Unravelling the Impact of Cyclic Mechanical Stretch in Keratoconus—A Transcriptomic Profiling Study
Abstract
:1. Introduction
2. Results
2.1. Identification of Differentially Expressed Genes
2.2. Droplet Digital PCR (ddPCR) Validation of Differentially Expressed Genes
3. Discussion
3.1. Overview
3.2. Differentially Expressed Genes in HKCs vs. HCFs and KC-Altered Genes Responsive to TGFβ1 Treatment
3.3. KC-Altered Genes Responsive to CMS
3.4. Limitations of Our Study
3.5. Conclusions
4. Materials and Methods
4.1. Culture of Primary Human Corneal Fibroblasts (HCFs) and KC-Derived Cells (HKCs)
4.2. RNA Extraction, Quality Check, Sequencing and Analysis
4.3. Validation of Differentially Expressed Genes Using ddPCR
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sarezky, D.; Orlin, S.E.; Pan, W.M.; VanderBeek, B.L.M. Trends in Corneal Transplantation in Keratoconus. Cornea 2017, 36, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Gupta, N.; Vanathi, M.; Tandon, R. Corneal transplantation in the modern era. Indian J. Med. Res. 2019, 150, 7–22. [Google Scholar] [CrossRef] [PubMed]
- Arnalich-Montiel, F.; Del Barrio, J.L.A.; Alió, J.L. Corneal surgery in keratoconus: Which type, which technique, which outcomes? Eye Vis. 2016, 3, 2. [Google Scholar] [CrossRef] [PubMed]
- Armitage, W.J.; Goodchild, C.; Griffin, M.D.; Gunn, D.J.; Hjortdal, J. High-risk Corneal Transplantation: Recent Developments and Future Possibilities. Transplantation 2019, 103, 2468. [Google Scholar] [CrossRef] [PubMed]
- Teo, A.W.J.; Mansoor, H.; Sim, N.; Lin, M.T.-Y.; Liu, Y.-C. In Vivo Confocal Microscopy Evaluation in Patients with Keratoconus. J. Clin. Med. 2022, 11, 393. [Google Scholar] [CrossRef]
- Xue, J.; Wang, H.; Wang, M.; Zeng, Q.; Jhanji, V.; Kim, A.D.; Wang, M.T.M.; Xu, Y.; Jin, X.; Chen, W. Comparison of deep anterior lamellar keratoplasty and corneal cross-linking in patients with advanced keratoconus. Jpn. J. Ophthalmol. 2022, 66, 52–57. [Google Scholar] [CrossRef]
- Wajnsztajn, D.; Shmueli, O.; Zur, K.; Frucht-Pery, J.; Solomon, A. Predicting factors for the efficacy of cross-linking for keratoconus. PLoS ONE 2022, 17, e0263528. [Google Scholar] [CrossRef]
- Tuft, S.J.; Moodaley, L.C.; Gregory, W.M.; Davison, C.R.; Buckley, R.J. Prognostic Factors for the Progression of Keratoconus. Ophthalmology 1994, 101, 439–447. [Google Scholar] [CrossRef]
- Sharif, R.; Bak-Nielsen, S.; Hjortdal, J.; Karamichos, D. Pathogenesis of Keratoconus: The intriguing therapeutic potential of Prolactin-inducible protein. Prog. Retin. Eye Res. 2018, 67, 150–167. [Google Scholar] [CrossRef]
- Torres-Netto, E.; Al-Otaibi, W.M.; Hafezi, N.L.; Kling, S.; Al-Farhan, H.M.; Randleman, J.B.; Hafezi, F. Prevalence of keratoconus in paediatric patients in Riyadh, Saudi Arabia. Br. J. Ophthalmol. 2018, 102, 1436–1441. [Google Scholar] [CrossRef]
- Jonas, J.B.; Nangia, V.; Matin, A.; Kulkarni, M.; Bhojwani, K. Prevalence and Associations of Keratoconus in Rural Maharashtra in Central India: The Central India Eye and Medical Study. Am. J. Ophthalmol. 2009, 148, 760–765. [Google Scholar] [CrossRef]
- Millodot, M.; Shneor, E.; Albou, S.; Atlani, E.; Gordon-Shaag, A. Prevalence and Associated Factors of Keratoconus in Jerusalem: A Cross-sectional Study. Ophthalmic Epidemiol. 2011, 18, 91–97. [Google Scholar] [CrossRef]
- Waked, N.; Fayad, A.; Fadlallah, A.; El Rami, H. Keratoconus screening in a Lebanese students’ population. J. Fr. Ophtalmol. 2012, 35, 23–29. [Google Scholar] [CrossRef]
- Gomes, J.A.P.; Rodrigues, P.F.; Lamazales, L.L. Keratoconus epidemiology: A review. Saudi. J. Ophthalmol. 2022, 36, 3–6. [Google Scholar] [CrossRef]
- Hashemi, H.; Khabazkhoob, M.; Yazdani, N.; Ostadimoghaddam, H.; Norouzirad, R.; Amanzadeh, K.; Miraftab, M.; Derakhshan, A.; Yekta, A. The prevalence of keratoconus in a young population in Mashhad, Iran. Ophthalmic Physiol. Opt. 2014, 34, 519–527. [Google Scholar] [CrossRef]
- Santodomingo-Rubido, J.; Carracedo, G.; Suzaki, A.; Villa-Collar, C.; Vincent, S.J.; Wolffsohn, J.S. Keratoconus: An updated review. Contact Lens Anterior Eye 2022, 45, 101559. [Google Scholar] [CrossRef]
- Omer, K. Epidemiology of Keratoconus Worldwide. Open Ophthalmol. J. 2018, 12, 289–299. [Google Scholar] [CrossRef]
- Delic, N.C.; Lyons, J.G.; Di Girolamo, N.; Halliday, G.M. Damaging Effects of Ultraviolet Radiation on the Cornea. Photochem. Photobiol. 2017, 93, 920–929. [Google Scholar] [CrossRef]
- Crawford, A.Z.M.; Zhang, J.; Gokul, A.B.; McGhee, C.N.D.; Ormonde, S.E.M. The Enigma of Environmental Factors in Keratoconus. Asia Pac. J. Ophthalmol. 2020, 9, 549–556. [Google Scholar] [CrossRef]
- Ghosh, S.; A Mutalib, H.; Kaur, S.; Ghoshal, R.; Retnasabapathy, S. Effects of contact lens wearing on keratoconus: A confocal microscopy observation. Int. J. Ophthalmol. 2017, 10, 228–234. [Google Scholar] [CrossRef]
- Macsai, M.S.; Varley, G.A.; Krachmer, J.H. Development of Keratoconus After Contact Lens Wear. Arch. Ophthalmol. 1990, 108, 534–538. [Google Scholar] [CrossRef] [PubMed]
- Sahebjada, S.; Chan, E.; Xie, J.; Snibson, G.R.; Daniell, M.; Baird, P.N. Risk factors and association with severity of keratoconus: The Australian study of Keratoconus. Int. Ophthalmol. 2020, 41, 891–899. [Google Scholar] [CrossRef] [PubMed]
- Elubous, K.A.; Al Bdour, M.; Alshammari, T.; Jeris, I.; AlRyalat, S.A.; Roto, A.; Abu Ameerha, M. Environmental Risk Factors Associated With the Need for Penetrating Keratoplasty in Patients With Keratoconus. Cureus 2021, 13, e16506. [Google Scholar] [CrossRef] [PubMed]
- Almusawi, L.A.; Hamied, F.M. Risk Factors for Development of Keratoconus: A Matched Pair Case-Control Study. Clin. Ophthalmol. 2021, 15, 3473–3479. [Google Scholar] [CrossRef]
- Edwards, M.; McGhee, C.N.; Dean, S. The genetics of keratoconus. Clin. Exp. Ophthalmol. 2001, 29, 345–351. [Google Scholar] [CrossRef]
- Stabuc-Silih, M.; Strazisar, M.; Ravnik-Glavac, M.; Hawlina, M.; Glavac, D. Genetics and clinical characteristics of keratoconus. Acta Derm. APA 2010, 19, 3–10. [Google Scholar]
- Coco, G.; Kheirkhah, A.; Foulsham, W.; Dana, R.; Ciolino, J.B. Keratoconus progression associated with hormone replacement therapy. Am. J. Ophthalmol. Case Rep. 2019, 15, 100519. [Google Scholar] [CrossRef]
- McKay, T.B.; Hjortdal, J.; Sejersen, H.; Asara, J.M.; Wu, J.; Karamichos, D. Endocrine and Metabolic Pathways Linked to Keratoconus: Implications for the Role of Hormones in the Stromal Microenvironment. Sci. Rep. 2016, 6, 25534. [Google Scholar] [CrossRef]
- Priyadarsini, S.; McKay, T.B.; Sarker-Nag, A.; Karamichos, D. Keratoconus in vitro and the key players of the TGF-β pathway. Mol. Vis. 2015, 21, 577–588. [Google Scholar]
- Verrecchia, F.; Mauviel, A. Transforming growth factor-beta signaling through the Smad pathway: Role in extracellular matrix gene expression and regulation. J. Investig. Dermatol. 2002, 118, 211–215. [Google Scholar] [CrossRef]
- Engler, C.; Chakravarti, S.; Doyle, J.; Eberhart, C.G.; Meng, H.; Stark, W.J.; Kelliher, C.; Jun, A.S. Transforming Growth Factor-β Signaling Pathway Activation in Keratoconus. Am. J. Ophthalmol. 2011, 151, 752–759.e2. [Google Scholar] [CrossRef]
- YiKim, I.; Kim, M.M.; Kim, S.-J. Transforming Growth Factor-β: Biology and Clinical Relevance. BMB Rep. 2005, 38, 1–8. [Google Scholar] [CrossRef]
- Schiller, M.; Javelaud, D.; Mauviel, A. TGF-β-induced SMAD signaling and gene regulation: Consequences for extracellular matrix remodeling and wound healing. J. Dermatol. Sci. 2004, 35, 83–92. [Google Scholar] [CrossRef]
- Chaudhury, A.; Howe, P.H. The tale of transforming growth factor-beta (TGFβ) signaling: A soigné enigma. IUBMB Life 2009, 61, 929–939. [Google Scholar] [CrossRef]
- Tzavlaki, K.; Moustakas, A. TGF-β Signaling. Biomolecule 2020, 10, 487. [Google Scholar] [CrossRef]
- Tandon, A.; Tovey, J.C.K.; Sharma, A.; Gupta, R.; Mohan, R.R. Role of Transforming Growth Factor Beta in Corneal Function, Biology and Pathology. Curr. Mol. Med. 2010, 10, 565–578. [Google Scholar] [CrossRef]
- Wilson, S.E. TGF beta −1, −2 and −3 in the modulation of fibrosis in the cornea and other organs. Exp. Eye Res. 2021, 207, 108594. [Google Scholar] [CrossRef]
- Kabza, M.; Karolak, J.A.; Rydzanicz, M.; Szcześniak, M.W.; Nowak, D.M.; Ginter-Matuszewska, B.; Polakowski, P.; Ploski, R.; Szaflik, J.P.; Gajecka, M. Collagen synthesis disruption and downregulation of core elements of TGF-β, Hippo, and Wnt pathways in keratoconus corneas. Eur. J. Hum. Genet. 2017, 25, 582–590. [Google Scholar] [CrossRef]
- Sharif, R.; Fowler, B.; Karamichos, D. Collagen cross-linking impact on keratoconus extracellular matrix. PLoS ONE 2018, 13, e0200704. [Google Scholar] [CrossRef]
- Tai, Y.; Woods, E.L.; Dally, J.; Kong, D.; Steadman, R.; Moseley, R.; Midgley, A.C. Myofibroblasts: Function, Formation, and Scope of Molecular Therapies for Skin Fibrosis. Biomolecules 2021, 11, 1095. [Google Scholar] [CrossRef]
- Lyon, D.; McKay, T.B.; Sarkar-Nag, A.; Priyadarsini, S.; Karamichos, D. Human Keratoconus Cell Contractility is Mediated by Transforming Growth Factor-Beta Isoforms. J. Funct. Biomater. 2015, 6, 422–438. [Google Scholar] [CrossRef] [PubMed]
- Asher, R.; Gefen, A.; Moisseiev, E.; Varssano, D. Etiology of Keratoconus: Proposed biomechanical pathogenesis. Silico Cell Tissue Sci. 2014, 1, 3. [Google Scholar] [CrossRef]
- Blackburn, B.J.; Jenkins, M.W.; Rollins, A.M.; Dupps, W.J. A Review of Structural and Biomechanical Changes in the Cornea in Aging, Disease, and Photochemical Crosslinking. Front. Bioeng. Biotechnol. 2019, 7, 66. [Google Scholar] [CrossRef] [PubMed]
- Mlyniuk, P.; Maczynska-Walkowiak, E.; Rzeszewska-Zamiara, J.; Grulkowski, I.; Kaluzny, B.J. Probing biomechanical properties of the cornea with air-puff-based techniques—An overview. Adv. Opt. Technol. 2021, 10, 375–391. [Google Scholar] [CrossRef]
- Li, X.; Wang, S.; Song, J.; Qin, Y.; Yang, J.; He, R.; Chen, W. Cyclic Stretch Induces Inflammatory Cytokines via the Oxidative Stress and NF-ΚB Pathways Activation in Human Keratoconic Fibroblasts. Mol. Cell. Biomech. 2021, 18, 11–19. [Google Scholar] [CrossRef]
- Bawazeer, A.M.; Hodge, W.G.; Lorimer, B. Atopy and keratoconus: A multivariate analysis. Br. J. Ophthalmol. 2000, 84, 834–836. [Google Scholar] [CrossRef]
- Shneor, E.; Millodot, M.; Blumberg, S.; Ortenberg, I.; Behrman, S.; Gordon-Shaag, A. Characteristics of 244 patients with keratoconus seen in an optometric contact lens practice. Clin. Exp. Optom. 2013, 96, 219–224. [Google Scholar] [CrossRef]
- Zadnik, K.; Barr, J.T.; Edrington, T.B.; Everett, D.F.; Jameson, M.; McMahon, T.T.; Shin, J.A.; Sterling, J.L.; Wagner, H.; Gordon, M.O. Baseline findings in the Collaborative Longitudinal Evaluation of Keratoconus (CLEK) Study. Investig. Opthalmol. Vis. Sci. 1998, 39, 2537–2546. [Google Scholar]
- Weed, K.H.; MacEwen, C.J.; Giles, T.; Low, J.; McGhee, C.N. The Dundee University Scottish Keratoconus study: Demographics, corneal signs, associated diseases, and eye rubbing. Eye 2007, 22, 534–541. [Google Scholar] [CrossRef]
- Balasubramanian, S.A.; Pye, D.C.; Willcox, M.D. Effects of eye rubbing on the levels of protease, protease activity and cytokines in tears: Relevance in keratoconus. Clin. Exp. Optom. 2013, 96, 214–218. [Google Scholar] [CrossRef]
- McMonnies, C.W. Mechanisms of Rubbing-Related Corneal Trauma in Keratoconus. Cornea 2009, 28, 607–615. [Google Scholar] [CrossRef]
- Turner, D.C.; Girkin, C.A.; Downs, J.C. The Magnitude of Intraocular Pressure Elevation Associated with Eye Rubbing. Ophthalmology 2018, 126, 171–172. [Google Scholar] [CrossRef]
- Ben-Eli, H.; Erdinest, N.; Solomon, A. Pathogenesis and complications of chronic eye rubbing in ocular allergy. Curr. Opin. Allergy Clin. Immunol. 2019, 19, 526–534. [Google Scholar] [CrossRef]
- Turner, D.C.; Edmiston, A.M.; Zohner, Y.E.; Byrne, K.J.; Seigfreid, W.P.; Girkin, C.A.; Morris, J.S.; Downs, J.C. Transient Intraocular Pressure Fluctuations: Source, Magnitude, Frequency, and Associated Mechanical Energy. Investig. Opthalmol. Vis. Sci. 2019, 60, 2572–2582. [Google Scholar] [CrossRef]
- Lee, M.S.; Harrison, A.R.; Grossman, D.S.; Sloan, F.A. Risk of Glaucoma Among Patients With Benign Essential Blepharospasm. Ophthalmic Plast. Reconstr. Surg. 2010, 26, 434–437. [Google Scholar] [CrossRef]
- Hawkes, E.; Nanavaty, M. Eye Rubbing and Keratoconus: A Literature Review. Int. J. Keratoconus Ectatic Corneal Dis. 2014, 3, 118–121. [Google Scholar] [CrossRef]
- Du, G.-L.; Chen, W.-Y.; Li, X.-N.; He, R.; Feng, P.-F. Induction of MMP-1 and -3 by cyclical mechanical stretch is mediated by IL-6 in cultured fibroblasts of keratoconus. Mol. Med. Rep. 2017, 15, 3885–3892. [Google Scholar] [CrossRef]
- Feng, P.; Li, X.; Chen, W.; Liu, C.; Rong, S.; Wang, X.; Du, G. Combined effects of interleukin-1β and cyclic stretching on metalloproteinase expression in corneal fibroblasts in vitro. Biomed. Eng. Online 2016, 15, 63. [Google Scholar] [CrossRef]
- Papatheodorou, I.; Oellrich, A.; Smedley, D. Linking gene expression to phenotypes via pathway information. J. Biomed. Semant. 2015, 6, 17. [Google Scholar] [CrossRef]
- Rao, M.S.; Van Vleet, T.R.; Ciurlionis, R.; Buck, W.R.; Mittelstadt, S.W.; Blomme, E.A.G.; Liguori, M.J. Comparison of RNA-Seq and Microarray Gene Expression Platforms for the Toxicogenomic Evaluation of Liver From Short-Term Rat Toxicity Studies. Front. Genet. 2018, 9, 636. [Google Scholar] [CrossRef]
- Finotello, F.; Di Camillo, B. Measuring differential gene expression with RNA-seq: Challenges and strategies for data analysis. Briefings Funct. Genom. 2014, 14, 130–142. [Google Scholar] [CrossRef] [PubMed]
- Hao, X.-D.; Chen, X.-N.; Zhang, Y.-Y.; Chen, P.; Wei, C.; Shi, W.-Y.; Gao, H. Multi-level consistent changes of the ECM pathway identified in a typical keratoconus twin’s family by multi-omics analysis. Orphanet J. Rare Dis. 2020, 15, 227. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Wang, Y.; Yang, L.; Wang, Y.; Chen, P.; Wang, Y.; Dong, X.; Xie, L. Histone deacetylase inhibitors blocked activation and caused senescence of corneal stromal cells. Mol. Vis. 2008, 14, 2556–2565. [Google Scholar] [PubMed]
- Chaerkady, R.; Shao, H.; Scott, S.-G.; Pandey, A.; Jun, A.S.; Chakravarti, S. The keratoconus corneal proteome: Loss of epithelial integrity and stromal degeneration. J. Proteom. 2013, 87, 122–131. [Google Scholar] [CrossRef] [PubMed]
- Karolak, J.A.; Gajecka, M. Genomic strategies to understand causes of keratoconus. Mol. Genet. Genom. 2016, 292, 251–269. [Google Scholar] [CrossRef]
- Nakatsu, M.N.; Ding, Z.; Ng, M.Y.; Truong, T.T.; Yu, F.; Deng, S.X. Wnt/β-Catenin Signaling Regulates Proliferation of Human Cornea Epithelial Stem/Progenitor Cells. Investig. Opthalmol. Vis. Sci. 2011, 52, 4734–4741. [Google Scholar] [CrossRef]
- Medeiros, C.S.; Marino, G.K.; Santhiago, M.R.; Wilson, S.E. The Corneal Basement Membranes and Stromal Fibrosis. Investig. Opthalmol. Vis. Sci. 2018, 59, 4044–4053. [Google Scholar] [CrossRef]
- Kanamoto, T.; Hikida, M.; Sato, S.; Oyama, S.; Tachi, Y.; Kuroda, S.; Mazuka, T.; Ebina, K.; Nakai, T.; Nakata, K. Integrin α2β1 plays an important role in the interaction between human articular cartilage-derived chondrocytes and atelocollagen gel. Sci. Rep. 2021, 11, 1757. [Google Scholar] [CrossRef]
- Schey, K.L.; Wang, Z.; Wenke, J.L.; Qi, Y. Aquaporins in the eye: Expression, function, and roles in ocular disease. Biochim. Biophys. Acta Gen. Subj. 2013, 1840, 1513–1523. [Google Scholar] [CrossRef]
- Foster, J.W.; Parikh, R.N.; Wang, J.; Bower, K.S.; Matthaei, M.; Chakravarti, S.; Jun, A.S.; Eberhart, C.G.; Soiberman, U.S. Transcriptomic and Immunohistochemical Analysis of Progressive Keratoconus Reveal Altered WNT10A in Epithelium and Bowman’s Layer. Investig. Opthalmol. Vis. Sci. 2021, 62, 16. [Google Scholar] [CrossRef]
- Cuellar-Partida, G.; Springelkamp, H.; Lucas, S.E.M.; Yazar, S.; Hewitt, A.W.; Iglesias, A.I.; Montgomery, G.W.; Martin, N.G.; Pennell, C.E.; van Leeuwen, E.M.; et al. WNT10A exonic variant increases the risk of keratoconus by decreasing corneal thickness. Hum. Mol. Genet. 2015, 24, 5060–5068. [Google Scholar] [CrossRef]
- Kaldawy, R.M.; Wagner, J.; Ching, S.; Seigel, G.M. Evidence of Apoptotic Cell Death in Keratoconus. Cornea 2002, 21, 206–209. [Google Scholar] [CrossRef]
- Meek, K.; Tuft, S.J.; Huang, Y.; Gill, P.S.; Hayes, S.; Newton, R.H.; Bron, A.J. Changes in Collagen Orientation and Distribution in Keratoconus Corneas. Investig. Opthalmol. Vis. Sci. 2005, 46, 1948–1956. [Google Scholar] [CrossRef]
- di Martino, E.; Ali, M.; Inglehearn, C.F. Matrix metalloproteinases in keratoconus–Too much of a good thing? Exp. Eye Res. 2019, 182, 137–143. [Google Scholar] [CrossRef]
- Ziff, J.L.; Crompton, M.; Powell, H.R.; Lavy, J.A.; Aldren, C.P.; Steel, K.P.; Saeed, S.R.; Dawson, S.J. Mutations and altered expression of SERPINF1 in patients with familial otosclerosis. Hum. Mol. Genet. 2016, 25, 2393–2403. [Google Scholar] [CrossRef]
- He, X.; Cheng, R.; Benyajati, S.; Ma, J.-X. PEDF and its roles in physiological and pathological conditions: Implication in diabetic and hypoxia-induced angiogenic diseases. Clin. Sci. 2015, 128, 805–823. [Google Scholar] [CrossRef]
- Dawson, D.W.; Volpert, O.V.; Gillis, P.; Crawford, S.E.; Xu, H.-J.; Benedict, W.; Bouck, N.P. Pigment Epithelium-Derived Factor: A Potent Inhibitor of Angiogenesis. Science 1999, 285, 245–248. [Google Scholar] [CrossRef]
- Hosomichi, J.; Yasui, N.; Koide, T.; Soma, K.; Morita, I. Involvement of the collagen I-binding motif in the anti-angiogenic activity of pigment epithelium-derived factor. Biochem. Biophys. Res. Commun. 2005, 335, 756–761. [Google Scholar] [CrossRef]
- Alberdi, E.; Hyde, C.C.; Becerra, S.P. Pigment Epithelium-Derived Factor (PEDF) Binds to Glycosaminoglycans: Analysis of the Binding Site. Biochemistry 1998, 37, 10643–10652. [Google Scholar] [CrossRef]
- Meyer, C.; Notari, L.; Becerra, S.P. Mapping the Type I Collagen-binding Site on Pigment Epithelium-derived Factor. J. Biol. Chem. 2002, 277, 45400–45407. [Google Scholar] [CrossRef]
- Meek, K.M. Corneal collagen—Its role in maintaining corneal shape and transparency. Biophys. Rev. 2009, 1, 83–93. [Google Scholar] [CrossRef]
- Jufri, N.F.; Mohamedali, A.; Avolio, A.; Baker, M.S. Mechanical stretch: Physiological and pathological implications for human vascular endothelial cells. Vasc. Cell 2015, 7, 8. [Google Scholar] [CrossRef]
- Quan, T.; He, T.; Kang, S.; Voorhees, J.J.; Fisher, G.J. Ultraviolet Irradiation Alters Transforming Growth Factor β/Smad Pathway in Human Skin In Vivo. J. Investig. Dermatol. 2002, 119, 499–506. [Google Scholar] [CrossRef] [PubMed]
- Fisher, G.J.; Datta, S.; Wang, Z.; Li, X.-Y.; Quan, T.; Chung, J.H.; Kang, S.; Voorhees, J.J. c-Jun–dependent inhibition of cutaneous procollagen transcription following ultraviolet irradiation is reversed by all-trans retinoic acid. J. Clin. Investig. 2000, 106, 663–670. [Google Scholar] [CrossRef]
- Sharif, R.; Hjortdal, J.; Sejersen, H.; Frank, G.; Karamichos, D. Human in vitro Model Reveals the Effects of Collagen Cross-linking on Keratoconus Pathogenesis. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Chan, C. Corneal Cross-Linking for Keratoconus: Current Knowledge and Practice and Future Trends. Asia Pacific J. Ophthalmol. 2020, 9, 557–564. [Google Scholar] [CrossRef]
- Yan, X.; Liu, Z.; Chen, Y. Regulation of TGF-β signaling by Smad7. Acta Biochim. Biophys. Sin. 2009, 41, 263–272. [Google Scholar] [CrossRef]
- Notara, M.; Refaian, N.; Braun, G.; Steven, P.; Bock, F.; Cursiefen, C. Short-term uvb-irradiation leads to putative limbal stem cell damage and niche cell-mediated upregulation of macrophage recruiting cytokines. Stem Cell Res. 2015, 15, 643–654. [Google Scholar] [CrossRef]
- Maurizi, E.; Schiroli, D.; Atkinson, S.D.; Mairs, L.; Courtney, D.G.; O’Hagan, B.; McGilligan, V.E.; Pagnamenta, A.T.; Taylor, J.C.; Vasquez, J.J.; et al. A novel role for CRIM1 in the corneal response to UV and pterygium development. Exp. Eye Res. 2019, 179, 75–92. [Google Scholar] [CrossRef]
- Ardan, T.; Čejková, J. Immunohistochemical expression of matrix metalloproteinases in the rabbit corneal epithelium upon UVA and UVB irradiation. Acta Histochem. 2012, 114, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Raman, N.; Imran, S.A.M.; Noordin, K.B.A.A.; Zaman, W.S.W.K.; Nordin, F. Mechanotransduction in Mesenchymal Stem Cells (MSCs) Differentiation: A Review. Int. J. Mol. Sci. 2022, 23, 4580. [Google Scholar] [CrossRef]
- Youngblood, H.; Cai, J.; Drewry, M.D.; Helwa, I.; Hu, E.; Liu, S.; Yu, H.; Mu, H.; Hu, Y.; Perkumas, K.; et al. Expression of mRNAs, miRNAs, and lncRNAs in Human Trabecular Meshwork Cells Upon Mechanical Stretch. Investig. Opthalmol. Vis. Sci. 2020, 61, 2. [Google Scholar] [CrossRef]
- Kirwan, R.P.; Fenerty, C.H.; Crean, J.; Wordinger, R.J.; Clark, A.F.; O’Brien, C.J. Influence of cyclical mechanical strain on extracellular matrix gene expression in human lamina cribrosa cells in vitro. Mol. Vis. 2005, 11, 798–810. [Google Scholar] [PubMed]
- Shelton, L.; Rada, J.S. Effects of cyclic mechanical stretch on extracellular matrix synthesis by human scleral fibroblasts. Exp. Eye Res. 2007, 84, 314–322. [Google Scholar] [CrossRef]
- Kong, F.; Binas, B.; Moon, J.H.; Kang, S.S.; Kim, H.J. Differential expression of adenylate kinase 4 in the context of disparate stress response strategies of HEK293 and HepG2 cells. Arch. Biochem. Biophys. 2013, 533, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Ström, A.-L.; Zhai, J.; Gal, J.; Bao, S.; Gong, W.; Zhu, H. Enzymatically inactive adenylate kinase 4 interacts with mitochondrial ADP/ATP translocase. Int. J. Biochem. Cell Biol. 2009, 41, 1371–1380. [Google Scholar] [CrossRef] [PubMed]
- Wojcik, K.A.; Kaminska, A.; Blasiak, J.; Szaflik, J.; Szaflik, J.P. Oxidative Stress in the Pathogenesis of Keratoconus and Fuchs Endothelial Corneal Dystrophy. Int. J. Mol. Sci. 2013, 14, 19294–19308. [Google Scholar] [CrossRef]
- Arnal, E.; Peris-Martínez, C.; Menezo, J.L.; Johnsen-Soriano, S.; Romero, F.J. Oxidative Stress in Keratoconus? Investig. Opthalmol. Vis. Sci. 2011, 52, 8592–8597. [Google Scholar] [CrossRef]
- Toprak, I.; Kucukatay, V.; Yildirim, C.; Kilic-Toprak, E.; Kilic-Erkek, O. Increased systemic oxidative stress in patients with keratoconus. Eye 2013, 28, 285–289. [Google Scholar] [CrossRef]
- Heuberger, D.M.; Schuepbach, R.A. Protease-activated receptors (PARs): Mechanisms of action and potential therapeutic modulators in PAR-driven inflammatory diseases. Thromb. J. 2019, 17, 1–24. [Google Scholar] [CrossRef]
- Vliagoftis, H.; Schwingshackl, A.; Milne, C.D.; Duszyk, M.; Hollenberg, M.D.; Wallace, J.L.; Befus, A.; Moqbel, R. Proteinase-activated receptor-2–mediated matrix metalloproteinase-9 release from airway epithelial cells. J. Allergy Clin. Immunol. 2000, 106, 537–545. [Google Scholar] [CrossRef] [PubMed]
- Shetty, R.; Ghosh, A.; Lim, R.R.; Subramani, M.; Mihir, K.; Reshma, A.R.; Ranganath, A.; Nagaraj, S.; Nuijts, R.M.M.A.; Beuerman, R.; et al. Elevated Expression of Matrix Metalloproteinase-9 and Inflammatory Cytokines in Keratoconus Patients Is Inhibited by Cyclosporine A. Investig. Opthalmol. Vis. Sci. 2015, 56, 738–750. [Google Scholar] [CrossRef] [PubMed]
- West-Mays, J.A.; Dwivedi, D.J. The keratocyte: Corneal stromal cell with variable repair phenotypes. Int. J. Biochem. Cell Biol. 2006, 38, 1625–1631. [Google Scholar] [CrossRef]
- Kim, A.; Lakshman, N.; Karamichos, D.; Petroll, W.M. Growth Factor Regulation of Corneal Keratocyte Differentiation and Migration in Compressed Collagen Matrices. Investig. Opthalmol. Vis. Sci. 2010, 51, 864–875. [Google Scholar] [CrossRef]
- Scott, S.-G.; Jun, A.S.; Chakravarti, S. Sphere formation from corneal keratocytes and phenotype specific markers. Exp. Eye Res. 2011, 93, 898–905. [Google Scholar] [CrossRef]
- Dreier, B.; Thomasy, S.M.; Mendonsa, R.; Raghunathan, V.K.; Russell, P.; Murphy, C.J. Substratum Compliance Modulates Corneal Fibroblast to Myofibroblast Transformation. Investig. Opthalmol. Vis. Sci. 2013, 54, 5901–5907. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Backman, L.J.; Zhang, W.; Ling, C.; Danielson, P. Regulation of Keratocyte Phenotype and Cell Behavior by Substrate Stiffness. ACS Biomater. Sci. Eng. 2020, 6, 5162–5171. [Google Scholar] [CrossRef]
- Karamichos, D.; Rich, C.B.; Zareian, R.; Hutcheon, A.E.K.; Ruberti, J.W.; Trinkaus-Randall, V.; Zieske, J.D. TGF-β3 Stimulates Stromal Matrix Assembly by Human Corneal Keratocyte-Like Cells. Investig. Opthalmol. Vis. Sci. 2013, 54, 6612–6619. [Google Scholar] [CrossRef]
- Sharif, R.; Khaled, M.L.; McKay, T.B.; Liu, Y.; Karamichos, D. Transcriptional profiling of corneal stromal cells derived from patients with keratoconus. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Phu, D.; Wray, L.S.; Warren, R.V.; Haskell, R.C.; Orwin, E.J. Effect of Substrate Composition and Alignment on Corneal Cell Phenotype. Tissue Eng. Part A 2011, 17, 799–807. [Google Scholar] [CrossRef]
- Karamichos, D.; Guo, X.Q.; Hutcheon, A.E.K.; Zieske, J.D. Human Corneal Fibrosis: An In Vitro Model. Investig. Opthalmol. Vis. Sci. 2010, 51, 1382–1388. [Google Scholar] [CrossRef] [PubMed]
- Thomas, P.D.; Ebert, D.; Muruganujan, A.; Mushayahama, T.; Albou, L.; Mi, H. PANTHER: Making genome-scale phylogenetics accessible to all. Protein Sci. 2021, 31, 8–22. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Thomas, P. PANTHER Pathway: An Ontology-Based Pathway Database Coupled with Data Analysis Tools. Methods Mol. Biol. 2009, 563, 123–140. [Google Scholar]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 923. [Google Scholar] [CrossRef]
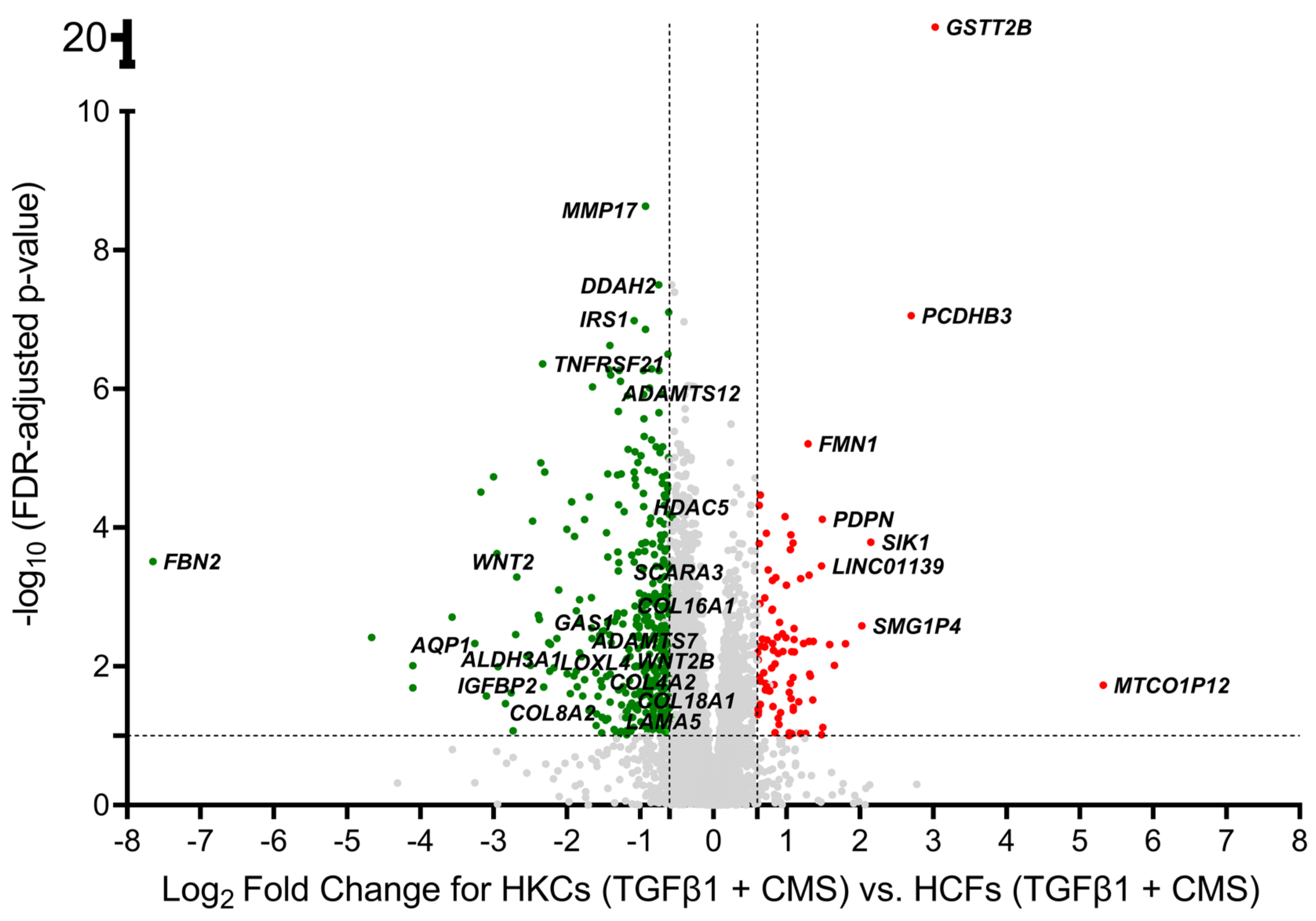


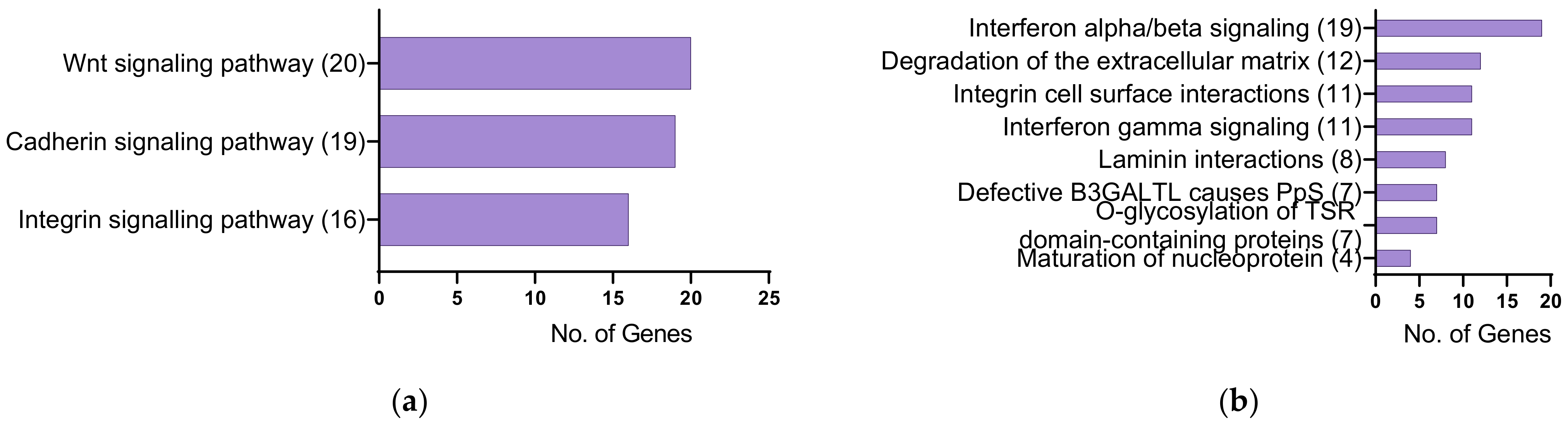


| Gene Symbol | Gene Description | Fold Change | FDR-Adjusted p-Value |
|---|---|---|---|
| Upregulated DEGs | |||
| MTCO1P12 | Mitochondrially Encoded Cytochrome C Oxidase I Pseudogene 12 | 39.9 | 1.87 × 10−2 |
| GSTT2B | Glutathione S-Transferase Theta-2B | 8.1 | 9.78 × 10−24 |
| PCDHB3 | Protocadherin Beta-3 | 6.5 | 8.83 × 10−8 |
| SIK1 | Salt Inducible Kinase 1 | 4.4 | 1.63 × 10−4 |
| SMG1P4 | SMG1 Pseudogene 4 | 4.1 | 2.62 × 10−3 |
| CDH10 | Cadherin 10 | 3.5 | 4.75 × 10−3 |
| RP11-649E7.5 | No description yet | 3.1 | 9.74 × 10−3 |
| LAMC2 | Laminin Subunit Gamma 2 | 3.0 | 4.87 × 10−3 |
| NOTCH2NLB | Notch 2 N-Terminal Like B | 2.8 | 7.57 × 10−2 |
| PDPN | Podoplanin | 2.8 | 7.59 × 10−5 |
| Downregulated DEGs | |||
| FBN2 | Fibrillin 2 | −200.9 | 3.09 × 10−4 |
| SLC8A1 | Solute Carrier Family 8 Member A1 | −25.4 | 3.83 × 10−3 |
| SORT1 | Sortilin 1 | −17.2 | 9.82 × 10−3 |
| PKD1P2 | Polycystin 1, Transient Receptor Potential Channel Interacting Pseudogene 2 | −17.2 | 2.03 × 10−2 |
| ITGA7 | Integrin Subunit Alpha 7 | −11.8 | 1.96 × 10−3 |
| AQP1 | Aquaporin 1 (Colton blood group) | −9.6 | 4.69 × 10−3 |
| DUSP4 | Dual Specificity Phosphatase 1 | −9.0 | 3.08 × 10−5 |
| ELN | Elastin | −8.6 | 2.69 × 10−2 |
| OXTR | Oxytocin Receptor | −8.0 | 1.85 × 10−5 |
| EPHB6 | EPH Receptor B6 | −7.8 | 2.37 × 10−4 |
| Gene Symbol | Gene Description | HKCs (TGFβ1 + CMS) vs. HCFs (TGFβ1 + CMS) | HCF (+/− 10 ng/mL TGFβ1 and CMS) | ||
|---|---|---|---|---|---|
| Fold Change | FDR-Adjusted p-Value | Fold Change | FDR-Adjusted p-Value | ||
| Upregulated DEGs | |||||
| LIPG | Lipase G, Endothelial Type | 2.1 | 1.46 × 10−2 | 9.4 | 4.27 × 10−5 |
| LAMC2 | Laminin Subunit Gamma 2 | 3.0 | 4.87 × 10−3 | 4.5 | 2.55 × 10−6 |
| MYO1D | Myosin 1D | 1.7 | 1.87 × 10−2 | 3.4 | 1.72 × 10−2 |
| BHLHE40 | Basic Helix-Loop-Helix Family Member E40 | 1.7 | 1.55 × 10−3 | 3.1 | 2.67 × 10−9 |
| SYT15 | Synaptotagmin 15 | 1.7 | 2.27 × 10−2 | 2.7 | 5.11 × 10−2 |
| CSMD2 | CUB And Sushi Multiple Domains 2 | 1.6 | 1.64 × 10−2 | 2.5 | 3.41 × 10−2 |
| Downregulated DEGs | |||||
| NPTX1 | Neuronal pentraxin 1 | −2.0 | 1.01 × 10−2 | −10.1 | 1.44 × 10−14 |
| CEMIP | Cell migration inducing hyaluronidase 1 | −2.3 | 4.52 × 10−2 | −8.7 | 2.27 × 10−26 |
| IFI35 | Interferon induced protein 35 | −1.7 | 3.01 × 10−2 | −8.0 | 5.52 × 10−5 |
| SECTM1 | Secreted And Transmembrane 1 | −3.7 | 1.59 × 10−3 | −7.9 | 1.10 × 10−4 |
| CLDN11 | Claudin 11 | −1.6 | 4.78 × 10−3 | −5.3 | 8.50 × 10−19 |
| KCNJ2 | Potassium inwardly rectifying channel subfamily J member 2 | −1.6 | 5.40 × 10−2 | −4.8 | 1.65 × 10−6 |
| ALDH3A1 | Aldehyde Dehydrogenase 3 Family Member A1 | −3.9 | 2.47 × 10−2 | −4.7 | 5.49 × 10−5 |
| PLPP3 | Phospholipid Phosphatase 3 | −1.6 | 4.01 × 10−2 | −4.6 | 4.44 × 10−17 |
| GREM2 | Gremlin 2, DAN family BMP antagonist | −2.0 | 4.97 × 10−3 | −4.5 | 2.42 × 10−11 |
| GJD3 | Gap Junction Protein Delta 3 | −2.1 | 2.54 × 10−2 | −4.3 | 5.06 × 10−2 |
| Gene Symbol | Gene Description/Homology | Fold Change | FDR-Adjusted p-Value (HKC vs. HCF) | Correlation (Conc (ng/mL)) | FDR-Adjusted p-Value (Conc (ng/mL)) |
|---|---|---|---|---|---|
| Positive correlation | |||||
| SEMA7A | Semaphorin 7A (John Milton Hagen Blood Group) | −1.6 | 3.74 × 10−2 | 0.7 | 1.19 × 10−5 |
| TENM4 | Teneurin Transmembrane Protein 4 | −1.6 | 1.14 × 10−2 | 0.7 | 1.95 × 10−5 |
| PSD4 | Pleckstrin and Sec7 Domain Containing 4 | −2.4 | 5.46 × 10−7 | 0.7 | 3.97 × 10−5 |
| POU2F2 | POU Class 2 Homeobox 2 | −1.9 | 2.71 × 10−6 | 0.6 | 9.02 × 10−5 |
| COL7A1 | Collagen Type VII Alpha 1 Chain | −2.0 | 1.59 × 10−3 | 0.6 | 9.31 × 10−5 |
| ZNF365 | Zinc Finger Protein 365 | −3.2 | 1.02 × 10−3 | 0.6 | 9.88 × 10−5 |
| CNN1 | Calponin 1 | −2.5 | 1.93 × 10−3 | 0.6 | 9.88 × 10−5 |
| UCN2 | Urocortin 2 | −2.0 | 3.58 × 10−3 | 0.6 | 1.11 × 10−4 |
| PFKFB4 | 6-Phosphofructo-2-Kinase/Fructose-2,6-Biphosphatase 4 | −1.9 | 1.91 × 10−3 | 0.6 | 1.59 × 10−4 |
| BTBD11 | Homo sapiens KBTBD11 antisense RNA 1 (KBTBD11-AS1) | −1.8 | 6.40 × 10−4 | 0.6 | 2.22 × 10−4 |
| Negative correlation | |||||
| SSH3 | Slingshot Protein Phosphatase 3 | −1.5 | 2.92 × 10−4 | −0.8 | 1.35 × 10−6 |
| IFI35 | Interferon Induced Protein 35 | −1.7 | 3.01 × 10−2 | −0.7 | 2.63 × 10−6 |
| CLDN11 | Claudin 11 | −1.6 | 4.78 × 10−3 | −0.7 | 2.83 × 10−6 |
| BCAM | Basal Cell Adhesion Molecule (Lutheran Blood Group) | −2.1 | 8.10 × 10−6 | −0.7 | 1.26 × 10−5 |
| CCN3 | Cellular Communication Network Factor 3 | 1.7 | 1.06 × 10−2 | −0.7 | 1.64 × 10−5 |
| CD248 | CD248 Molecule | −1.5 | 1.47 × 10−3 | −0.7 | 1.72 × 10−5 |
| PLPP3 | Phospholipid Phosphatase 3 | −1.6 | 4.01 × 10−2 | −0.7 | 2.25 × 10−5 |
| SECTM1 | Secreted and Transmembrane 1 | −3.7 | 1.59 × 10−3 | −0.7 | 2.76 × 10−5 |
| AHRR | Aryl Hydrocarbon Receptor Repressor | −1.9 | 3.99 × 10−4 | −0.7 | 5.36 × 10−5 |
| GREM2 | Gremlin 2, DAN Family BMP Antagonist | −2.0 | 4.97 × 10−3 | −0.7 | 5.74 × 10−5 |
| Gene Symbol | Gene Description or Homology | Fold Change | FDR-Adjusted p-Value (HKC vs. HCF) | FDR-Adjusted p-Value (Stretch) |
|---|---|---|---|---|
| DDAH2 | Dimethylarginine Dimethylaminohydrolase 2 | −1.7 | 3.17 × 10−8 | 1.96 × 10−2 |
| OBSCN | Obscurin, Cytoskeletal Calmodulin And Titin-Interacting RhoGEF | −2.7 | 2.37 × 10−7 | 7.36 × 10−2 |
| CLU | Clusterin | −1.9 | 1.49 × 10−5 | 4.53 × 10−2 |
| LINC00565 | Homo sapiens chromosome 13 open reading frame 46 (C13orf46) | −1.6 | 1.86 × 10−5 | 2.30 × 10−2 |
| RP11-1334A24.5 | No description yet | −1.9 | 3.20 × 10−5 | 1.92 × 10−2 |
| HDAC5 | Histone Deacetylase 5 | −1.5 | 6.45 × 10−5 | 9.94 × 10−3 |
| AK4 | Adenylate Kinase 4 | 1.7 | 5.78 × 10−4 | 1.12 × 10−2 |
| GSN-AS1 | GSN Antisense RNA 1 | −1.7 | 9.95 × 10−4 | 9.65 × 10−2 |
| RPLP0P2 | Ribosomal Protein Lateral Stalk Subunit P0 Pseudogene 2 | −2.1 | 1.36 × 10−3 | 7.98 × 10−2 |
| ITGA10 | Integrin Subunit Alpha 10 | −2.1 | 2.12 × 10−3 | 1.84 × 10−3 |
| KCNS3 | Potassium Voltage-Gated Channel Modifier Subfamily S Member 3 | −1.5 | 2.33 × 10−3 | 5.18 × 10−2 |
| CLDN11 | Claudin 11 | −1.6 | 4.78 × 10−3 | 4.04 × 10−2 |
| F2RL1 | F2R Like Trypsin Receptor 1 | −2.2 | 7.18 × 10−3 | 6.15 × 10−2 |
| Gene Symbol | Gene Description/Homology | Fold Change | FDR-Adjusted p-Value (HKC vs. HCF) | Correlation (Conc (ng/mL)) | FDR-Adjusted p-Value (Conc (ng/mL)) | FDR-Adjusted p-Value (Stretch) |
|---|---|---|---|---|---|---|
| DDAH2 | Dimethylarginine Dimethylaminohydrolase 2 | −1.7 | 3.17 × 10−8 | −0.4 | 3.49 × 10−2 | 1.96 × 10−2 |
| CLU | Clusterin | −1.9 | 1.49 × 10−5 | −0.2 | 3.75 × 10−3 | 4.53 × 10−2 |
| LINC00565 | Homo sapiens chromosome 13 open reading frame 46 (C13orf46) | −1.6 | 1.86 × 10−5 | 0.5 | 5.64 × 10−4 | 2.30 × 10−2 |
| RP11-1334A24.5 | No description yet | −1.9 | 3.20 × 10−5 | 0.6 | 1.02 × 10−3 | 1.92 × 10−2 |
| CLDN11 | Claudin 11 | −1.6 | 4.78 × 10−3 | 0.6 | 2.83 × 10−6 | 4.04 × 10−2 |
| F2RL1 | F2R Like Trypsin Receptor 1 | −2.2 | 7.18 × 10−3 | −0.1 | 1.00 × 10−2 | 6.15 × 10−2 |
| Gene Symbol | Gene Name | ddPCR | RNA-Seq | ||
|---|---|---|---|---|---|
| Effect Size (HKCs vs. HCFs) | p-Value (HKCs vs. HCFs) | Fold Change (HKCs vs. HCFs) | FDR-Adjusted p-Value (HKCs vs. HCFs) | ||
| COL7A1 | Collagen type VII alpha 1 chain | 11.7 | 6.39 × 10−4 | −2.0 | 1.59 × 10−3 |
| SCARA3 | Scavenger receptor class A member 3 | 18.3 | 7.57 × 10−6 | −2.0 | 2.62 × 10−3 |
| SERPINF1 | Serpin family F member 1 | −23.4 | 4.90 × 10−4 | −2.4 | 3.19 × 10−4 |
| FBN2 | Fibrillin 2 | −4.2 | 1.28 × 10−1 | −200.9 | 3.09 × 10−4 |
| Gene Symbol | Gene Name | ddPCR | RNA-Seq | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Effect Size (HKCs vs. HCFs) | p-Value (HKCs vs. HCFs) | Effect Size (TGFβ1 Treatment) | p-Value (TGFβ1 Treatment) | Fold Change (HKCs vs. HCFs) | FDR- Adjusted p-Value (HKCs vs. HCFs) | Correlation (Conc (ng/mL)) | FDR- Adjusted p-Value (TGFβ1 Treatment) | ||
| COL7A1 | Collagen type VII alpha 1 chain | 11.7 | 6.39 × 10−4 | 1.8 | 3.75 × 10−5 | −2.0 | 1.59 × 10−3 | 0.6 | 9.31 × 10−5 |
| SERPINF1 | Serpin family F member 1 | −23.4 | 4.90 × 10−4 | −2.3 | 7.21 × 10−3 | −2.0 | 3.19 × 10−4 | −0.4 | 1.64 × 10−2 |
| Gene Symbol | Gene Name | ddPCR | RNA-Seq | ||||
|---|---|---|---|---|---|---|---|
| Effect Size (HKCs vs. HCFs) | p-Value (HKCs vs. HCFs) | p-Value (Stretch) | Fold Change (HKCs vs. HCFs) | FDR-Adjusted p-Value (HKCs vs. HCFs) | FDR-Adjusted p-Value (Stretch) | ||
| CLU | Clusterin | −5.4 | 0.1695 | 0.0952 | −1.9 | 1.49 × 10−5 | 4.53 × 10−2 |
| Gene Symbol | Gene Name | ddPCR | RNA-Seq | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Fold Change (HKCs vs. HCFs) | p-Value (HKCs vs. HCFs) | Effect Size (TGFβ1 Treatment) | p-Value (TGFβ1 Treatment) | p-Value (Stretch) | Fold Change (HKCs vs. HCFs) | FDR-Adjusted p-Value (HKCs vs. HCFs) | Correlation (Conc (ng/mL)) | FDR- Adjusted p-Value (TGFβ1 Treatment) | FDR- Adjusted p-Value (Stretch) | ||
| CLU | Clusterin | −5.4 | 0.1695 | 0.8 | 0.0979 | 0.0952 | −1.9 | 1.49 × 10−5 | −0.2 | 3.75 × 10−3 | 4.53 × 10−2 |
| Sample ID | Age (y/o) | Gender | Ethnicity | Cause of Death |
|---|---|---|---|---|
| HCF-1 | 65 | Male | Caucasian | Cardiopulmonary Arrest (CPA) |
| HCF-2 | 63 | Male | Caucasian | Acute STEMI |
| HCF-3 | 72 | Female | Caucasian | Motor vehicle accident (MVA) |
| HCF-4 | 71 | Male | Caucasian | Cardiorespiratory Failure |
| HKC-1 | 43 | Female | Unknown | N/A (transplant patient) |
| HKC-2 | 19 | Male | Unknown | N/A (transplant patient) |
| HKC-3 | 69 | Male | Unknown | N/A (transplant patient) |
| HKC-4 | 31 | Male | African American | N/A (transplant patient) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Akoto, T.; Cai, J.; Nicholas, S.; McCord, H.; Estes, A.J.; Xu, H.; Karamichos, D.; Liu, Y. Unravelling the Impact of Cyclic Mechanical Stretch in Keratoconus—A Transcriptomic Profiling Study. Int. J. Mol. Sci. 2023, 24, 7437. https://doi.org/10.3390/ijms24087437
Akoto T, Cai J, Nicholas S, McCord H, Estes AJ, Xu H, Karamichos D, Liu Y. Unravelling the Impact of Cyclic Mechanical Stretch in Keratoconus—A Transcriptomic Profiling Study. International Journal of Molecular Sciences. 2023; 24(8):7437. https://doi.org/10.3390/ijms24087437
Chicago/Turabian StyleAkoto, Theresa, Jingwen Cai, Sarah Nicholas, Hayden McCord, Amy J. Estes, Hongyan Xu, Dimitrios Karamichos, and Yutao Liu. 2023. "Unravelling the Impact of Cyclic Mechanical Stretch in Keratoconus—A Transcriptomic Profiling Study" International Journal of Molecular Sciences 24, no. 8: 7437. https://doi.org/10.3390/ijms24087437








