Investigation of the Key Genes Associated with Anthocyanin Accumulation during Inner Leaf Reddening in Ornamental Kale (Brassica oleracea L. var. acephala)
Abstract
:1. Introduction
2. Results
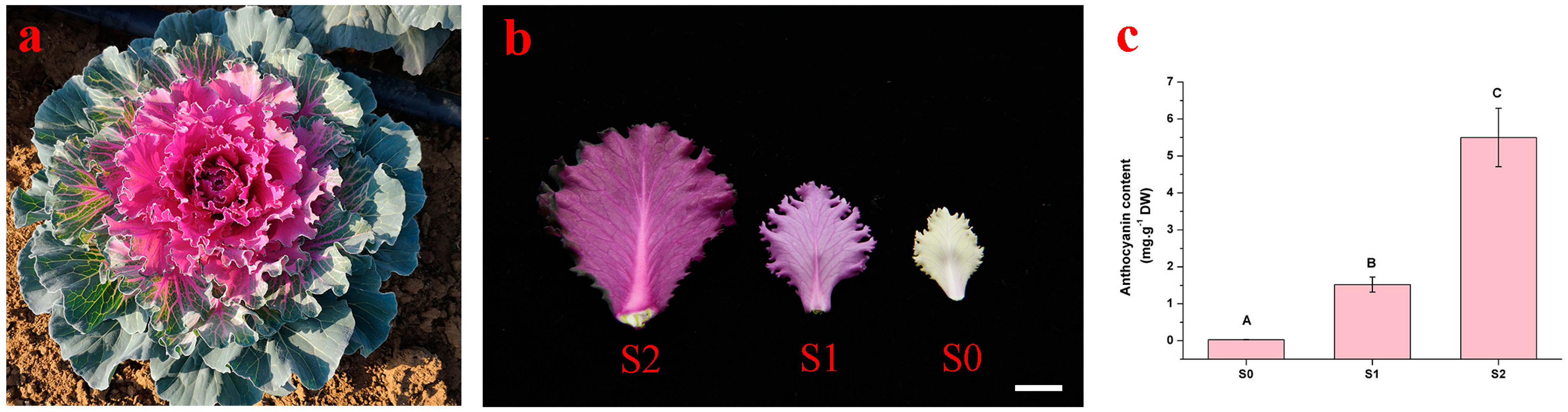
2.1. Anthocyanin Levels in the New Leaves of Red-Leaf Kale at Three Developmental Stages
2.2. Anthocyanin Metabolite Analysis
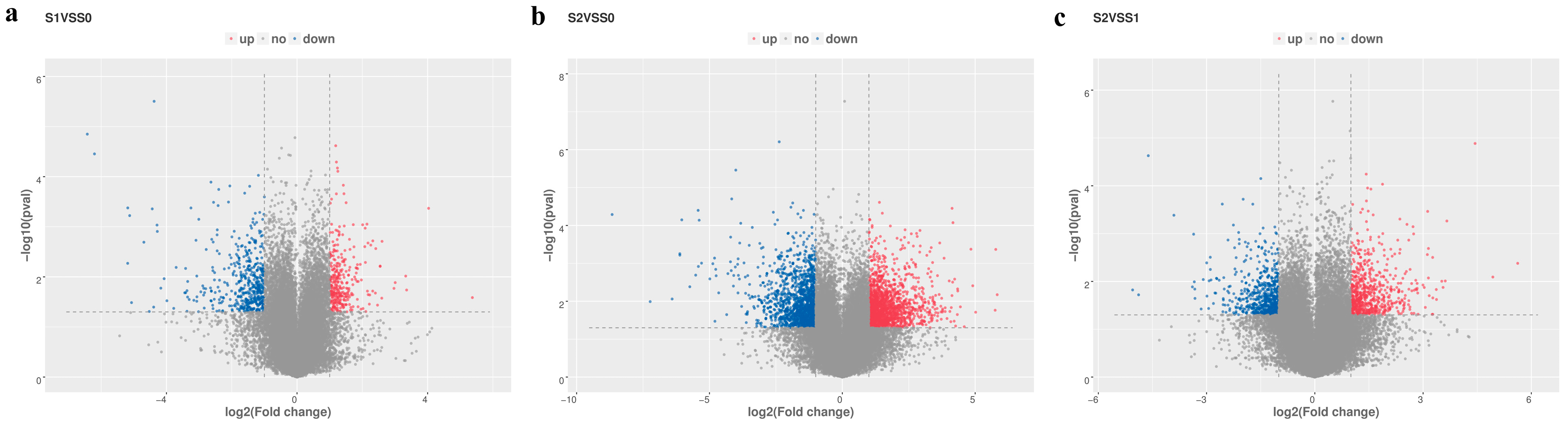
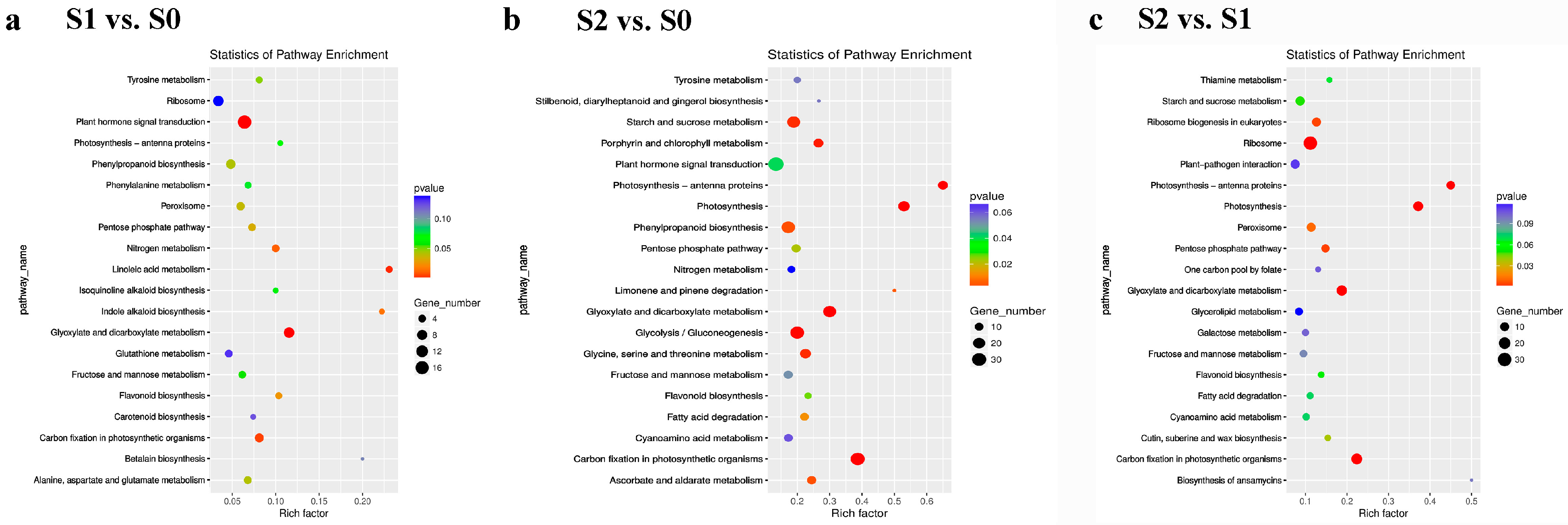
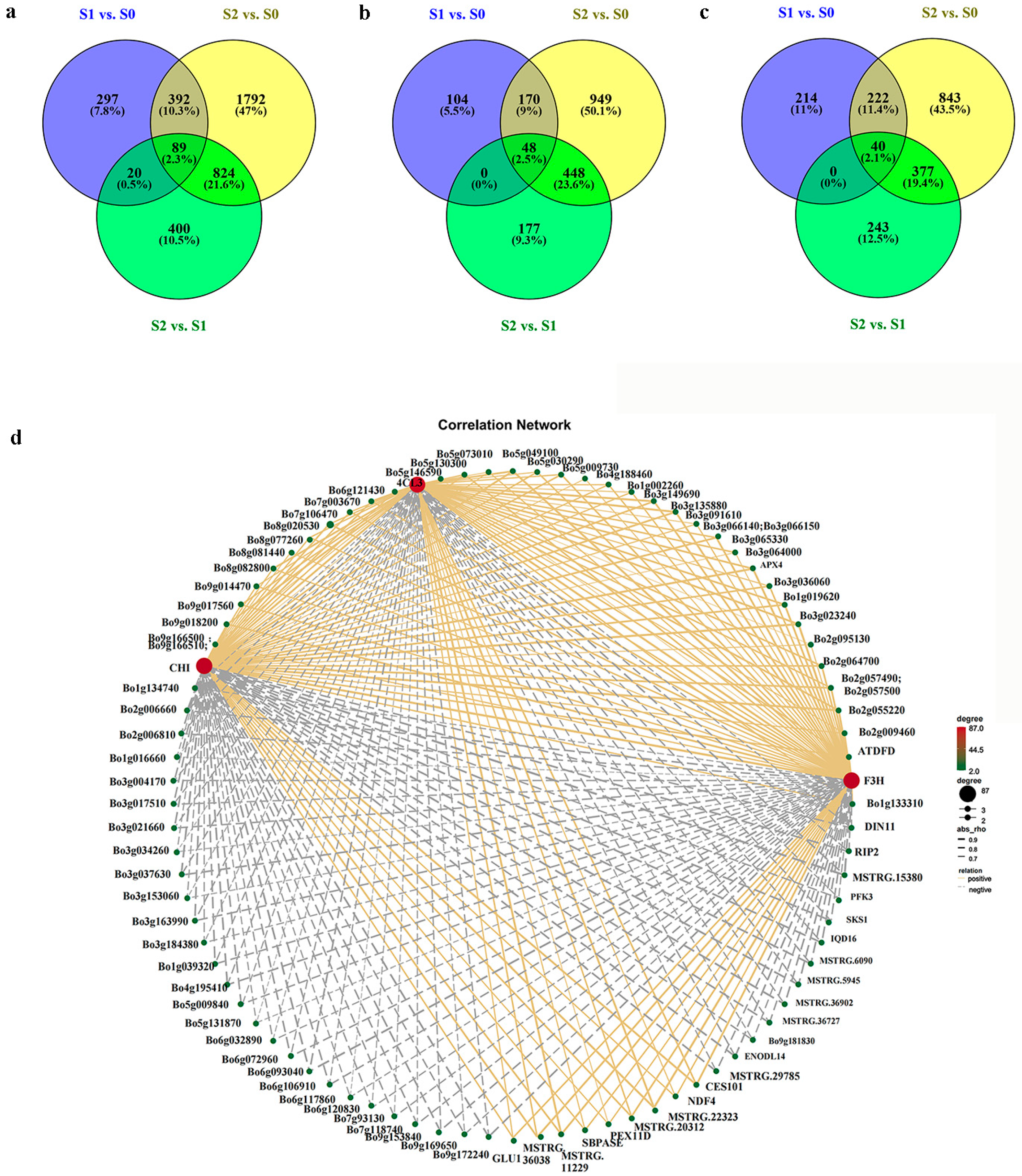
2.3. Library Construction and DEG Analysis
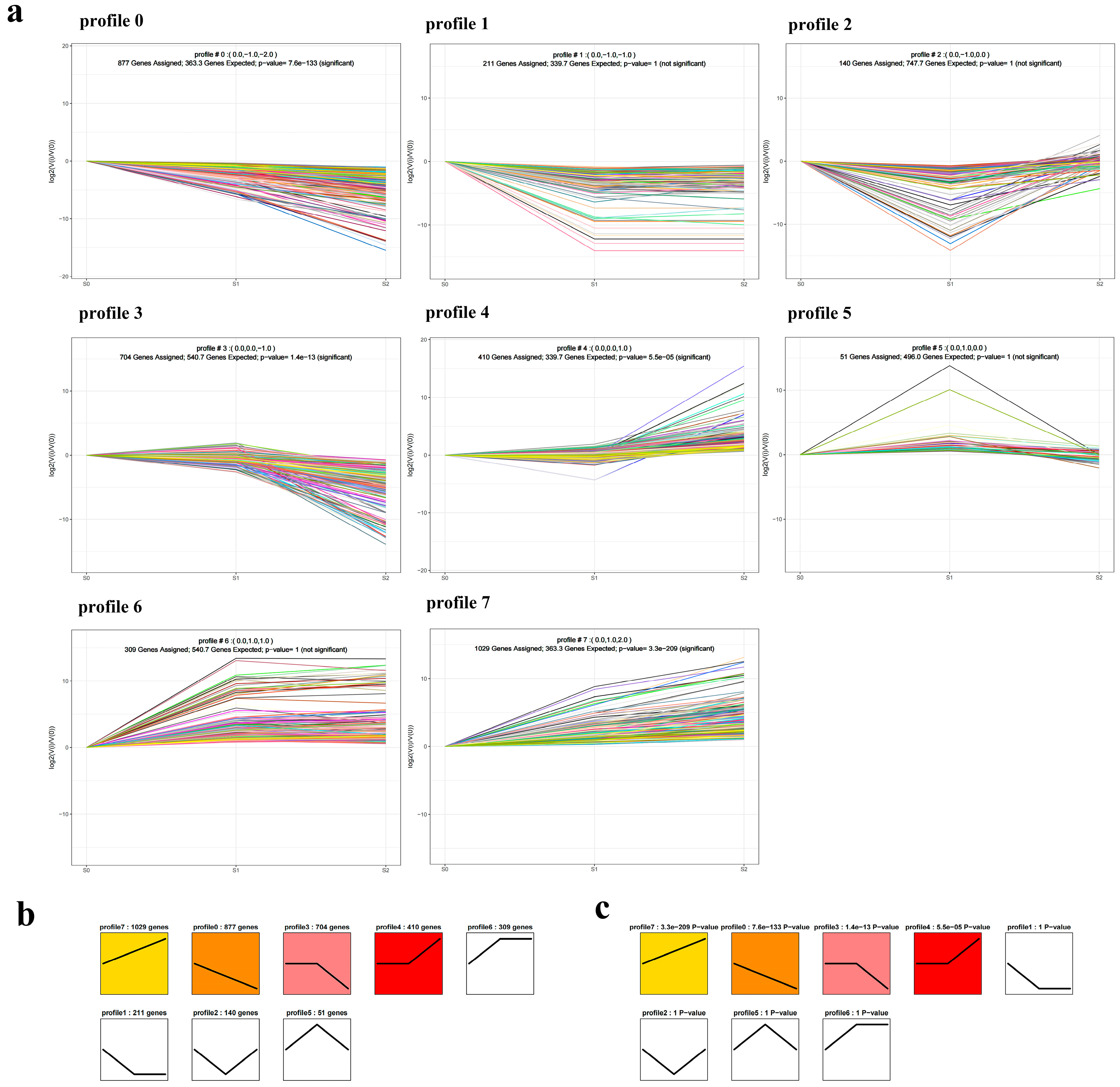
2.4. Trend Analysis
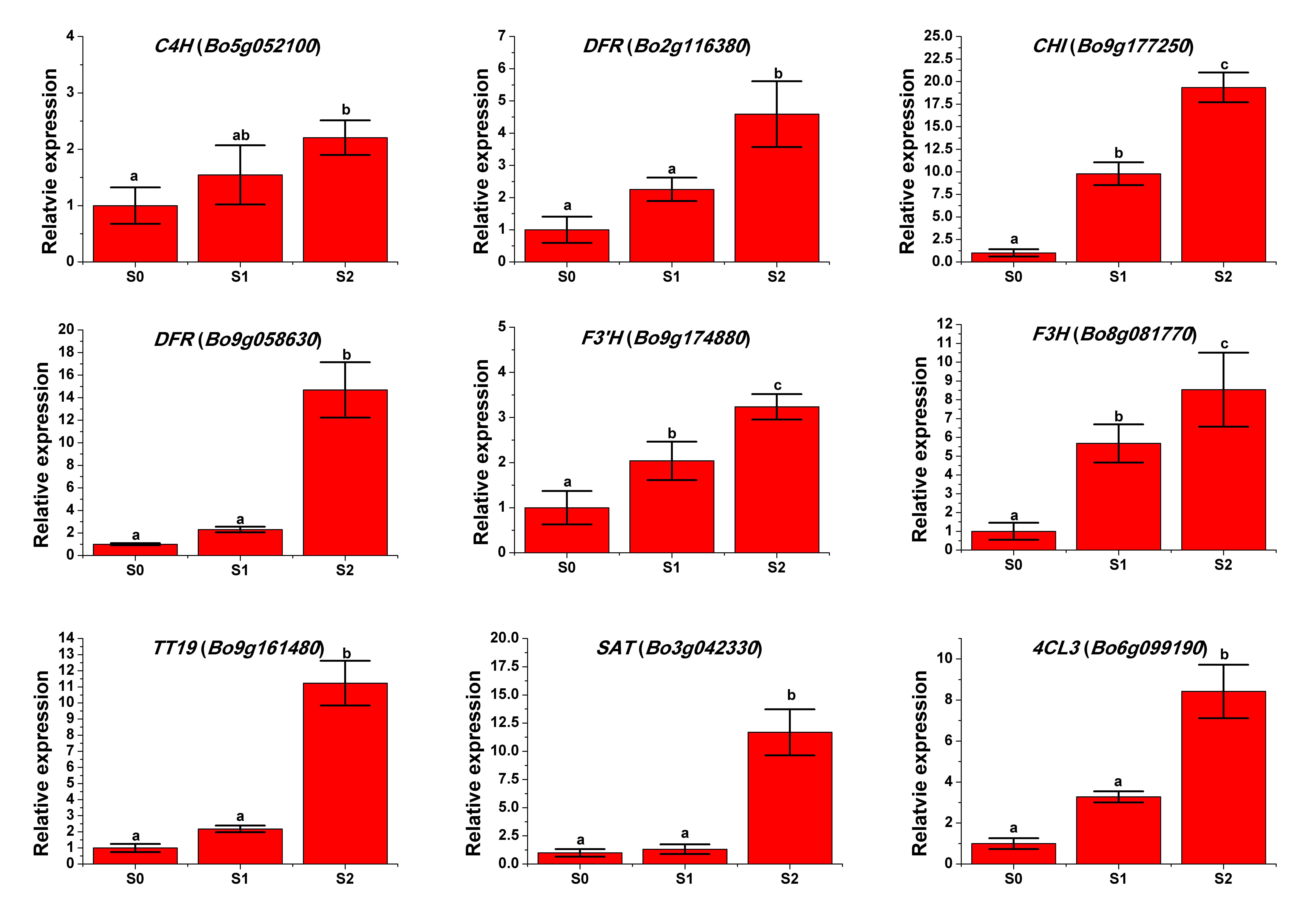
2.5. qRT-PCR to Determine Gene Expression Patterns
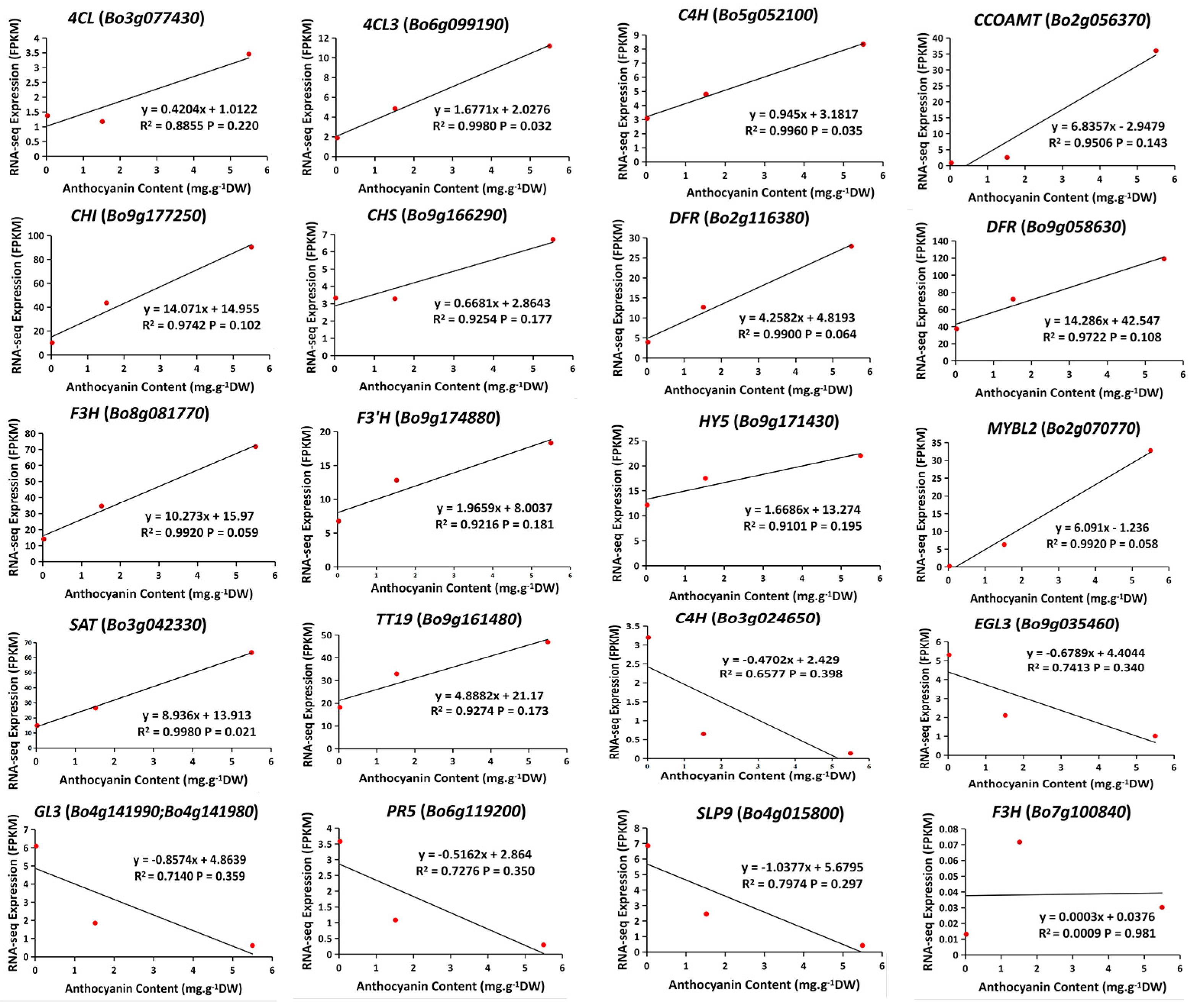
2.6. Anthocyanin Biosynthesis Genes at S0–S2
2.7. Identification of Differentially Co-Expressed Genes
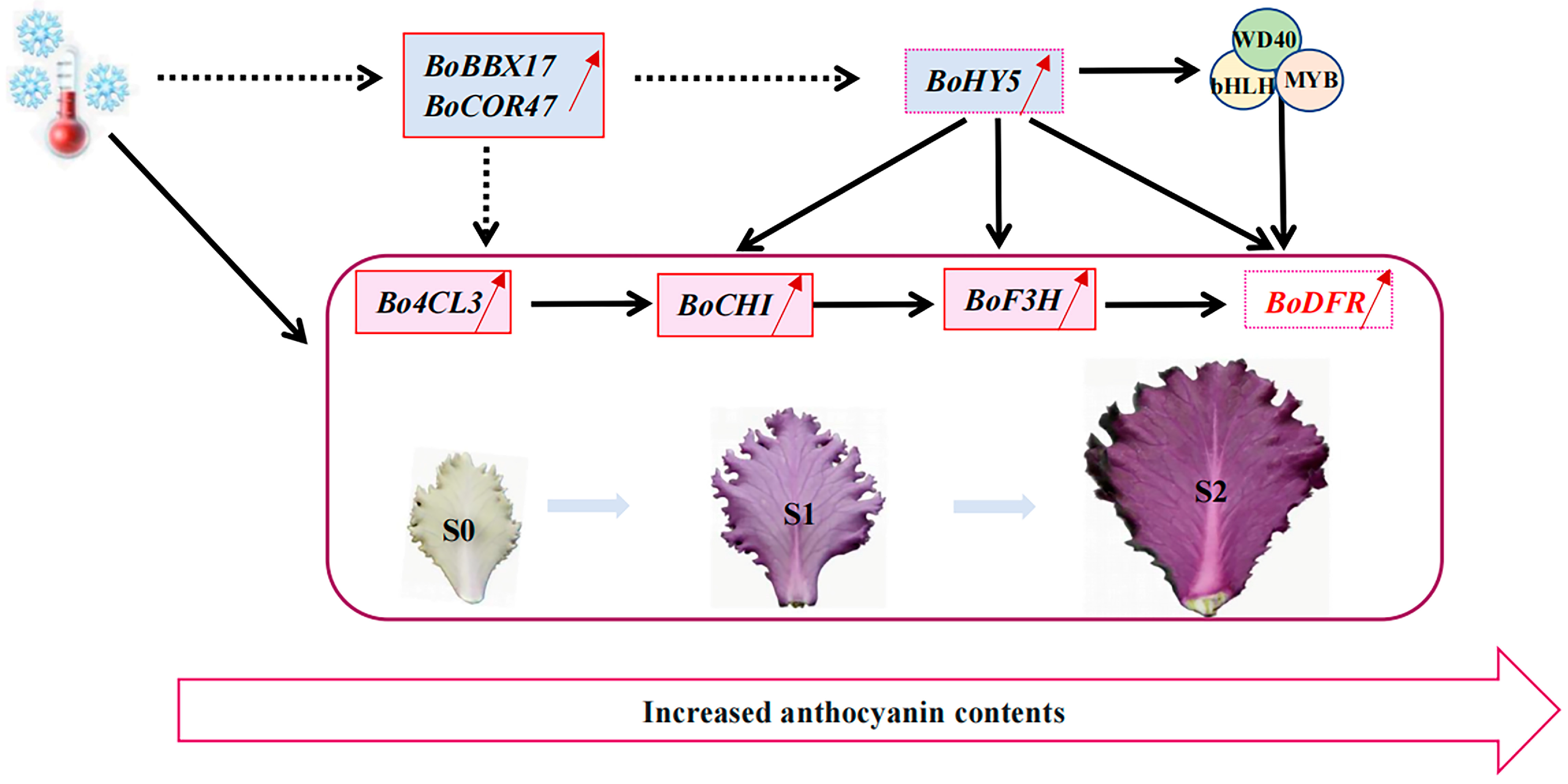
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Measurement of Anthocyanin Content
4.3. Anthocyanins Extraction and Multiple Reaction Monitoring
4.4. Identification of Anthocyanin Biosynthesis Genes in Brassica Oleracea
4.5. RNA Extraction and Library Construction
4.6. Read Mapping onto the Reference Genome
4.7. Differential Expression Analysis
4.8. Trend Analysis and Correlation Analysis
4.9. qRT-PCR
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hughes, N.M. Winter leaf reddening in ‘evergreen’ species. New Phytol. 2011, 190, 573–581. [Google Scholar] [CrossRef]
- Hughes, N.M.; Carpenter, K.L.; Cannon, J.G. Estimating contribution of anthocyanin pigments to osmotic adjustment during winter leaf reddening. J. Plant Physiol. 2012, 170, 230–233. [Google Scholar] [CrossRef]
- Zhang, S.M.; Gao, H.; Cai, L.X.; Jin, C.X.; Cui, L.H. Study on growing techniques of ornamental kale’s introduction. Agric. Sci. J. Yanbian Univ. 1998, 2, 67–69. [Google Scholar]
- Li, Y.C.; Wang, J.L. Discussion on cultivation management and landscape application of collard. Mod. Landsc. Archit. 2010, 12, 63–64. [Google Scholar]
- Ren, J.; Fu, W.; Du, J.T.; Hou, A.L.; Liu, Z.Y.; Feng, H. Identification of a candidate gene for Re, the factor determining the red leaf phenotype in ornamental kale using fine mapping and transcriptome analysis. Plant Breed. 2017, 136, 738–748. [Google Scholar] [CrossRef]
- Ren, J.; Liu, Z.Y.; Niu, R.Q.; Feng, H. Mapping of Re, a gene conferring the red leaf trait in ornamental kale (Brassica oleracea L. var. acephala). Plant Breed. 2015, 134, 494–500. [Google Scholar] [CrossRef]
- Field, T.S.; Lee, D.W.; Holbrook, N.M. Why leaves turn red in autumn. The role of anthocyanins in senescing leaves of redosier dogwood. Plant Physiol. 2001, 127, 566–574. [Google Scholar] [CrossRef]
- Honda, T.; Saito, N. ChemInform abstract: Recent progress in the chemistry of poly acylated anthocyanins as flower color pigments. ChemInform 2002, 33, 566–574. [Google Scholar] [CrossRef]
- Tena, N.; Martin, J.; Asuero, A.G. State of the art of anthocyanins: Antioxidant activity, sources, bioavailability, and therapeutic effect in human health. Antioxidants 2020, 9, 451. [Google Scholar] [CrossRef] [PubMed]
- Koes, R.; Verweij, W.; Quattrocchio, F. Flavonoids: A colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci. 2005, 10, 236–242. [Google Scholar] [CrossRef]
- Iwashina, T. Contribution to flower colors of flavonoids including anthocyanins: A review. Nat. Prod. Commun. 2015, 10, 529–544. [Google Scholar] [CrossRef] [PubMed]
- Ramsay, N.A.; Glover, B.J. MYB-bHLH-WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci. 2005, 10, 63–70. [Google Scholar] [CrossRef]
- Yang, Y.N.; Yao, G.F.; Yue, W.Q.; Zhang, S.L.; Wu, J. Transcriptome profiling reveals differential gene expression in proanthocyanidin biosynthesis associated with red/green skin color mutant of pear (Pyrus communis L.). Front. Plant Sci. 2015, 6, 795. [Google Scholar] [CrossRef]
- Chalker-Scott, L. Environmental significance of anthocyanins in plant stress responses. Photochem Photobiol. 1999, 70, 1–9. [Google Scholar] [CrossRef]
- Hughes, N.M.; Reinhardt, K.; Field, T.S.; Gerardi, A.R.; Smith, W.K. Association between winter anthocyanin production and drought stress in angiosperm evergreen species. J. Exp. Bot. 2010, 61, 1699–1709. [Google Scholar] [CrossRef] [PubMed]
- An, J.-P.; Wang, X.-F.; Zhang, X.-W.; Xu, H.-F.; Bi, S.-Q.; You, C.-X.; Hao, Y.-J. An apple MYB transcription factor regulates cold tolerance and anthocyanin accumulation and undergoes MIEL1-mediated degradation. Plant Biotechnol. J. 2020, 18, 337–353. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.; He, J.X. Protective role of anthocyanins in plants under low nitrogen stress. Biochem. Biophys. Res. Commun. 2018, 498, 946–953. [Google Scholar] [CrossRef]
- Mbarki, S.; Sytar, O.; Zivcak, M.; Abdelly, C.; Cerda, A.; Brestic, M. Anthocyanins of coloured wheat genotypes in specific response to salstress. Molecules 2018, 23, 1518. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.L.; Zhai, J.J.; Shao, L.; Lin, W.; Peng, C.L. Accumulation of anthocyanins: An adaptation strategy of Mikania micrantha to low temperature in winter. Front. Plant Sci. 2019, 10, 1049. [Google Scholar] [CrossRef] [PubMed]
- Castañeda-Ovando, A.; Pacheco-Hernández, M.D.L.; Páez-Hernández, M.E.; Rodríguez, J.A.; Galán-Vidal, C.A. Chemical studies of anthocyanins: A review. Food Chem. 2009, 113, 859–871. [Google Scholar] [CrossRef]
- Jeon, J.; Kim, J.K.; Kim, H.; Kim, Y.J.; Park, Y.J.; Kim, S.J.; Kim, C.; Park, S.U. Transcriptome analysis and metabolic profiling of green and red kale (Brassica oleracea var. acephala) seedlings. Food Chem. 2018, 241, 7–13. [Google Scholar] [CrossRef] [PubMed]
- He, Q.; Zhang, Z.F.; Zhang, L.G. Anthocyanin accumulation, antioxidant ability and stability, and a transcriptional analysis of anthocyanin biosynthesis in purple heading Chinese cabbage (Brassica rapa L. ssp. pekinensis). J. Agric. Food Chem. 2016, 64, 132–145. [Google Scholar] [CrossRef] [PubMed]
- Olsen, H.; Aaby, K.; Borge, G.I. Characterization, quantification, and yearly variation of the naturally occurring polyphenols in a common red variety of curly kale (Brassica oleracea L. convar. acephala var. sabellica cv. ‘Redbor’). J. Agric. Food Chem. 2010, 58, 11346–11354. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.Q.; Zheng, W.W.; Zang, Y.X.; Shao, Y.F.; Zhu, Z.J. Studies on change of pigment contents during different developmental stages among several flowering kale cultivars. Chin. Agric. Sci. Bull. 2013, 29, 54–161. [Google Scholar]
- Zhou, H.; Lin-Wang, K.; Wang, H.; Gu, C.; Dare, A.P.; Espley, R.V.; He, H.; Allan, A.C.; Han, Y. Molecular genetics of blood-fleshed peach reveals activation of anthocyanin biosynthesis by NAC transcription factors. Plant J. 2015, 82, 105–121. [Google Scholar] [CrossRef]
- Liu, Y.Q.; Jiang, T.; Zhao, X.E.; Yang, G.D.; Du, J.H.; Wang, X. Analysis of anthocyanins in Brassica oleracea L. var. capitata L. and Brassica oleracea var. acephala D.C. by high performance liquid chromatography-electrospray ionization/quadrupole-time of flight-mass/mass spectrometry. Chin. J. Anal. Chem. 2011, 39, 419–424. [Google Scholar]
- Guo, N.; Han, S.; Zong, M.; Wang, G.X.; Zheng, S.N.; Liu, F. Identification and differential expression analysis of anthocyanin biosynthetic genes in leaf color variants of ornamental kale. BMC Genom. 2019, 20, 564. [Google Scholar] [CrossRef]
- Gu, W.H.; Zheng, H.J.; Zhang, Y.; Liu, Z.Y. A preliminary study on selection and breeding of new lines and main genetic characteristics of ornamental kale. J. Shanghai Jiaotong Univ. Agric. Sci. 2002, 20, 129–132. [Google Scholar]
- Xie, L.N. Genetic Analysis of Leaf Color and Shape and Mechanism of Self-Incompatibility of Brassica oleracea var. acephalea. Master’s Thesis, Northeast Forestry University, Harbin, China, 2003. [Google Scholar]
- Wang, Y.S. Isolated Microspore Culture and Identification of Markers Linked to Red Leaf Traits in Kale. Ph.D. Thesis, Shenyang Agricultural University, Shenyang, China, 2011. [Google Scholar]
- Zhu, P.F. Breeding and Cultivation Techniques of Ornamental Kale, 1st ed.; Liaoning Science and Technology Publishing House: Shenyang, China, 2014; pp. 10–11. [Google Scholar]
- Liu, X.-P.; Gao, B.-Z.; Han, F.-Q.; Fang, Z.-Y.; Yang, L.-M.; Zhuang, M.; Lv, H.-H.; Liu, Y.-M.; Li, Z.-S.; Cai, C.-C.; et al. Genetics and fine mapping of a purple leaf gene, BoPr, in ornamental kale (Brassica oleracea L. var. acephala). BMC Genom. 2017, 18, 230. [Google Scholar] [CrossRef]
- Feng, X.; Zhang, Y.T.; Wang, H.; Tian, Z.D.; Xin, S.Y.; Zhu, P.F. The dihydroflavonol 4-reductase BoDFR1 drives anthocyanin accumulation in pink-leaved ornamental kale. Theor. Appl. Genet. 2021, 134, 159–169. [Google Scholar] [CrossRef]
- Yan, C.; An, G.; Zhu, T.; Zhang, W.; Zhang, L.; Peng, L.; Chen, J.; Kuang, H. Independent activation of the BoMYB2 gene leading to purple traits in Brassica oleracea. Theor. Appl. Genet. 2019, 132, 895–906. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.W.; Rahim, M.A.; Afrin, K.S.; Park, J.I.; Kang, J.G.; Nou, I.S. Transcriptome profiling of two contrasting ornamental cabbage (Brassica oleracea var. acephala) lines provides insights into purple and white inner leaf pigmentation. BMC Genom. 2018, 19, 797. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Ohmiya, A. Seeing is believing: Engineering anthocyanin and carotenoid biosynthetic pathways. Curr. Opin. Biotechnol. 2008, 19, 190–197. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.Y.; Li, C.Y.; Hwang, I.F. Characterisation of the pigment components in red cabbage (Brassica oleracea L. var.) juice and their anti-inflammatory effects on LPS-stimulated murine splenocytes. Food Chem. 2008, 109, 771–781. [Google Scholar] [CrossRef]
- McDougall, G.J.; Fyffe, S.; Dobson, P.; Stewart, D. Anthocyanins from red cabbage--stability to simulated gastrointestinal digestion. Phytochemistry 2007, 68, 1285–1294. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.-F.; Zhao, D.-H.; Zhang, J.-Q.; Chen, J.-S.; Li, J.-L.; Weng, Z.; Rong, L.-P. De novo transcriptome sequencing and anthocyanin metabolite analysis reveals leaf color of Acer pseudosieboldianum in autumn. BMC Genom. 2021, 22, 383. [Google Scholar] [CrossRef] [PubMed]
- Quattrocchio, F.; Wing, J.F.; Leppen, H.T.C.; Mol, J.N.M.; Koes, R.E. Regulatory genes controlling anthocyanin pigmentation are functionally conserved among plant species and have distinct sets of target genes. Plant Cell 1993, 5, 1497–1512. [Google Scholar] [CrossRef]
- Nabavi, S.M.; Šamec, D.; Tomczyk, M.; Milella, L.; Russo, D.; Habtemariam, S.; Suntar, I.; Rastrelli, L.; Daglia, M.; Xiao, J.; et al. Flavonoid biosynthetic pathways in plants: Versatile targets for metabolic engineering. Biotechnol. Adv. 2020, 38, 107316. [Google Scholar] [CrossRef]
- Cheng, F.; Wu, J.; Wang, X.W. Genome triplication drove the diversification of Brassica plants. Hortic. Res. 2014, 1, 14024. [Google Scholar] [CrossRef]
- Li, Y.; Kim, J.I.; Pysh, L.; Chapple, C. Four isoforms of Arabidopsis 4-coumarate: CoA ligase have overlapping yet distinct Roles in phenylpropanoid metabolism. Plant Physiol. 2015, 169, 2409–2421. [Google Scholar] [CrossRef]
- Sun, Y.Y.; Wang, J.L.; Qiu, Y.; Liu, T.J.; Song, J.P.; Li, X.X. Identification of ‘Xinlimei’ radish candidate genes associated with anthocyanin biosynthesis based on a transcriptome analysis. Gene 2018, 657, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Raza, A.; Su, W.; Hussain, M.A.; Mehmood, S.S.; Zhang, X.; Cheng, Y.; Zou, X.; Lv, Y. Integrated analysis of metabolome and transcriptome reveals insights for cold tolerance in rapeseed (Brassica napus L.). Front. Plant Sci. 2021, 12, 721681. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, J.; Xia, R.X. Expression of chalcone synthase and chalcone isomerase genes and accumulation of corresponding flavonoids during fruit maturation of Guoqing No. 4 satsuma mandarin (Citrus unshiu Marcow). Sci. Hortic. 2010, 125, 110–116. [Google Scholar] [CrossRef]
- Zhou, X.W.; Li, J.Y.; Fan, Z.Q. Cloning and sequence analysis of chalcone synthase gene cDNA from Camellia nitidissima. For. Res. 2011, 21, 58–64. [Google Scholar]
- Zhao, D.Q.; Tao, J.; Han, C.X.; Ge, J.T. Flower color diversity revealed by differential expression of flavonoid biosynthetic genes and flavonoid accumulation in herbaceous peony (Paeonia lactiflora Pall.). Mol. Biol. Rep. 2012, 39, 11263–11275. [Google Scholar] [CrossRef]
- Zhou, Y.H.; Zhou, J.; Huang, L.F.; Ding, X.T.; Shi, K.; Yu, J.Q. Grafting of Cucumis sativus onto Cucurbita ficifolia leads to improved plant growth, increased light utilization and reduced accumulation of reactive oxygen species in chilled plants. J. Plant Res. 2009, 122, 529–540. [Google Scholar] [CrossRef]
- Wu, Y.Q.; Liu, L.; Zhao, D.Q.; Tao, J. Age-associated methylation change of CHI promoter in herbaceous peony (Paeonia lactiflora Pall.). Biosci Rep. 2018, 38, BSR20180482. [Google Scholar] [CrossRef]
- Forkmann, G.; Heller, W.; Grisebach, H. Anthocyanin Biosynthesis in Flowers of Matthiola incana Flavanone 3-and Flavonoid 3′-Hydroxylases. Z. Für Nat. C 1980, 35, 691–695. [Google Scholar] [CrossRef]
- Wisman, E.; Hartmann, U.; Sagasser, M.; Baumann, E.; Palme, K.; Hahlbrock, K.; Saedler, H.; Weisshaar, B. Knock-out mutants from an En-1 mutagenized Arabidopsis thaliana population generate phenylpropanoid biosynthesis phenotypes. Proc. Natl. Acad. Sci. USA 1998, 95, 12432–12437. [Google Scholar] [CrossRef]
- Armitage, A.; Carlson, W. The effect of quantum flux density, day and night temperature and phosphorus and potassium status on anthocyanin and chlorophyll content in marigold leaves [Tagetes patula]. J. Am. Soc. Horticult. Sci. 1981, 106, 639–642. [Google Scholar] [CrossRef]
- Lo Piero, A.R.; Puglisi, I.; Rapisarda, P.; Petrone, G. Anthocyanins accumulation and related gene expression in red orange fruit induced by low temperature storage. J. Agric. Food Chem. 2005, 53, 9083–9088. [Google Scholar] [CrossRef] [PubMed]
- Naing, A.H.; Ai, T.N.; Lim, K.B.; Lee, I.J.; Kim, C.K. Overexpression of rosea1 from snapdragon enhances anthocyanin accumulation and abiotic stress tolerance in transgenic tobacco. Front. Plant Sci. 2018, 9, 1070. [Google Scholar] [CrossRef]
- Zhang, Y.Q.; Zheng, S.; Liu, Z.J.; Wang, L.G.; Bi, Y.R. Both HY5 and HYH are necessary regulators for low temperature-induced anthocyanin accumulation in Arabidopsis seedlings. J. Plant Physiol. 2011, 168, 367–374. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.W.; Ward, R.W.; Thomashow, M.F. Characterization of a cold-regulated wheat gene related to Arabidopsis cor47. Plant Physiol. 1992, 100, 915–922. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Sánchez, I.E.; Maruri-López, I.; Graether, S.P.; Jiménez-Bremont, J.F. In vivo evidence for homo- and heterodimeric interactions of Arabidopsis thaliana dehydrins AtCOR47, AtERD10, and AtRAB18. Sci. Rep. 2017, 7, 17036. [Google Scholar] [CrossRef]
- Puhakainen, T.; Hess, M.W.; Mäkelä, P.; Svensson, J.; Heino, P.; Palva, E.T. Overexpression of multiple dehydrin genes enhances tolerance to freezing stress in Arabidopsis. Plant Mol. Biol. 2004, 54, 743–753. [Google Scholar] [CrossRef]
- Gangappa, S.N.; Botto, J.F. The BBX family of plant transcription factors. Trends Plant Sci. 2016, 19, 460–470. [Google Scholar] [CrossRef]
- Song, K.; Li, B.; Wu, H.; Sha, Y.; Qin, L.; Chen, X.; Liu, Y.; Tang, H.; Yang, L. The function of BBX gene family under multiple stresses in Nicotiana tabacum. Genes 2022, 13, 1841. [Google Scholar] [CrossRef]
- Bai, S.L.; Saito, T.; Honda, C.; Hatsuyama, Y.; Ito, A.; Moriguchi, T. An apple B-box protein, MdCOL11, is involved in UV-B- and temperature-induced anthocyanin biosynthesis. Planta 2014, 240, 1051–1062. [Google Scholar] [CrossRef]
- An, J.P.; Wang, X.F.; Zhang, X.W.; You, C.X.; Hao, Y.J. Apple B-box protein BBX37 regulates jasmonic acid mediated cold tolerance through the JAZ-BBX37-ICE1-CBF pathway and undergoes MIEL1-mediated ubiquitination and degradation. New Phytol. 2021, 229, 2707–2729. [Google Scholar] [CrossRef]
- Bursch, K.; Toledo-Ortiz, G.; Pireyre, M.; Lohr, M.; Braatz, C.; Johansson, H. Identification of BBX proteins as rate-limiting cofactors of HY5. Nat. Plants 2020, 6, 921–928. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.F. Experimental Guidance of Plant Physiology; Beijing Higher Education Press: Beijing, China, 2006; pp. 74–76. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-DDCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]


| Compounds | Class | Content (μg·g−1) |
|---|---|---|
| Cyanidin-3-(6″-caffeylsophoroside)-5-glucoside | Cyanidin | 3.889 ± 0.304 |
| Cyanidin-3-O-sambubioside-5-O-glucoside | Cyanidin | 0.436 ± 0.077 |
| Cyanidin-3-O-glucoside | Cyanidin | 64.825 ± 4.807 |
| Cyanidin-3-O-5-O-(6-O-coumaroyl)-diglucoside | Cyanidin | 36.004 ± 2.653 |
| Cyanidin-3,5,3-O-triglucoside | Cyanidin | 5.515 ± 0.258 |
| Cyanidin-3-O-sophoroside | Cyanidin | 10.300 ± 0.954 |
| Delphinidin-3-O-(6-O-malonyl-beta-D-glucoside) | Delphinidin | 0.062 ± 0.006 |
| Delphinidin-3-O-galactoside | Delphinidin | 1.360 ± 0.208 |
| Delphinidin-3-O-(6-O-acetyl)-glucoside | Delphinidin | 0.017 ± 0.005 |
| Delphinidin-3-O-sophoroside | Delphinidin | 0.964 ± 0.073 |
| Delphinidin-3-O-sambubioside | Delphinidin | 0.015 ± 0.0005 |
| Delphinidin-3-O-rhamnoside | Delphinidin | 0.138 ± 0.142 |
| Delphinidin-3,5-O-diglucoside | Delphinidin | 2.365 ± 0.223 |
| Malvidin-3-O-glucoside | Malvidin | 0.030 ± 0.003 |
| Pelargonidin-3-(6″-caffeylsophoroside)-5-glucoside | Pelargonidin | 0.008 ± 0.002 |
| Pelargonidin-3-sophoroside-5-glucoside | Pelargonidin | 1.577 ± 0.258 |
| Peonidin-3-(caffeoyl-glucosyl-glucoside)-5-glucoside | Peonidin | 0.010 ± 0.003 |
| Peonidin-3-O-(6″-ferulylsophoroside)-5-glucoside | Peonidin | 0.003 ± 0.001 |
| Peonidin-3-O-glucoside | Peonidin | 0.131 ± 0.019 |
| Peonidin-3,5-O-diglucoside | Peonidin | 0.536 ± 0.042 |
| Peonidin-3-sophoroside-5-glucoside | Peonidin | 1.105 ± 0.224 |
| Petunidin-3-O-sophoroside | Petunidin | 0.877 ± 0.133 |
| Petunidin-3-O-(6-O-malonyl-beta-D-glucoside) | Petunidin | 0.277 ± 0.061 |
| Gene | Annotation | S1 vs. S0 | S2 vs. S1 | S2 vs. S0 |
|---|---|---|---|---|
| Bo5g052100 | C4H | up | up | up * |
| Bo3g024650 | C4H | down | down | down * |
| Bo6g099190 | 4CL3 | up * | up * | up * |
| Bo3g077430 | 4CL | up | up * | up * |
| Bo9g166290 | CHS | up | up * | up |
| Bo9g177250 | CHI | up * | up * | up * |
| Bo8g081770 | F3H | up * | up * | up * |
| Bo7g100840 | F3H | up | down | up * |
| Bo9g174880 | F3′H | - | up | up * |
| Bo9g058630 | DFR | up | up | up * |
| Bo2g116380 | DFR | up * | up | up * |
| Bo3g042330 | SAT | up * | up | up * |
| Bo4g141980 | GL3 | down | down | down * |
| Bo4g141990 | GL3 | down | down | down * |
| Bo9g035460 | EGL3 | down | down | down * |
| Bo9g171430 | HY5 | up | up | up * |
| Bo2g070770 | MYBL2 | up * | up | up * |
| Bo4g015800 | SLP9 | down | down | down * |
| Bo9g161480 | TT19 | up * | up | up |
| Bo6g119200 | PR5 | down | down * | down * |
| Bo2g056370 | CCOAMT | up | up * | up * |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zou, J.; Gong, Z.; Liu, Z.; Ren, J.; Feng, H. Investigation of the Key Genes Associated with Anthocyanin Accumulation during Inner Leaf Reddening in Ornamental Kale (Brassica oleracea L. var. acephala). Int. J. Mol. Sci. 2023, 24, 2837. https://doi.org/10.3390/ijms24032837
Zou J, Gong Z, Liu Z, Ren J, Feng H. Investigation of the Key Genes Associated with Anthocyanin Accumulation during Inner Leaf Reddening in Ornamental Kale (Brassica oleracea L. var. acephala). International Journal of Molecular Sciences. 2023; 24(3):2837. https://doi.org/10.3390/ijms24032837
Chicago/Turabian StyleZou, Jiaqi, Zhichao Gong, Zhiyong Liu, Jie Ren, and Hui Feng. 2023. "Investigation of the Key Genes Associated with Anthocyanin Accumulation during Inner Leaf Reddening in Ornamental Kale (Brassica oleracea L. var. acephala)" International Journal of Molecular Sciences 24, no. 3: 2837. https://doi.org/10.3390/ijms24032837





