Genome-Wide Landscape of mRNAs, lncRNAs, and circRNAs during Testicular Development of Yak
Abstract
1. Introduction
2. Results
2.1. Overview of the Sequencing Data
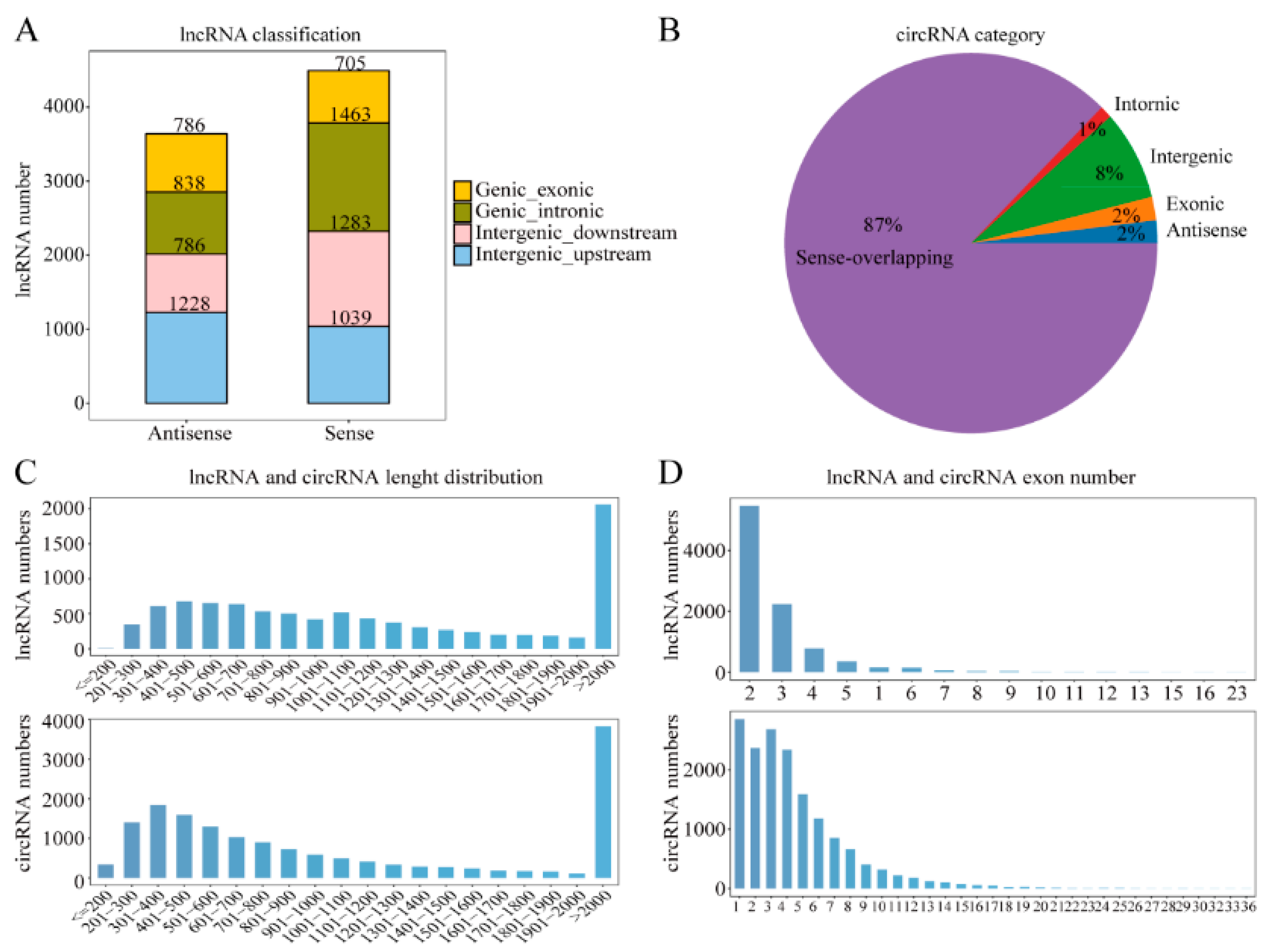
2.2. Predictions and Properties of ncRNAs in the Yak Testis
2.3. Differential Expression Analysis of mRNAs, lncRNAs, and circRNA
2.4. Gene ontology (GO) Analysis of DE Genes between M30 and M6
2.5. Gene Ontology (GO) Analysis of DE Genes between M18 and M6
2.6. Gene Ontology (GO) Analysis of DE Genes between M30 and M18
2.7. Function Analysis of Common DE Genes during Testicular Development
2.8. Construction of the lncRNA-mRNA Co-Expression Network
2.9. Validation of DE Genes by qRT-PCR
3. Discussion
4. Materials and Methods
4.1. Ethics Statement
4.2. Animals
4.3. RNA-Sequencing Data Analysis
4.4. Identification of lncRNA and circRNA
4.5. Analysis of Differentially Expressed (DE) Genes
4.6. Functional Enrichment Analysis
4.7. Co-Expression Network Construction
4.8. Gene Expression Validation by Quantitative Real-Time PCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| DE | differentially expressed |
| GO | Gene ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| qRT-PCR | Quantitative real-time PCR |
| M6 | 6-month-old |
| M18 | 18-month-old |
| M30 | 30-month-old |
| FPKM | fragments per kilobase of transcript per million reads mapped |
| SRPBM | spliced reads per billion mappings |
| LncRNAs | long non-coding RNAs |
| CircRNAs | circular RNAs |
References
- Qiu, Q.; Zhang, G.; Ma, T.; Qian, W.; Wang, J.; Ye, Z.; Cao, C.; Hu, Q.; Kim, J.; Larkin, D.M.; et al. The yak genome and adaptation to life at high altitude. Nat. Genet. 2012, 44, 946–949. [Google Scholar] [CrossRef] [PubMed]
- Ruan, C.M.; Wang, J.; Yang, Y.X.; Hu, J.J.; Ma, Y.J.; Zhang, Y.; Zhao, X.X. Proteomic analysis of Tianzhu White Yak (Bos grunniens) testis at different sexual developmental stages. Anim. Sci. J. 2019, 90, 333–343. [Google Scholar] [CrossRef] [PubMed]
- Prakash, B.S.; Sarkar, M.; Mondal, M. An update on reproduction in yak and mithun. Reprod. Domest. Anim. 2008, 43, 217–223. [Google Scholar] [CrossRef]
- Dahariya, S.; Paddibhatla, I.; Kumar, S.; Raghuwanshi, S.; Pallepati, A.; Gutti, R.K. Long non-coding RNA: Classification, biogenesis and functions in blood cells. Mol. Immunol. 2019, 112, 82–92. [Google Scholar] [CrossRef] [PubMed]
- Lorenzen, J.M.; Thum, T. Long noncoding RNAs in kidney and cardiovascular diseases. Nat. Rev. Nephrol. 2016, 12, 360–373. [Google Scholar] [CrossRef]
- Zhang, L.; Zhao, X.; Wang, W. lncRNA and mRNA sequencing of the left testis in experimental varicocele rats treated with Morinda officinalis polysaccharide. Exp. Ther. Med. 2021, 22, 1136. [Google Scholar] [CrossRef]
- Joshi, M.; Rajender, S. Long non-coding RNAs (lncRNAs) in spermatogenesis and male infertility. Reprod. Biol. Endocrinol. 2020, 18, 103. [Google Scholar] [CrossRef]
- Zhao, W.; Ahmed, S.; Ahmed, S.; Yangliu, Y.; Wang, H.; Cai, X. Analysis of long non-coding RNAs in epididymis of cattleyak associated with male infertility. Theriogenology 2021, 160, 61–71. [Google Scholar] [CrossRef]
- An, S.-Y.; Liu, Z.-F.; El-Samahy, M.A.; Deng, M.-T.; Gao, X.-X.; Liang, Y.-X.; Shi, C.-B.; Lei, Z.-H.; Wang, F.; Zhang, G.-M. LncRNA LOC102176306 plays important roles in goat testicular development. Reproduction 2021, 161, 523–537. [Google Scholar] [CrossRef]
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.V.W.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691. [Google Scholar] [CrossRef]
- Ebbesen, K.K.; Kjems, J.; Hansen, T.B. Circular RNAs: Identification, biogenesis and function. Biochim. Biophys. Acta 2016, 1859, 163–168. [Google Scholar] [CrossRef]
- Ou, Y.; Dores, C.; Rodriguez-Sosa, J.R.; van der Hoorn, F.A.; Dobrinski, I. Primary cilia in the developing pig testis. Cell Tissue Res. 2014, 358, 597–605. [Google Scholar] [CrossRef]
- Yan, P.; Pan, H.Q.; Zhi, D.J. Growth and development of testes in domesticated and hybrid (wild X domesticated) yak bulls. Anim. Sci. 2000, 70, 481–485. [Google Scholar]
- Zhu, Z.; Li, C.; Yang, S.; Tian, R.; Wang, J.; Yuan, Q.; Dong, H.; He, Z.; Wang, S.; Li, Z. Dynamics of the transcriptome during human spermatogenesis: Predicting the potential key genes regulating male gametes generation. Sci. Rep. 2016, 6, 19069. [Google Scholar] [CrossRef]
- Wang, J.; Ren, Q.L.; Hua, L.S.; Chen, J.F.; Zhang, J.Q.; Bai, H.J.; Li, H.L.; Xu, B.; Shi, Z.H.; Cao, H.; et al. Comprehensive analysis of differentially expressed mRNA, lncRNA and circRNA and their ceRNA networks in the longissimus dorsi muscle of two different pig breeds. Int. J. Mol. Sci. 2019, 20, 1107. [Google Scholar] [CrossRef]
- Liu, J.; Liu, T.; Wang, X.; He, A. Circles reshaping the RNA world: From waste to treasure. Mol. Cancer 2017, 16, 58. [Google Scholar] [CrossRef]
- Paquin, J.; Danalache, B.A.; Jankowski, M.; McCann, S.M.; Gutkowska, J. Oxytocin induces differentiation of P19 embryonic stem cells to cardiomyocytes. Proc. Natl. Acad. Sci. USA 2002, 99, 9550–9555. [Google Scholar] [CrossRef]
- Gao, Y.; Li, S.; Lai, Z.; Zhou, Z.; Wu, F.; Huang, Y.; Lan, X.; Lei, C.; Chen, H.; Dang, R. Analysis of long non-coding RNA and mRNA expression profiling in immature and mature bovine (Bos taurus) testes. Front. Genet. 2019, 10, 646. [Google Scholar] [CrossRef]
- Mäkelä, J.A.; Koskenniemi, J.J.; Virtanen, H.E.; Toppari, J. Testis development. Endocr. Rev. 2019, 40, 857–905. [Google Scholar] [CrossRef]
- Anguera, M.C.; Ma, W.; Clift, D.; Namekawa, S.; Lee, J.T.J.P.G. Tsx produces a long noncoding RNA and has general functions in the germline, stem cells, and brain. PLoS Genet. 2011, 7, e1002248. [Google Scholar] [CrossRef]
- Valentine, A.A.; Tao, S.X.; Lei, N.; Leslie, L.H. A Wt1-Dmrt1 transgene restores DMRT1 to sertoli cells of Dmrt1/testis: A novel model of DMRT1-Deficient germ cells. Biol. Reprod. 2013, 88, 51. [Google Scholar]
- Miller, W.L. Mitochondrial specificity of the early steps in steroidogenesis. J. Steroid Biochem. Mol. Biol. 1995, 55, 607–616. [Google Scholar] [CrossRef] [PubMed]
- Hu, M.C.; Hsu, H.J.; Guo, I.C.; Chung, B.C. Function of Cyp11a1 in animal models. Mol. Cell. Endocrinol. 2004, 215, 95–100. [Google Scholar] [CrossRef] [PubMed]
- Kyronlahti, A.; Euler, R.; Bielinska, M.; Schoeller, E.L.; Moley, K.H.; Toppari, J.; Heikinheimo, M.; Wilson, D.B. GATA4 regulates Sertoli cell function and fertility in adult male mice. Mol. Cell. Endocrinol. 2011, 333, 85–95. [Google Scholar] [CrossRef] [PubMed]
- Bielinska, M.; Seehra, A.; Toppari, J.; Heikinheimo, M.; Wilson, D.B. GATA-4 is required for sex steroidogenic cell development in the fetal mouse. Dev. Dyn. 2007, 236, 203–213. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Ghosh, S.; Chan, C.K.K. Analysis of RNA-Seq Data Using TopHat and Cufflinks. Methods Mol. Biol. 2016, 1374, 339–361. [Google Scholar]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Li, Y.; Zheng, Q.; Bao, C.; Li, S.; Guo, W.; Zhao, J.; Chen, D.; Gu, J.; He, X.; Huang, S. Circular RNA is enriched and stable in exosomes: A promising biomarker for cancer diagnosis. Cell Res. 2015, 25, 981–984. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Kanehisa, M.; Araki, M.; Goto, S.; Hattori, M.; Hirakawa, M.; Itoh, M.; Katayama, T.; Kawashima, S.; Okuda, S.; Tokimatsu, T. Kegg for linking genomes to life and the environment. Nucleic Acids Res. 2008, 36, D480–D484. [Google Scholar] [CrossRef]
- Saito, R.; Smoot, M.E.; Ono, K.; Ruscheinski, J.; Wang, P.L.; Lotia, S.; Pico, A.R.; Bader, G.D.; Ideker, T. A travel guide to Cytoscape plugins. Nat. Methods 2012, 9, 1069–1076. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(T) (-Delta Delta C) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]






| Sample | Raw Reads | Clean Reads | Q30 (%) | GC Content (%) | Mapped Reads | Uniquely Mapped |
|---|---|---|---|---|---|---|
| M18-1 | 101.07M | 99.83M | 95.11 | 48.28 | 89.95M (94.48%) | 80.21M (84.26%) |
| M18-2 | 91.07M | 89.98M | 95.09 | 48.33 | 81.17M (94.60%) | 72.12M (84.05%) |
| M18-4 | 97.24M | 96.07M | 95.14 | 48.87 | 86.66M (94.59%) | 76.68M (83.69%) |
| M6-2 | 88.87M | 87.66M | 94.89 | 47.93 | 78.59M (94.01%) | 70.38M (84.19%) |
| M6-3 | 91.57M | 90.39M | 94.83 | 46.47 | 81.29M (94.31%) | 74.12M (85.98%) |
| M6-4 | 97.52M | 96.20M | 94.68 | 47.49 | 86.61M (94.41%) | 78.23M (85.27%) |
| M30-1 | 100.51M | 99.26M | 94.96 | 48.20 | 89.58M (94.63%) | 79.83M (84.34%) |
| M30-2 | 103.01M | 101.81M | 95.06 | 48.13 | 92.03M (94.78%) | 81.81M (84.25%) |
| M30-3 | 99.81M | 98.60M | 95.05 | 48.06 | 89.03M (94.67%) | 79.24M (84.27%) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
La, Y.; Ma, X.; Bao, P.; Chu, M.; Yan, P.; Liang, C.; Guo, X. Genome-Wide Landscape of mRNAs, lncRNAs, and circRNAs during Testicular Development of Yak. Int. J. Mol. Sci. 2023, 24, 4420. https://doi.org/10.3390/ijms24054420
La Y, Ma X, Bao P, Chu M, Yan P, Liang C, Guo X. Genome-Wide Landscape of mRNAs, lncRNAs, and circRNAs during Testicular Development of Yak. International Journal of Molecular Sciences. 2023; 24(5):4420. https://doi.org/10.3390/ijms24054420
Chicago/Turabian StyleLa, Yongfu, Xiaoming Ma, Pengjia Bao, Min Chu, Ping Yan, Chunnian Liang, and Xian Guo. 2023. "Genome-Wide Landscape of mRNAs, lncRNAs, and circRNAs during Testicular Development of Yak" International Journal of Molecular Sciences 24, no. 5: 4420. https://doi.org/10.3390/ijms24054420
APA StyleLa, Y., Ma, X., Bao, P., Chu, M., Yan, P., Liang, C., & Guo, X. (2023). Genome-Wide Landscape of mRNAs, lncRNAs, and circRNAs during Testicular Development of Yak. International Journal of Molecular Sciences, 24(5), 4420. https://doi.org/10.3390/ijms24054420







