Bipartite Genomes in Enterobacterales: Independent Origins of Chromids, Elevated Openness and Donors of Horizontally Transferred Genes
Abstract
:1. Introduction
2. Results
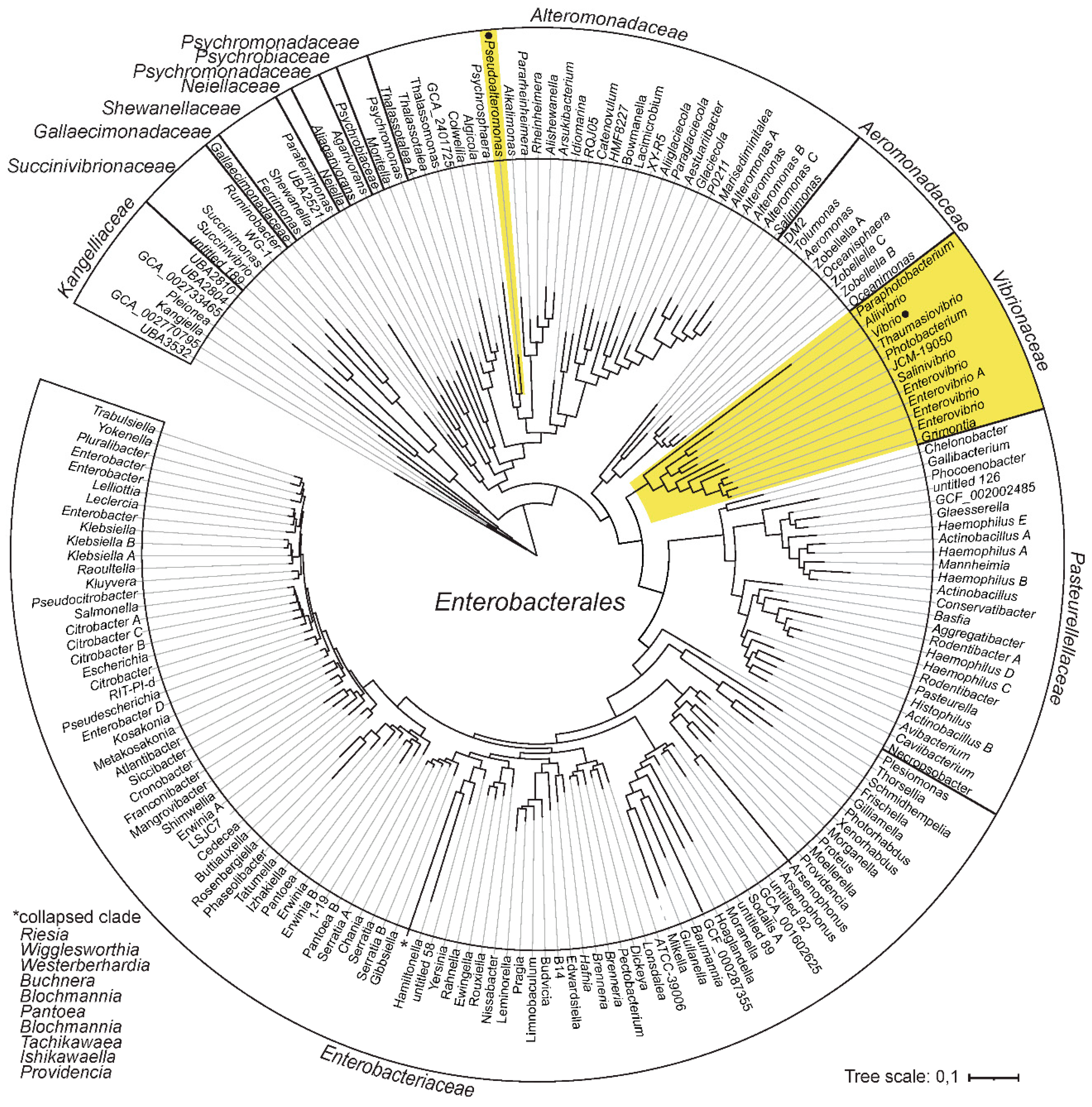
2.1. Vibrio and Pseudoalteromonas Belong to the Same Bacterial Order
2.2. Separate Origin of Chromids in Vibrionaceae and Pseudoalteromonas
2.3. The Chromids in Pseudoalteromonas and Vibrio Play a Significant Role in the Openness of the Two Genomes
2.4. Bipartite Genomes Are More Open Compared to Monopartite Genomes
2.5. Codon Usage Is Specific for Each Pangene Category Rather Than for Each Replicon Type
2.6. Shewanella Represents the Top Donor of HTGs to Vibrio and Pseudoalteromonas
3. Discussion
4. Material and methods
4.1. Enterobacterales Reference Tree
4.2. ParAB phylogenetic tree
4.3. Genome Retrieval and Gene Annotation
4.4. Pangenome Calculation
4.5. Calculation of Codon Usage
4.6. Prediction of Horizontally Transferred Genes
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Harrison, P.W.; Lower, R.P.J.; Kim, N.K.D.; Young, J.P.W. Introducing the Bacterial “Chromid”: Not a Chromosome, not a Plasmid. Trends Microbiol. 2010, 18, 141–148. [Google Scholar] [CrossRef]
- DiCenzo, G.C.; Finan, T.M. The Divided Bacterial Genome. Microbiol. Mol. Biol. Rev. 2017, 81, e00019-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Misra, H.S.; Maurya, G.K.; Kota, S.; Charaka, V.K. Maintenance of Multipartite Genome System and Its Functional Significance in Bacteria. J. Genet. 2018, 97, 1013–1038. [Google Scholar] [CrossRef] [PubMed]
- Almalki, F.; Choudhary, M.; Azad, R.K. Analysis of Multipartite Bacterial Genomes Using Alignment Free and Alignment-Based Pipelines. Arch. Microbiol. 2023, 205, 25. [Google Scholar] [CrossRef] [PubMed]
- Oren, A.; Garrity, G. Valid publication of the names of forty-two phyla of prokaryotes. Int. J. Syst. Evol. Microbiol. 2021, 71, 004851. [Google Scholar] [CrossRef]
- Egan, E.S.; Fogel, M.A.; Waldor, M.K. MicroReview: Divided Genomes: Negotiating the Cell Cycle in Prokaryotes with Multiple Chromosomes. Mol. Microbiol. 2005, 56, 1129–1138. [Google Scholar] [CrossRef]
- Choudhary, M.; Cho, H.; Bavishi, A.; Trahan, C.; Myagmarjav, B. Evolution of Multipartite Genomes in Prokaryotes. In Evolutionary Biology: Mechanisms and Trends; Pontarotti, P., Ed.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 301–323. [Google Scholar]
- Dicenzo, G.C.; Mengoni, A.; Perrin, E. Chromids Aid Genome Expansion and Functional Diversification in the Family Burkholderiaceae. Mol. Biol. Evol. 2019, 36, 562–574. [Google Scholar] [CrossRef] [PubMed]
- Galardini, M.; Pini, F.; Bazzicalupo, M.; Biondi, E.G.; Mengoni, A. Replicon-Dependent Bacterial Genome Evolution: The Case of Sinorhizobium Meliloti. Genome Biol. Evol. 2013, 5, 542–558. [Google Scholar] [CrossRef]
- diCenzo, G.C.; MacLean, A.M.; Milunovic, B.; Golding, G.B.; Finan, T.M. Examination of Prokaryotic Multipartite Genome Evolution through Experimental Genome Reduction. PLoS Genet. 2014, 10, e1004742. [Google Scholar] [CrossRef] [Green Version]
- Cooper, V.S.; Vohr, S.H.; Wrocklage, S.C.; Hatcher, P.J. Why Genes Evolve Faster on Secondary Chromosomes in Bacteria. PLoS Comput. Biol. 2010, 6, e1000732. [Google Scholar] [CrossRef]
- Feng, Z.; Zhang, Z.; Liu, Y.; Gu, J.; Cheng, Y.; Hu, W.; Li, Y.; Han, W. The Second Chromosome Promotes the Adaptation of the Genus Flammeovirga to Complex Environments. Microbiol. Spectr. 2021, 9, e00980-21. [Google Scholar] [CrossRef]
- Dryselius, R.; Izutsu, K.; Honda, T.; Iida, T. Differential Replication Dynamics for Large and Small Vibrio Chromosomes Affect Gene Dosage, Expression and Location. BMC Genom. 2008, 9, 559. [Google Scholar] [CrossRef] [Green Version]
- Couturier, E.; Rocha, E.P.C. Replication-Associated Gene Dosage Effects Shape the Genomes of Fast-Growing Bacteria but Only for Transcription and Translation Genes. Mol. Microbiol. 2006, 59, 1506–1518. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, P.; Chattoraj, D.K. Selective Chromosome Amplification in Vibrio Cholerae. Mol. Microbiol. 2007, 66, 1016–1028. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, T.; Jensen, R.B.; Skovgaard, O. The Two Chromosomes of Vibrio Cholerae Are Initiated at Different Time Points in the Cell Cycle. EMBO J. 2007, 26, 3124–3131. [Google Scholar] [CrossRef] [Green Version]
- Slater, S.C.; Goldman, B.S.; Goodner, B.; Setubal, J.C.; Farrand, S.K.; Nester, E.W.; Burr, T.J.; Banta, L.; Dickerman, A.W.; Paulsen, I.; et al. Genome Sequences of Three Agrobacterium Biovars Help Elucidate the Evolution of Multichromosome Genomes in Bacteria. J. Bacteriol. 2009, 191, 2501–2511. [Google Scholar] [CrossRef] [Green Version]
- Tettelin, H.; Masignani, V.; Cieslewicz, M.J.; Donati, C.; Medini, D.; Ward, N.L.; Angiuoli, S.V.; Crabtree, J.; Jones, A.L.; Durkin, A.S.; et al. Genome Analysis of Multiple Pathogenic Isolates of Streptococcus Agalactiae: Implications for the Microbial “Pan-Genome. ” Proc. Natl. Acad. Sci. USA 2005, 102, 13950–13955. [Google Scholar] [CrossRef] [Green Version]
- Tettelin, H.; Riley, D.; Cattuto, C.; Medini, D. Comparative Genomics: The Bacterial Pan-Genome. Curr. Opin. Microbiol. 2008, 11, 472–477. [Google Scholar] [CrossRef]
- Ikemura, T. Codon Usage and TRNA Content in Unicellular and Multicellular Organisms. Mol. Biol. Evol. 1985, 2, 13–34. [Google Scholar] [CrossRef]
- Plotkin, J.B.; Kudla, G. Synonymous but Not the Same. Natl. Rev. Genet. 2011, 12, 32–42. [Google Scholar] [CrossRef] [Green Version]
- Tuller, T.; Girshovich, Y.; Sella, Y.; Kreimer, A.; Freilich, S.; Kupiec, M.; Gophna, U.; Ruppin, E. Association between Translation Efficiency and Horizontal Gene Transfer within Microbial Communities. Nucleic Acids Res. 2011, 39, 4743–4755. [Google Scholar] [CrossRef] [Green Version]
- Komar, A.A. The Yin and Yang of Codon Usage. Hum. Mol. Genet. 2016, 25, R77–R85. [Google Scholar] [CrossRef]
- Tuller, T. Codon Bias, TRNA Pools, and Horizontal Gene Transfer. Mob. Genet. Elem. 2011, 1, 75–77. [Google Scholar] [CrossRef] [Green Version]
- Parks, D.H.; Chuvochina, M.; Rinke, C.; Mussig, A.J.; Chaumeil, P.A.; Hugenholtz, P. GTDB: An ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 2022, 50, D785–D794. [Google Scholar] [CrossRef]
- Liao, L.; Liu, C.; Zeng, Y.; Zhao, B.; Zhang, J.; Chen, B. Multipartite Genomes and the SRNome in Response to Temperature Stress of an Arctic Pseudoalteromonas Fuliginea BSW20308. Environ. Microbiol. 2019, 21, 272–285. [Google Scholar] [CrossRef] [Green Version]
- Xie, B.B.; Rong, J.C.; Tang, B.L.; Wang, S.; Liu, G.; Qin, Q.L.; Zhang, X.Y.; Zhang, W.; She, Q.; Chen, Y.; et al. Evolutionary Trajectory of the Replication Mode of Bacterial Replicons. MBio 2021, 12, e02745-20. [Google Scholar] [CrossRef]
- Fournes, F.; Val, M.E.; Skovgaard, O.; Mazel, D. Replicate Once per Cell Cycle: Replication Control of Secondary Chromosomes. Front. Microbiol. 2018, 9, 1833. [Google Scholar] [CrossRef] [Green Version]
- Heidelberg, J.F.; Elsen, J.A.; Nelson, W.C.; Clayton, R.A.; Gwinn, M.L.; Dodson, R.J.; Haft, D.H.; Hickey, E.K.; Peterson, J.D.; Umayam, L.; et al. DNA Sequence of Both Chromosomes of the Cholera Pathogen Vibrio Cholerae. Nature 2000, 406, 477–483. [Google Scholar] [CrossRef] [Green Version]
- Médigue, C.; Krin, E.; Pascal, G.; Barbe, V.; Bernsel, A.; Bertin, P.N.; Cheung, F.; Cruveiller, S.; D’Amico, S.; Duilio, A.; et al. Coping with Cold: The Genome of the Versatile Marine Antarctica Bacterium Pseudoalteromonas Haloplanktis TAC125. Genome Res. 2005, 15, 1325–1335. [Google Scholar] [CrossRef] [Green Version]
- Rong, J.C.; Liu, M.; Li, Y.; Sun, T.Y.; Pang, X.H.; Qin, Q.L.; Chen, X.L.; Xie, B. Bin Complete Genome Sequence of a Marine Bacterium with Two Chromosomes, Pseudoalteromonas Translucida KMM 520T. Mar. Genom. 2016, 26, 17–20. [Google Scholar] [CrossRef] [Green Version]
- Sonnenberg, C.B.; Haugen, P. The Pseudoalteromonas Multipartite Genome: Distribution and Expression of Pangene Categories, and a Hypothesis for the Origin and Evolution of the Chromid. G3 2021, 11, jkab256. [Google Scholar] [CrossRef]
- Kemter, F.S.; Messerschmidt, S.J.; Schallopp, N.; Sobetzko, P.; Lang, E.; Bunk, B.; Spröer, C.; Teschler, J.K.; Yildiz, F.H.; Overmann, J.; et al. Synchronous Termination of Replication of the Two Chromosomes Is an Evolutionary Selected Feature in Vibrionaceae. PLoS Genet. 2018, 14, e1007251. [Google Scholar] [CrossRef]
- Val, M.-E.; Marbouty, M.; de Lemos Martins, F.; Kennedy, S.P.; Kemble, H.; Bland, M.J.; Possoz, C.; Koszul, R.; Skovgaard, O.; Mazel, D. A Checkpoint Control Orchestrates the Replication of the Two Chromosomes of Vibrio Cholerae. Sci. Adv. 2016, 2, e1501914. [Google Scholar] [CrossRef] [Green Version]
- Sonnenberg, C.B.; Kahlke, T.; Haugen, P. Vibrionaceae Core, Shell and Cloud Genes Are Non-Randomly Distributed on Chr 1: An Hypothesis That Links the Genomic Location of Genes with Their Intracellular Placement. BMC Genom. 2020, 21, 695. [Google Scholar] [CrossRef]
- David, A.; Demarre, G.; Muresan, L.; Paly, E.; Barre, F.X.; Possoz, C. The Two Cis-Acting Sites, ParS1 and OriC1, Contribute to the Longitudinal Organisation of Vibrio Cholerae Chromosome I. PLoS Genet. 2014, 10, e1004448. [Google Scholar] [CrossRef] [Green Version]
- Fogel, M.A.; Waldor, M.K. Distinct Segregation Dynamics of the Two Vibrio Cholerae Chromosomes. Mol. Microbiol. 2005, 55, 125–136. [Google Scholar] [CrossRef]
- Jalal, A.S.; Tran, N.T.; Le, T.B. ParB Spreading on DNA Requires Cytidine Triphosphate in Vitro. Elife 2020, 20, e53515. [Google Scholar] [CrossRef]
- Zhu, Q.; Kosoy, M.; Dittmar, K. HGTector: An Automated Method Facilitating Genome-Wide Discovery of Putative Horizontal Gene Transfers. BMC Genom. 2014, 15, 717. [Google Scholar] [CrossRef] [Green Version]
- Medini, D.; Donati, C.; Tettelin, H.; Masignani, V.; Rappuoli, R. The Microbial Pan-Genome. Curr. Opin. Genet. Dev. 2005, 15, 589–594. [Google Scholar] [CrossRef]
- López, J.L.; Lozano, M.J.; Lagares, J.A.; Fabre, M.L.; Draghi, W.O.; Del Papa, M.F.; Pistorio, M.; Becker, A.; Wibberg, D.; Schlüter, A.; et al. Codon Usage Heterogeneity in the Multipartite Prokaryote Genome: Selection-Based Coding Bias Associated with Gene Location, Expression Level, and Ancestry. MBio 2019, 10, e00505-19. [Google Scholar] [CrossRef] [Green Version]
- Kopejtka, K.; Lin, Y.; Jakubovičová, M.; Koblízek, M.; Tomasch, J.; Moran, N. Clustered Core- And Pan-Genome Content on Rhodobacteraceae Chromosomes. Genome Biol. Evol. 2019, 11, 2208–2217. [Google Scholar] [CrossRef] [Green Version]
- Oliveira, P.H.; Touchon, M.; Cury, J.; Rocha, E.P.C. The Chromosomal Organization of Horizontal Gene Transfer in Bacteria. Nat. Commun. 2017, 8, 841. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mendler, K.; Chen, H.; Parks, D.H.; Lobb, B.; Hug, L.A.; Doxey, A.C. AnnoTree: Visualization and Exploration of a Functionally Annotated Microbial Tree of Life. Nucleic Acids Res. 2019, 47, 4442–4448. [Google Scholar] [CrossRef] [Green Version]
- López-Pérez, M.; Ramon-Marco, N.; Rodriguez-Valera, F. Networking in Microbes: Conjugative Elements and Plasmids in the Genus Alteromonas. BMC Genom. 2017, 18, 36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adam, Y.; Brezellec, P.; Espinosa, E.; Besombes, A.; Naquin, D.; Paly, E.; Possoz, C.; van Dijk, E.; Barre, F.X.; Ferat, J.L. Plesiomonas Shigelloides, an Atypical Enterobacterales with a Vibrio-Related Secondary Chromosome. Genome Biol. Evol. 2022, 14, evac011. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [Green Version]
- Hall, T.A. BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT.–ScienceOpen. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Stecher, G.; Tamura, K.; Kumar, S. Molecular Evolutionary Genetics Analysis (MEGA) for MacOS. Mol. Biol. Evol. 2020, 37, 1237–1239. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- O’leary, N.A.; Wright, M.W.; Brister, J.R.; Ciufo, S.; Haddad, D.; Mcveigh, R.; Rajput, B.; Robbertse, B.; Smith-White, B.; Ako-Adjei, D.; et al. Reference Sequence (RefSeq) Database at NCBI: Current Status, Taxonomic Expansion, and Functional Annotation. Nucleic Acids Res. 2015, 44, D733–D745. [Google Scholar] [CrossRef] [Green Version]
- Val, M.-E.; Soler-Bistué, A.; Bland, M.J.; Mazel, D. Management of Multipartite Genomes: The Vibrio Cholerae Model. Curr. Opin. Microbiol. 2014, 22, 120–126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid Annotations Using Subsystems Technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jain, C.; Rodriguez-R, L.M.; Phillippy, A.M.; Konstantinidis, K.T.; Aluru, S. High Throughput ANI Analysis of 90K Prokaryotic Genomes Reveals Clear Species Boundaries. Nat. Commun. 2018, 9, 5114. [Google Scholar] [CrossRef] [Green Version]
- Contreras-Moreira, B.; Vinuesa, P. GET_HOMOLOGUES, a Versatile Software Package for Scalable and Robust Microbial Pangenome Analysis. Appl. Environ. Microbiol. 2013, 79, 7696–7701. [Google Scholar] [CrossRef] [Green Version]
- Costa, S.S.; Guimarães, L.C.; Silva, A.; Soares, S.C.; Baraúna, R.A. First Steps in the Analysis of Prokaryotic Pan-Genomes. Bioinform Biol Insights 2020, 14, 1177932220938064. [Google Scholar] [CrossRef]
- Onofri, A. The Broken Bridge between Biologists and Statisticians: A Blog and R Package. 2020. Available online: https://www.statforbiology.com (accessed on 1 September 2022).
- Peden, J.F. Analysis of Codon Usage. Ph.D. Thesis, University of Nottingham, Nottingham, UK, 2000. [Google Scholar]
- Anwar, A.M.; Soudy, M. vhcub: Virus-Host Codon Usage Co-Adaptation Analysis. 2019. Available online: https://CRAN.r-project.org/package=vhcub (accessed on 1 September 2022).
- Wright, F. The “effective Number of Codons” Used in a Gene. Gene 1990, 87, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and Sensitive Protein Alignment Using DIAMOND. Nat. Methods 2014, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- RStudio Team. RStudio: Integrated Development for R. 2021. Available online: http://www.rstudio.com/ (accessed on 6 September 2022).






Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sonnenberg, C.B.; Haugen, P. Bipartite Genomes in Enterobacterales: Independent Origins of Chromids, Elevated Openness and Donors of Horizontally Transferred Genes. Int. J. Mol. Sci. 2023, 24, 4292. https://doi.org/10.3390/ijms24054292
Sonnenberg CB, Haugen P. Bipartite Genomes in Enterobacterales: Independent Origins of Chromids, Elevated Openness and Donors of Horizontally Transferred Genes. International Journal of Molecular Sciences. 2023; 24(5):4292. https://doi.org/10.3390/ijms24054292
Chicago/Turabian StyleSonnenberg, Cecilie Bækkedal, and Peik Haugen. 2023. "Bipartite Genomes in Enterobacterales: Independent Origins of Chromids, Elevated Openness and Donors of Horizontally Transferred Genes" International Journal of Molecular Sciences 24, no. 5: 4292. https://doi.org/10.3390/ijms24054292





