Bacillus subtilis RadA/Sms-Mediated Nascent Lagging-Strand Unwinding at Stalled or Reversed Forks Is a Two-Step Process: RadA/Sms Assists RecA Nucleation, and RecA Loads RadA/Sms
Abstract
1. Introduction
2. Results
2.1. RadA/Sms Partially Counters the Negative Effect of SsbA on RecA Nucleation onto cssDNA
2.2. The RecO-SsbA Mediator Poorly Counteracts the Negative Effect of RadA/Sms on RecA ATPase
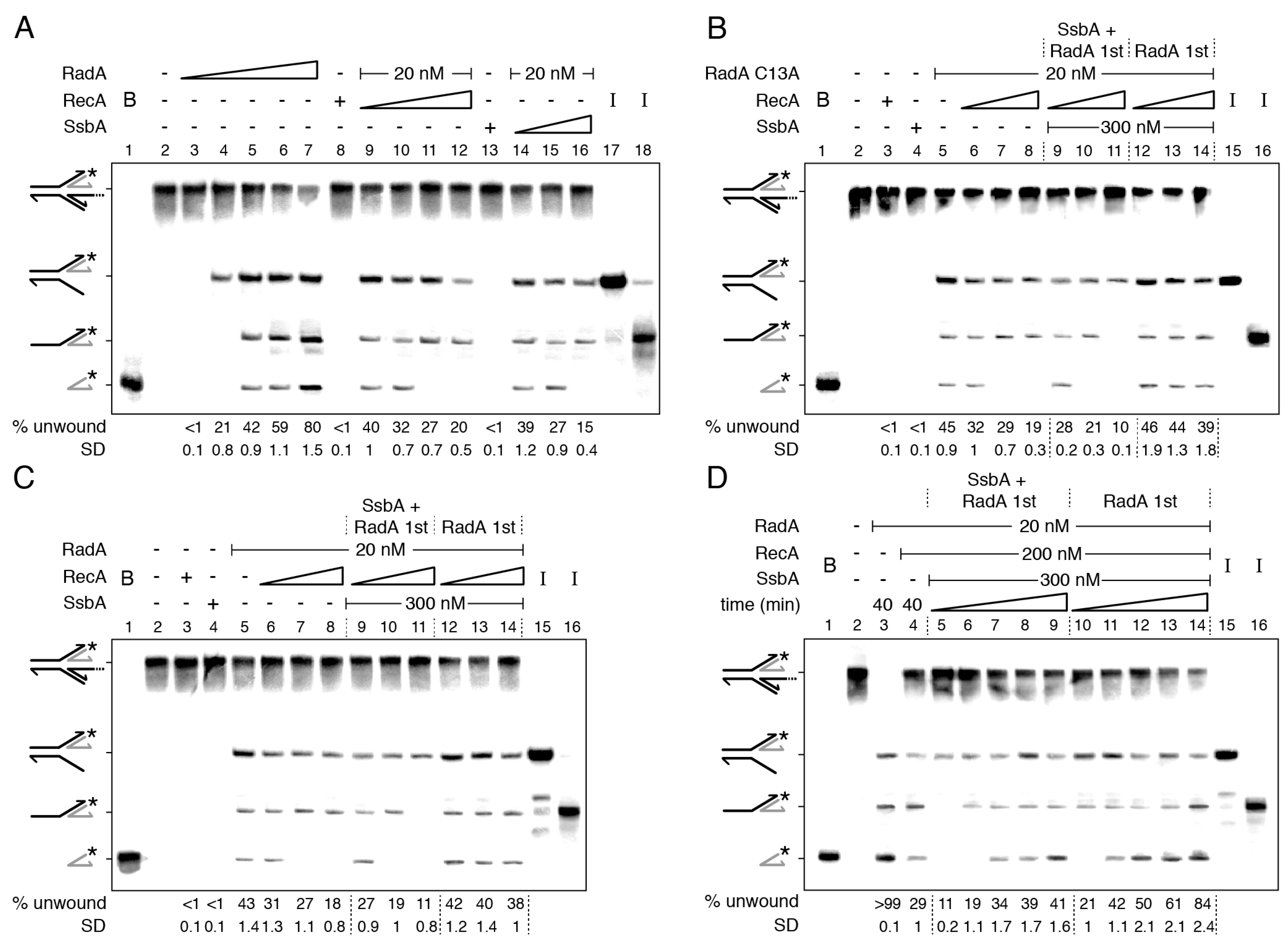
2.3. RadA/Sms Catalyzes Unwinding of Reversed Forks with Longer Nascent Lagging-Strand, but RecA and SsbA Reduce Unwinding
2.4. How RecA Activates RadA/Sms to Unwind a HJ Structure with a Nascent Leading-Strand Longer Than the Nascent Lagging-Strand
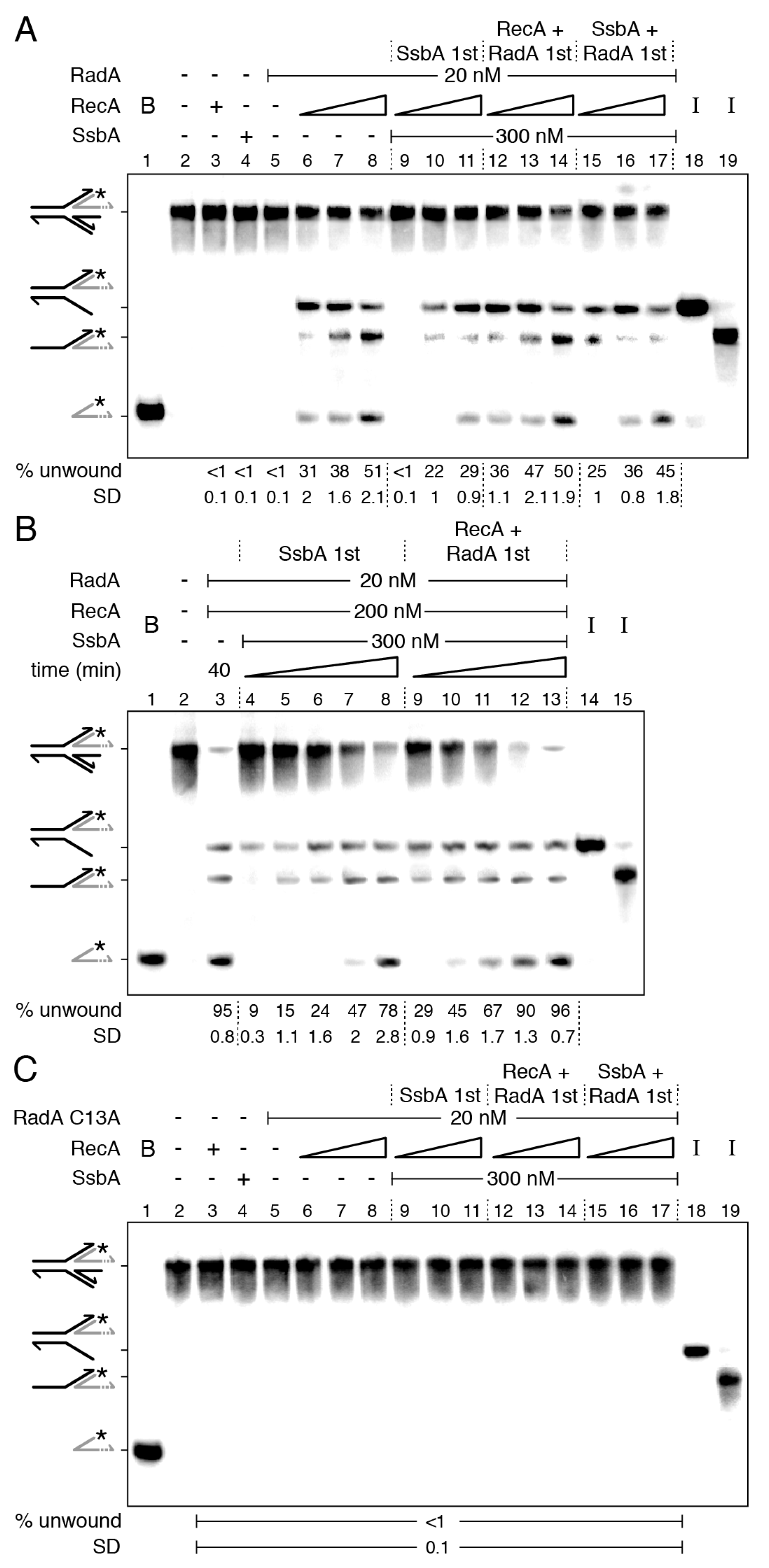
2.5. Two Step RadA/Sms Loading on the Nascent-Lagging Strand of HJ-Lead DNA
2.6. RecA Mediators Increase RecA Dynamics and Limit RadA/Sms Unwinding of Reversed Forks
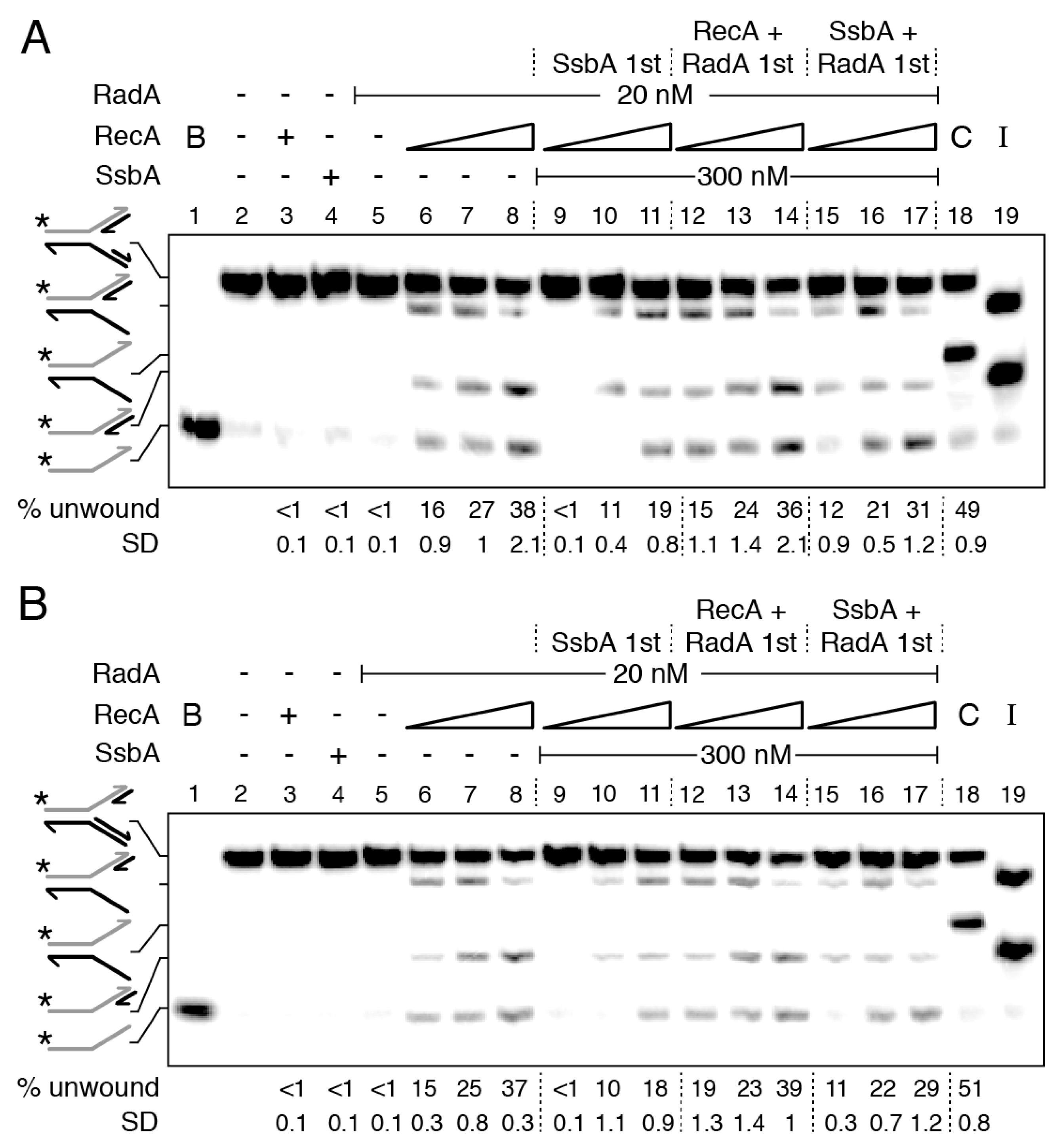
2.7. RecA Activates RadA/Sms Unwinding of the Nascent Lagging-Strand of Synthetic Stalled Forks
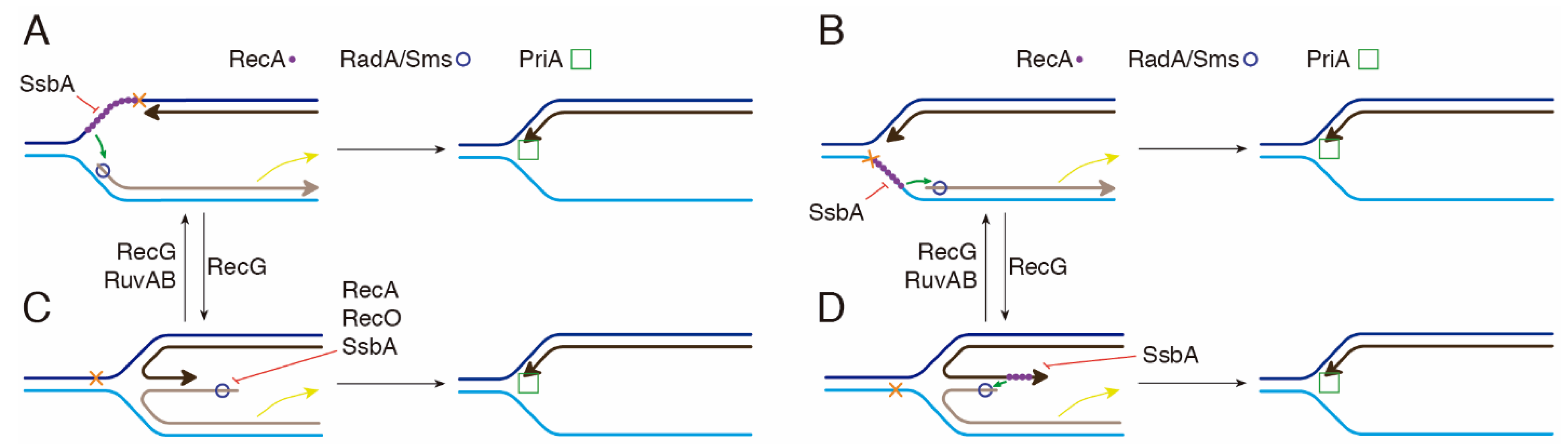
3. Discussion
4. Materials and Methods
4.1. Strains and Plasmids
4.2. Enzymes, Reagents, Protein and DNA Purification, Protein–Protein Interaction
4.3. ATP Hydrolysis Assays
4.4. DNA Unwinding Assays
4.5. RecA-Mediated DNA Strand Exchange
4.6. Protein–Protein Interaction Assays
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mirkin, E.V.; Mirkin, S.M. Replication fork stalling at natural impediments. Microbiol. Mol. Biol. Rev. 2007, 71, 13–35. [Google Scholar] [CrossRef] [PubMed]
- Zeman, M.K.; Cimprich, K.A. Causes and consequences of replication stress. Nat. Cell Biol. 2014, 16, 2–9. [Google Scholar] [CrossRef]
- Quinet, A.; Lemacon, D.; Vindigni, A. Replication fork reversal: Players and guardians. Mol. Cell 2017, 68, 830–833. [Google Scholar] [CrossRef] [PubMed]
- Gaillard, H.; Aguilera, A. Transcription as a threat to genome integrity. Annu. Rev. Biochem. 2016, 85, 291–317. [Google Scholar] [CrossRef]
- Marians, K.J. Lesion bypass and the reactivation of stalled replication forks. Annu. Rev. Biochem. 2018, 87, 217–238. [Google Scholar] [CrossRef]
- Cortez, D. Replication-Coupled DNA repair. Mol. Cell 2019, 74, 866–876. [Google Scholar] [CrossRef] [PubMed]
- Berti, M.; Cortez, D.; Lopes, M. The plasticity of DNA replication forks in response to clinically relevant genotoxic stress. Nat. Rev. Mol. Cell Biol. 2020, 21, 633–651. [Google Scholar] [CrossRef]
- Ghodke, H.; Paudel, B.P.; Lewis, J.S.; Jergic, S.; Gopal, K.; Romero, Z.J.; Wood, E.A.; Woodgate, R.; Cox, M.M.; van Oijen, A.M. Spatial and temporal organization of RecA in the Escherichia coli DNA-damage response. Elife 2019, 8, e42761. [Google Scholar] [CrossRef] [PubMed]
- Soubry, N.; Wang, A.; Reyes-Lamothe, R. Replisome activity slowdown after exposure to ultraviolet light in Escherichia coli. Proc. Natl. Acad. Sci. USA 2019, 116, 11747–11753. [Google Scholar] [CrossRef]
- Henrikus, S.S.; Henry, C.; Ghodke, H.; Wood, E.A.; Mbele, N.; Saxena, R.; Basu, U.; van Oijen, A.M.; Cox, M.M.; Robinson, A. RecFOR epistasis group: RecF and RecO have distinct localizations and functions in Escherichia coli. Nucleic Acids Res. 2019, 47, 2946–2965. [Google Scholar] [CrossRef]
- Rupp, W.D.; Howard-Flanders, P. Discontinuities in the DNA synthesized in an excision-defective strain of Escherichia coli following ultraviolet irradiation. J. Mol. Biol. 1968, 31, 291–304. [Google Scholar] [CrossRef] [PubMed]
- Courcelle, J.; Hanawalt, P.C. RecA-dependent recovery of arrested DNA replication forks. Annu. Rev. Genet. 2003, 37, 611–646. [Google Scholar] [CrossRef]
- Pages, V.; Fuchs, R.P. Uncoupling of leading- and lagging-strand DNA replication during lesion bypass in vivo. Science 2003, 300, 1300–1303. [Google Scholar] [CrossRef]
- Mangiameli, S.M.; Merrikh, C.N.; Wiggins, P.A.; Merrikh, H. Transcription leads to pervasive replisome instability in bacteria. eLife 2017, 6, e19848. [Google Scholar] [CrossRef]
- Atkinson, J.; McGlynn, P. Replication fork reversal and the maintenance of genome stability. Nucleic Acids Res. 2009, 37, 3475–3492. [Google Scholar] [CrossRef]
- De Septenville, A.L.; Duigou, S.; Boubakri, H.; Michel, B. Replication fork reversal after replication-transcription collision. PLoS Genet. 2012, 8, e1002622. [Google Scholar] [CrossRef] [PubMed]
- Lecointe, F.; Serena, C.; Velten, M.; Costes, A.; McGovern, S.; Meile, J.C.; Errington, J.; Ehrlich, S.D.; Noirot, P.; Polard, P. Anticipating chromosomal replication fork arrest: SSB targets repair DNA helicases to active forks. EMBO J. 2007, 26, 4239–4251. [Google Scholar] [CrossRef] [PubMed]
- Costes, A.; Lecointe, F.; McGovern, S.; Quevillon-Cheruel, S.; Polard, P. The C-terminal domain of the bacterial SSB protein acts as a DNA maintenance hub at active chromosome replication forks. PLoS Genet. 2010, 6, e1001238. [Google Scholar] [CrossRef]
- Simmons, L.A.; Grossman, A.D.; Walker, G.C. Replication is required for the RecA localization response to DNA damage in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 2007, 104, 1360–1365. [Google Scholar] [CrossRef]
- Bernard, R.; Marquis, K.A.; Rudner, D.Z. Nucleoid occlusion prevents cell division during replication fork arrest in Bacillus subtilis. Mol. Microbiol. 2010, 78, 866–882. [Google Scholar] [CrossRef]
- Wang, J.D.; Sanders, G.M.; Grossman, A.D. Nutritional control of elongation of DNA replication by (p)ppGpp. Cell 2007, 128, 865–875. [Google Scholar] [CrossRef]
- Lenhart, J.S.; Brandes, E.R.; Schroeder, J.W.; Sorenson, R.J.; Showalter, H.D.; Simmons, L.A. RecO and RecR are necessary for RecA loading in response to DNA damage and replication fork stress. J. Bacteriol. 2014, 196, 2851–2860. [Google Scholar] [CrossRef]
- Million-Weaver, S.; Samadpour, A.N.; Merrikh, H. Replication restart after replication-transcription conflicts requires RecA in Bacillus subtilis. J. Bacteriol. 2015, 197, 2374–2382. [Google Scholar] [CrossRef]
- Vlasic, I.; Mertens, R.; Seco, E.M.; Carrasco, B.; Ayora, S.; Reitz, G.; Commichau, F.M.; Alonso, J.C.; Moeller, R. Bacillus subtilis RecA and its accessory factors, RecF, RecO, RecR and RecX, are required for spore resistance to DNA double-strand break. Nucleic Acids Res. 2014, 42, 2295–2307. [Google Scholar] [CrossRef] [PubMed]
- Carrasco, B.; Seco, E.M.; Lopez-Sanz, M.; Alonso, J.C.; Ayora, S. Bacillus subtilis RarA modulates replication restart. Nucleic Acids Res. 2018, 46, 7206–7220. [Google Scholar] [CrossRef] [PubMed]
- Gándara, C.; Torres, R.; Carrasco, B.; Ayora, S.; Alonso, J.C. DisA restrains the processing and cleavage of reversed replication forks by the RuvAB-RecU resolvasome. Int. J. Mol. Sci. 2021, 22, 11323. [Google Scholar] [CrossRef]
- Ramos, C.; Hernandez-Tamayo, R.; Lopez-Sanz, M.; Carrasco, B.; Serrano, E.; Alonso, J.C.; Graumann, P.L.; Ayora, S. The RecD2 helicase balances RecA activities. Nucleic Acids Res. 2022, 50, 3432–3444. [Google Scholar] [CrossRef]
- Yadav, T.; Carrasco, B.; Serrano, E.; Alonso, J.C. Roles of Bacillus subtilis DprA and SsbA in RecA-mediated genetic recombination. J. Biol. Chem. 2014, 289, 27640–27652. [Google Scholar] [CrossRef] [PubMed]
- Carrasco, B.; Yadav, T.; Serrano, E.; Alonso, J.C. Bacillus subtilis RecO and SsbA are crucial for RecA-mediated recombinational DNA repair. Nucleic Acids Res. 2015, 43, 5984–5997. [Google Scholar] [CrossRef]
- Manfredi, C.; Carrasco, B.; Ayora, S.; Alonso, J.C. Bacillus subtilis RecO nucleates RecA onto SsbA-coated single-stranded DNA. J. Biol. Chem. 2008, 283, 24837–24847. [Google Scholar] [CrossRef] [PubMed]
- Kidane, D.; Carrasco, B.; Manfredi, C.; Rothmaier, K.; Ayora, S.; Tadesse, S.; Alonso, J.C.; Graumann, P.L. Evidence for different pathways during horizontal gene transfer in competent Bacillus subtilis cells. PLoS Genet. 2009, 5, e1000630. [Google Scholar] [CrossRef] [PubMed]
- Marie, L.; Rapisarda, C.; Morales, V.; Berge, M.; Perry, T.; Soulet, A.L.; Gruget, C.; Remaut, H.; Fronzes, R.; Polard, P. Bacterial RadA is a DnaB-type helicase interacting with RecA to promote bidirectional D-loop extension. Nat. Commun. 2017, 8, 15638. [Google Scholar] [CrossRef] [PubMed]
- Torres, R.; Gándara, C.; Carrasco, B.; Baquedano, I.; Ayora, S.; Alonso, J.C. DisA limits RecG activities at stalled or reversed replication forks. Cells 2021, 10, 1357. [Google Scholar] [CrossRef]
- Torres, R.; Serrano, E.; Alonso, J.C. Bacillus subtilis RecA interacts with and loads RadA/Sms to unwind recombination intermediates during natural chromosomal transformation. Nucleic Acids Res. 2019, 47, 9198–9215. [Google Scholar] [PubMed]
- Slade, D.; Lindner, A.B.; Paul, G.; Radman, M. Recombination and replication in DNA repair of heavily irradiated Deinococcus radiodurans. Cell 2009, 136, 1044–1055. [Google Scholar] [CrossRef]
- Inoue, M.; Fukui, K.; Fujii, Y.; Nakagawa, N.; Yano, T.; Kuramitsu, S.; Masui, R. The Lon protease-like domain in the bacterial RecA paralog RadA is required for DNA binding and repair. J. Biol. Chem. 2017, 292, 9801–9814. [Google Scholar] [CrossRef]
- Botos, I.; Lountos, G.T.; Wu, W.; Cherry, S.; Ghirlando, R.; Kudzhaev, A.M.; Rotanova, T.V.; de Val, N.; Tropea, J.E.; Gustchina, A.; et al. Cryo-EM structure of substrate-free E. coli Lon protease provides insights into the dynamics of Lon machinery. Curr. Res. Struct. Biol. 2019, 1, 13–20. [Google Scholar] [CrossRef]
- Gándara, C.; de Lucena, D.K.C.; Torres, R.; Serrano, E.; Altenburger, S.; Graumann, P.L.; Alonso, J.C. Activity and in vivo dynamics of Bacillus subtilis DisA are affected by RadA/Sms and by Holliday junction-processing proteins. DNA Repair 2017, 55, 17–30. [Google Scholar] [CrossRef] [PubMed]
- Raguse, M.; Torres, R.; Seco, E.M.; Gándara, C.; Ayora, S.; Moeller, R.; Alonso, J.C. Bacillus subtilis DisA helps to circumvent replicative stress during spore revival. DNA Repair 2017, 59, 57–68. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, H.; Carrasco, B.; Cozar, M.C.; Alonso, J.C. Bacillus subtilis RecG branch migration translocase is required for DNA repair and chromosomal segregation. Mol. Microbiol. 2007, 65, 920–935. [Google Scholar] [CrossRef]
- Ayora, S.; Carrasco, B.; Cárdenas, P.P.; César, C.E.; Cañas, C.; Yadav, T.; Marchisone, C.; Alonso, J.C. Double-strand break repair in bacteria: A view from Bacillus subtilis. FEMS Microbiol. Rev. 2011, 35, 1055–1081. [Google Scholar] [CrossRef]
- Torres, R.; Serrano, E.; Tramm, K.; Alonso, J.C. Bacillus subtilis RadA/Sms contributes to chromosomal transformation and DNA repair in concert with RecA and circumvents replicative stress in concert with DisA. DNA Repair 2019, 77, 45–57. [Google Scholar] [CrossRef] [PubMed]
- Cooper, D.L.; Lovett, S.T. Recombinational branch migration by the RadA/Sms paralog of RecA in Escherichia coli. eLife 2016, 5, e10807. [Google Scholar] [CrossRef]
- Hertzog, M.; Perry, T.N.; Dupaigne, P.; Serres, S.; Morales, V.; Soulet, A.L.; Bell, J.C.; Margeat, E.; Kowalczykowski, S.C.; Le Cam, E.; et al. Assembly mechanism and cryoEM structure of RecA recombination nucleofilaments from Streptococcus pneumoniae. bioRxiv 2022. [Google Scholar] [CrossRef]
- Torres, R.; Alonso, J.C. Bacillus subtilis RecA, DisA, and RadA/Sms interplay prevents replication stress by regulating fork remodeling. Front. Microbiol. 2021, 12, 766897. [Google Scholar] [CrossRef] [PubMed]
- Windgassen, T.A.; Wessel, S.R.; Bhattacharyya, B.; Keck, J.L. Mechanisms of bacterial DNA replication restart. Nucleic Acids Res. 2018, 46, 504–519. [Google Scholar] [CrossRef]
- Yadav, T.; Carrasco, B.; Myers, A.R.; George, N.P.; Keck, J.L.; Alonso, J.C. Genetic recombination in Bacillus subtilis: A division of labor between two single-strand DNA-binding proteins. Nucleic Acids Res. 2012, 40, 5546–5559. [Google Scholar] [CrossRef] [PubMed]
- Bianco, P.R.; Lu, Y. Single-molecule insight into stalled replication fork rescue in Escherichia coli. Nucleic Acids Res. 2021, 49, 4220–4238. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, P.; Camerini-Otero, C.S.; Camerini-Otero, R.D. The synapsis event in the homologous pairing of DNAs: RecA recognizes and pairs less than one helical repeat of DNA. Proc. Natl. Acad. Sci. USA 1992, 89, 6492–6496. [Google Scholar] [CrossRef]
- Sussman, R.; Sharma, S.K.; Kuzirian, A. Catalytic activities of recA protein are dependent on the lattice length of the single-strand DNA ligand. Cell Cycle 2008, 7, 89–95. [Google Scholar] [CrossRef]
- Yang, H.; Zhou, C.; Dhar, A.; Pavletich, N.P. Mechanism of strand exchange from RecA-DNA synaptic and D-loop structures. Nature 2020, 586, 801–806. [Google Scholar] [CrossRef]
- Cañas, C.; Suzuki, Y.; Marchisone, C.; Carrasco, B.; Freire-Beneitez, V.; Takeyasu, K.; Alonso, J.C.; Ayora, S. Interaction of branch migration translocases with the Holliday junction-resolving enzyme and their implications in Holliday junction resolution. J. Biol. Chem. 2014, 289, 17634–17646. [Google Scholar] [CrossRef]
- Chen, Z.; Yang, H.; Pavletich, N.P. Mechanism of homologous recombination from the RecA-ssDNA/dsDNA structures. Nature 2008, 453, 489–494. [Google Scholar] [CrossRef]
- Arias-Palomo, E.; O’Shea, V.L.; Hood, I.V.; Berger, J.M. The bacterial DnaC helicase loader is a DnaB ring breaker. Cell 2013, 153, 438–448. [Google Scholar] [CrossRef] [PubMed]
- Kelch, B.A. Review: The lord of the rings: Structure and mechanism of the sliding clamp loader. Biopolymers 2016, 105, 532–546. [Google Scholar] [CrossRef]
- Joo, C.; McKinney, S.A.; Nakamura, M.; Rasnik, I.; Myong, S.; Ha, T. Real-time observation of RecA filament dynamics with single monomer resolution. Cell 2006, 126, 515–527. [Google Scholar] [CrossRef]
- Carrasco, B.; Cozar, M.C.; Lurz, R.; Alonso, J.C.; Ayora, S. Genetic recombination in Bacillus subtilis 168: Contribution of Holliday junction-processing functions in chromosome segregation. J. Bacteriol. 2004, 186, 5557–5566. [Google Scholar] [CrossRef] [PubMed]
- Bell, J.C.; Kowalczykowski, S.C. RecA: Regulation and Mechanism of a Molecular Search Engine. Trends Biochem. Sci. 2016, 41, 491–507. [Google Scholar] [CrossRef]
- Ayora, S.; Carrasco, B.; Doncel-Perez, E.; Lurz, R.; Alonso, J.C. Bacillus subtilis RecU protein cleaves Holliday junctions and anneals single-stranded DNA. Proc. Natl. Acad. Sci. USA 2004, 101, 452–457. [Google Scholar] [CrossRef]
- Le, S.; Serrano, E.; Kawamura, R.; Carrasco, B.; Yan, J.; Alonso, J.C. Bacillus subtilis RecA with DprA-SsbA antagonizes RecX function during natural transformation. Nucleic Acids Res. 2017, 45, 8873–8885. [Google Scholar] [CrossRef] [PubMed]
- Serrano, E.; Carrasco, B.; Gilmore, J.L.; Takeyasu, K.; Alonso, J.C. RecA regulation by RecU and DprA during Bacillus subtilis natural plasmid transformation. Front. Microbiol. 2018, 9, 1514. [Google Scholar] [CrossRef] [PubMed]
- Gassel, M.; Alonso, J.C. Expression of the recE gene during induction of the SOS response in Bacillus subtilis recombination-deficient strains. Mol. Microbiol. 1989, 3, 1269–1276. [Google Scholar] [CrossRef] [PubMed]
- Carrasco, B.; Manfredi, C.; Ayora, S.; Alonso, J.C. Bacillus subtilis SsbA and dATP regulate RecA nucleation onto single-stranded DNA. DNA Repair 2008, 7, 990–996. [Google Scholar] [CrossRef]
- Carrasco, B.; Ayora, S.; Lurz, R.; Alonso, J.C. Bacillus subtilis RecU Holliday-junction resolvase modulates RecA activities. Nucleic Acids Res. 2005, 33, 3942–3952. [Google Scholar] [CrossRef] [PubMed]
- Ayora, S.; Missich, R.; Mesa, P.; Lurz, R.; Yang, S.; Egelman, E.H.; Alonso, J.C. Homologous-pairing activity of the Bacillus subtilis bacteriophage SPP1 replication protein G35P. J. Biol. Chem. 2002, 277, 35969–35979. [Google Scholar] [CrossRef]
- Carrasco, B.; Moreno-Del Alamo, M.; Torres, R.; Alonso, J.C. PcrA Dissociates RecA filaments and the SsbA and RecO mediators counterbalance such activity. Front. Mol. Biosci. 2022, 9, 836211. [Google Scholar] [CrossRef]
- Moreno-Del Alamo, M.; Carrasco, B.; Torres, R.; Alonso, J.C. Bacillus subtilis PcrA helicase removes trafficking barriers. Cells 2021, 10, 935. [Google Scholar] [CrossRef]
- Lovett, C.M., Jr.; Roberts, J.W. Purification of a RecA protein analogue from Bacillus subtilis. J. Biol. Chem. 1985, 260, 3305–3313. [Google Scholar] [CrossRef]
- Steffen, S.E.; Katz, F.S.; Bryant, F.R. Complete inhibition of Streptococcus pneumoniae RecA protein catalyzed ATP hydrolysis by single-stranded DNA-binding protein (SSB protein): Implications for the mechanism of SSB protein-stimulated DNA strand exchange. J. Biol. Chem. 2002, 277, 14493–14500. [Google Scholar] [CrossRef]
- Robu, M.E.; Inman, R.B.; Cox, M.M. RecA protein promotes the regression of stalled replication forks in vitro. Proc. Natl. Acad. Sci. USA 2001, 98, 8211–8218. [Google Scholar] [CrossRef]
- Soultanas, P. Loading mechanisms of ring helicases at replication origins. Mol. Microbiol. 2012, 84, 6–16. [Google Scholar] [CrossRef] [PubMed]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Torres, R.; Carrasco, B.; Alonso, J.C. Bacillus subtilis RadA/Sms-Mediated Nascent Lagging-Strand Unwinding at Stalled or Reversed Forks Is a Two-Step Process: RadA/Sms Assists RecA Nucleation, and RecA Loads RadA/Sms. Int. J. Mol. Sci. 2023, 24, 4536. https://doi.org/10.3390/ijms24054536
Torres R, Carrasco B, Alonso JC. Bacillus subtilis RadA/Sms-Mediated Nascent Lagging-Strand Unwinding at Stalled or Reversed Forks Is a Two-Step Process: RadA/Sms Assists RecA Nucleation, and RecA Loads RadA/Sms. International Journal of Molecular Sciences. 2023; 24(5):4536. https://doi.org/10.3390/ijms24054536
Chicago/Turabian StyleTorres, Rubén, Begoña Carrasco, and Juan C. Alonso. 2023. "Bacillus subtilis RadA/Sms-Mediated Nascent Lagging-Strand Unwinding at Stalled or Reversed Forks Is a Two-Step Process: RadA/Sms Assists RecA Nucleation, and RecA Loads RadA/Sms" International Journal of Molecular Sciences 24, no. 5: 4536. https://doi.org/10.3390/ijms24054536
APA StyleTorres, R., Carrasco, B., & Alonso, J. C. (2023). Bacillus subtilis RadA/Sms-Mediated Nascent Lagging-Strand Unwinding at Stalled or Reversed Forks Is a Two-Step Process: RadA/Sms Assists RecA Nucleation, and RecA Loads RadA/Sms. International Journal of Molecular Sciences, 24(5), 4536. https://doi.org/10.3390/ijms24054536







