Comprehensive Analysis of the INDETERMINATE DOMAIN (IDD) Gene Family and Their Response to Abiotic Stress in Zea mays
Abstract
:1. Introduction
2. Results
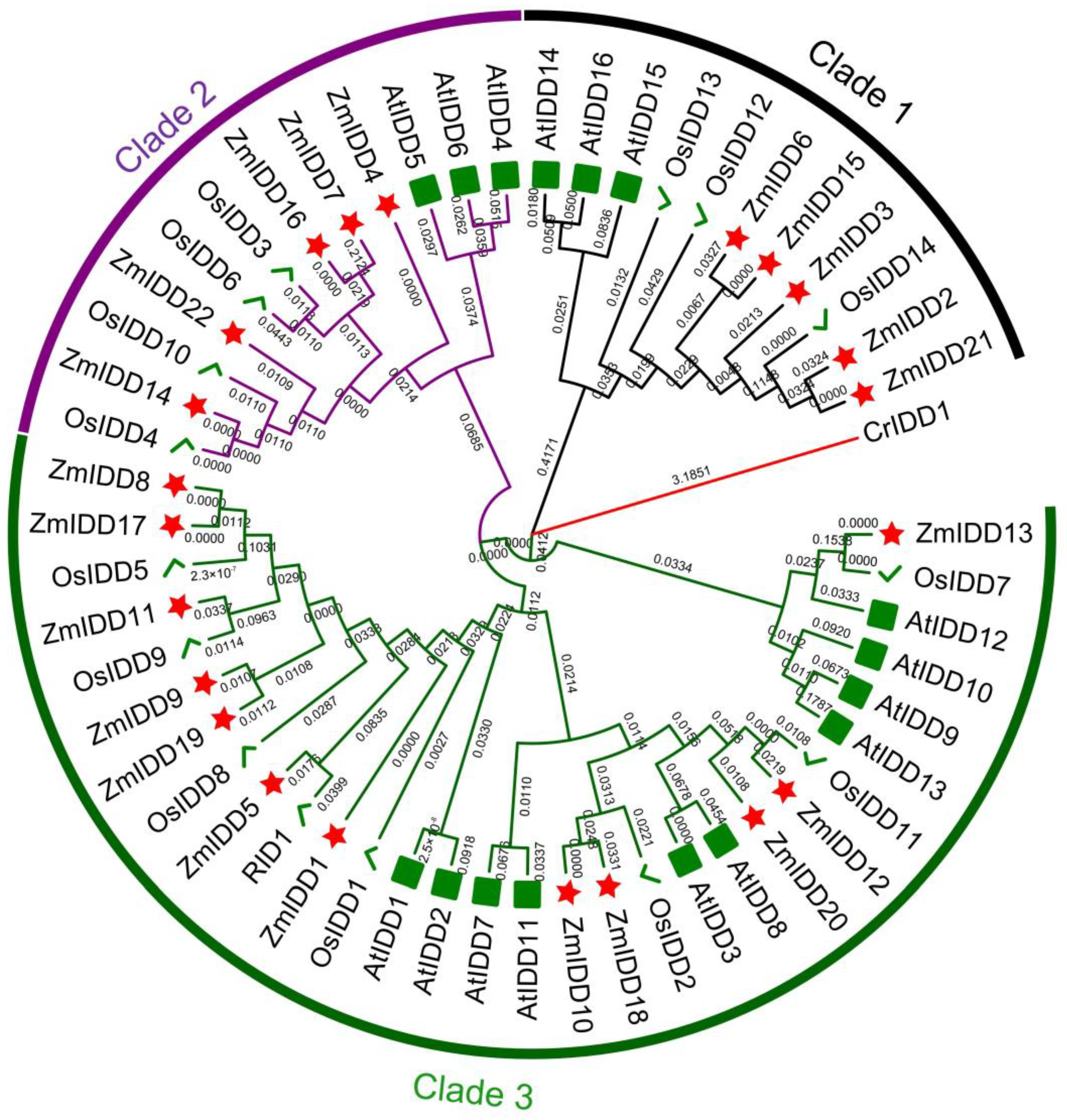
2.1. Identification and Evolution Analysis of ZmIDD Family Genes in Zea mays
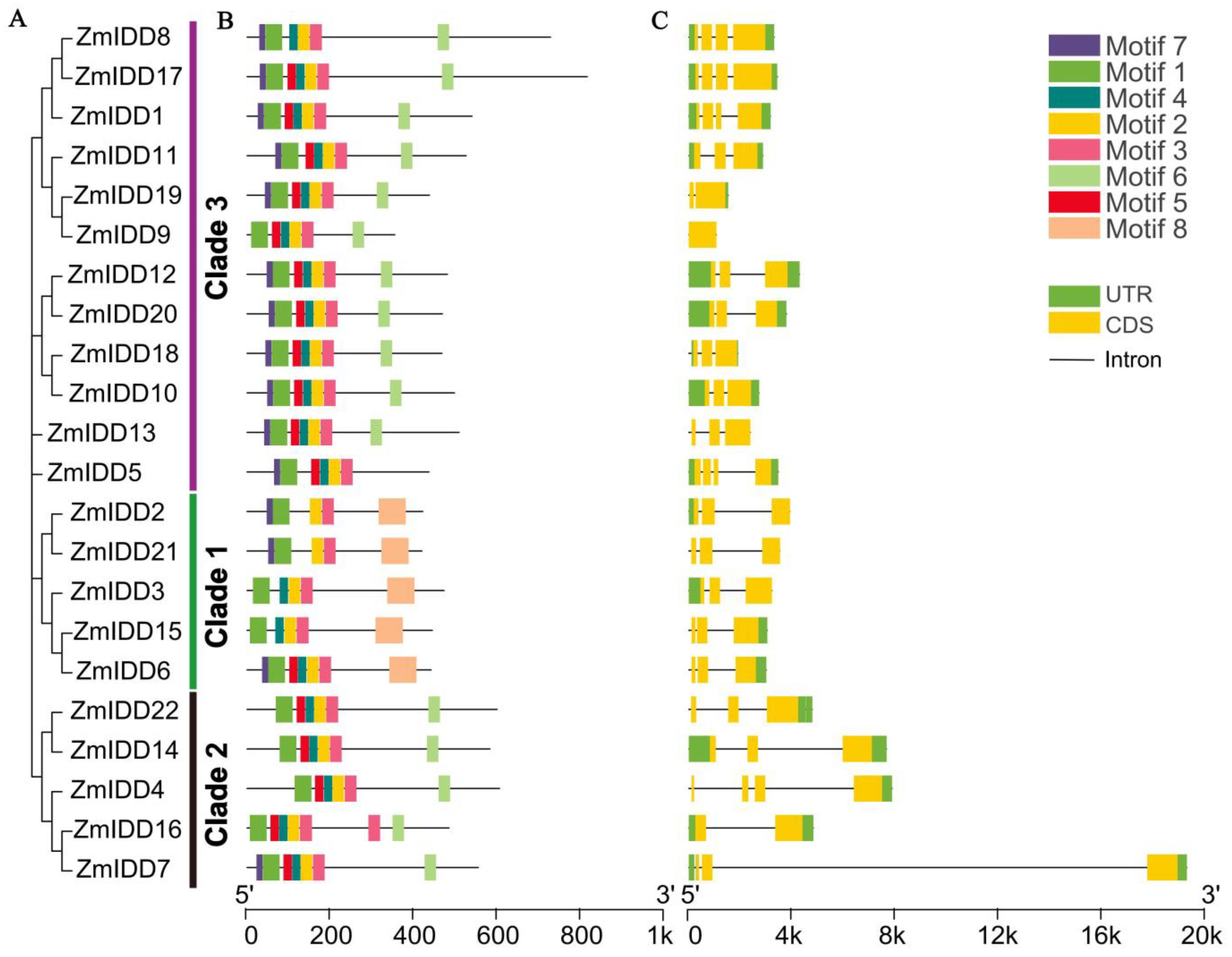
2.2. Structure Analysis of ZmIDD
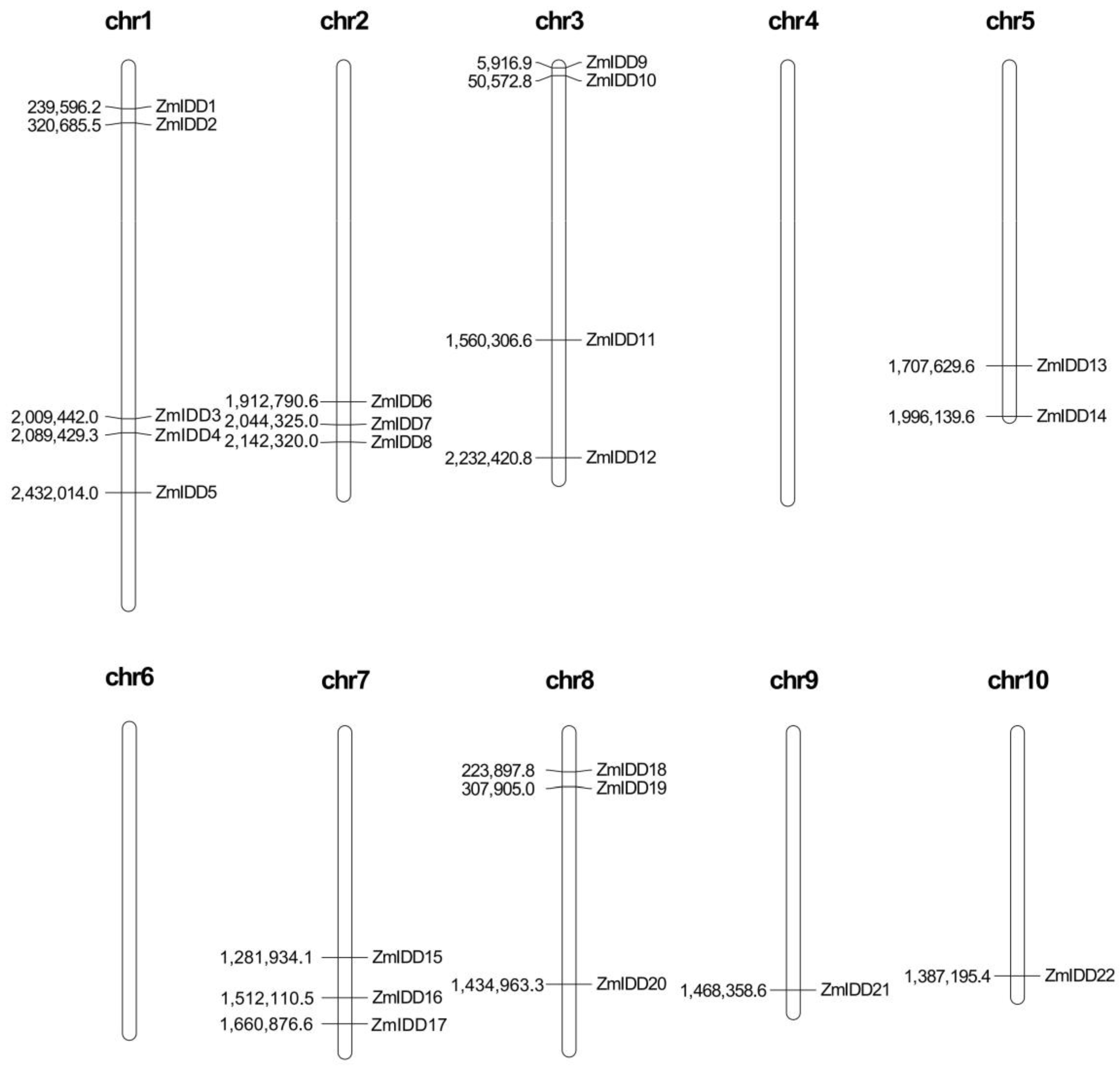
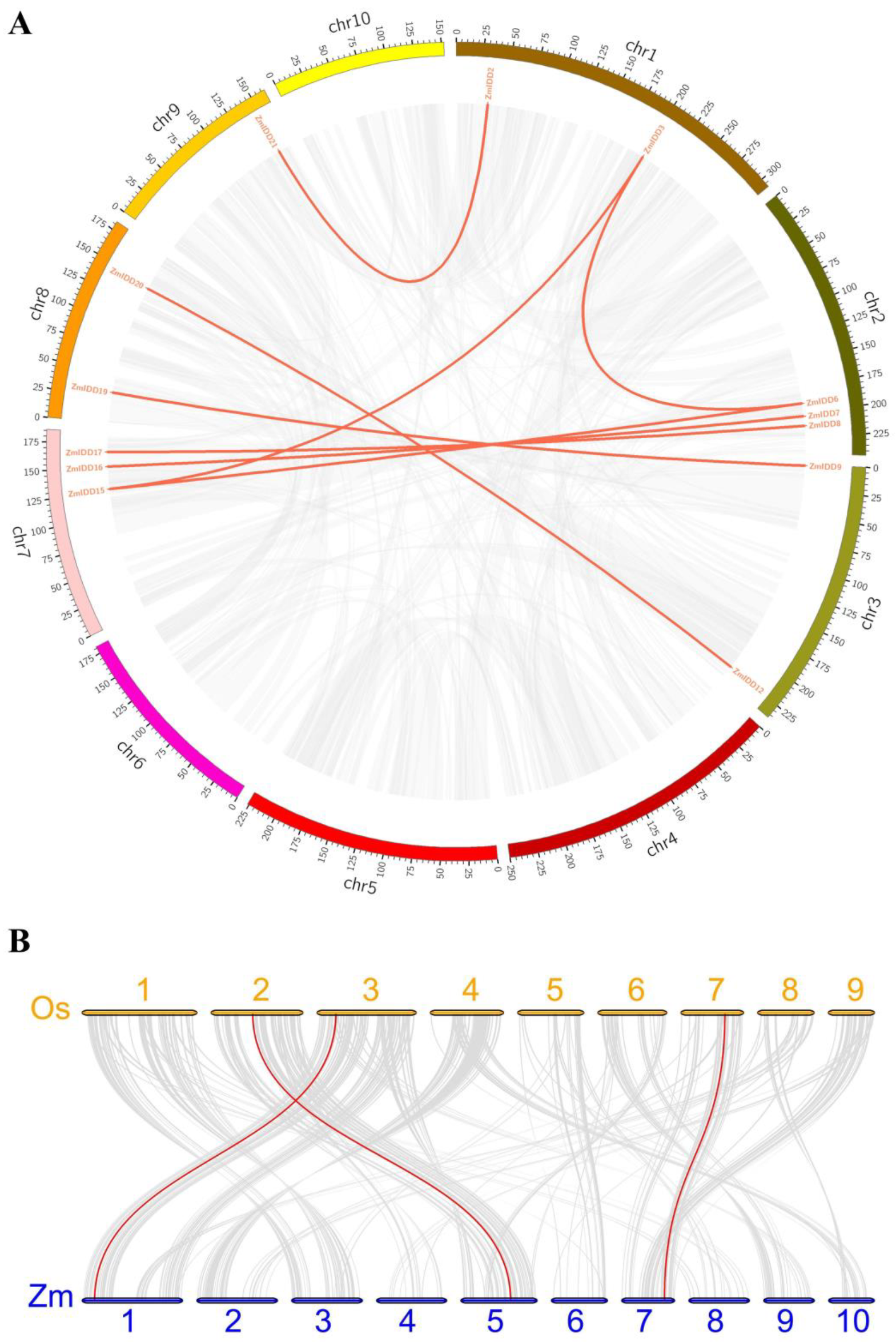
2.3. Location and Duplication of ZmIDDs
2.4. Cis-Elements Analysis in ZmIDDs Promoters
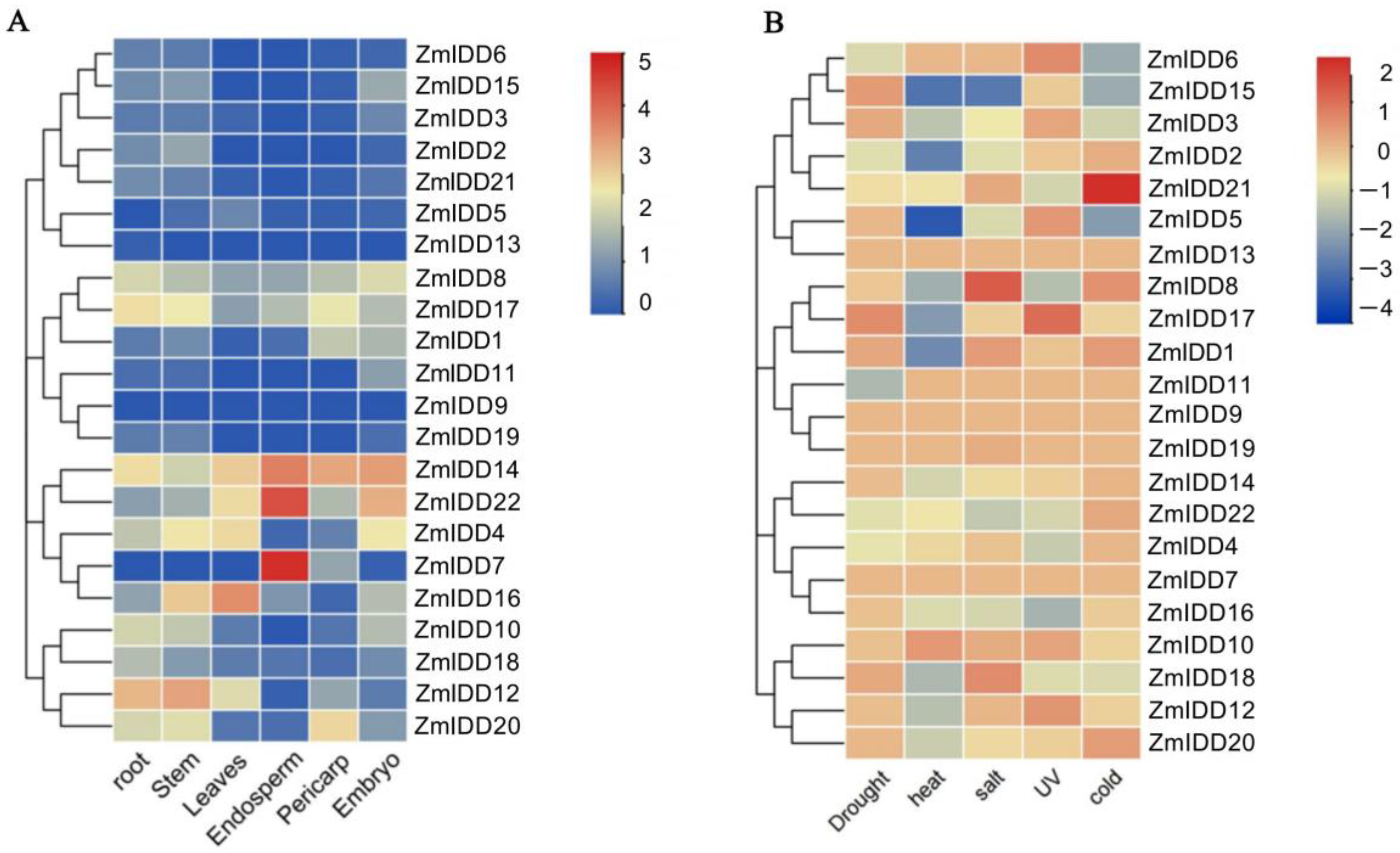
2.5. Tissue-Specific Expression Patterns of ZmIDDs
2.6. Stress-Induced Expression Patterns of ZmIDDs
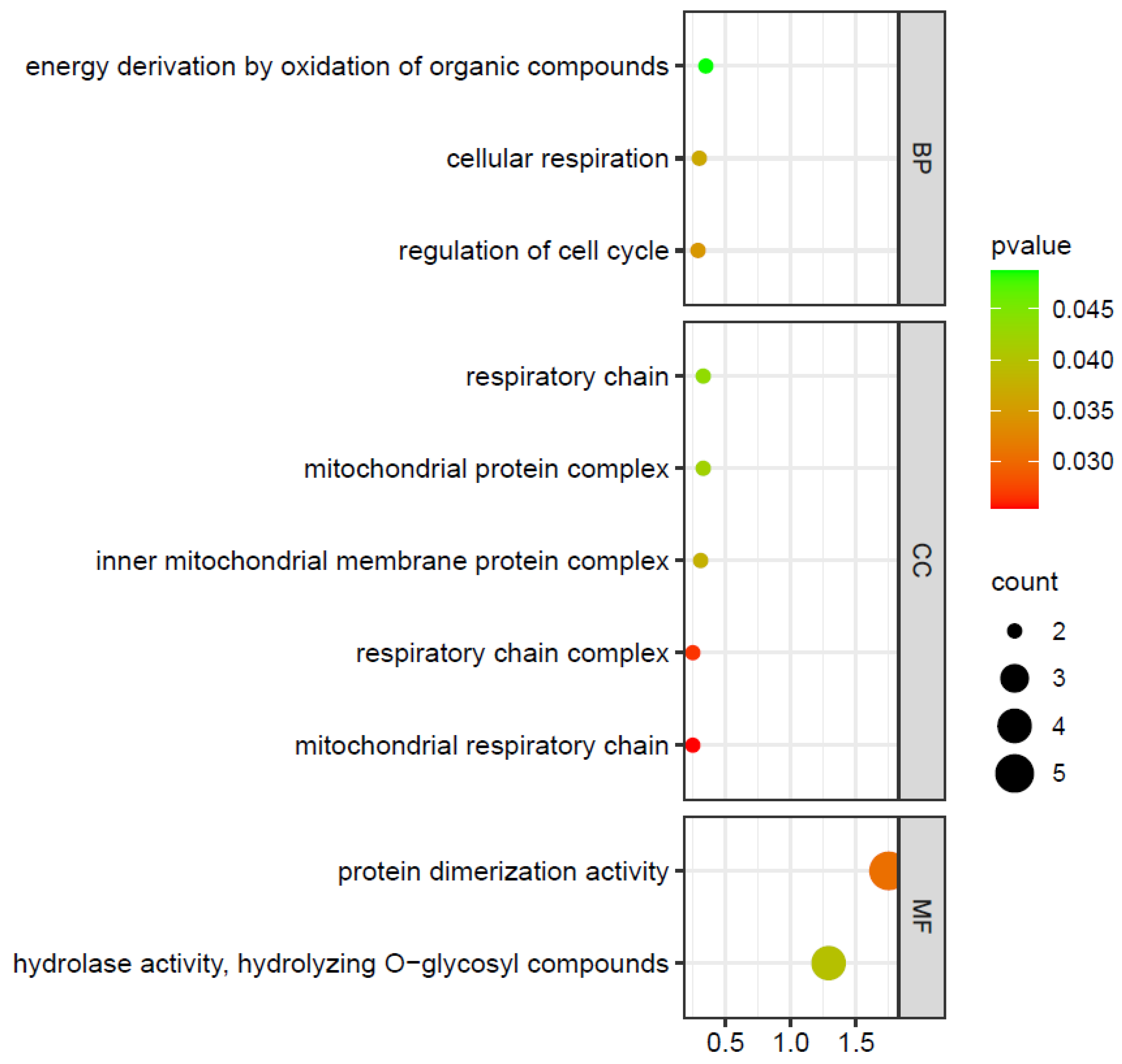
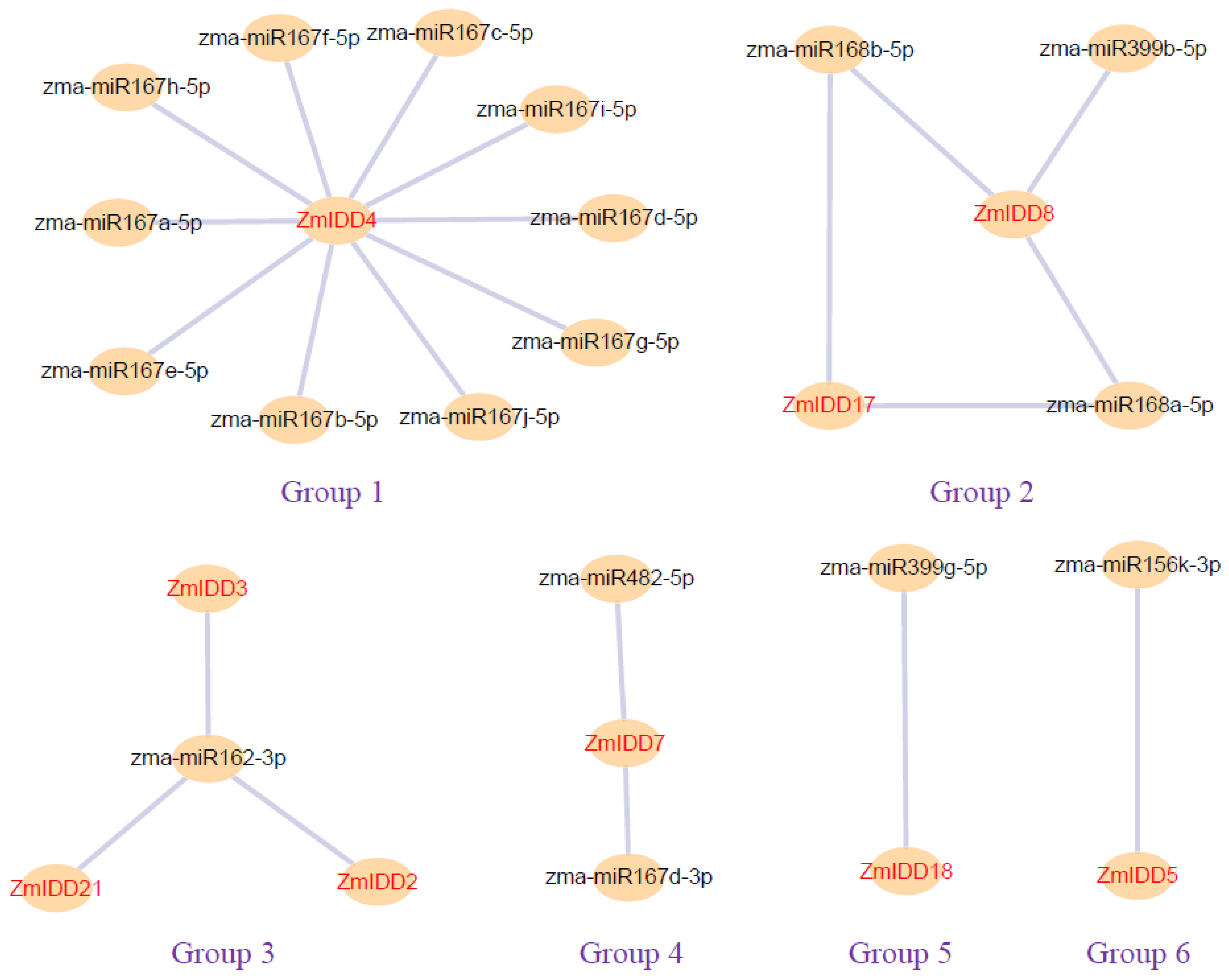
2.7. ZmIDDs-Regulated Genes and miRNA-Targeted ZmIDDs
3. Discussion
4. Materials and Methods
4.1. Identification of ZmIDDs in Zea mays
4.2. Phylogenetic Tree, Gene Structure, Conserved Motifs, and Cis-Elements Analysis
4.3. Chromosome Distribution and Synteny Block of ZmIDDs
4.4. Modeling of 3D Structures of ZmIDDs and Zinc Finger Alignment
4.5. Prediction of ZmIDDs-Regulated Genes
4.6. Prediction of miRNA-ZmIDD Regulatory Networks
4.7. Expression Profile Analysis of ZmIDDs
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Liu, T.; Chen, T.; Kan, J.; Yao, Y.; Guo, D.; Yang, Y.; Ling, X.; Wang, J.; Zhang, B. The GhMYB36 transcription factor confers resistance to biotic and abiotic stress by enhancing PR1 gene expression in plants. Plant Biotechnol. J. 2022, 20, 722–735. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Huang, S.C.; Wise, A.; Castanon, R.; Nery, J.R.; Chen, H.; Watanabe, M.; Thomas, J.; Bar-Joseph, Z.; Ecker, J.R. A transcription factor hierarchy defines an environmental stress response network. Science 2016, 354, aag1550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franco-Zorrilla, J.M.; López-Vidriero, I.; Carrasco, J.L.; Godoy, M.; Vera, P.; Solano, R. DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc. Natl. Acad. Sci. USA 2014, 111, 2367–2372. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, G.; Lu, C.; Guo, J.; Qiao, Z.; Sui, N.; Qiu, N.; Wang, B. C2H2 Zinc finger proteins: Master tegulators of abiotic stress responses in plants. Front. Plant Sci. 2020, 20, 115. [Google Scholar] [CrossRef] [Green Version]
- Khoso, M.A.; Hussain, A.; Ritonga, F.N.; Ali, Q.; Channa, M.M.; Alshegaihi, R.M.; Meng, Q.; Ali, M.; Zaman, W.; Brohi, R.D.; et al. WRKY transcription factors (TFs): Molecular switches to regulate drought, temperature, and salinity stresses in plants. Front. Plant Sci. 2022, 13, 1039329. [Google Scholar] [CrossRef]
- Lu, R.; Li, Y.; Zhang, J.; Wang, Y.; Zhang, J.; Li, Y.; Zheng, Y.; Li, X.B. The bHLH/HLH transcription factors GhFP2 and GhACE1 antagonistically regulate fiber elongation in cotton. Plant Physiol. 2022, 189, 628–643. [Google Scholar] [CrossRef]
- Ma, D.; Constabel, C.P. MYB Repressors as regulators of phenylpropanoid metabolism in plants. Trends Plant Sci. 2019, 24, 275–289. [Google Scholar] [CrossRef]
- Joo, H.; Baek, W.; Lim, C.W.; Lee, S.C. Post-translational modifications of bZIP transcription factors in abscisic acid signaling and drought responses. Curr. Genom. 2021, 22, 4–15. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, J.; Hu, Z.; Guo, X.; Tian, S.; Chen, G. Genome-wide analysis of the MADS-Box transcription factor family in Solanum lycopersicum. Int. J. Mol. Sci. 2019, 20, 2961. [Google Scholar] [CrossRef] [Green Version]
- Colasanti, J.; Tremblay, R.; Wong, A.Y.; Coneva, V.; Kozaki, A.; Mable, B.K. The maize INDETERMINATE1 flowering time regulator defines a highly conserved zinc finger protein family in higher plants. BMC Genom. 2006, 7, 158. [Google Scholar] [CrossRef] [Green Version]
- Coneva, V.; Zhu, T.; Colasanti, J. Expression differences between normal and indeterminate1 maize suggest downstream targets of ID1, a floral transition regulator in maize. J. Exp. Bot. 2007, 58, 3679–3693. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, H.; Hirano, K.; Sato, T.; Mitsuda, N.; Nomoto, M.; Maeo, K.; Koketsu, E.; Mitani, R.; Kawamura, M.; Ishiguro, S.; et al. DELLA protein functions as a transcriptional activator through the DNA binding of the indeterminate domain family proteins. Proc. Natl. Acad. Sci. USA 2014, 111, 7861–7866. [Google Scholar] [CrossRef] [Green Version]
- Hirano, Y.; Nakagawa, M.; Suyama, T.; Murase, K.; Shirakawa, M.; Takayama, S.; Sun, T.; Hakoshima, T. Structure of the SHR-SCR heterodimer bound to the BIRD/IDD transcriptional factor JKD. Nat. Plants 2017, 3, 17010. [Google Scholar] [CrossRef] [Green Version]
- Colasanti, J.; Yuan, Z.; Sundaresan, V. The indeterminate gene encodes a zinc finger protein and regulates a leaf-generated signal required for the transition to flowering in maize. Curr. Opin. Plant Biol. 1998, 1, 276. [Google Scholar] [CrossRef] [Green Version]
- Kumar, M.; Le, D.T.; Hwang, S.; Seo, P.J.; Kim, H.U. Role of the INDETERMINATE DOMAIN genes in Plants. Int. J. Mol. Sci. 2019, 20, 2286. [Google Scholar] [CrossRef] [Green Version]
- Zhang, T.; Tan, M.; Geng, L.; Li, J.; Xiang, Y.; Zhang, B.; Zhao, Y. New insight into comprehensive analysis of INDETERMINATE DOMAIN (IDD) gene family in rice. Plant Physiol. Biochem. 2020, 154, 547–556. [Google Scholar] [CrossRef]
- Guo, X.; Zhou, M.; Chen, J.; Shao, M.; Zou, L.; Ying, Y.; Liu, S. Genome-wide identification of the highly conserved INDETERMINATE DOMAIN (IDD) zinc finger gene family in Moso Bamboo (Phyllostachys edulis). Int. J. Mol. Sci. 2022, 23, 13952. [Google Scholar] [CrossRef]
- Coelho, C.P.; Huang, P.; Lee, D.Y.; Brutnell, T.P. Making roots, shoots, and seeds: IDD gene family diversification in plants. Trends Plant Sci. 2018, 23, 66–78. [Google Scholar] [CrossRef]
- Prochetto, S.; Reinheimer, R. Step by step evolution of Indeterminate Domain (IDD) transcriptional regulators: From algae to angiosperms. Ann. Bot. 2020, 126, 85–101. [Google Scholar] [CrossRef]
- Fukazawa, J.; Teramura, H.; Murakoshi, S.; Nasuno, K.; Nishida, N.; Ito, T.; Yoshida, M.; Kamiya, Y.; Yamaguchi, S.; Takahashi, Y. DELLAs function as coactivators of GAI-ASSOCIATED FACTOR1 in regulation of gibberellin homeostasis and signaling in Arabidopsis. Plant Cell 2014, 26, 2920–2938. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Shu, D.; Tan, Z.; Ma, M.; Guo, N.; Gao, S.; Duan, G.; Kuai, B.; Hu, Y.; Li, S.; et al. The Arabidopsis IDD14 transcription factor interacts with bZIP-type ABFs/AREBs and cooperatively regulates ABA-mediated drought tolerance. New Phytol. 2022, 236, 929–942. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Dong, G.; Xu, J.; Su, Y.; Shi, Z.; Ye, W.; Li, Y.; Li, G.; Zhang, B.; Hu, J.; et al. A point mutation in the zinc finger motif of RID1/EHD2/OsID1 protein leads to outstanding yield-related traits in japonica rice variety Wuyunjing 7. Rice 2013, 6, 24. [Google Scholar] [CrossRef] [Green Version]
- Takehara, S.; Huang, J.; Hirano, K.; Ordonio, R.L.; Matsuoka, M.; UeguchiTanaka, M. OsIDD2, a zinc finger and Indeterminate Domain protein, regulates secondary cell wall formation. J. Integr. Plant Biol. 2018, 60, 130–143. [Google Scholar]
- Dou, M.; Cheng, S.; Zhao, B.; Xuan, Y.; Shao, M. The Indeterminate Domain Protein ROC1 regulates chilling tolerance via activation of DREB1B/CBF1 in rice. Int. J. Mol. Sci. 2016, 17, 233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xuan, Y.H.; Priatama, R.A.; Huang, J.; Je, B.I.; Liu, J.M.; Park, S.J.; Piao, H.L.; Son, D.Y.; Lee, J.J.; Park, S.H.; et al. INDETERMINATE DOMAIN 10 regulates ammonium-mediated gene expression in rice roots. New Phytol. 2013, 197, 791–804. [Google Scholar] [CrossRef]
- Jöst, M.; Hensel, G.; Kappel, C.; Druka, A.; Sicard, A.; Hohmann, U.; Beier, S.; Himmelbach, A.; Waugh, R.; Kumlehn, J.; et al. The INDETERMINATE DOMAIN protein BROAD LEAF1 limits barley leaf width by restricting lateral proliferation. Curr. Biol. 2016, 26, 903–909. [Google Scholar] [CrossRef] [Green Version]
- Yi, G.; Neelakandan, A.K.; Gontarek, B.C.; Vollbrecht, E.; Becraft, P.W. The naked endosperm genes encode duplicate INDETERMINATE domain transcription factors required for maize endosperm cell patterning and differentiation. Plant Physiol. 2015, 167, 443–456. [Google Scholar] [CrossRef] [Green Version]
- Gontarek, B.C.; Neelakandan, A.K.; Wu, H.; Becraft, P.W. NKD transcription factors are central regulators of maize endosperm development. Plant Cell 2016, 28, 2916–2936. [Google Scholar] [CrossRef] [Green Version]
- Deng, L.; Li, L.; Zhang, S.; Shen, J.; Li, S.; Hu, S.; Peng, Q.; Xiao, J.; Wu, C. Suppressor of rid1 (SID1) shares common targets with RID1 on florigen genes to initiate floral transition in rice. PLoS Genet. 2017, 13, e1006642. [Google Scholar] [CrossRef] [Green Version]
- Seo, P.J.; Kim, M.J.; Ryu, J.Y.; Jeong, E.Y.; Park, C.M. Two splice variants of the IDD14 transcription factor competitively form nonfunctional heterodimers which may regulate starch metabolism. Nat. Commun. 2011, 2, 303. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.C.; Lim, M.H.; Kim, J.A.; Lee, S.I.; Kim, J.S.; Jin, M.; Kwon, S.J.; Mun, J.H.; Kim, Y.K.; Kim, H.U.; et al. Transcriptome analysis in Brassica rapa under the abiotic stresses using Brassica 24K oligo microarray. Mol. Cells 2008, 26, 595–605. [Google Scholar]
- Wong, A.Y.; Colasanti, J. Maize floral regulator protein INDETERMINATE1 is localized to developing leaves and is not altered by light or the sink/source transition. J. Exp. Bot. 2007, 5, 403–414. [Google Scholar] [CrossRef]
- Xu, G.; Guo, C.; Shan, H.; Kong, H. Divergence of duplicate genes in exon-intron structure. Proc. Natl. Acad. Sci. USA 2012, 109, 1187–1192. [Google Scholar] [CrossRef] [Green Version]
- Su, X.; Meng, T.; Zhao, Y.; Li, G.; Cheng, X.; Abdullah, M.; Sun, X.; Cai, Y.; Lin, Y. Comparative genomic analysis of the IDD genes in five Rosaceae species and expression analysis in Chinese white pear (Pyrus bretschneideri). PeerJ 2019, 7, e6628. [Google Scholar] [CrossRef] [Green Version]
- Fan, S.; Zhang, D.; Xing, L.; Qi, S.; Du, L.; Wu, H.; Shao, H.; Li, Y.; Ma, J.; Han, M. Phylogenetic analysis of IDD gene family and characterization of its expression in response to flower induction in Malus. Mol. Genet. Genom. 2017, 292, 755–771. [Google Scholar] [CrossRef]
- Waadt, R.; Seller, C.A.; Hsu, P.K.; Takahashi, Y.; Munemasa, S.; Schroeder, J.I. Plant hormone regulation of abiotic stress responses. Nat. Rev. Mol. Cell Biol. 2022, 23, 680–694. [Google Scholar] [CrossRef]
- Jeong, E.Y.; Seo, P.J.; Woo, J.C.; Park, C.M. AKIN10 delays flowering by inactivating IDD8 transcription factor through protein phosphorylation in Arabidopsis. BMC Plant Biol. 2015, 15, 110. [Google Scholar] [CrossRef] [Green Version]
- Völz, R.; Rayapuram, N.; Hirt, H. Phosphorylation regulates the activity of INDETERMINATE-DOMAIN (IDD/BIRD) proteins in response to diverse environmental conditions. Plant Signal. Behav. 2019, 14, e1642037. [Google Scholar] [CrossRef]
- Cui, D.; Zhao, J.; Jing, Y.; Fan, M.; Liu, J.; Wang, Z.; Xin, W.; Hu, Y. The Arabidopsis IDD14, IDD15 and IDD16 cooperatively regulate lateral organ morphogenesis and gravitropism by promoting auxin biosynthesis and transport. PLoS Genet. 2013, 9, e1003759. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Li, K.; Chen, L.; Zou, Y.; Liu, H.; Tian, Y.; Li, D.; Wang, R.; Zhao, F.; Ferguson, B.J.; et al. MicroRNA167-Directed Regulation of the Auxin Response Factors GmARF8a and GmARF8b Is Required for Soybean Nodulation and Lateral Root Development. Plant Physiol. 2015, 168, 984–999. [Google Scholar] [CrossRef] [Green Version]
- Sasidharan, R.; Keuskamp, D.H.; Kooke, R.; Voesenek, L.A.; Pierik, R. Interactions between auxin, microtubules and XTHs mediate green shade-induced petiole elongation in arabidopsis. PLoS ONE 2014, 9, e90587. [Google Scholar] [CrossRef] [PubMed]
- Gullner, G.; Komives, T.; Király, L.; Schröder, P. Glutathione S-Transferase Enzymes in Plant-Pathogen Interactions. Front. Plant Sci. 2018, 9, 1836. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, M.; Tang, W. MicroRNA156 amplifies transcription factor-associated cold stress tolerance in plant cells. Mol. Genet. Genom. 2019, 294, 379–393. [Google Scholar] [CrossRef] [PubMed]
- Arshad, M.; Hannoufa, A. Alfalfa transcriptome profiling provides insight into miR156-mediated molecular mechanisms of heat stress tolerance. Genome 2022, 65, 315–330. [Google Scholar] [CrossRef] [PubMed]
- Horrer, D.; Flutsch, S.; Pazmino, D.; Matthews, J.S.; Thalmann, M.; Nigro, A.; Leonhardt, N.; Lawson, T.; Santelia, D. Blue light induces a distinct starch degradation pathway in guard cells for stomatal opening. Curr. Biol. 2016, 26, 362–370. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Wong, D.C.J.; Wang, Y.; Xu, G.; Ren, C.; Liu, Y.; Kuang, Y.; Fan, P.; Li, S.; Xin, H.; et al. GRAS-domain transcription factor PAT1 regulates jasmonic acid biosynthesis in grape cold stress response. Plant Physiol. 2021, 186, 1660–1678. [Google Scholar] [CrossRef]
- Finn, R.D.; Clements, J.; Eddy, S.R. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 2011, 39, 29–37. [Google Scholar] [CrossRef] [Green Version]
- Kersey, P.J.; Allen, J.E.; Allot, A.; Barba, M.; Boddu, S.; Bolt, B.J.; Carvalho-Silva, D.; Christensen, M.; Davis, P.; Grabmueller, C.; et al. Ensemble Genomes 2018: An integrated omics infrastructure for non-vertebrate species. Nucleic Acids Res. 2018, 46, 802–808. [Google Scholar] [CrossRef]
- Letunic, I.; Doerks, T.; Bork, P. SMART 7: Recent updates to the protein domain annotation resource. Nucleic Acids Res. 2012, 40, 302–305. [Google Scholar] [CrossRef]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, 597–603. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, 256–259. [Google Scholar] [CrossRef] [Green Version]
- Hu, B.; Jin, J.; Guo, A.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [Green Version]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, 39–49. [Google Scholar] [CrossRef] [Green Version]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Gu, Z.; Gu, L.; Eils, R.; Schlesner, M.; Brors, B. Circlize Implements and enhances circular visualization in R. Bioinformatics 2014, 30, 2811–2812. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, 49. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Z.; Xiao, J.; Wu, J.; Zhang, H.; Liu, G.; Wang, X.; Dai, L. ParaAT: A parallel tool 665 for constructing multiple protein-coding DNA alignments. Biochem. Biophys. Res. Commun. 2012, 419, 779–781. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Zhang, Z.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A toolkit incorporating gamma-series methods and sliding window strategies. Genom. Proteom. Bioinform. 2010, 8, 77–80. [Google Scholar] [CrossRef] [Green Version]
- Koch, M.A.; Haubold, B.; Mitchell-Olds, T. Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis, Arabis, and related genera (Brassicaceae). Mol. Biol. Evol. 2000, 17, 1483–1498. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arnold, K.; Bordoli, L.; Kopp, J.; Schwede, T. The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics 2006, 22, 195–201. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tian, F.; Yang, D.C.; Meng, Y.Q.; Jin, J.; Gao, G. PlantRegMap: Charting functional regulatory maps in plants. Nucleic Acids Res. 2020, 48, 1104–1113. [Google Scholar] [CrossRef]
- Su, W.; Raza, A.; Zeng, L.; Gao, A.; Lv, Y.; Ding, X.; Cheng, Y.; Zou, X. Genome-wide analysis and expression patterns of lipid phospholipid phospholipase gene family in Brassica napus L. BMC Genom. 2021, 22, 548. [Google Scholar] [CrossRef]
- Opitz, N.; Paschold, A.; Marcon, C.; Malik, W.A.; Lanz, C.; Piepho, H.P.; Hochholdinger, F. Transcriptomic complexity in young maize primary roots in response to low water potentials. BMC Genom. 2014, 15, 741. [Google Scholar] [CrossRef] [Green Version]
- Stelpflug, S.C.; Sekhon, R.S.; Vaillancourt, B.; Hirsch, C.N.; Buell, C.R.; de Leon, N.; Kaeppler, S.M. An Expanded Maize Gene Expression Atlas Based on RNA Sequencing and Its Use to Explore Root Development. Plant Genome 2016, 9, plantgenome2015.04.0025. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Feng, X.; Yu, Q.; Zeng, J.; He, X.; Ma, W.; Ge, L.; Liu, W. Comprehensive Analysis of the INDETERMINATE DOMAIN (IDD) Gene Family and Their Response to Abiotic Stress in Zea mays. Int. J. Mol. Sci. 2023, 24, 6185. https://doi.org/10.3390/ijms24076185
Feng X, Yu Q, Zeng J, He X, Ma W, Ge L, Liu W. Comprehensive Analysis of the INDETERMINATE DOMAIN (IDD) Gene Family and Their Response to Abiotic Stress in Zea mays. International Journal of Molecular Sciences. 2023; 24(7):6185. https://doi.org/10.3390/ijms24076185
Chicago/Turabian StyleFeng, Xue, Qian Yu, Jianbin Zeng, Xiaoyan He, Wujun Ma, Lei Ge, and Wenxing Liu. 2023. "Comprehensive Analysis of the INDETERMINATE DOMAIN (IDD) Gene Family and Their Response to Abiotic Stress in Zea mays" International Journal of Molecular Sciences 24, no. 7: 6185. https://doi.org/10.3390/ijms24076185
APA StyleFeng, X., Yu, Q., Zeng, J., He, X., Ma, W., Ge, L., & Liu, W. (2023). Comprehensive Analysis of the INDETERMINATE DOMAIN (IDD) Gene Family and Their Response to Abiotic Stress in Zea mays. International Journal of Molecular Sciences, 24(7), 6185. https://doi.org/10.3390/ijms24076185





