Chromosomal-Scale Genome Assemblies of Two Coastal Plant Species, Scaevola taccada and S. hainanensis—Insight into Adaptation Outside of the Common Range
Abstract
:1. Introduction
2. Results
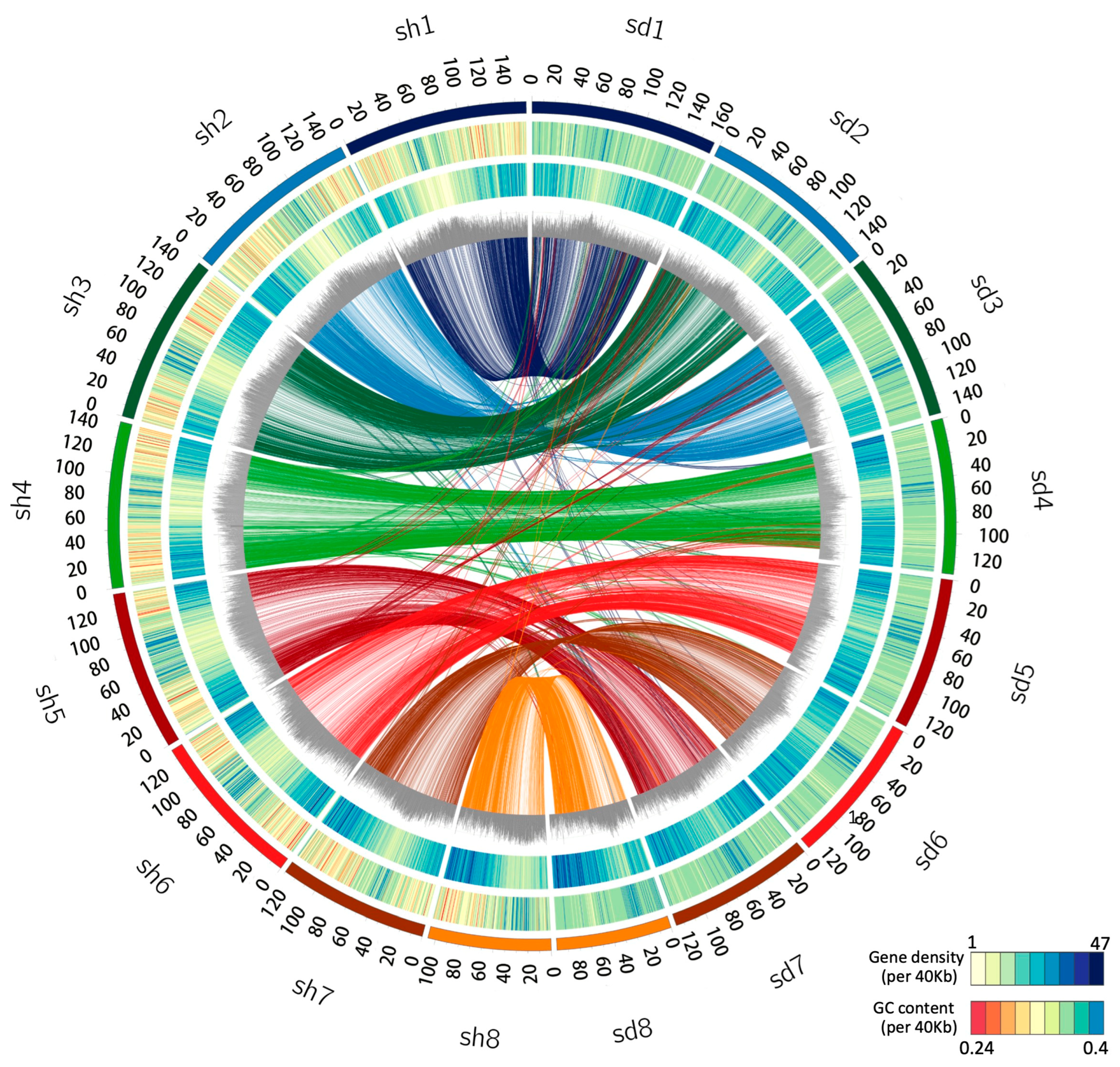
2.1. Genome Assembly and Annotation
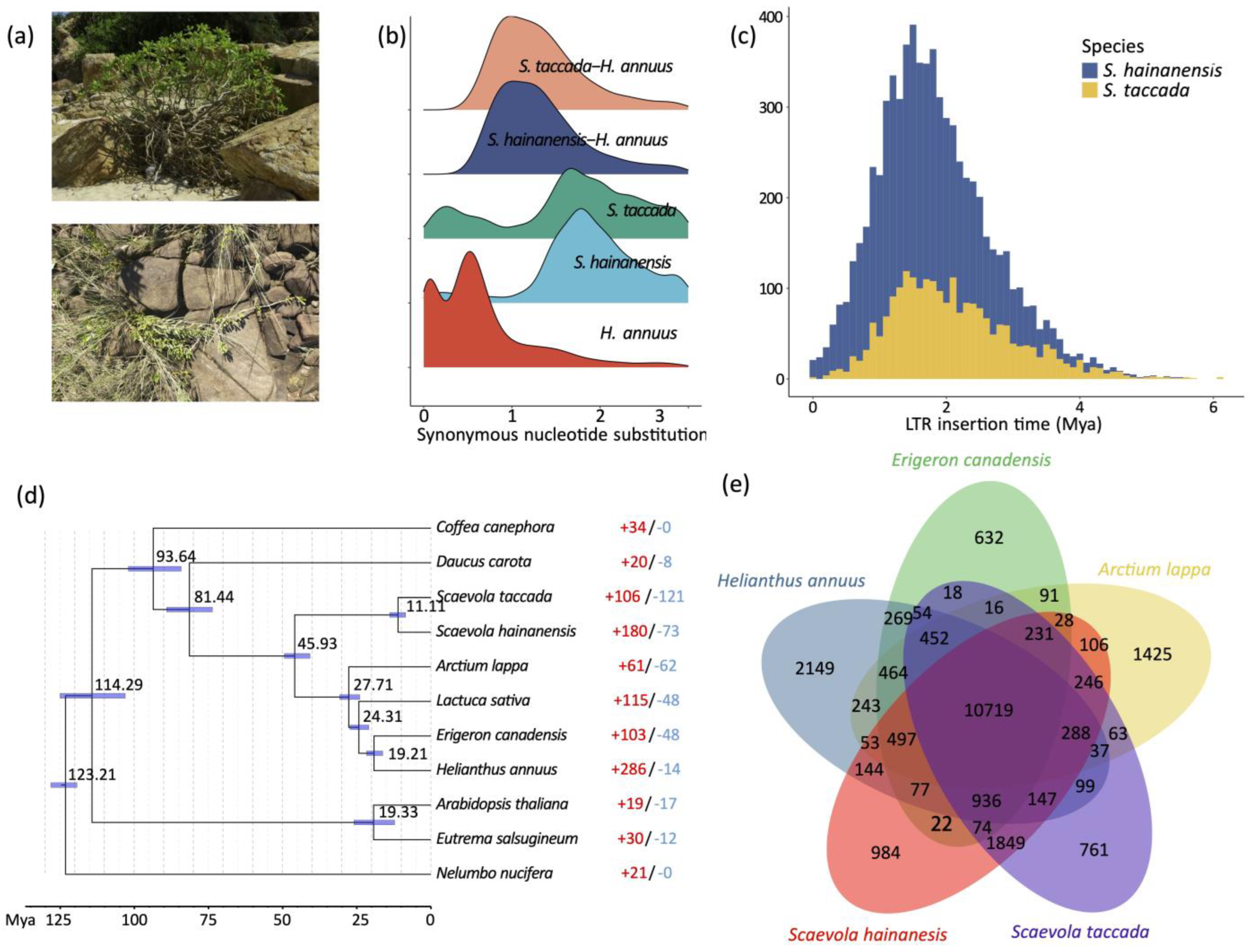
2.2. Whole Genome Duplication and LTR Retrotransposon Insertion
2.3. Phylogeny Construction and Gene Copy Number Evolution
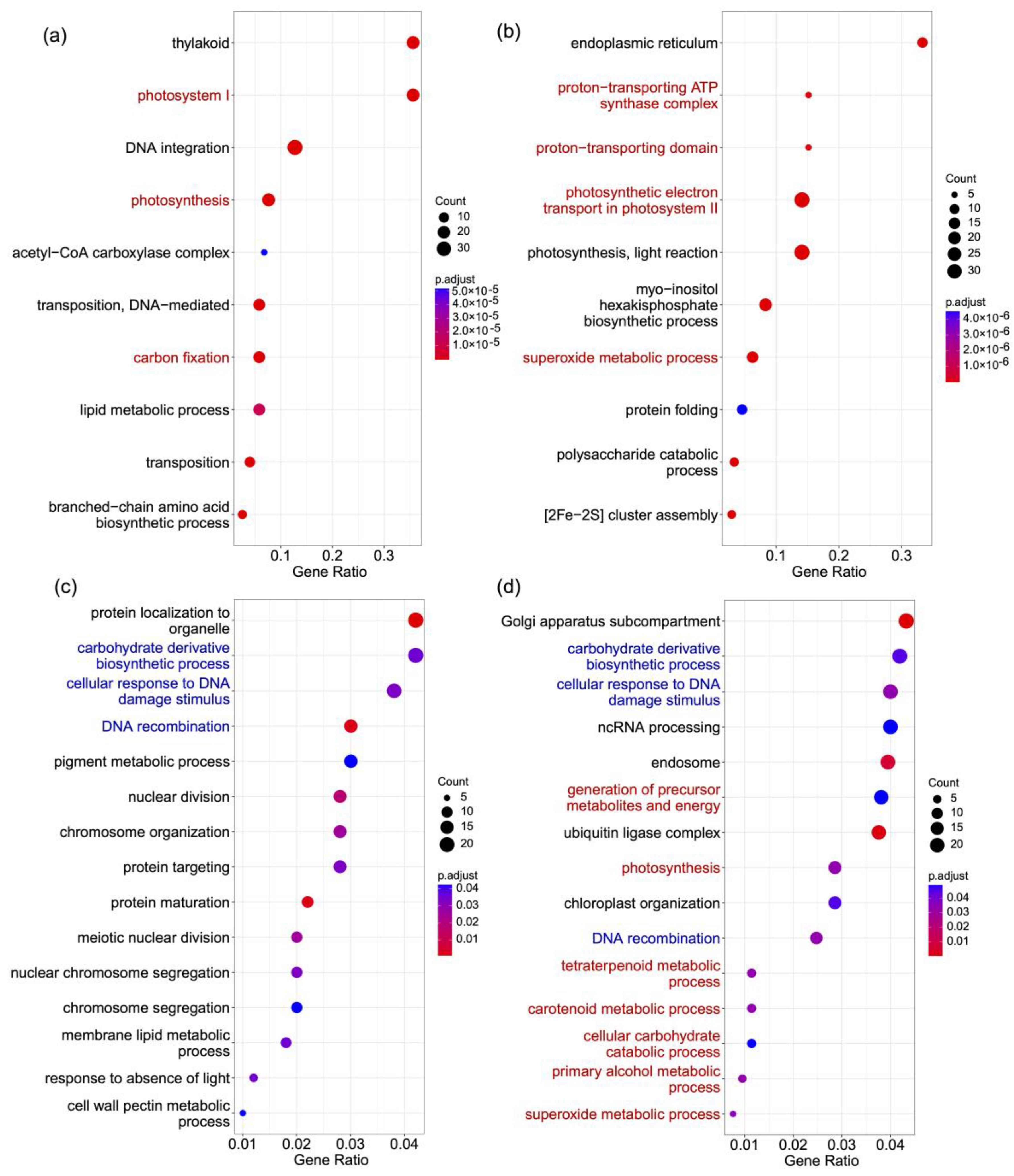
2.4. Genes under Positive Selection
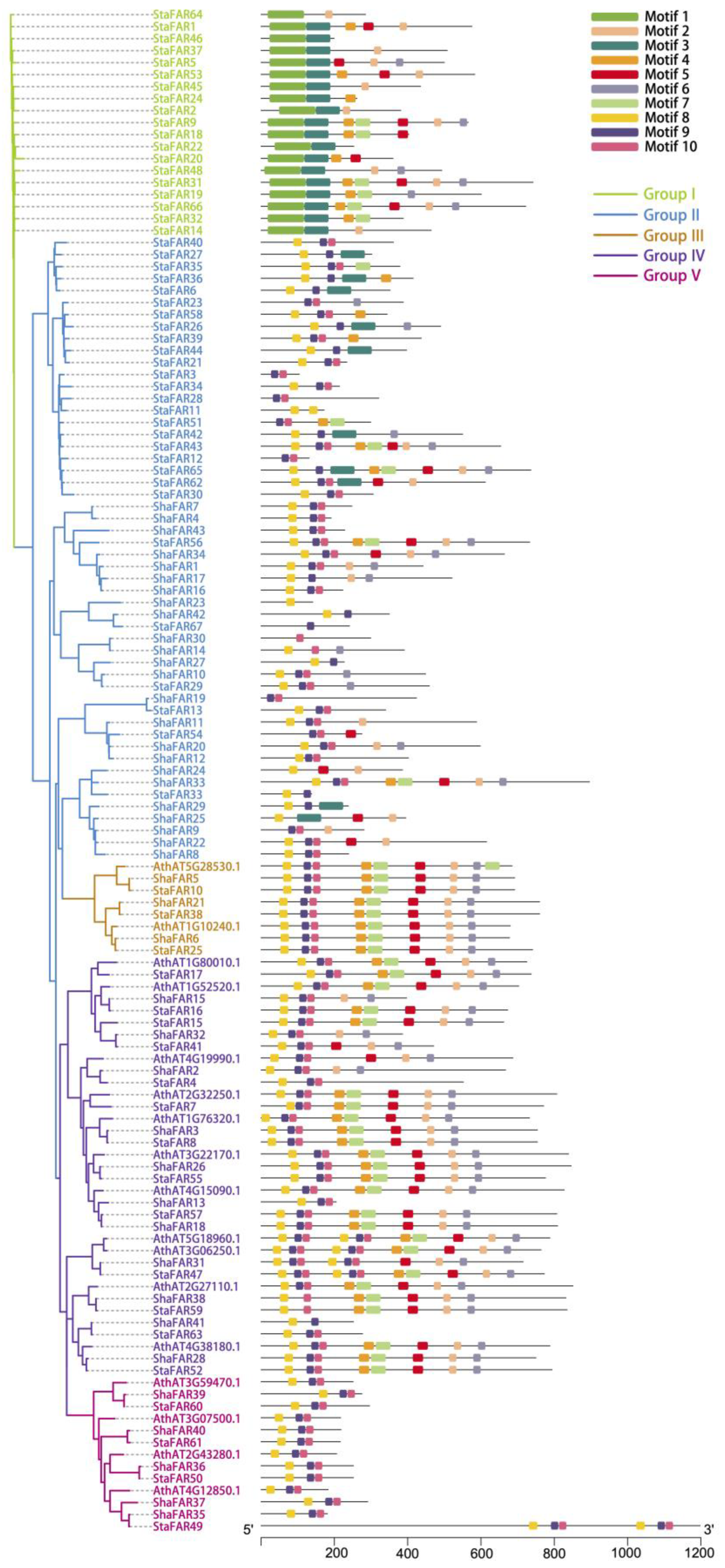
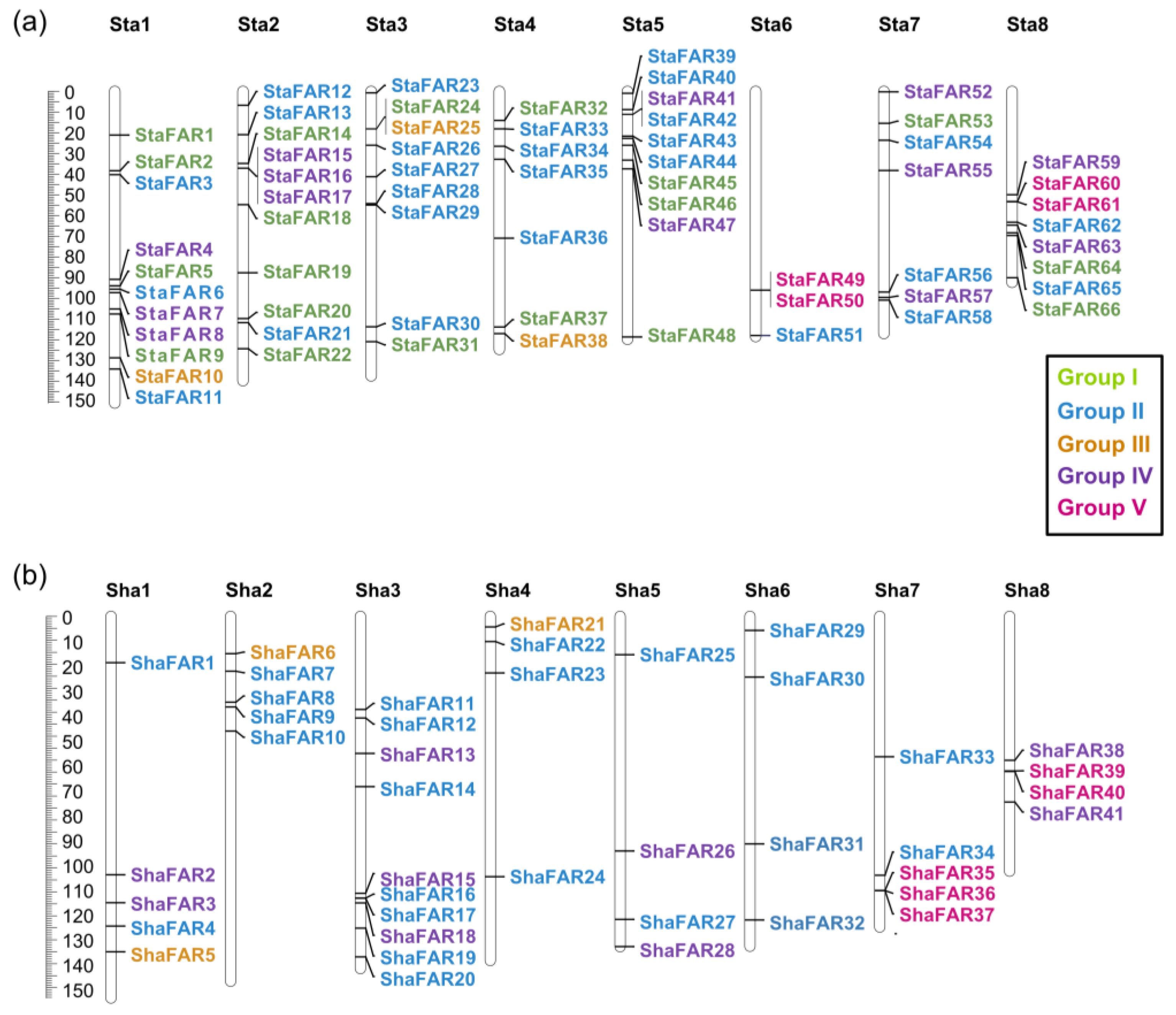
2.5. Evolution of the FAR1 Gene Family in S. taccada and S. hainanensis
3. Discussion
4. Material and Methods
4.1. Plant Material
4.2. 10× Genomics Sequencing
4.3. Hi-C Sequencing
4.4. Short-Read Sequencing
4.5. De Novo Genome Assembling
4.6. Genome Annotation
4.7. Collinearity Analysis and Whole-Genome Duplication (WGD) Analysis
4.8. Phylogeny Construction and Divergence Time Estimation
4.9. Gene Group Evolution and Positive Selection Detection
4.10. Analyses of the FAR1 Gene Family
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Carolin, R.C. Brunoniaceae, Goodeniaceae. In Flora of Australia; Australian Government Publishing Service: Canberra, Australia, 1992; Volume 35, pp. 147–281. ISBN 0-644-14553-6. [Google Scholar]
- Howarth, D.G.; Gustafsson, M.H.G.; Baum, D.A.; Motley, T.J. Phylogenetics of the Genus Scaevola (Goodeniaceae): Implication for Dispersal Patterns across the Pacific Basin and Colonization of the Hawaiian Islands. Am. J. Bot. 2003, 90, 915–923. [Google Scholar] [CrossRef] [PubMed]
- Jabaily, R.S.; Shepherd, K.A.; Gardner, A.G.; Gustafsson, M.H.G.; Howarth, D.G.; Motley, T.J. Historical Biogeography of the Predominantly Australian Plant Family Goodeniaceae. J. Biogeogr. 2014, 41, 2057–2067. [Google Scholar] [CrossRef]
- Howarth, D.G.; Baum, D.A. Genealogical Evidence of Homoploid Hybrid Speciation in an Adaptive Radiation of Scaevola (Goodeniaceae) in the Hawaiian Islands. Evolution 2005, 59, 948–961. [Google Scholar] [CrossRef]
- Castillo-Campos, G.; García-Franco, J.; Martínez, L. First Record of Naturalization of Scaevola Taccada (Gaertn.) Roxb. (Goodeniaceae) in Southeastern Mexico. BIR 2021, 10, 425–435. [Google Scholar] [CrossRef]
- Emura, N.; Denda, T.; Sakai, M.; Ueda, K. Dimorphism of the Seed-Dispersing Organ in a Pantropical Coastal Plant, Scaevola Taccada: Heterogeneous Population Structures across Islands. Ecol. Res. 2014, 29, 733–740. [Google Scholar] [CrossRef]
- Goldstein, G.; Drake, D.R.; Alpha, C.; Melcher, P.; Heraux, J.; Azocar, A. Growth and Photosynthetic Responses of Scaevola Sericea, a Hawaiian Coastal Shrub, to Substrate Salinity and Salty Spray. Int. J. Plant Sci. 1996, 157, 171–179. [Google Scholar] [CrossRef]
- Hong, D.; Howarth, D.G. Goodeniaceae. In Flora of China; eFloras.org: Beijing, China, 1983; Volume 19, pp. 568–569. [Google Scholar]
- Huang, T.-C. Goodeniaceae. In Flora of Taiwan, 2nd ed.; Institute of Ecology and Evolutionary Biology, NTU: Singapore, 1998; Volume 4, pp. 803–805. [Google Scholar]
- Su, M.; He, S.; Li, W.; Wang, S.; Tan, F.; Huang, Y. Systematic Position of a Rare Plant Species Scaevola Hainanensis Based on ITS Sequence Analysis. Acta Sci. Nat. Univ. Sunyatseni 2016, 55, 16–23. [Google Scholar] [CrossRef]
- Ho, K.Y.; Deng, S.L.; Chang, Y.H.; Tsai, C.S.; Kao, M.F.; Hsiao, J.Y. Genetic Variation of the Endangered Scaevola Hainanensis (Goodeniaceae) in the Jiangjun Stream Mouth, Taiwan. Taiwan J. For. Sci. 2005, 20, 193–202. [Google Scholar] [CrossRef]
- Huang, X.; Ouyang, X.; Yang, P.; Lau, O.S.; Li, G.; Li, J.; Chen, H.; Deng, X.W. Arabidopsis FHY3 and HY5 Positively Mediate Induction of COP1 Transcription in Response to Photomorphogenic UV-B Light. Plant Cell 2012, 24, 4590–4606. [Google Scholar] [CrossRef]
- Wang, H.; Wang, H. Multifaceted Roles of FHY3 and FAR1 in Light Signaling and Beyond. Trends Plant Sci. 2015, 20, 453–461. [Google Scholar] [CrossRef]
- Krzeszowiec, W.; Novokreshchenova, M.; Gabryś, H. Chloroplasts in C3 Grasses Move in Response to Blue-Light. Plant Cell Rep. 2020, 39, 1331–1343. [Google Scholar] [CrossRef]
- Huai, J.; Zhang, X.; Li, J.; Ma, T.; Zha, P.; Jing, Y.; Lin, R. SEUSS and PIF4 Coordinately Regulate Light and Temperature Signaling Pathways to Control Plant Growth. Mol. Plant 2020, 13, 1825. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Wang, W.; Chen, D.; Ji, Q.; Jing, Y.; Wang, H.; Lin, R. Transposase-Derived Proteins FHY3/FAR1 Interact with PHYTOCHROME-INTERACTING FACTOR1 to Regulate Chlorophyll Biosynthesis by Modulating HEMB1 during Deetiolation in Arabidopsis. Plant Cell 2012, 24, 1984–2000. [Google Scholar] [CrossRef] [PubMed]
- Lin, R.; Wang, H. Arabidopsis FHY3/FAR1 Gene Family and Distinct Roles of Its Members in Light Control of Arabidopsis Development. Plant Physiol. 2004, 136, 4010–4022. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Siddiqui, H.; Teng, Y.; Lin, R.; Wan, X.; Li, J.; Lau, O.-S.; Ouyang, X.; Dai, M.; Wan, J.; et al. Coordinated Transcriptional Regulation Underlying the Circadian Clock in Arabidopsis. Nat. Cell Biol. 2011, 13, 616–622. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ma, M.; Li, G.; Yuan, L.; Xie, Y.; Wei, H.; Ma, X.; Li, Q.; Devlin, P.F.; Xu, X.; et al. Transcription Factors FHY3 and FAR1 Regulate Light-Induced CIRCADIAN CLOCK ASSOCIATED1 Gene Expression in Arabidopsis. Plant Cell 2020, 32, 1464–1478. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup The Sequence Alignment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Seppey, M.; Manni, M.; Zdobnov, E.M. BUSCO: Assessing Genome Assembly and Annotation Completeness. Methods Mol. Biol. 2019, 1962, 227–245. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and Applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Zhang, C.; Zhang, T.; Luebert, F.; Xiang, Y.; Huang, C.-H.; Hu, Y.; Rees, M.; Frohlich, M.W.; Qi, J.; Weigend, M.; et al. Asterid Phylogenomics/Phylotranscriptomics Uncover Morphological Evolutionary Histories and Support Phylogenetic Placement for Numerous Whole-Genome Duplications. Mol. Biol. Evol. 2020, 37, 3188–3210. [Google Scholar] [CrossRef]
- Bennetzen, J.L. Mechanisms of Recent Genome Size Variation in Flowering Plants. Ann. Bot. 2005, 95, 127–132. [Google Scholar] [CrossRef] [PubMed]
- Dai, S.; Zhu, X.; Hutang, G.; Li, J.; Tian, J.; Jiang, X.; Zhang, D.; Gao, L. Genome Size Variation and Evolution Driven by Transposable Elements in the Genus Oryza. Front. Plant Sci. 2022, 13, 921937. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Shi, C.; Shi, C.-C.; Li, W.; Zhang, Q.-J.; Zhang, Y.; Li, K.; Lu, H.-F.; Shi, C.; Zhu, S.-T.; et al. The Chromosome-Based Rubber Tree Genome Provides New Insights into Spurge Genome Evolution and Rubber Biosynthesis. Mol. Plant 2020, 13, 336–350. [Google Scholar] [CrossRef]
- Albalat, R.; Cañestro, C. Evolution by Gene Loss. Nat. Rev. Genet 2016, 17, 379–391. [Google Scholar] [CrossRef]
- Fang, Y.; Jiang, J.; Hou, X.; Guo, J.; Li, X.; Zhao, D.; Xie, X. Plant Protein-Coding Gene Families: Their Origin and Evolution. Front. Plant Sci. 2022, 13, 995746. [Google Scholar] [CrossRef]
- Jin, J.; Tian, F.; Yang, D.-C.; Meng, Y.-Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a Central Hub for Transcription Factors and Regulatory Interactions in Plants. Nucleic Acids Res. 2017, 45, D1040–D1045. [Google Scholar] [CrossRef]
- Borges, A.; Rosa, M.S.; Recchia, G.H.; de Queiroz-Silva, J.R.; de Andrade Bressan, E.; Veasey, E.A. CTAB Methods for DNA Extraction of Sweetpotato for Microsatellite Analysis. Sci. Agric. 2009, 66, 529–534. [Google Scholar] [CrossRef]
- Yang, G.; Zhou, R.; Tang, T.; Shi, S. Simple and Efficient Isolation of High-Quality Total RNA from Hibiscus Tiliaceus, a Mangrove Associate and Its Relatives. Prep. Biochem. Biotechnol. 2008, 38, 257–264. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Feng, X.; Chen, Q.; Li, L.; Li, S.; Han, K.; Guo, Z.; Wang, J.; Liu, M.; Shi, C.; et al. Evolution of Coastal Forests Based on a Full Set of Mangrove Genomes. Nat. Ecol. Evol. 2022, 6, 738–749. [Google Scholar] [CrossRef]
- Chen, Y.; Chen, Y.; Shi, C.; Huang, Z.; Zhang, Y.; Li, S.; Li, Y.; Ye, J.; Yu, C.; Li, Z.; et al. SOAPnuke: A MapReduce Acceleration-Supported Software for Integrated Quality Control and Preprocessing of High-Throughput Sequencing Data. GigaScience 2018, 7, gix120. [Google Scholar] [CrossRef]
- Marçais, G.; Kingsford, C. A Fast, Lock-Free Approach for Efficient Parallel Counting of Occurrences of k-Mers. Bioinformatics 2011, 27, 764–770. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Shi, Y.; Yuan, J.; Hu, X.; Zhang, H.; Li, N.; Li, Z.; Chen, Y.; Mu, D.; Fan, W. Estimation of Genomic Characteristics by Analyzing K-Mer Frequency in de Novo Genome Projects. arXiv 2013. [Google Scholar] [CrossRef]
- Weisenfeld, N.I.; Kumar, V.; Shah, P.; Church, D.M.; Jaffe, D.B. Direct Determination of Diploid Genome Sequences. Genome Res. 2017, 27, 757–767. [Google Scholar] [CrossRef]
- Xu, M.; Guo, L.; Gu, S.; Wang, O.; Zhang, R.; Peters, B.A.; Fan, G.; Liu, X.; Xu, X.; Deng, L.; et al. TGS-GapCloser: A Fast and Accurate Gap Closer for Large Genomes with Low Coverage of Error-Prone Long Reads. GigaScience 2020, 9, giaa094. [Google Scholar] [CrossRef] [PubMed]
- Servant, N.; Varoquaux, N.; Lajoie, B.R.; Viara, E.; Chen, C.-J.; Vert, J.-P.; Heard, E.; Dekker, J.; Barillot, E. HiC-Pro: An Optimized and Flexible Pipeline for Hi-C Data Processing. Genome Biol. 2015, 16, 259. [Google Scholar] [CrossRef] [PubMed]
- Durand, N.C.; Shamim, M.S.; Machol, I.; Rao, S.S.P.; Huntley, M.H.; Lander, E.S.; Aiden, E.L. Juicer Provides a One-Click System for Analyzing Loop-Resolution Hi-C Experiments. Cell Systems 2016, 3, 95–98. [Google Scholar] [CrossRef]
- Durand, N.C.; Robinson, J.T.; Shamim, M.S.; Machol, I.; Mesirov, J.P.; Lander, E.S.; Aiden, E.L. Juicebox Provides a Visualization System for Hi-C Contact Maps with Unlimited Zoom. Cell Systems 2016, 3, 99–101. [Google Scholar] [CrossRef]
- Dudchenko, O.; Batra, S.S.; Omer, A.D.; Nyquist, S.K.; Hoeger, M.; Durand, N.C.; Shamim, M.S.; Machol, I.; Lander, E.S.; Aiden, A.P.; et al. De Novo Assembly of the Aedes Aegypti Genome Using Hi-C Yields Chromosome-Length Scaffolds. Science 2017, 328, 119–130. [Google Scholar] [CrossRef]
- Jurka, J.; Kapitonov, V.V.; Pavlicek, A.; Klonowski, P.; Kohany, O.; Walichiewicz, J. Repbase Update, a Database of Eukaryotic Repetitive Elements. Cytogenet. Genome Res. 2005, 110, 462–467. [Google Scholar] [CrossRef]
- Tarailo-Graovac, M.; Chen, N. Using RepeatMasker to Identify Repetitive Elements in Genomic Sequences. CP Bioinform. 2009, 25, 4–10. [Google Scholar] [CrossRef]
- Abrusán, G.; Grundmann, N.; DeMester, L.; Makalowski, W. TEclass—A Tool for Automated Classification of Unknown Eukaryotic Transposable Elements. Bioinformatics 2009, 25, 1329–1330. [Google Scholar] [CrossRef] [PubMed]
- Benson, G. Tandem Repeats Finder: A Program to Analyze DNA Sequences. Nucleic Acids Res. 1999, 27, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Wang, H. LTR-FINDER: An Efficient Tool for the Prediction of Full-Length LTR Retrotransposons. Nucleic Acids Res. 2007, 35, W265–W268. [Google Scholar] [CrossRef] [PubMed]
- Stanke, M.; Keller, O.; Gunduz, I.; Hayes, A.; Waack, S.; Morgenstern, B. AUGUSTUS: Ab Initio Prediction of Alternative Transcripts. Nucleic Acids Res. 2006, 34, W435–W439. [Google Scholar] [CrossRef]
- Majoros, W.H.; Pertea, M.; Salzberg, S.L. TigrScan and GlimmerHMM: Two Open Source Ab Initio Eukaryotic Gene-Finders. Bioinformatics 2004, 20, 2878–2879. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast Gapped-Read Alignment with Bowtie 2. Nat Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate Alignment of Transcriptomes in the Presence of Insertions, Deletions and Gene Fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential Gene and Transcript Expression Analysis of RNA-Seq Experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef]
- Cantarel, B.L.; Korf, I.; Robb, S.M.C.; Parra, G.; Ross, E.; Moore, B.; Holt, C.; Sánchez Alvarado, A.; Yandell, M. MAKER: An Easy-to-Use Annotation Pipeline Designed for Emerging Model Organism Genomes. Genome Res. 2008, 18, 188–196. [Google Scholar] [CrossRef]
- Zheng, Y.; Jiao, C.; Sun, H.; Rosli, H.G.; Pombo, M.A.; Zhang, P.; Banf, M.; Dai, X.; Martin, G.B.; Giovannoni, J.J.; et al. ITAK: A Program for Genome-Wide Prediction and Classification of Plant Transcription Factors, Transcriptional Regulators, and Protein Kinases. Mol. Plant 2016, 9, 1667–1670. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A Toolkit for Detection and Evolutionary Analysis of Gene Synteny and Collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, İ.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An Information Aesthetic for Comparative Genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed]
- Suyama, M.; Torrents, D.; Bork, P. PAL2NAL: Robust Conversion of Protein Sequence Alignments into the Corresponding Codon Alignments. Nucleic Acids Res. 2006, 34, W609–W612. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A Toolkit Incorporating Gamma-Series Methods and Sliding Window Strategies. Genom. Proteom. Bioinform. 2010, 8, 77–80. [Google Scholar] [CrossRef]
- Castresana, J. Selection of Conserved Blocks from Multiple Alignments for Their Use in Phylogenetic Analysis. Mol. Biol. Evol. 2000, 17, 540–552. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. JModelTest 2: More Models, New Heuristics and Parallel Computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Guindon, S.; Dufayard, J.-F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef]
- Morris, J.L.; Puttick, M.N.; Clark, J.W.; Edwards, D.; Kenrick, P.; Pressel, S.; Wellman, C.H.; Yang, Z.; Schneider, H.; Donoghue, P.C.J. The Timescale of Early Land Plant Evolution. Proc. Natl. Acad. Sci. USA 2018, 115, E2274–E2283. [Google Scholar] [CrossRef]
- Han, M.V.; Thomas, G.W.C.; Lugo-Martinez, J.; Hahn, M.W. Estimating Gene Gain and Loss Rates in the Presence of Error in Genome Assembly and Annotation Using CAFE 3. Mol. Biol. Evol. 2013, 30, 1987–1997. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. ClusterProfiler 4.0: A Universal Enrichment Tool for Interpreting Omics Data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef] [PubMed]
- Mistry, J.; Finn, R.D.; Eddy, S.R.; Bateman, A.; Punta, M. Challenges in Homology Search: HMMER3 and Convergent Evolution of Coiled-Coil Regions. Nucleic Acids Res. 2013, 41, e121. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef] [PubMed]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Qiao, X.; Li, Q.; Yin, H.; Qi, K.; Li, L.; Wang, R.; Zhang, S.; Paterson, A.H. Gene Duplication and Evolution in Recurring Polyploidization–Diploidization Cycles in Plants. Genome Biol. 2019, 20, 38. [Google Scholar] [CrossRef]
| S. taccada | S. hainanensis | |
|---|---|---|
| Assembly summary (Scaffold length ≥ 10 kb) | ||
| Number of scaffolds | 430 | 1014 |
| Total length (bp) | 1,011,548,119 | 1,141,139,833 |
| GC content (%) | 34.78% | 32.71% |
| N50 (scaffolds) (bp) | 124,591,045 | 143,667,403 |
| N90 (scaffolds) (bp) | 92,220,726 | 106,356,671 |
| No. of scaffolds ≥ N50 | 4 | 4 |
| No. of scaffolds ≥ N90 | 8 | 8 |
| BUSCO evaluation of genome assembly | ||
| Complete BUSCOs | 1525 (94.49%) | 1514 (93.80%) |
| Single-copy BUSCOs | 1510 (93.56%) | 1485 (92.01%) |
| Duplicated BUSCOs | 15 (0.93%) | 29 (1.80%) |
| Fragmented BUSCOs | 40 (2.48%) | 41 (2.54%) |
| Missing BUSCOs | 49 (3.04%) | 59 (3.66%) |
| BUSCO evaluation of predicted gene models | ||
| Complete BUSCOs | 1878 (88.5%) | 1974 (93.1%) |
| Single-copy BUSCOs | 1844 (86.9%) | 1922 (90.6%) |
| Duplicated BUSCOs | 34 (1.6%) | 52 (2.5%) |
| Fragmented BUSCOs | 78 (3.7%) | 57 (2.7%) |
| Fragmented BUSCOs | 165 (7.8%) | 90 (4.2%) |
| Annotation summary | ||
| Number of genes | 34,560 | 34,033 |
| Mean gene length (bp) | 2949 | 2902 |
| Mean exon length (bp) | 218 | 228 |
| Mean exon counts per gene | 4.7 | 4.6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.; Mao, X.; He, Z.; Xu, S.; Guo, Z.; Shi, S. Chromosomal-Scale Genome Assemblies of Two Coastal Plant Species, Scaevola taccada and S. hainanensis—Insight into Adaptation Outside of the Common Range. Int. J. Mol. Sci. 2023, 24, 7355. https://doi.org/10.3390/ijms24087355
Li S, Mao X, He Z, Xu S, Guo Z, Shi S. Chromosomal-Scale Genome Assemblies of Two Coastal Plant Species, Scaevola taccada and S. hainanensis—Insight into Adaptation Outside of the Common Range. International Journal of Molecular Sciences. 2023; 24(8):7355. https://doi.org/10.3390/ijms24087355
Chicago/Turabian StyleLi, Sen, Xiaomeng Mao, Ziwen He, Shaohua Xu, Zixiao Guo, and Suhua Shi. 2023. "Chromosomal-Scale Genome Assemblies of Two Coastal Plant Species, Scaevola taccada and S. hainanensis—Insight into Adaptation Outside of the Common Range" International Journal of Molecular Sciences 24, no. 8: 7355. https://doi.org/10.3390/ijms24087355





