photosystem I | 1.5 |
| 12 | P37107 | Signal recognition particle 54 kDa protein, chloroplastic | Required for light-harvesting chlorophyll a/b-binding protein integration into thylakoids | 4.3 |
| 13 | P27140 | Beta carbonic anhydrase 1, chloroplastic/AtbCA1 | Photosynthesis; carbon utilization | 3.6 |
| 14 | O80796 | Membrane-associated protein VIPP1, chloroplastic | Thylakoid membrane organization | 1.9 |
| General metabolism | ||||
| 1 | P42737 | Beta carbonic anhydrase 2, chloroplastic/AtbCA2 | Reversible hydration of carbon dioxide; plays an important role in amino acid biosynthesis | 9.8 |
| 2 | Q9FYE3 | Alpha carbonic anhydrase 3/AtaCA3 | Reversible hydration of carbon dioxide | 2.5 |
| 3 | Q9LR30 | Glutamate--glyoxylate aminotransferase 1 | Glycine biosynthetic process | 2.2 |
| 4 | Q56YA5 | Serine-glyoxylate aminotransferase | Glycine biosynthetic process | 1.9 |
| 5 | Q94B78 | Glycine dehydrogenase (decarboxylating) 1, mitochondrial | Glycine catabolic process | 2.8 |
| 6 | Q8W593 | Probable lactoylglutathione lyase, chloroplastic | Methylglyoxal catabolic process; response to cold | 3.3 |
| 7 | O50008 | 5-methyltetrahydropteroyltriglutamate homocysteine methyltransferase 1 | Methionine biosynthesis | 3.0 |
| 8 | Q9SYM5 | Trifunctional UDP-glucose 4,6-dehydratase/UDP-4-keto-6-deoxy-D-glucose 3,5-epimerase/UDP-4-keto-L-rhamnose-reductase RHM1 | UDP-rhamnose biosynthetic process; plays a major role in supplying UDP-rhamnose for flavonol biosynthesis | 1.7 |
| 9 | P56757 | ATP synthase subunit alpha, chloroplastic | ATP biosynthesis; response to cold | 3.0 |
| 10 | Q9C7N5 | GDSL esterase/lipase At1g29660 | Lipid metabolism | 3.0 |
| 11 | O23553 | Beta-amylase 3, chloroplastic | Carbohydrate metabolism; response to cold | 3.0 |
| 12 | Q84TF0 | Aldo-keto reductase family 4-member C10 | Metabolism of ketosteroids and aldehydes | 1.5 |
| 13 | Q9LD43 | Acetyl-coenzyme A carboxylase carboxyl transferase subunit alpha, chloroplastic | Fatty acid biosynthesis | 2.5 |
| 14 | P52410 | 3-oxoacyl-[acyl-carrier-protein] synthase I, chloroplastic | Fatty acid biosynthesis | 1.5 |
| 15 | P42734 | Probable cinnamyl alcohol dehydrogenase 9 | Lignin biosynthesis | 1.7 |
| 16 | O64767 | AICARFT/IMPCHase bienzyme family protein | Purine biosynthesis | 1.5 |
| 17 | Q96533 | Alcohol dehydrogenase class-3 | Ethanol oxidation; formaldehyde catabolic process | 1.6 |
| 18 | Q949Y0 | Ubiquitin carboxyl-terminal hydrolase 6 | Proteasome-mediated ubiquitin-dependent protein catabolic process | 3.4 |
| 19 | Q8H0S9 | Puromycin-sensitive aminopeptidase/PSA | Proteolysis; essential for cell growth and viability | 3.0 |
| 20 | Q1EBV4 | Protein DNA-DAMAGE INDUCIBLE 1 | Proteolysis | 2.3 |
| Protein biosynthesis | ||||
| 1-2 | P49227 Q8LBI1 | 60S ribosomal protein L5-2 60S ribosomal protein L5-1 | Protein biosynthesis; leaf morphogenesis; root morphogenesis | 2.3 |
| 3-4 | Q9SIP7 Q9FJA6 | 40S ribosomal protein S3-1 40S ribosomal protein S3-3 | Translation | 2.3 |
| 5 | Q93VC7 | 30S ribosomal protein S1, chloroplastic | Translation; required for optimal plastid performance in terms of photosynthesis | 1.8 |
| 6 | O50061 | 50S ribosomal protein L4, chloroplastic | Translation | 1.7 |
| 7 | P36210 | 50S ribosomal protein L12-1, chloroplastic | Translation | 1.6 |
| 8 | P51412 | 50S ribosomal protein L21, chloroplastic | Translation; embryo development ending in seed dormancy | 1.8 |
| 9 | Q9LY66 | 50S ribosomal protein L1, chloroplastic | Translation | 1.5 |
| 10 | Q9SI75 | Elongation factor G, chloroplastic | Protein biosynthesis; post-embryonic development | 2.3 |
| 11 | Q9ASR1 | Elongation factor 2 | Protein biosynthesis; cold acclimation | 2.1 |
| 12 | Q8VZW6 | Elongation factor P (EF-P) family protein | Protein biosynthesis | 2.0 |
| Other proteins | ||||
| 1 | Q941D3 | Probable plastid-lipid-associated protein 8, chloroplastic | Unknown function | 2.4 |
| 2 | Q8L606 | Tetratricopeptide repeat (TPR)-like superfamily protein | Unknown function | 2.9 |
| 3 | Q9LVT8 | RGG repeats nuclear RNA binding protein C | Unknown function | 1.5 |
| UniProtKB Code | Protein | Function or Biological Process | Reference |
|---|---|---|---|
| Upregulated | |||
| Q42406 | Peptidyl-prolyl cis-trans isomerase CYP18-4/Rotamase cyclophilin-5/ROC5 | The closest analogue AtCYP18-3 is involved in plant-pathogen interactions | [56] |
| Q9ZRW8 | Glutathione S-transferase U19, cytosolic | Response to oxidative stress | [57] |
| O49195 | Vegetative storage protein1/VSP1 | Defense response. Induced by mechanical wounding, jasmonic acid (JA), insect herbivory, osmotic and nutritional stresses | [40] |
| Q93XW5 | Nitrile-specifier protein 5 | Defense against herbivory and pathogen attacks; component of the glucosinolate-myrosinase system | [58] |
| Q9LK72 | Lectin-like protein At3g16530 | Plant defense response; homologs are induced by fungal elicitors | [59] |
| Q9SR37 | Beta-glucosidase 23/PYK10 | Defense against pests and fungi | [33] |
| Q9S7E4 | Formate dehydrogenase, chloroplastic/mitochondrial | Formaldehyde metabolism; induced by various stresses | [60] |
| P21276 | Superoxide dismutase [Fe] 1, chloroplastic/FSD1 | Response to oxidative stress: response to light intensity | [61] |
| Q9M5J9 | Polygalacturonase inhibitor 1/ PGIP1 | Restricts the growth of invasive fungal pathogens; defense against cyst nematodes | [41,42] |
| Q05431 | L-ascorbate peroxidase 1, cytosolic | Cellular response to oxidative stress; phenylpropanoid biosynthetic process; lignin biosynthesis | [62] |
| P24806 | Xyloglucan endotransglucosylase/ hydrolase protein 24 | Cell wall biogenesis | [63] |
| Downregulated | |||
| P27140 | Beta carbonic anhydrase 1, chloroplastic | Photosynthesis; stomatal development; defense response | [64] |
| P36210 | 50S ribosomal protein L12-1, chloroplastic | Translation | UniProtKB |
| Q9LVT8 | RGG repeats nuclear RNA binding protein C | mRNA and RNA binding | UniProtKB |
| O80796 | Membrane-associated protein VIPP1, chloroplastic | Thylakoid membrane organization and vesicle organization | [65] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vereshchagina, Y.V.; Mironova, A.A.; Bulgakov, D.V.; Bulgakov, V.P. Proteomic Analysis of Proteins Related to Defense Responses in Arabidopsis Plants Transformed with the rolB Oncogene. Int. J. Mol. Sci. 2023, 24, 1880. https://doi.org/10.3390/ijms24031880
Vereshchagina YV, Mironova AA, Bulgakov DV, Bulgakov VP. Proteomic Analysis of Proteins Related to Defense Responses in Arabidopsis Plants Transformed with the rolB Oncogene. International Journal of Molecular Sciences. 2023; 24(3):1880. https://doi.org/10.3390/ijms24031880
Chicago/Turabian StyleVereshchagina, Yulia V., Anastasiya A. Mironova, Dmitry V. Bulgakov, and Victor P. Bulgakov. 2023. "Proteomic Analysis of Proteins Related to Defense Responses in Arabidopsis Plants Transformed with the rolB Oncogene" International Journal of Molecular Sciences 24, no. 3: 1880. https://doi.org/10.3390/ijms24031880
APA StyleVereshchagina, Y. V., Mironova, A. A., Bulgakov, D. V., & Bulgakov, V. P. (2023). Proteomic Analysis of Proteins Related to Defense Responses in Arabidopsis Plants Transformed with the rolB Oncogene. International Journal of Molecular Sciences, 24(3), 1880. https://doi.org/10.3390/ijms24031880






