Cyanidin-3-O-glucoside Contributes to Leaf Color Change by Regulating Two bHLH Transcription Factors in Phoebe bournei
Abstract
:1. Introduction
2. Results
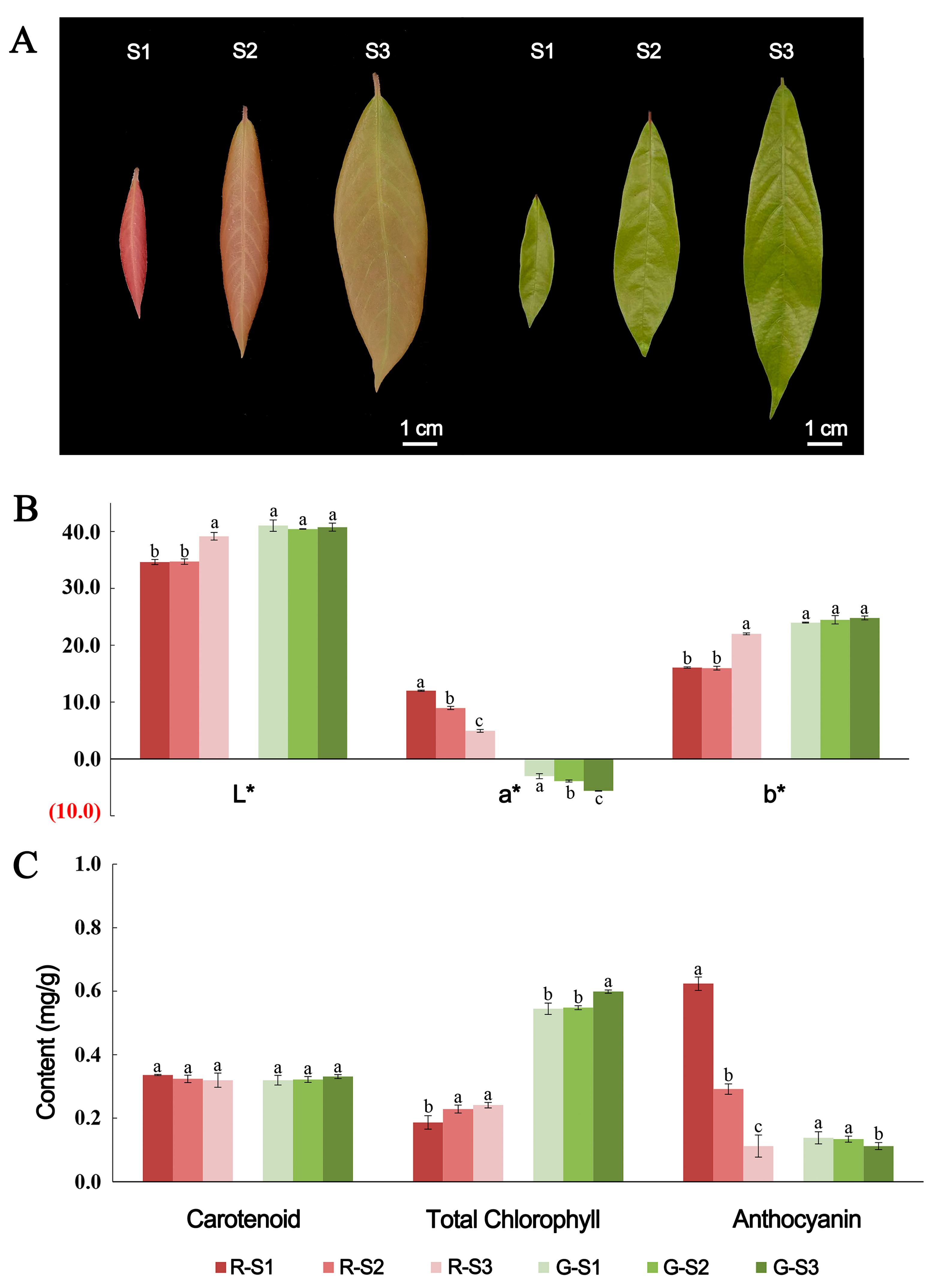
2.1. Phenotypic Characteristics of P. bournei Leaves
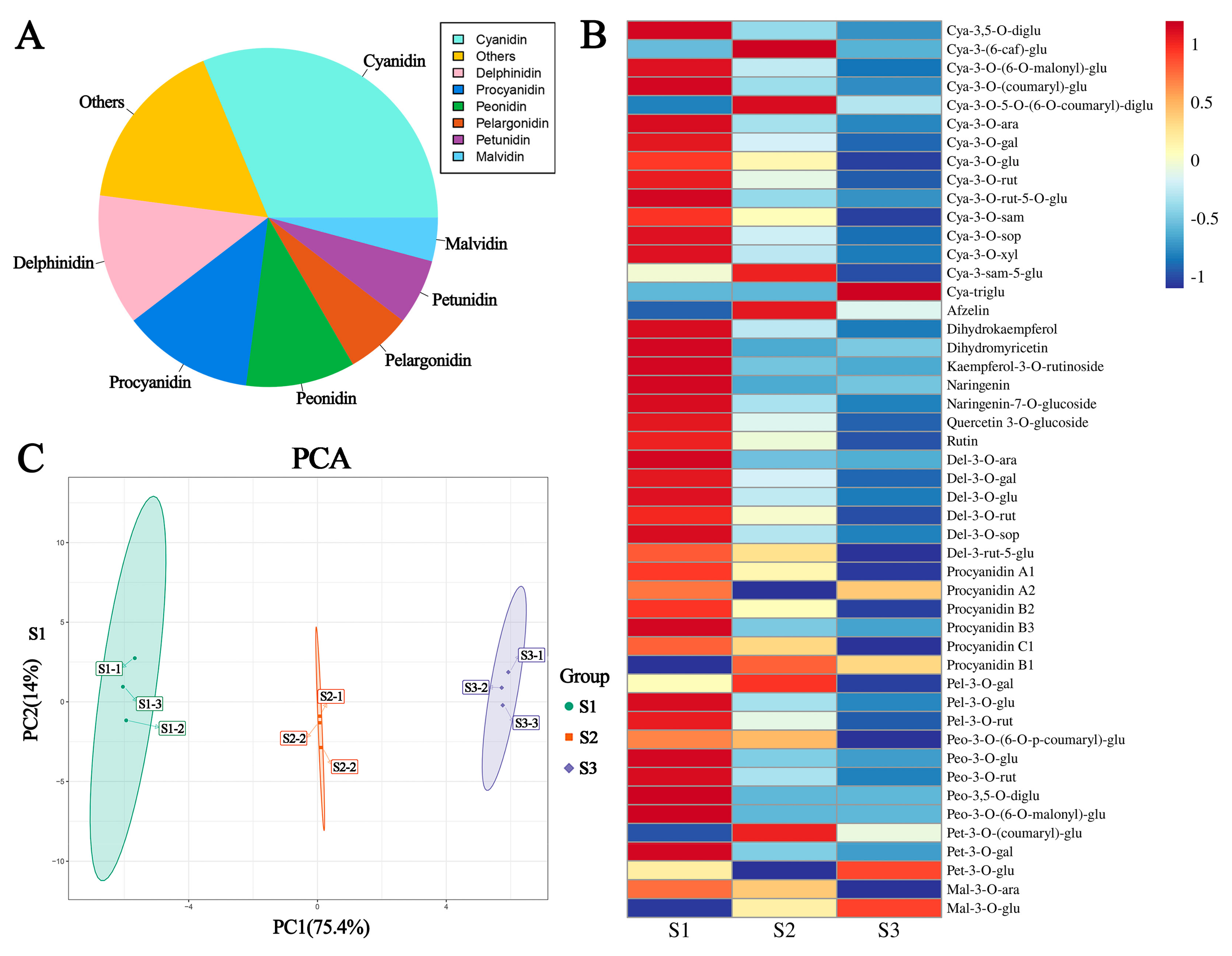
2.2. Anthocyanin Metabolites in the Red Leaves of P. bournei at Different Developmental Stages
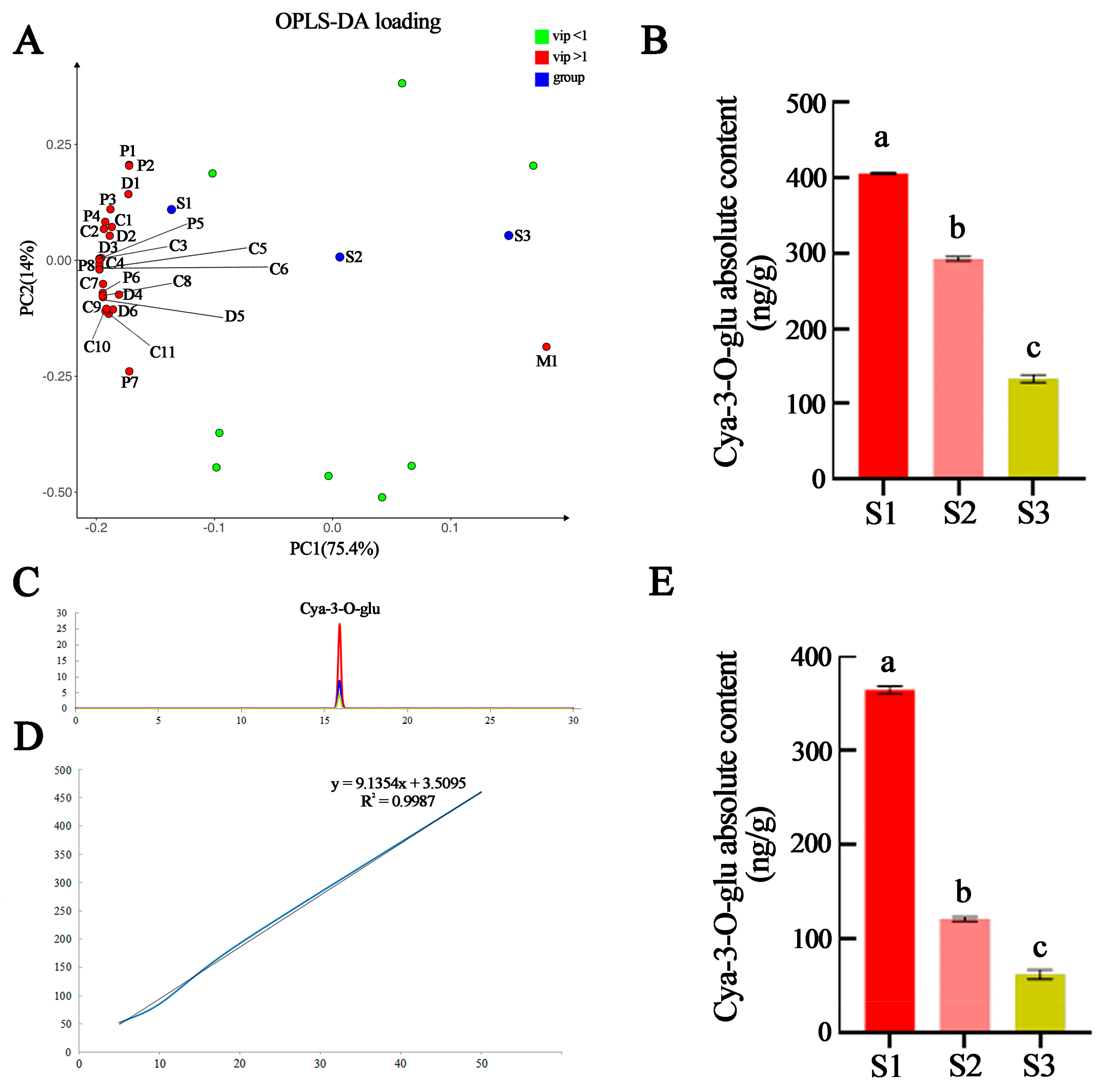
2.3. Identification of Key Metabolites for Leaf Coloration
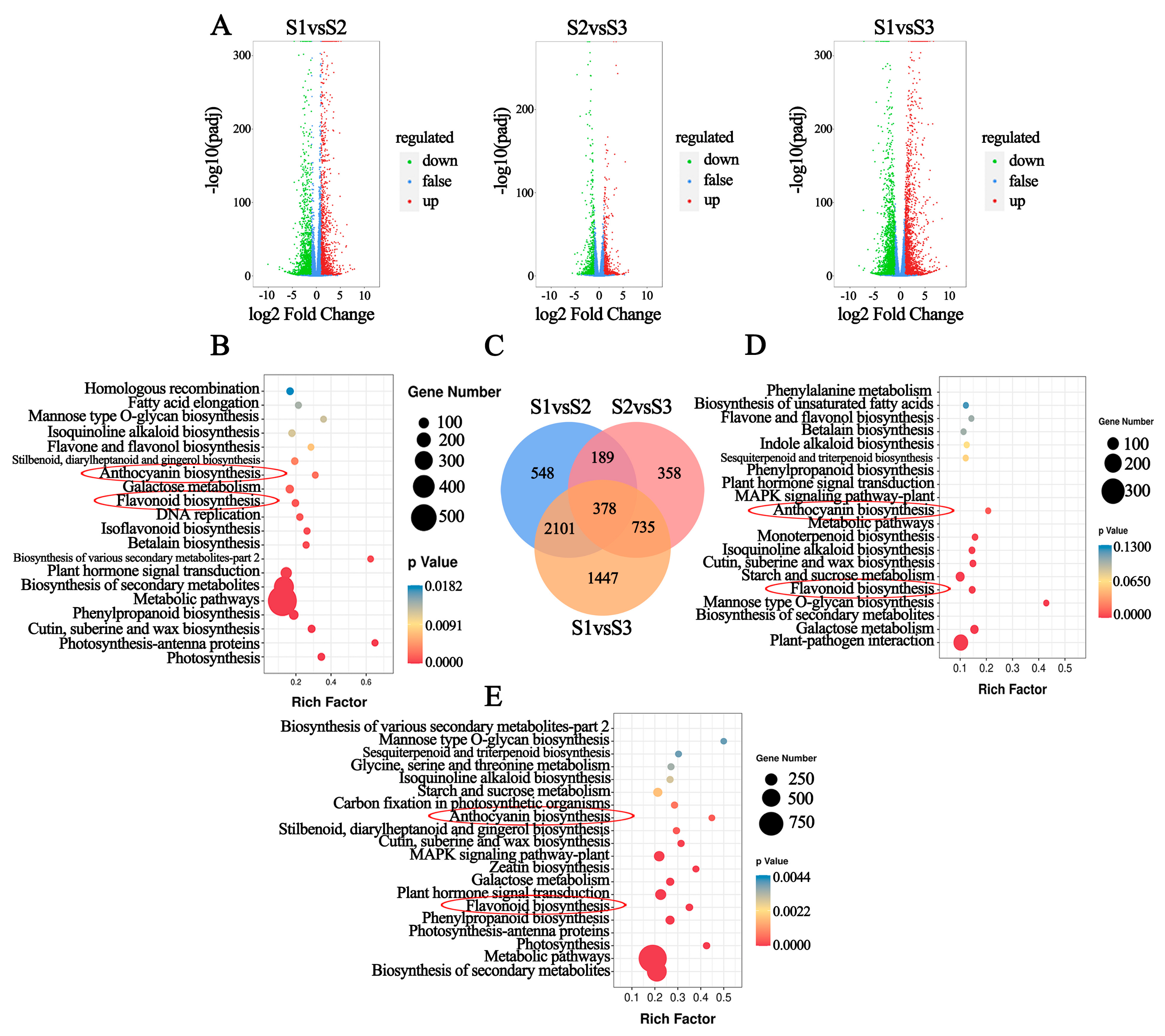
2.4. Transcriptomic Analysis of Leaves during Development
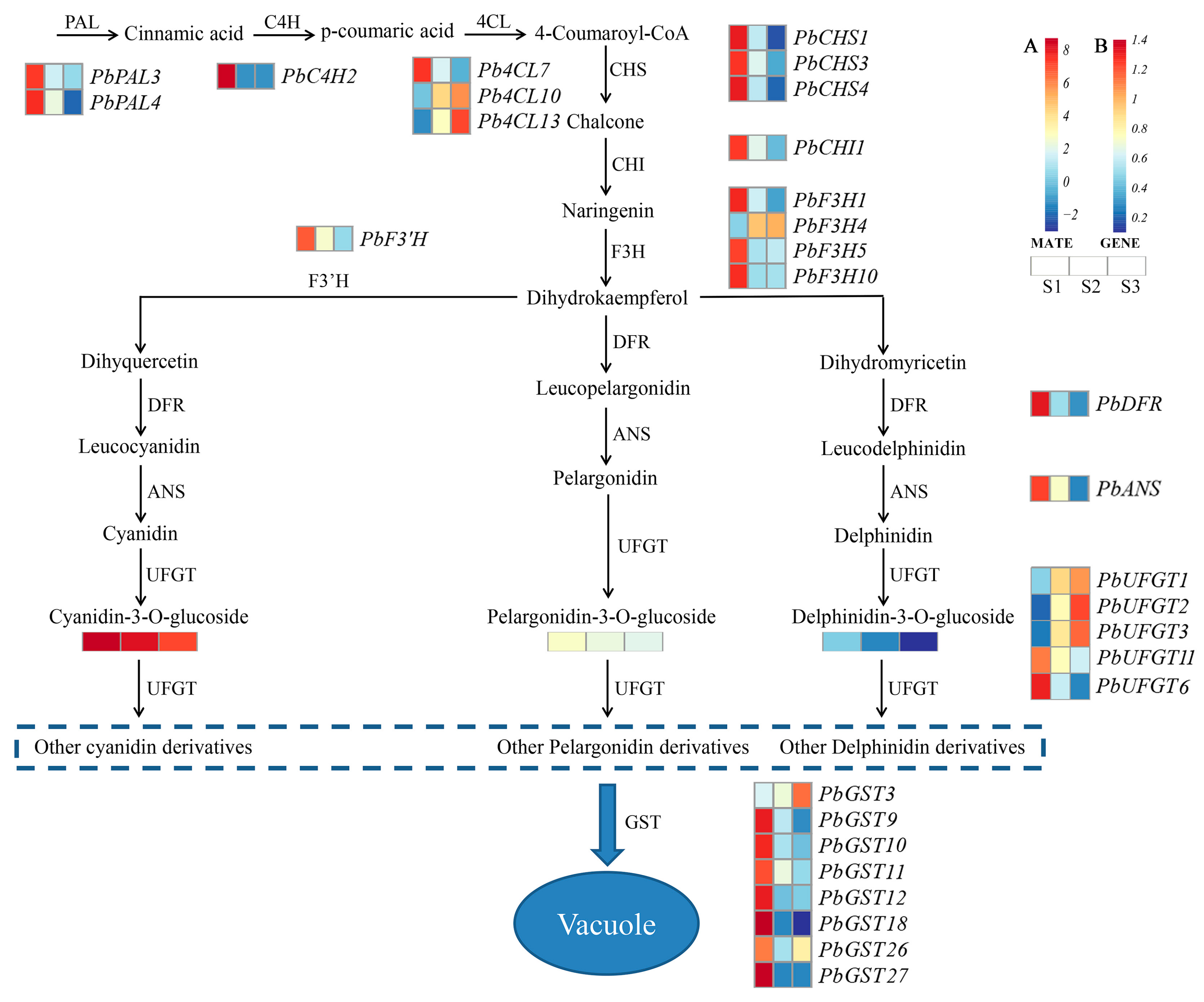
2.5. DEGs Related to Anthocyanin Biosynthesis
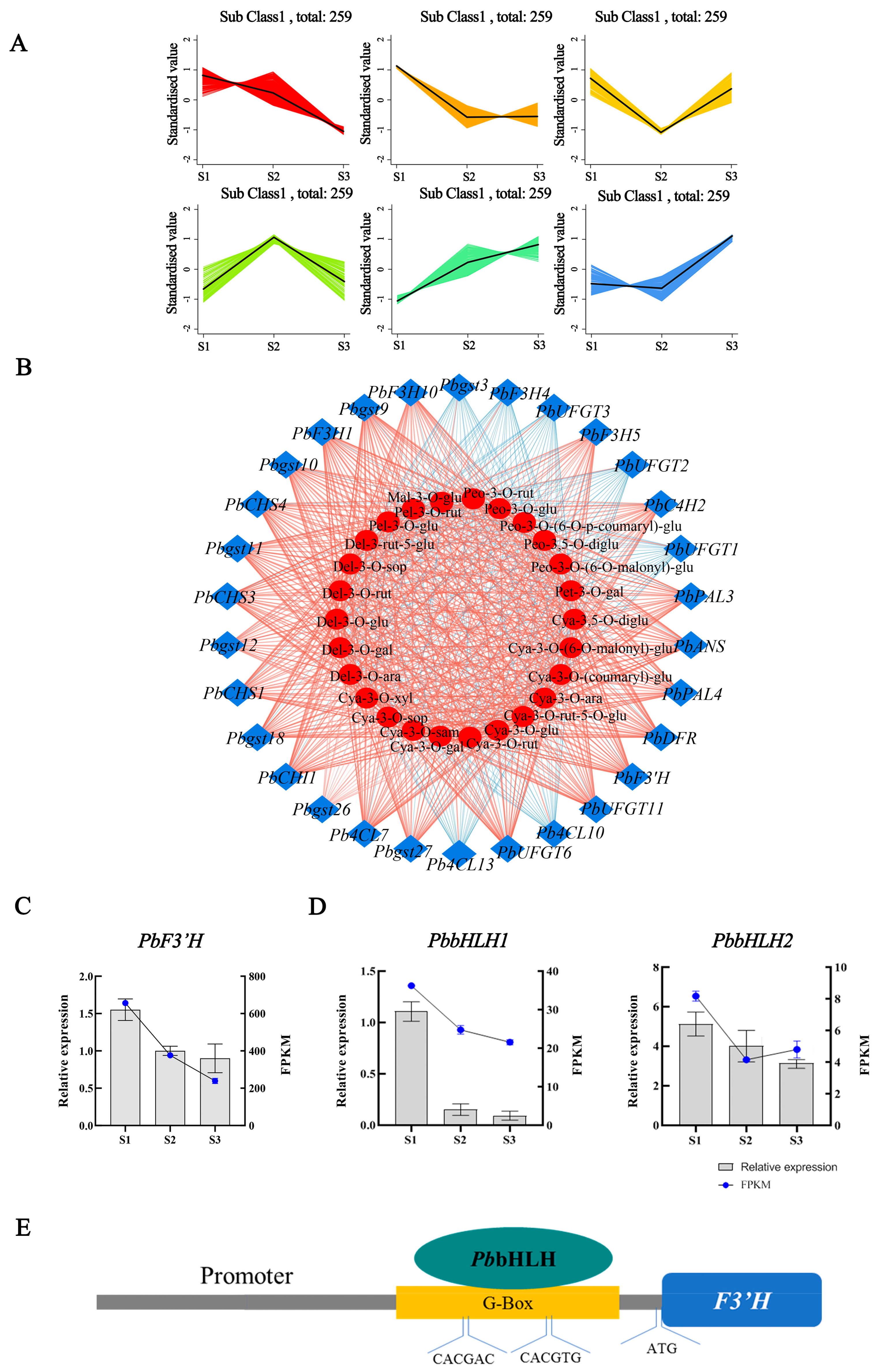
2.6. Integrated Analysis of the Transcriptome and Metabolome
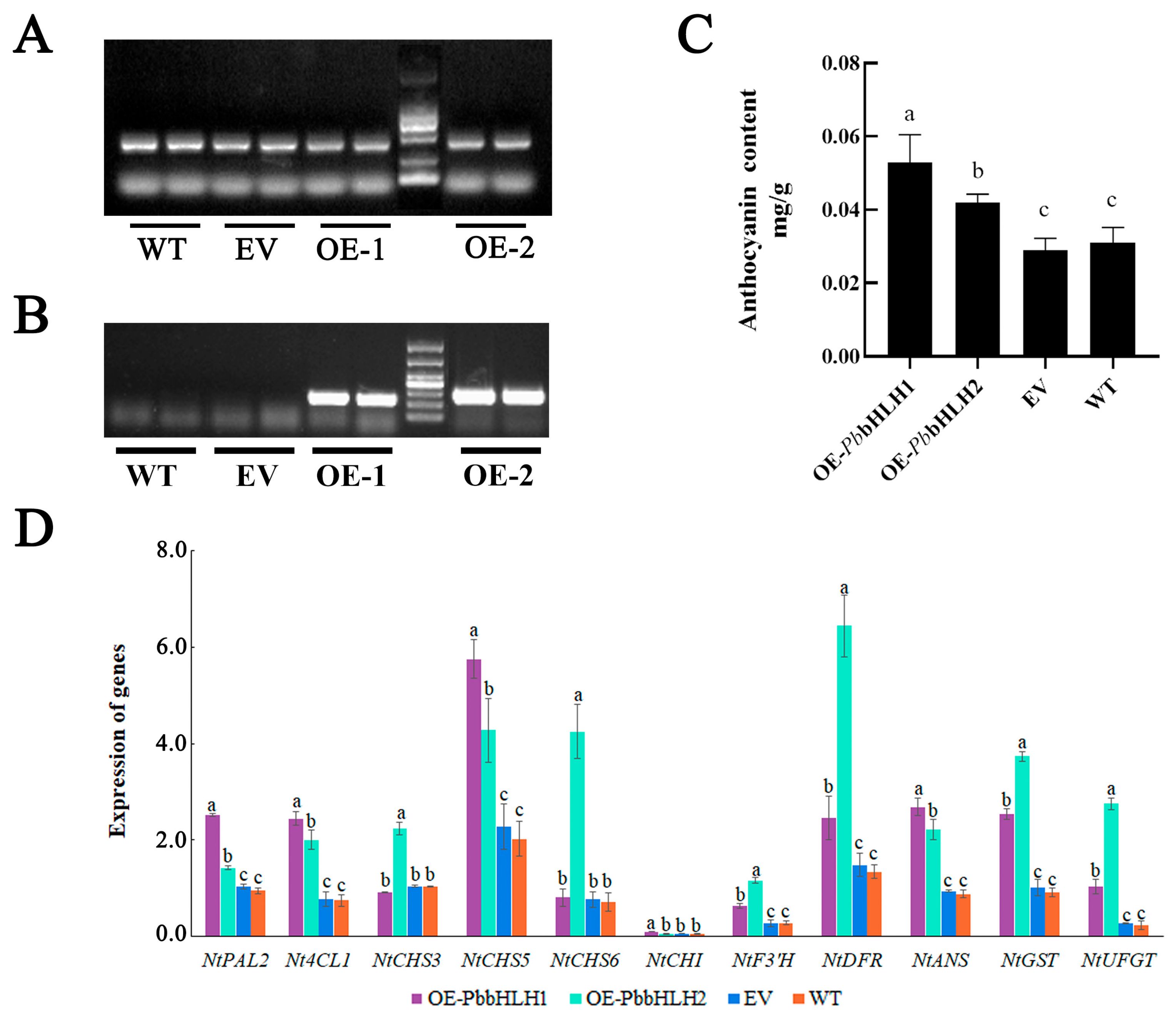
2.7. The Potential Regulatory Roles of PbbHLH1 and PbbHLH2
3. Discussion
3.1. Anthocyanin Characteristics Varied with Leaf Development in P. bournei
3.2. The Genes Involved in Anthocyanin Biosynthesis in Red Leaves of P. bournei
3.3. Transcription Factors Related to Anthocyanin Biosynthesis
4. Materials and Methods
4.1. Plant Materials
4.2. Determination of Plant Pigments
4.2.1. Determination of Total Anthocyanin Content (TAC)
4.2.2. Determination of Photosynthetic Pigments
4.3. Metabolomic Analysis of Anthocyanins
4.4. RNA Extraction, Library Construction, and Sequencing
4.5. RNA-Seq Analysis and Differentially Expressed Gene (DEG) Identification
4.6. Molecular Cloning of PbbHLH1 and PbbHLH2
4.7. Transient Expression Assay Using Nicotiana tabacum
4.8. qPCR Analysis
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, H.N.; Li, W.C.; Wang, H.C.; Shi, S.Y.; Shu, B.; Liu, L.Q.; Wei, Y.Z.; Xie, J.H. Transcriptome Profiling of Light-Regulated Anthocyanin Biosynthesis in the Pericarp of Litchi. Front. Plant Sci. 2016, 7, 963. [Google Scholar] [CrossRef] [PubMed]
- Qiao, Z.; Liu, S.; Zeng, H.; Li, Y.; Wang, X.; Chen, Y.; Wang, X.; Cai, N. Exploring the Molecular Mechanism underlying the Stable Purple-Red Leaf Phenotype in Lagerstroemia indica cv. Ebony Embers. Int. J. Mol. Sci. 2019, 20, 5636. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Huang, H.; Wang, L.; Sun, Y.; Dai, S. Transcriptomics and Metabolite Analysis Reveals the Molecular Mechanism of Anthocyanin Biosynthesis Branch Pathway in Different Senecio cruentus Cultivars. Front. Plant Sci. 2016, 7, 1307. [Google Scholar] [CrossRef] [PubMed]
- Andersen, Ø.M.; Jordheim, M. Anthocyanins. In Encyclopedia of Life Sciences (ELS); eLS: New York, NY, USA, 2010. [Google Scholar]
- Ono, E.; Fukuchi-Mizutani, M.; Nakamura, N.; Fukui, Y.; Yonekura-Sakakibara, K.; Yamaguchi, M.; Nakayama, T.; Tanaka, T.; Kusumi, T.; Tanaka, Y. Yellow flowers generated by expression of the aurone biosynthetic pathway. Proc. Natl. Acad. Sci. USA 2006, 103, 11075–11080. [Google Scholar] [CrossRef]
- Han, L.; Zou, H.; Zhou, L.; Wang, Y. Transcriptome-Based Identification and Expression Analysis of the Glutathione S-Transferase (GST) Family in Tree Peony Reveals a Likely Role in Anthocyanin Transport. Hortic. Plant J. 2022, 8, 787–802. [Google Scholar] [CrossRef]
- Alappat, B.; Alappat, J. Anthocyanin Pigments: Beyond Aesthetics. Molecules 2020, 25, 5500. [Google Scholar] [CrossRef]
- Huang, D.; Ming, R.; Yao, S.; Li, L.; Huang, R.; Tan, Y. Identification of anthocyanins in the fruits of Kadsura coccinea using UPLC-MS/MS-based metabolomics. Biochem. Syst. Ecol. 2021, 98, 104324. [Google Scholar] [CrossRef]
- Bao, Y.; Nie, T.; Wang, D.; Chen, Q. Anthocyanin regulatory networks in Solanum tuberosum L. leaves elucidated via integrated metabolomics, transcriptomics, and StAN1 overexpression. BMC Plant Biol. 2022, 22, 228. [Google Scholar] [CrossRef]
- Petroni, K.; Tonelli, C. Recent advances on the regulation of anthocyanin synthesis in reproductive organs. Plant Sci. 2011, 181, 219–229. [Google Scholar] [CrossRef]
- Jaakola, L. New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant Sci. 2013, 18, 477–483. [Google Scholar] [CrossRef] [Green Version]
- Springob, K.; Nakajima, J.; Yamazaki, M.; Saito, K. Recent advances in the biosynthesis and accumulation of anthocyanins. Nat. Prod. Rep. 2003, 20, 288–303. [Google Scholar] [CrossRef] [PubMed]
- Mouradov, A.; Spangenberg, G. Modification of Flavonoid Biosynthesis in Plants. US20090113573, 30 April 2009. [Google Scholar]
- Liu, Y.; Ma, K.; Qi, Y.; Lv, G.; Ren, X.; Liu, Z.; Ma, F. Transcriptional Regulation of Anthocyanin Synthesis by MYB-bHLH-WDR Complexes in Kiwifruit (Actinidia chinensis). J. Agric. Food Chem. 2021, 69, 3677–3691. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Feng, C.; Zhang, M.; Yin, X.; Xu, C.; Chen, K. The MrWD40-1 Gene of Chinese Bayberry (Myrica rubra) Interacts with MYB and bHLH to Enhance Anthocyanin Accumulation. Plant Mol. Biol. Rep. 2013, 31, 1474–1484. [Google Scholar] [CrossRef]
- Sun, Q.; Jiang, S.; Zhang, T.; Xu, H.; Fang, H.; Zhang, J.; Su, M.; Wang, Y.; Zhang, Z.; Wang, N. Apple NAC transcription factor MdNAC52 regulates biosynthesis of anthocyanin and proanthocyanidin through MdMYB9 and MdMYB11. Plant Sci. 2019, 289, 110286. [Google Scholar] [CrossRef] [PubMed]
- An, J.P.; Yao, J.F.; Xu, R.R.; You, C.X.; Wang, X.F.; Hao, Y.J. Apple bZIP transcription factor MdbZIP44 regulates abscisic acid-promoted anthocyanin accumulation. Plant Cell Environ. 2018, 41, 2678–2692. [Google Scholar] [CrossRef]
- Ma, H.; Yang, T.; Li, Y.; Zhang, J.; Wu, T.; Song, T.; Yao, Y.; Tian, J. The long noncoding RNA MdLNC499 bridges MdWRKY1 and MdERF109 function to regulate early-stage light-induced anthocyanin accumulation in apple fruit. Plant Cell 2021, 33, 3309–3330. [Google Scholar] [CrossRef]
- Ji, X.; Ren, J.; Zhang, Y.; Lang, S.; Wang, D.; Song, X. Integrated Analysis of the Metabolome and Transcriptome on Anthocyanin Biosynthesis in Four Developmental Stages of Cerasus humilis Peel Coloration. Int. J. Mol. Sci. 2021, 22, 11880. [Google Scholar] [CrossRef]
- Zhang, A.; Yang, H.; Ji, S.; Tian, C.; Chen, N.; Gong, H.; Li, J. Metabolome and Transcriptome Analyses of Anthocyanin Accumulation Mechanisms Reveal Metabolite Variations and Key Candidate Genes Involved in the Pigmentation of Prunus tomentosa Thunb. Cherry Fruit. Front. Plant Sci. 2022, 13, 938908. [Google Scholar] [CrossRef]
- Gao, Y.F.; Zhao, D.H.; Zhang, J.Q.; Chen, J.S.; Li, J.L.; Weng, Z.; Rong, L.P. De novo transcriptome sequencing and anthocyanin metabolite analysis reveals leaf color of Acer pseudosieboldianum in autumn. BMC Genom. 2021, 22, 383. [Google Scholar] [CrossRef]
- Anran, S.; Xiaona, P.; Shikai, Z.; Zhiming, H.; Ying, X.; Guanzheng, Q.; Xiaoqing, H.; Mulualem, T.; Xiyang, Z. Metabolome and Transcriptome Analyses Unravels Molecular Mechanisms of Leaf Color Variation by Anthocyanidin Biosynthesis in Acer triflorum. Horticulturae 2022, 8, 635. [Google Scholar] [CrossRef]
- Mei, Y.; Xie, H.; Liu, S.; Zhu, J.; Zhao, S.; Wei, C. Metabolites and Transcriptional Profiling Analysis Reveal the Molecular Mechanisms of the Anthocyanin Metabolism in the “Zijuan” Tea Plant (Camellia sinensis var. assamica). J. Agric. Food Chem. 2021, 69, 414–427. [Google Scholar] [CrossRef]
- Ge, Y.J.; Liu, Y.J.; Shen, A.H.; Lin, X.C. Fengshui forests conserve genetic diversity: A case study of Phoebe bournei (Hemsl.) Yang in southern China. Genet. Mol. Res. 2015, 14, 1986–1993. [Google Scholar] [CrossRef]
- Ye, Z.; Huang, S.; Zhu, J.; Zuo, H.; Tan, G. Propagation, maintenance and garden applications of four ornamental plants of the genus Camphoraceae. Guangdong Gard. 2016, 38, 48–51. [Google Scholar] [CrossRef]
- Yuan, L.; Niu, H.; Yun, Y.; Tian, J.; Lao, F.; Liao, X.; Gao, Z.; Ren, D.; Zhou, L. Analysis of coloration characteristics of Tunisian soft-seed pomegranate arils based on transcriptome and metabolome. Food Chem. 2022, 370, 131270. [Google Scholar] [CrossRef]
- Han, X.; Zhang, J.; Han, S.; Chong, S.L.; Meng, G.; Song, M.; Wang, Y.; Zhou, S.; Liu, C.; Lou, L. The chromosome-scale genome of Phoebe bournei reveals contrasting fates of terpene synthase (TPS)-a and TPS-b subfamilies. Plant Commun. 2022, 3, 100410. [Google Scholar] [CrossRef]
- Verediano, T.A.; Stampini Duarte Martino, H.; Dias Paes, M.C.; Tako, E. Effects of Anthocyanin on Intestinal Health: A Systematic Review. Nutrients 2021, 13, 1331. [Google Scholar] [CrossRef]
- Bi, X.; Zhang, J.; Chen, C.; Zhang, D.; Li, P.; Ma, F. Anthocyanin contributes more to hydrogen peroxide scavenging than other phenolics in apple peel. Food Chem. 2014, 152, 205–209. [Google Scholar] [CrossRef]
- Fu, M.; Yang, X.; Zheng, J.; Wang, L.; Yang, X.; Tu, Y.; Ye, J.; Zhang, W.; Liao, Y.; Cheng, S.; et al. Unraveling the Regulatory Mechanism of Color Diversity in Camellia japonica Petals by Integrative Transcriptome and Metabolome Analysis. Front. Plant Sci. 2021, 12, 685136. [Google Scholar] [CrossRef]
- Feng, X.; Gao, G.; Yu, C.; Zhu, A.; Chen, J.; Chen, K.; Wang, X.; Abubakar, A.S.; Chen, P. Transcriptome and metabolome analysis reveals anthocyanin biosynthesis pathway associated with ramie (Boehmeria nivea (L.) Gaud.) leaf color formation. BMC Genom. 2021, 22, 684. [Google Scholar] [CrossRef]
- Pan, L.; Li, J.; Yin, H.; Fan, Z.; Li, X. Integrated Physiological and Transcriptomic Analyses Reveal a Regulatory Network of Anthocyanin Metabolism Contributing to the Ornamental Value in a Novel Hybrid Cultivar of Camellia Japonica. Plants 2020, 9, 1724. [Google Scholar] [CrossRef]
- Zhong, Y.; Chen, C.; Gong, X.; Luan, X.; Wu, Z.; Li, H.; Liu, Q.; Xu, M.; Yu, F. Transcriptome and metabolome analyses reveal a key role of the anthocyanin biosynthetic pathway cascade in the pigmentation of a Cinnamomum camphora red bark mutant (‘Gantong 1′). Ind. Crops Prod. 2022, 175, 114236. [Google Scholar] [CrossRef]
- Veberic, R.; Slatnar, A.; Bizjak, J.; Stampar, F.; Mikulic-Petkovsek, M. Anthocyanin composition of different wild and cultivated berry species. LWT Food Sci. Technol. 2015, 60, 509–517. [Google Scholar] [CrossRef]
- Rao, M.J.; Duan, M.; Yang, M.; Fan, H.; Shen, S.; Hu, L.; Wang, L. Novel Insights into Anthocyanin Metabolism and Molecular Characterization of Associated Genes in Sugarcane Rinds Using the Metabolome and Transcriptome. Int. J. Mol. Sci. 2021, 23, 338. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Zhang, Q.; Lang, L.; Jiang, C.; Wang, X.; Sun, H. Integrated metabolome and transcriptome analysis of the anthocyanin biosynthetic pathway in relation to color mutation in miniature roses. BMC Plant Biol. 2021, 21, 257. [Google Scholar] [CrossRef]
- Parchman, T.L.; Geist, K.S.; Grahnen, J.A.; Benkman, C.W.; Buerkle, C.A. Transcriptome sequencing in an ecologically important tree species: Assembly, annotation, and marker discovery. BMC Genom. 2010, 11, 180. [Google Scholar] [CrossRef]
- Jiao, Y.; Ma, R.J.; Shen, Z.J.; Yan, J.; Yu, M.L. Gene regulation of anthocyanin biosynthesis in two blood-flesh peach (Prunus persica (L.) Batsch) cultivars during fruit development. J. Zhejiang Univ. Sci. B 2014, 15, 809–819. [Google Scholar] [CrossRef]
- Chen, Z.; Lu, X.; Xuan, Y.; Tang, F.; Ren, J. Transcriptome analysis based on a combination of sequencing platforms provides insights into leaf pigmentation in Acer rubrum. BMC Plant Biol. 2019, 19, 240. [Google Scholar] [CrossRef]
- Teng, K.; Liu, L.; Fan, X.; Zhang, H.; Han, C.; Wu, J.; Chang, Z. Combination analysis of single-molecule long-read and Illumina sequencing provides insights into the anthocyanin accumulation mechanism in an ornamental grass, Pennisetum setaceum cv. Rubrum. Plant Mol. Biol. 2022, 109, 159–175. [Google Scholar] [CrossRef]
- Yu, M.; Man, Y.; Lei, R.; Lu, X.; Wang, Y. Metabolomics Study of Flavonoids and Anthocyanin-Related Gene Analysis in Kiwifruit (Actinidia chinensis) and Kiwiberry (Actinidia arguta). Plant Mol. Biol. Report. 2020, 38, 353–369. [Google Scholar] [CrossRef]
- Sun, H.; Li, Y.; Feng, S.; Zou, W.; Guo, K.; Fan, C.; Si, S.; Peng, L. Analysis of five rice 4-coumarate:coenzyme A ligase enzyme activity and stress response for potential roles in lignin and flavonoid biosynthesis in rice. Biochem. Biophys. Res. Commun. 2013, 430, 1151–1156. [Google Scholar] [CrossRef]
- Chen, Y.; Mao, Y.; Liu, H.; Yu, F.; Li, S.; Yin, T. Transcriptome analysis of differentially expressed genes relevant to variegation in peach flowers. PLoS ONE 2014, 9, e90842. [Google Scholar] [CrossRef]
- Jeong, S.T.; Goto-Yamamoto, N.; Hashizume, K.; Esaka, M. Expression of the flavonoid 3′-hydroxylase and flavonoid 3′,5′-hydroxylase genes and flavonoid composition in grape (Vitis vinifera). Plant Sci. 2005, 170, 61–69. [Google Scholar] [CrossRef]
- Russo, M.P.; Cotroneo, P.S.; Lo, P.A.R.; Consoli, A.; Reforgiato, R.G. Expression levels of CHS, DFR and ANS and anthocyanin content in 19 genotypes of Citrus. In Proceedings of the XLVII Italian Society of Agricultural Genetics-SIGA Annual Congress, Verona, Italy, 24–27 September 2003. [Google Scholar]
- Chapple, C.; Shirley, B.W.; Zook, M.; Hammerschmidt, R.; Somerville, S.C. 36 Secondary Metabolism in Arabidopsis. Cold Spring Harb. Monogr. Arch. 1994, 27. [Google Scholar] [CrossRef]
- Zhuang, H.; Lou, Q.; Liu, H.; Han, H.; Wang, Q.; Tang, Z.; Ma, Y.; Wang, H. Differential Regulation of Anthocyanins in Green and Purple Turnips Revealed by Combined De Novo Transcriptome and Metabolome Analysis. Int. J. Mol. Sci. 2019, 20, 4387. [Google Scholar] [CrossRef]
- Silan, D.; Yan, H. Molecular Breeding for Flower Colors Modification on Ornamental Plants Based on the Mechanism of Anthocyanins Biosynthesis and Coloration. Sci. Agric. Sin. 2016, 49, 529–542. [Google Scholar] [CrossRef]
- Jie, Z.; Han, Z.Y.; Ji, T. The expression level of anthocyanidin synthase determines the anthocyanin content of crabapple (Malus sp.) petals. Acta Physiol. Plant. 2015, 37, 109. [Google Scholar] [CrossRef]
- Wu, X.; Gong, Q.; Ni, X.; Zhou, Y.; Gao, Z. UFGT: The Key Enzyme Associated with the Petals Variegation in Japanese Apricot. Front. Plant Sci. 2017, 8, 108. [Google Scholar] [CrossRef]
- Luo, H.; Dai, C.; Li, Y.; Feng, J.; Liu, Z.; Kang, C. Reduced Anthocyanins in Petioles codes for a GST anthocyanin transporter that is essential for the foliage and fruit coloration in strawberry. J. Exp. Bot. 2018, 69, 2595–2608. [Google Scholar] [CrossRef]
- Moglia, A.; Florio, F.E.; Iacopino, S.; Guerrieri, A.; Milani, A.M.; Comino, C.; Barchi, L.; Marengo, A.; Cagliero, C.; Rubiolo, P.; et al. Identification of a new R3 MYB type repressor and functional characterization of the members of the MBW transcriptional complex involved in anthocyanin biosynthesis in eggplant (S. melongena L.). PLoS ONE 2020, 15, e0232986. [Google Scholar] [CrossRef]
- Outchkourov, N.S.; Carollo, C.A.; Gomez-Roldan, V.; De Vos, R.C.; Bosch, D.; Hall, R.D.; Beekwilder, J. Control of anthocyanin and non-flavonoid compounds by anthocyanin-regulating MYB and bHLH transcription factors in Nicotiana benthamiana leaves. Front. Plant Sci. 2014, 5, 519. [Google Scholar] [CrossRef] [Green Version]
- Ning, G.X.; Li, W.F.; Chu, M.Y.; Ma, Z.H.; Wang, P.; Mao, J.; Chen, B.H. MdbHLH51 plays a positive role in anthocyanin accumulation in ‘Red Delicious’ apples. Trees 2022, 36, 1687–1695. [Google Scholar] [CrossRef]
- Deng, J.; Li, J.; Su, M.; Lin, Z.; Chen, L.; Yang, P. A bHLH gene NnTT8 of Nelumbo nucifera regulates anthocyanin biosynthesis. Plant Physiol. Biochem. 2021, 158, 518–523. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Xing, M.; Cai, C.; Li, S. B-box Proteins in Arachis duranensis: Genome-Wide Characterization and Expression Profiles Analysis. Agronomy 2019, 10, 23. [Google Scholar] [CrossRef]
- Neugart, S.; Schreiner, M. UVB and UVA as eustressors in horticultural and agricultural crops. Sci. Hortic. 2018, 234, 370–381. [Google Scholar] [CrossRef]
- Ruiyan, T.; Wenjie, Y.; Yuhao, G.; Junbei, N.; Lei, Y.; Xiao, Z.; Hongxu, L.; Dongsheng, W.; Songling, B.; Yuanwen, T. Light-induced basic/helix-loop-helix 64 enhances anthocyanin biosynthesis and undergoes CONSTITUTIVELY PHOTOMORPHOGENIC 1-mediated degradation in pear. Plant Physiol. 2020, 184, 1684–1701. [Google Scholar] [CrossRef]
- Gaiotti, F.; Pastore, C.; Filippetti, I.; Lovat, L.; Belfiore, N.; Tomasi, D. Low night temperature at veraison enhances the accumulation of anthocyanins in Corvina grapes (Vitis Vinifera L.). Sci. Rep. 2018, 8, 8719. [Google Scholar] [CrossRef]
- He, Q.; Ren, Y.; Zhao, W.; Li, R.; Zhang, L. Low Temperature Promotes Anthocyanin Biosynthesis and Related Gene Expression in the Seedlings of Purple Head Chinese Cabbage (Brassica rapa L.). Genes 2020, 11, 81. [Google Scholar] [CrossRef]
- Gu, K.-D.; Wang, C.-K.; Hu, D.-G.; Hao, Y.-J. How do anthocyanins paint our horticultural products? Sci. Hortic. 2019, 249, 257–262. [Google Scholar] [CrossRef]
- An, J.P.; Li, H.H.; Song, L.Q.; Su, L.; Liu, X.; You, C.X.; Wang, X.F.; Hao, Y.J. The molecular cloning and functional characterization of MdMYC2, a bHLH transcription factor in apple. Plant Physiol. Biochem. 2016, 108, 24–31. [Google Scholar] [CrossRef]
- Shen, X.; Zhao, K.; Liu, L.; Zhang, K.; Yuan, H.; Liao, X.; Wang, Q.; Guo, X.; Li, F.; Li, T. A role for PacMYBA in ABA-regulated anthocyanin biosynthesis in red-colored sweet cherry cv. Hong Deng (Prunus avium L.). Plant Cell Physiol. 2014, 55, 862–880. [Google Scholar] [CrossRef]
- Ji, X.-H.; Zhang, R.; Wang, N.; Yang, L.; Chen, X.-S. Transcriptome profiling reveals auxin suppressed anthocyanin biosynthesis in red-fleshed apple callus (Malus sieversii f. niedzwetzkyana). Plant Cell Tissue Organ Cult. PCTOC 2015, 123, 389–404. [Google Scholar] [CrossRef]
- Jiang, T.; Mao, Y.; Sui, L.; Yang, N.; Li, S.; Zhu, Z.; Wang, C.; Yin, S.; He, J.; He, Y. Degradation of anthocyanins and polymeric color formation during heat treatment of purple sweet potato extract at different pH. Food Chem. 2018, 274, 460–470. [Google Scholar] [CrossRef]
- Adriana, C.M.; Gheorghe, C.; Gabriela, N. Studies concerning the extraction of chlorophyll and total carotenoids from vegetables. Rom. Biotechnol. Lett. 2012, 17, 7702–7708. [Google Scholar]
- De Ferrars, R.M.; Czank, C.; Saha, S.; Needs, P.W.; Zhang, Q.; Raheem, K.S.; Botting, N.P.; Kroon, P.A.; Kay, C.D. Methods for isolating, identifying, and quantifying anthocyanin metabolites in clinical samples. Anal. Chem. 2014, 86, 10052–10058. [Google Scholar] [CrossRef]
- Cruz, A.A.D.l.; Hilbert, G.; Rivière, C.; Mengin, V.; Ollat, N.; Bordenave, L.; Decroocq, S.; Delaunay, J.-C.; Delrot, S.; Mérillon, J.-M.; et al. Anthocyanin identification and composition of wild Vitis spp. accessions by using LC–MS and LC–NMR. Anal. Chim. Acta 2012, 732, 145–152. [Google Scholar] [CrossRef]
- Parkhomchuk, D.; Borodina, T.; Amstislavskiy, V.; Banaru, M.; Hallen, L.; Krobitsch, S.; Lehrach, H.; Soldatov, A. Transcriptome analysis by strand-specific sequencing of complementary DNA. Nucleic Acids Res. 2009, 37, e123. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Varet, H.; Brillet-Gueguen, L.; Coppee, J.Y.; Dillies, M.A. SARTools: A DESeq2- and EdgeR-Based R Pipeline for Comprehensive Differential Analysis of RNA-Seq Data. PLoS ONE 2016, 11, e0157022. [Google Scholar] [CrossRef]
- Yi, D.; Zhang, H.; Lai, B.; Liu, L.; Pan, X.; Ma, Z.; Wang, Y.; Xie, J.; Shi, S.; Wei, Y. Integrative Analysis of the Coloring Mechanism of Red Longan Pericarp through Metabolome and Transcriptome Analyses. J. Agric. Food Chem. 2021, 69, 1806–1815. [Google Scholar] [CrossRef]
- Liu, F.; Chen, G.; Zhang, Y.; Zhao, P.; Dong, D.; Wang, Y.; Wang, S.; Yang, Y. A Comparative Analysis of Metabolome Reveals the Regulation of the Anthocyanin Biosynthesis Branch in Potatoes. Potato Res. 2022, 1–20. [Google Scholar] [CrossRef]
- Otasek, D.; Morris, J.H.; Boucas, J.; Pico, A.R.; Demchak, B. Cytoscape Automation: Empowering workflow-based network analysis. Genome Biol. 2019, 20, 185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, L.; Wang, Q.; Fu, N.; Song, M.; Han, X.; Yang, Q.; Zhang, Y.; Tong, Z.; Zhang, J. Cyanidin-3-O-glucoside Contributes to Leaf Color Change by Regulating Two bHLH Transcription Factors in Phoebe bournei. Int. J. Mol. Sci. 2023, 24, 3829. https://doi.org/10.3390/ijms24043829
Wang L, Wang Q, Fu N, Song M, Han X, Yang Q, Zhang Y, Tong Z, Zhang J. Cyanidin-3-O-glucoside Contributes to Leaf Color Change by Regulating Two bHLH Transcription Factors in Phoebe bournei. International Journal of Molecular Sciences. 2023; 24(4):3829. https://doi.org/10.3390/ijms24043829
Chicago/Turabian StyleWang, Li, Qiguang Wang, Ningning Fu, Minyan Song, Xiao Han, Qi Yang, Yuting Zhang, Zaikang Tong, and Junhong Zhang. 2023. "Cyanidin-3-O-glucoside Contributes to Leaf Color Change by Regulating Two bHLH Transcription Factors in Phoebe bournei" International Journal of Molecular Sciences 24, no. 4: 3829. https://doi.org/10.3390/ijms24043829






