Role of Histone Tails and Single Strand DNA Breaks in Nucleosomal Arrest of RNA Polymerase
Abstract
:1. Introduction
2. Results
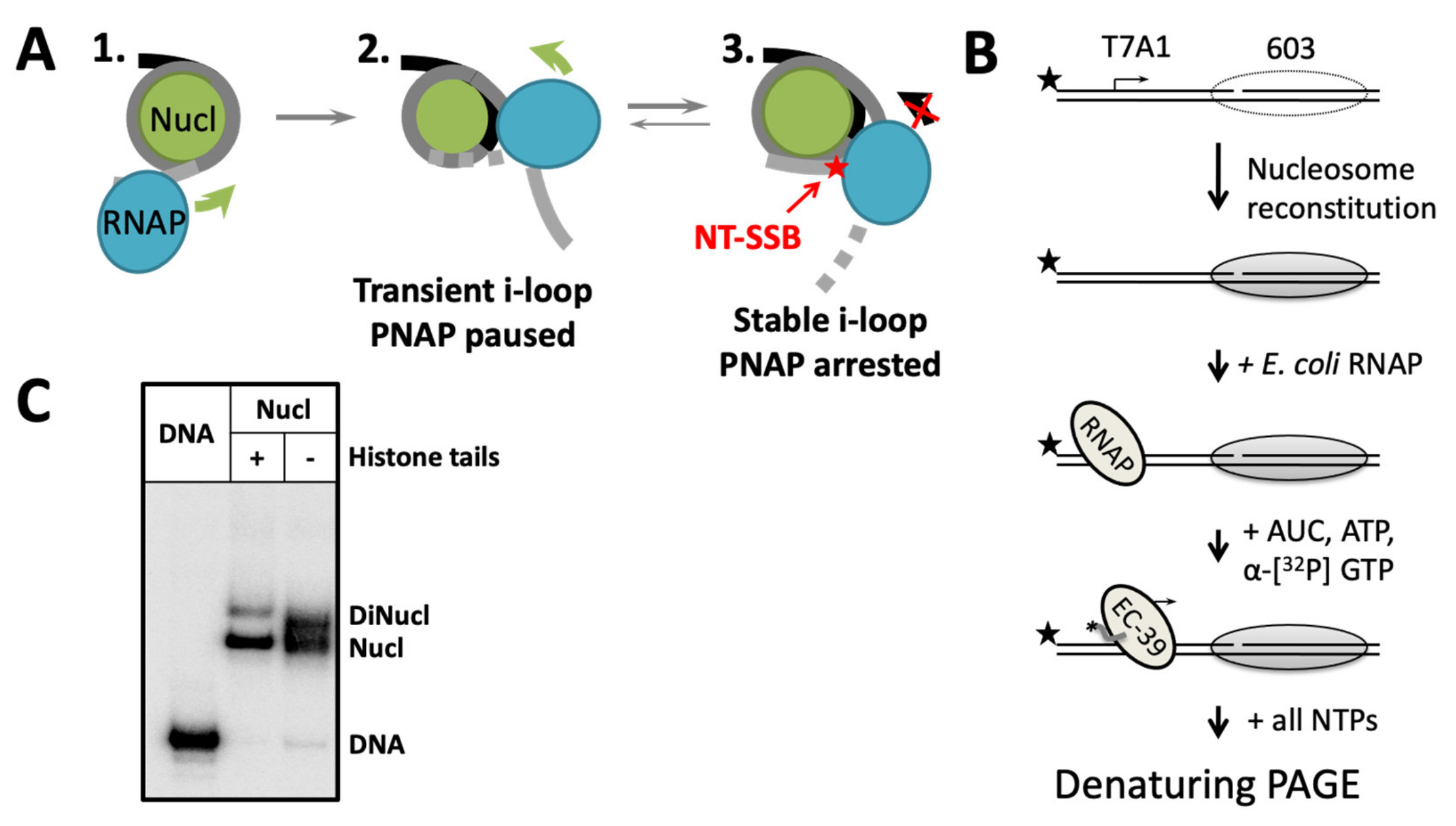
2.1. Experimental Approach
2.2. NT-SSB at the Position +2 Induces Arrest of RNAP at the +(10–20) Region
2.3. Extranuclesomal NT-SSBs Can Induce Nucleosome-Specific Arrest of RNAP
2.4. Histone Tails Increase the Efficiency of the Early Nucleosomal Pausing/Arrest
2.5. Histone Tails Contribute to Protection of Nucleosomal and Linker DNA from Hydroxyl Radicals
2.6. Formation of an i-loop Containing Backtracked RNAP at the Position +1
3. Discussion
4. Materials and Methods
4.1. Preparation of DNA Templates
4.1.1. Preparation of Templates with Single Nucleotide Gaps at the Positions −10, −11 or −12
4.1.2. Preparation of Template with Nick at Position +2
4.1.3. Preparation of Template to Obtain EC+2
4.2. Purification of Proteins and Trypsin Cleavage of the Donor Chromatin
4.3. Nucleosome Assembly
4.4. Transcription
4.5. Hydroxyl Radical Footprinting
4.6. Analysis of RNA Products and DNA Footprinting Probes
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tubbs, A.; Nussenzweig, A. Endogenous DNA Damage as a Source of Genomic Instability in Cancer. Cell 2017, 168, 644–656. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lindahl, T. Instability and decay of the primary structure of DNA. Nature 1993, 362, 709–715. [Google Scholar] [CrossRef] [PubMed]
- Pogozelski, W.K.; Tullius, T.D. Oxidative Strand Scission of Nucleic Acids: Routes Initiated by Hydrogen Abstraction from the Sugar Moiety. Chem. Rev. 1998, 98, 1089–1108. [Google Scholar] [CrossRef] [PubMed]
- Caldecott, K.W. DNA single-strand break repair and human genetic disease. Trends Cell Biol 2022, 32, 733–745. [Google Scholar] [CrossRef] [PubMed]
- Caldecott, K.W.; Ward, M.E.; Nussenzweig, A. The threat of programmed DNA damage to neuronal genome integrity and plasticity. Nat. Genet. 2022, 54, 115–120. [Google Scholar] [CrossRef] [PubMed]
- McKinnon, P.J.; Caldecott, K.W. DNA strand break repair and human genetic disease. Annu. Rev. Genomics Hum. Genet. 2007, 8, 37–55. [Google Scholar] [CrossRef]
- McKinnon, P.J. DNA repair deficiency and neurological disease. Nat. Rev. Neurosci. 2009, 10, 100–112. [Google Scholar] [CrossRef] [Green Version]
- Reynolds, J.J.; Stewart, G.S. A nervous predisposition to unrepaired DNA double strand breaks. DNA Repair. 2013, 12, 588–599. [Google Scholar] [CrossRef]
- Welch, G.; Tsai, L.H. Mechanisms of DNA damage-mediated neurotoxicity in neurodegenerative disease. EMBO Rep. 2022, 23, e54217. [Google Scholar] [CrossRef]
- Lin, Y.; Raj, J.; Li, J.; Ha, A.; Hossain, M.A.; Richardson, C.; Mukherjee, P.; Yan, S. APE1 senses DNA single-strand breaks for repair and signaling. Nucleic Acids Res. 2020, 48, 1925–1940. [Google Scholar] [CrossRef]
- Caldecott, K.W. Single-strand break repair and genetic disease. Nat. Rev. Genet. 2008, 9, 619–631. [Google Scholar] [CrossRef] [PubMed]
- Weinfeld, M.; Chaudhry, M.A.; D’Amours, D.; Pelletier, J.D.; Poirier, G.G.; Povirk, L.F.; Lees-Miller, S.P. Interaction of DNA-dependent protein kinase and poly(ADP-ribose) polymerase with radiation-induced DNA strand breaks. Radiat. Res. 1997, 148, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Le Cam, E.; Fack, F.; Menissier-de Murcia, J.; Cognet, J.A.; Barbin, A.; Sarantoglou, V.; Revet, B.; Delain, E.; de Murcia, G. Conformational analysis of a 139 base-pair DNA fragment containing a single-stranded break and its interaction with human poly(ADP-ribose) polymerase. J. Mol. Biol. 1994, 235, 1062–1071. [Google Scholar] [CrossRef]
- Caldecott, K.W. DNA single-strand break repair. Exp. Cell Res. 2014, 329, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Caldecott, K.W. Protein ADP-ribosylation and the cellular response to DNA strand breaks. DNA Repair. 2014, 19, 108–113. [Google Scholar] [CrossRef] [PubMed]
- Iyama, T.; Wilson, D.M. DNA repair mechanisms in dividing and non-dividing cells. DNA Repair. 2013, 12, 620–636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davis, L.; Maizels, N. Homology-directed repair of DNA nicks via pathways distinct from canonical double-strand break repair. Proc. Natl. Acad. Sci. USA 2014, 111, E924–E932. [Google Scholar] [CrossRef] [Green Version]
- Luger, K.; Mäder, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef]
- Biechele-Speziale, D.J.; Sutton, T.B.; Delaney, S. Obstacles and opportunities for base excision repair in chromatin. DNA Repair. 2022, 116, 103345. [Google Scholar] [CrossRef]
- Zhou, W.; Doetsch, P.W. Effects of abasic sites and DNA single-strand breaks on prokaryotic RNA polymerases. Proc. Natl. Acad. Sci. USA 1993, 90, 6601–6605. [Google Scholar] [CrossRef]
- Kathe, S.D.; Shen, G.P.; Wallace, S.S. Single-stranded breaks in DNA but not oxidative DNA base damages block transcriptional elongation by RNA polymerase II in HeLa cell nuclear extracts. J. Biol. Chem. 2004, 279, 18511–18520. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Neil, A.J.; Belotserkovskii, B.P.; Hanawalt, P.C. Transcription blockage by bulky end termini at single-strand breaks in the DNA template: Differential effects of 5′ and 3′ adducts. Biochemistry 2012, 51, 8964–8970. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, S.; Smerdon, M.J. Dissecting transcription-coupled and global genomic repair in the chromatin of yeast GAL1-10 genes. J. Biol. Chem. 2004, 279, 14418–14426. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Belotserkovskii, B.P.; Neil, A.J.; Saleh, S.S.; Shin, J.H.; Mirkin, S.M.; Hanawalt, P.C. Transcription blockage by homopurine DNA sequences: Role of sequence composition and single-strand breaks. Nucleic Acids Res. 2013, 41, 1817–1828. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bielas, J.H. Non-transcribed strand repair revealed in quiescent cells. Mutagenesis 2006, 21, 49–53. [Google Scholar] [CrossRef]
- Wellinger, R.E.; Thoma, F. Nucleosome structure and positioning modulate nucleotide excision repair in the non-transcribed strand of an active gene. EMBO J. 1997, 16, 5046–5056. [Google Scholar] [CrossRef] [Green Version]
- Pestov, N.A.; Gerasimova, N.S.; Kulaeva, O.I.; Studitsky, V.M. Structure of transcribed chromatin is a sensor of DNA damage. Sci. Adv. 2015, 1, e1500021. [Google Scholar] [CrossRef] [Green Version]
- Gerasimova, N.; Akhtar, M.; Student, V. Effect of single-strand DNA breaks on transcription of nucleosomes. Biology 2022, 77, 241–247. [Google Scholar] [CrossRef]
- Gerasimova, N.S.; Pestov, N.A.; Kulaeva, O.I.; Clark, D.J.; Studitsky, V.M. Transcription-induced DNA supercoiling: New roles of intranucleosomal DNA loops in DNA repair and transcription. Transcription 2016, 7, 91–95. [Google Scholar] [CrossRef] [Green Version]
- Gerasimova, N.S.; Volokh, O.I.; Pestov, N.A.; Armeev, G.A.; Kirpichnikov, M.P.; Shaytan, A.K.; Sokolova, O.S.; Studitsky, V.M. Structure of an Intranucleosomal DNA Loop That Senses DNA Damage during Transcription. Cells 2022, 11, 2678. [Google Scholar] [CrossRef]
- Kulaeva, O.I.; Gaykalova, D.A.; Pestov, N.A.; Golovastov, V.V.; Vassylyev, D.G.; Artsimovitch, I.; Studitsky, V.M. Mechanism of chromatin remodeling and recovery during passage of RNA polymerase II. Nat. Struct. Mol. Biol. 2009, 16, 1272–1278. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hartzog, G.A.; Speer, J.L.; Lindstrom, D.L. Transcript elongation on a nucleoprotein template. Biochim. Biophys. Acta 2002, 1577, 276–286. [Google Scholar] [CrossRef] [PubMed]
- Thastrom, A.; Lowary, P.T.; Widlund, H.R.; Cao, H.; Kubista, M.; Widom, J. Sequence motifs and free energies of selected natural and non-natural nucleosome positioning DNA sequences. J. Mol. Biol. 1999, 288, 213–229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lowary, P.T.; Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 1998, 276, 19–42. [Google Scholar] [CrossRef]
- Bondarenko, V.A.; Steele, L.M.; Ujvári, A.; Gaykalova, D.A.; Kulaeva, O.I.; Polikanov, Y.S.; Luse, D.S.; Studitsky, V.M. Nucleosomes can form a polar barrier to transcript elongation by RNA polymerase II. Mol. Cell 2006, 24, 469–479. [Google Scholar] [CrossRef]
- Chang, H.-W.; Kulaeva, O.I.; Shaytan, A.K.; Kibanov, M.; Kuznedelov, K.; Severinov, K.V.; Kirpichnikov, M.P.; Clark, D.J.; Studitsky, V.M. Analysis of the mechanism of nucleosome survival during transcription. Nucleic Acids Res. 2014, 42, 1619–1627. [Google Scholar] [CrossRef] [Green Version]
- Walter, W.; Kireeva, M.L.; Studitsky, V.M.; Kashlev, M. Bacterial polymerase and yeast polymerase II use similar mechanisms for transcription through nucleosomes. J. Biol. Chem. 2003, 278, 36148–36156. [Google Scholar] [CrossRef] [Green Version]
- Weber, C.M.; Ramachandran, S.; Henikoff, S. Nucleosomes are context-specific, H2A.Z-modulated barriers to RNA polymerase. Mol. Cell 2014, 53, 819–830. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.; Gabizon, R.; Brown, A.I.; Lee, A.; Song, A.; Diaz-Celis, C.; Kaplan, C.D.; Koslover, E.F.; Yao, T.; Bustamante, C. High-resolution and high-accuracy topographic and transcriptional maps of the nucleosome barrier. eLife 2019, 8, e48281. [Google Scholar] [CrossRef]
- Ujvári, A.; Hsieh, F.-K.; Luse, S.W.; Studitsky, V.M.; Luse, D.S. Histone N-terminal tails interfere with nucleosome traversal by RNA polymerase II. J. Biol. Chem. 2008, 283, 32236–32243. [Google Scholar] [CrossRef]
- Chang, H.W.; Feofanov, A.V.; Lyubitelev, A.V.; Armeev, G.A.; Kotova, E.Y.; Hsieh, F.K.; Kirpichnikov, M.P.; Shaytan, A.K.; Studitsky, V.M. N-Terminal Tails of Histones H2A and H2B Differentially Affect Transcription by RNA Polymerase II in Vitro. Cells 2022, 11, 2475. [Google Scholar] [CrossRef] [PubMed]
- Diaz, B.M.; Walker, I.O. Trypsin digestion of core chromatin. Biosci. Rep. 1983, 3, 283–292. [Google Scholar] [CrossRef] [PubMed]
- Gaykalova, D.A.; Kulaeva, O.I.; Volokh, O.; Shaytan, A.K.; Hsieh, F.K.; Kirpichnikov, M.P.; Sokolova, O.S.; Studitsky, V.M. Structural analysis of nucleosomal barrier to transcription. Proc. Natl. Acad. Sci. USA 2015, 112, E5787–E5795. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, H.; Ramachandran, S.; Fong, N.; Phang, T.; Lee, S.; Parsa, P.; Liu, X.; Harmacek, L.; Danhorn, T.; Song, T.; et al. JMJD5 couples with CDK9 to release the paused RNA polymerase II. Proc. Natl. Acad. Sci. USA 2020, 117, 19888–19895. [Google Scholar] [CrossRef] [PubMed]
- Kotova, E.Y.; Hsieh, F.K.; Chang, H.W.; Maluchenko, N.V.; Langelier, M.F.; Pascal, J.M.; Luse, D.S.; Feofanov, A.V.; Studitsky, V.M. Human PARP1 Facilitates Transcription through a Nucleosome and Histone Displacement by Pol II in Vitro. Int. J. Mol. Sci. 2022, 23, 7107. [Google Scholar] [CrossRef]
- Gibson, B.A.; Zhang, Y.; Jiang, H.; Hussey, K.M.; Shrimp, J.H.; Lin, H.; Schwede, F.; Yu, Y.; Kraus, W.L. Chemical genetic discovery of PARP targets reveals a role for PARP-1 in transcription elongation. Science 2016, 353, 45–50. [Google Scholar] [CrossRef] [Green Version]
- Kireeva, M.L.; Komissarova, N.; Waugh, D.S.; Kashlev, M. The 8-nucleotide-long RNA:DNA hybrid is a primary stability determinant of the RNA polymerase II elongation complex. J. Biol. Chem. 2000, 275, 6530–6536. [Google Scholar] [CrossRef] [Green Version]
- Artsimovitch, I.; Svetlov, V.; Murakami, K.S.; Landick, R. Co-overexpression of Escherichia coli RNA polymerase subunits allows isolation and analysis of mutant enzymes lacking lineage-specific sequence insertions. J. Biol. Chem. 2003, 278, 12344–12355. [Google Scholar] [CrossRef] [Green Version]
- Vassylyeva, M.N.; Svetlov, V.; Dearborn, A.D.; Klyuyev, S.; Artsimovitch, I.; Vassylyev, D.G. The carboxy-terminal coiled-coil of the RNA polymerase beta’-subunit is the main binding site for Gre factors. EMBO Rep. 2007, 8, 1038–1043. [Google Scholar] [CrossRef] [Green Version]
- Côté, J.; Utley, R.T.; Workman, J.L. Basic analysis of transcription factor binding to nucleosomes. In Methods in Molecular Genetics; Adolph, K.W., Ed.; Academic Press: Cambridge, MA, USA, 1995; Volume 6, pp. 108–128. [Google Scholar]
- Kireeva, M.L.; Walter, W.; Tchernajenko, V.; Bondarenko, V.; Kashlev, M.; Studitsky, V.M. Nucleosome remodeling induced by RNA polymerase II: Loss of the H2A/H2B dimer during transcription. Mol. Cell 2002, 9, 541–552. [Google Scholar] [CrossRef]
- Dixon, W.J.; Hayes, J.J.; Levin, J.R.; Weidner, M.F.; Dombroski, B.A.; Tullius, T.D. Hydroxyl radical footprinting. Methods Enzymol. 1991, 208, 380–413. [Google Scholar] [CrossRef] [PubMed]
- Gerasimova, N.S.; Studitsky, V.M. Hydroxyl radical footprinting of fluorescently labeled DNA. Mosc. Univ. Biol. Sci. Bull. 2016, 71, 93–96. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]






| Name | Oligonucleotide | Manufacturer |
|---|---|---|
| Forext603 | CCAACGCAGCCCAGTTCGCGCGCCC | IDT * |
| 282Revwhst | CGTCGGATCCGTACCCCAGGGACTT | IDT |
| s603-10 | AAGCCAACACCGGCACTGGGGCAACGCAGCCCAGTTCGCGCGCCCACCTA | IDT |
| s603-11 | AAGCCAACACCGGCACTGGGGCCAACGCAGCCCAGTTCGCGCGCCCACCT | IDT |
| s603-12 | AAGCCAACACCGGCACTGGGGCCCAACGCAGCCCAGTTCGCGCGCCCACC | IDT |
| T7A1up1 | GATCCCGAAAATTTATCAAAAAGAGTATTGACTTAAAGTCTAACCTATAGGATACTTACA | IDT |
| T7A1up2 | GCCATCGAGAGGGACACGGCGAAAAGCCAACACCGGCACTGG | IDT |
| T7A1dw1 | CTATAGGTTAGACTTTAAGTCAATACTCTTTTTGATAAATTTTCGGGATC | IDT |
| T7A1dw2 | GGTGTTGGCTTTTCGCCGTGTCCCTCTCGATGGCTGTAAGTATC | IDT |
| ROX-282Forwhst | ROX-CCGGGATCCAGATCCCGAAAATTTA | Lumiprobe ** |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gerasimova, N.S.; Pestov, N.A.; Studitsky, V.M. Role of Histone Tails and Single Strand DNA Breaks in Nucleosomal Arrest of RNA Polymerase. Int. J. Mol. Sci. 2023, 24, 2295. https://doi.org/10.3390/ijms24032295
Gerasimova NS, Pestov NA, Studitsky VM. Role of Histone Tails and Single Strand DNA Breaks in Nucleosomal Arrest of RNA Polymerase. International Journal of Molecular Sciences. 2023; 24(3):2295. https://doi.org/10.3390/ijms24032295
Chicago/Turabian StyleGerasimova, Nadezhda S., Nikolay A. Pestov, and Vasily M. Studitsky. 2023. "Role of Histone Tails and Single Strand DNA Breaks in Nucleosomal Arrest of RNA Polymerase" International Journal of Molecular Sciences 24, no. 3: 2295. https://doi.org/10.3390/ijms24032295





