Characterization of Angraecum (Angraecinae, Orchidaceae) Plastomes and Utility of Sequence Variability Hotspots
Abstract
:1. Introduction
2. Results
2.1. Characteristics of the Plastome
2.2. Repeated Analysis
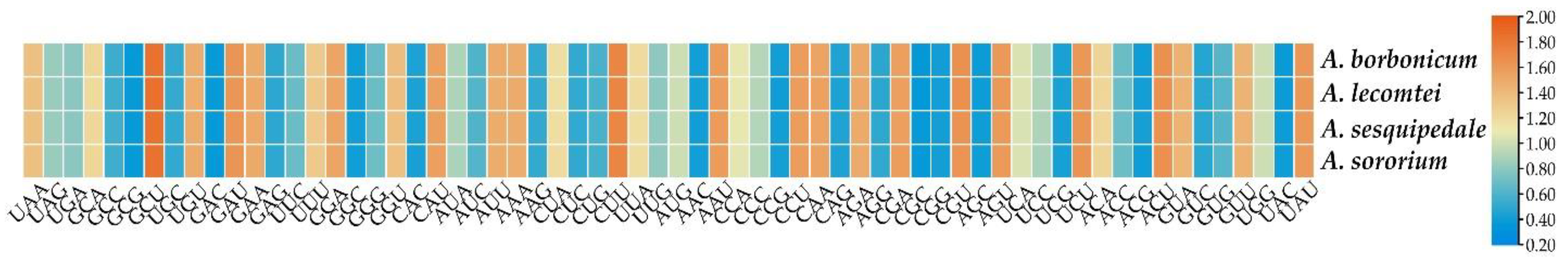
2.3. Codon Usage Analyses
2.4. Selective Pressure Analysis
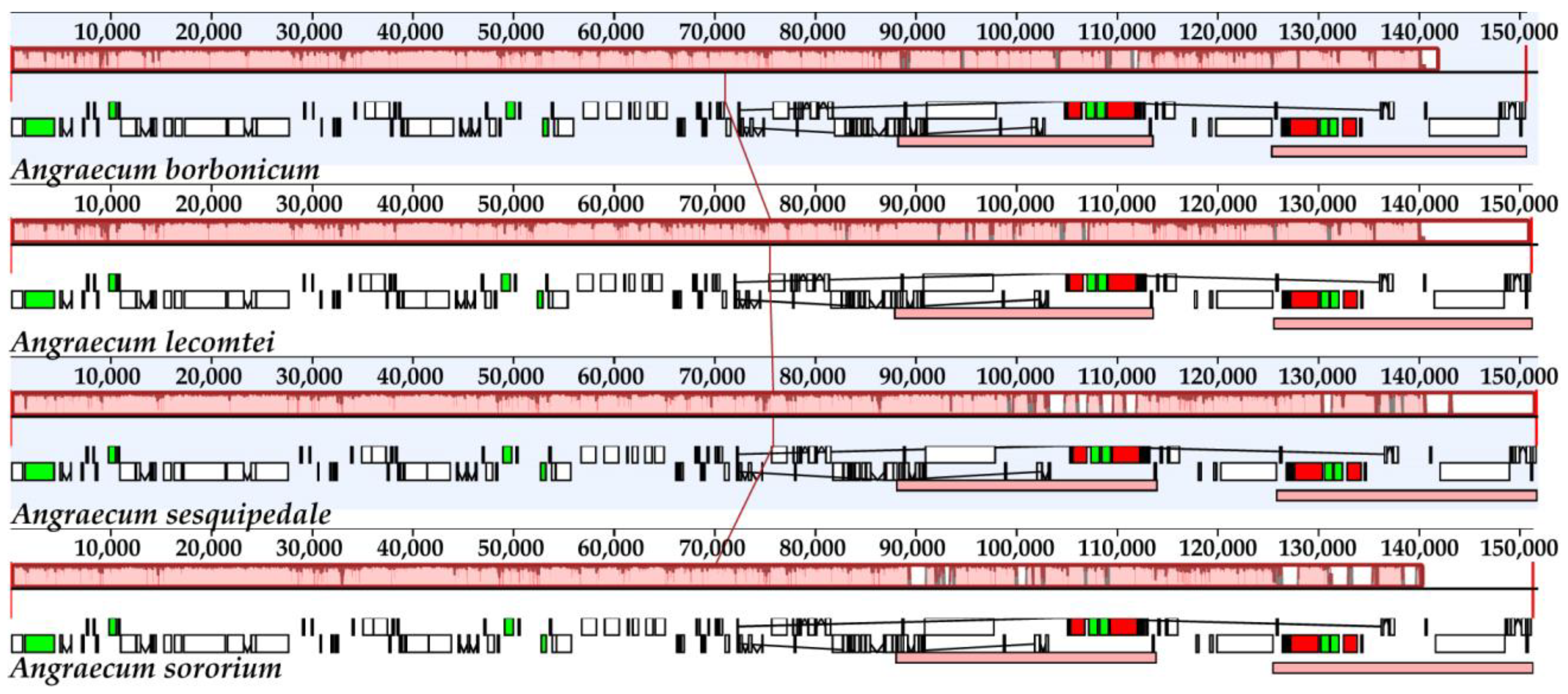
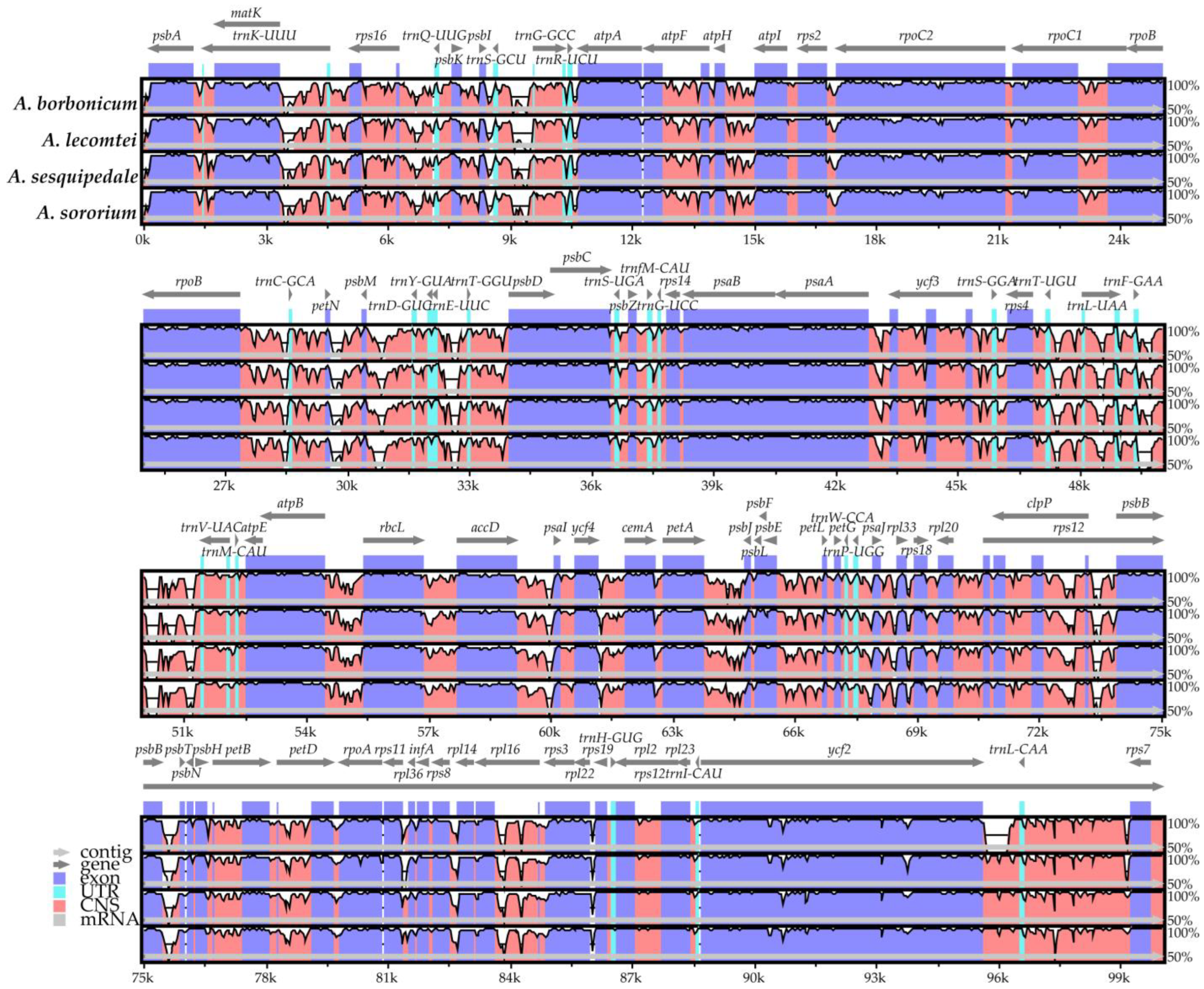
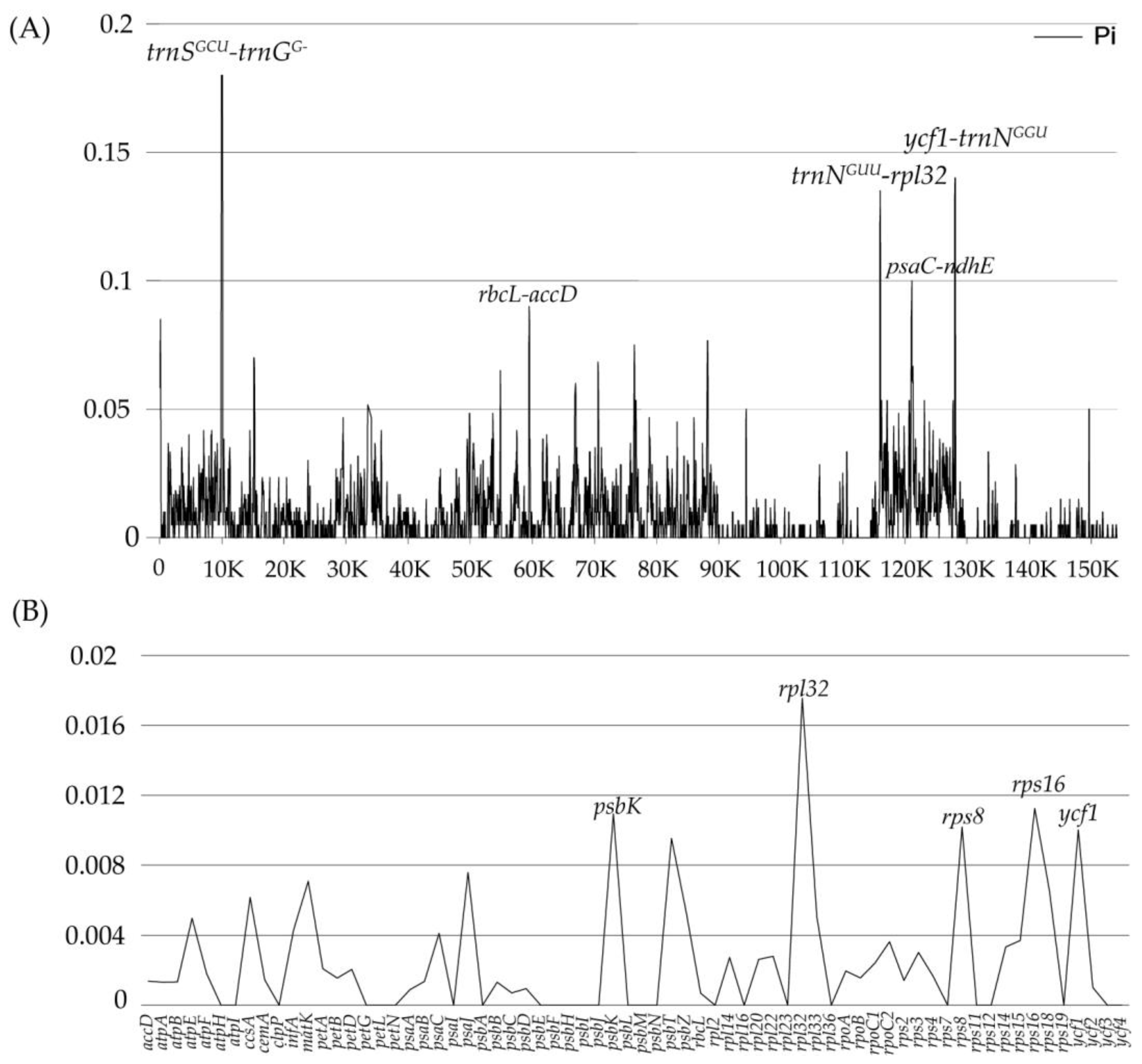
2.5. Plastome Sequence Divergence and Barcoding Investigation
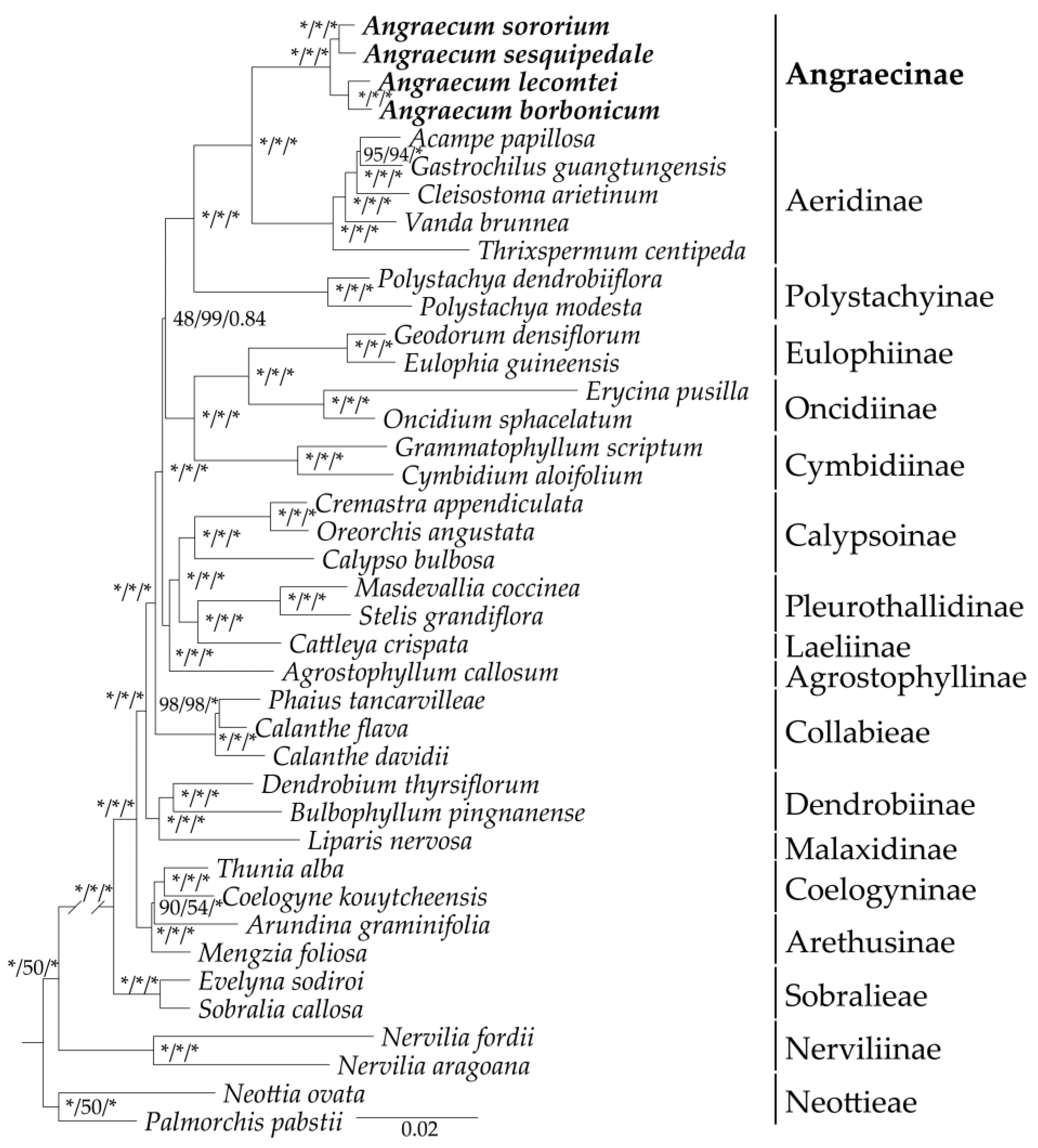
2.6. Phylogenetic Analysis
3. Discussion
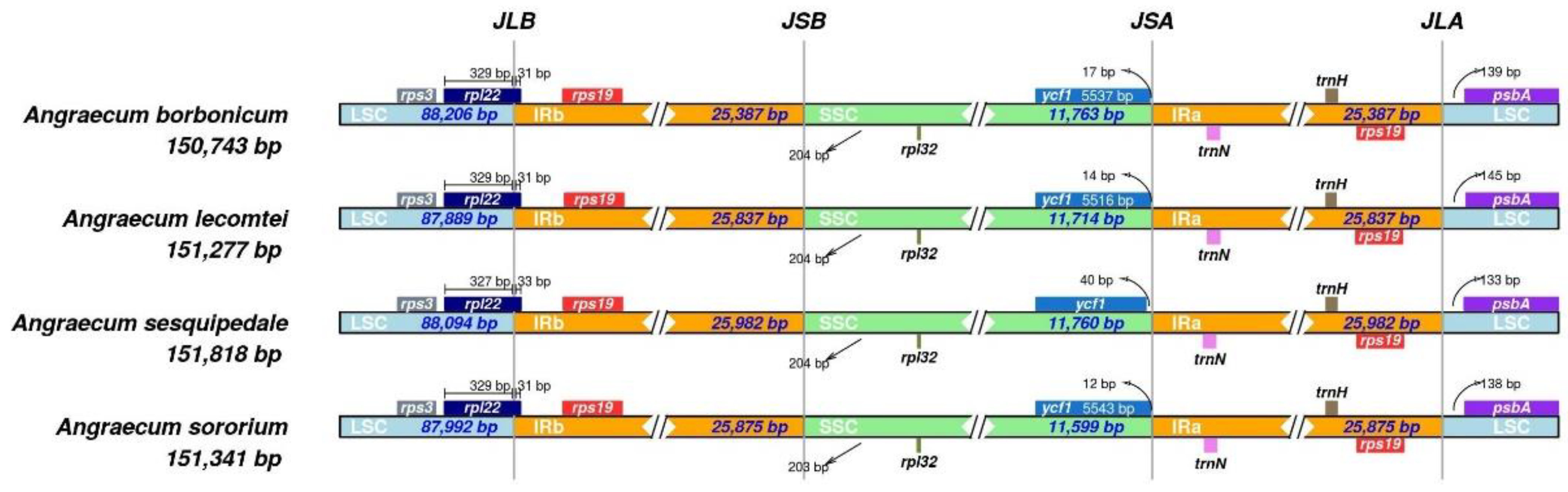
3.1. The Plastome Characteristics and Structural Evolution
3.2. Plastid Genomic Evolutionary Hotspots
3.3. Phylogenetic Analysis
4. Materials and Methods
4.1. Taxon Sampling and Sequencing
4.2. Plastome Assembly and Annotation
4.3. Plastome Comparative and Codon Usage Analysis
4.4. Selective Pressure Estimation
4.5. Sequence Divergence, Barcoding Investigation and Phylogeny
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Harder, L.D.; Johnson, S.D. Darwin’s beautiful contrivances: Evolutionary and functional evidence for floral adaptation. New Phytol. 2009, 183, 530–545. [Google Scholar] [CrossRef] [PubMed]
- Hermans, J.; Verlynde, S.; Rajaovelona, L.; Cribb, P.J.; Hervouet, J.M. New species and nomenclatural changes in Angraecum (Orchidaceae) from Madagascar. Kew Bull. 2020, 75, 49. [Google Scholar] [CrossRef]
- Hermans, J.; Rajaovelona, L.; Cribb, P. Angraecum inflatum, a new species in Angraecinae (Orchidaceae) from Madagascar. Kew Bull. 2021, 76, 513–517. [Google Scholar] [CrossRef]
- Simo-Droissart, M.; Micheneau, C.; Sonké, B.; Droissart, V.; Plunkett, G.M.; Lowry, P.P., II; Hardy, O.J.; Stévart, T. Morphometrics and molecular phylogenetics of the continental African species of Angraecum section Pectinaria (Orchidaceae). Plant Ecol. Evol. 2013, 146, 295–309. [Google Scholar] [CrossRef]
- Andriananjamanantsoa, H.N.; Engberg, S.; Louis, E.E., Jr.; Brouillet, L. Diversification of Angraecum (Orchidaceae, Vandeae) in Madagascar: Revised phylogeny reveals species accumulation through time rather than rapid radiation. PLoS ONE 2016, 11, e0163194. [Google Scholar] [CrossRef] [PubMed]
- Pridgeon, A.M.; Cribb, P.J.; Chase, M.W.; Rasmussen, F.N. Genera Orchidacearum Volume 6: Epidendroideae (Part 3); OUP Oxford Press: Oxford, UK, 2014. [Google Scholar]
- Nilsson, L.A.; Jonsson, L.; Rason, L.; Randrianjohany, E. Monophily and pollination mechanisms in Angraecum arachnites Schltr. (Orchidaceae) in a guild of long-tongued hawk-moths (Sphingidae) in Madagascar. Biol. J. Linn. Soc. 1985, 26, 1–19. [Google Scholar] [CrossRef]
- Micheneau, C.; Fournel, J.; Pailler, T. Bird Pollination in an Angraecoid Orchid on Reunion Island (Mascarene Archipelago, Indian Ocean). Ann. Bot. 2006, 97, 965–974. [Google Scholar] [CrossRef]
- Buyun, L.I.; Cherevchenko, T.M.; Kovalska, L.A.; Ivannikov, R.V. Reproductive biology of Angraecum eburneum subsp. superbum (Orchidaceae) under glasshouse conditions. Environ. Exp. Biol. 2015, 13, 33–39. [Google Scholar]
- Garay, L.A. Systematics of the genus Angraecum (Orchidaceae). Kew Bull. 1973, 28, 495–516. [Google Scholar] [CrossRef]
- Carlsward, B.S.; Whitten, W.M.; Williams, N.H.; Bytebier, B. Molecular phylogeny of Vandeae (Orchidaceae) and the evolution of leaflessness. Am. J. Bot. 2006, 93, 770–786. [Google Scholar] [CrossRef]
- Micheneau, C.; Carlsward, B.S.; Fay, M.F.; Bytebier, B.; Pailler, T.; Chase, M.W. Phylogenetics and biogeography of Mascarene angraecoid orchids (Vandeae, Orchidaceae). Mol. Phylogenet. Evol. 2008, 46, 908–922. [Google Scholar] [CrossRef] [PubMed]
- Simo-Droissart, M.; Plunkett, G.M.; Droissart, V.; Edwards, M.B.; Farminhão, J.N.M.; Ječmenica, V.; D’haijère, T.; Lowry, P.P.; Sonké, B.; Micheneau, C.; et al. New phylogenetic insights toward developing a natural generic classification of African angraecoid orchids (Vandeae, Orchidaceae). Mol. Phylogenet. Evol. 2018, 126, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Farminhão, J.N.M.; Verlynde, S.; Kaymak, E.; Droissart, V.; Simo-Droissart, M.; Collobert, G.; Martos, F.; Stévart, T. Rapid radiation of angraecoids (Orchidaceae, Angraecinae) in tropical Africa characterized by multiple karyotypic shifts under major environmental instability. Mol. Phylogenet. Evol. 2021, 159, 107105. [Google Scholar] [CrossRef]
- Niehuis, O.; Hartig, G.; Grath, S.; Pohl, H.; Lehmann, J.; Tafer, H.; Donath, A.; Krauss, V.; Eisenhardt, C.; Hertel, J.; et al. Genomic and morphological evidence converge to resolve the enigma of Strepsiptera. Curr. Biol. 2012, 22, 1309–1313. [Google Scholar] [CrossRef] [PubMed]
- Lyu, R.; Xiao, J.; Li, M.; Luo, Y.; He, J.; Cheng, J.; Xie, L. Phylogeny and Historical Biogeography of the East Asian Clematis Group, Sect. Tubulosae, Inferred from Phylogenomic Data. Int. J. Mol. Sci. 2023, 24, 3056. [Google Scholar] [CrossRef] [PubMed]
- Yan, R.; Gu, L.; Qu, L.; Wang, X.; Hu, G. New Insights into Phylogenetic Relationship of Hydrocotyle (Araliaceae) Based on Plastid Genomes. Int. J. Mol. Sci. 2023, 24, 16629. [Google Scholar] [CrossRef] [PubMed]
- Ohyama, K. Chloroplast and mitochondrial genomes from a liverwort, Marchantia polymorpha—Gene organization and molecular evolution. Biosci. Biotechnol. Biochem. 1996, 60, 16–24. [Google Scholar] [CrossRef] [PubMed]
- Givnish, T.J.; Spalink, D.; Ames, M.; Lyon, S.P.; Hunter, S.J.; Zuluaga, A.; Iles, W.J.; Clements, M.A.; Arroyo, M.T.; Leebens-Mack, J.; et al. Orchid phylogenomics and multiple drivers of their extraordinary diversification. Proc. Biol. Sci. B 2015, 282, 2108–2111. [Google Scholar] [CrossRef]
- Li, Y.X.; Li, Z.H.; Schuiteman, A.; Chase, M.W.; Li, J.W.; Huang, W.C.; Hidayat, A.; Wu, S.S.; Jin, X.H. Phylogenomics of Orchidaceae based on plastid and mitochondrial genomes. Mol. Phylogenet. Evol. 2019, 139, 106540. [Google Scholar] [CrossRef]
- Kim, Y.K.; Jo, S.; Cheon, S.H.; Kwak, M.; Kim, Y.D.; Kim, K.J. Plastome evolution and phylogeny of subtribe Aeridinae (Vandeae, Orchidaceae). Mol. Phylogenet. Evol. 2020, 144, 106721. [Google Scholar] [CrossRef]
- Liu, D.K.; Tu, X.D.; Zhao, Z.; Zeng, M.Y.; Zhang, S.; Ma, L.; Zhang, G.Q.; Wang, M.M.; Liu, Z.J.; Lan, S.R.; et al. Plastid phylogenomic data yield new and robust insights into the phylogeny of Cleisostoma-Gastrochilus clades (Orchidaceae, Aeridinae). Mol. Phylogenet. Evol. 2020, 145, 106729. [Google Scholar] [CrossRef] [PubMed]
- Tu, X.D.; Liu, D.K.; Xu, S.W.; Zhou, C.Y.; Gao, X.Y.; Zeng, M.Y.; Zhang, S.; Chen, J.L.; Ma, L.; Zhou, Z.; et al. Plastid phylogenomics improves resolution of phylogenetic relationship in the Cheirostylis and Goodyera clades of Goodyerinae (Orchidoideae, Orchidaceae). Mol. Phylogenet. Evol. 2021, 164, 107269. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.Y.; Yang, J.X.; Bai, M.Z.; Zhang, G.Q.; Liu, Z.J. The chloroplast genome evolution of Venus slipper (Paphiopedilum): IR expansion, SSC contraction, and highly rearranged SSC regions. BMC Plant Biol. 2021, 21, 248. [Google Scholar] [CrossRef] [PubMed]
- Niu, Z.; Zhu, S.; Pan, J.; Li, L.; Sun, J.; Ding, X. Comparative analysis of Dendrobium plastomes and utility of plastomic mutational hotspots. Sci. Rep. 2017, 7, 2073. [Google Scholar]
- Jiang, H.; Tian, J.; Yang, J.; Dong, X.; Zhong, Z.; Mwachala, G.; Zhang, C.; Hu, G.; Wang, Q. Comparative and phylogenetic analyses of six Kenya Polystachya (Orchidaceae) species based on the complete chloroplast genome sequences. BMC Plant Biol. 2022, 22, 177. [Google Scholar] [CrossRef]
- Chen, Y.; Zhong, H.; Zhu, Y.; Huang, Y.; Wu, S.; Liu, Z.; Lan, S.; Zhai, J. Plastome structure and adaptive evolution of Calanthe s.l. species. PeerJ 2020, 8, e10051. [Google Scholar] [CrossRef]
- Li, L.; Wu, Q.; Fang, L.; Wu, K.; Li, M.; Zeng, S. Comparative Chloroplast Genomics and Phylogenetic Analysis of Thuniopsis and Closely Related Genera within Coelogyninae (Orchidaceae). Front. Genet. 2022, 13, 850201. [Google Scholar] [CrossRef]
- Schelkunov, M.I.; Shtratnikova, V.Y.; Nuraliev, M.S.; Selosse, M.A.; Penin, A.A.; Logacheva, M.D. Exploring the limits for reduction of plastid genomes: A case study of the mycoheterotrophic orchids Epipogium aphyllum and Epipogium roseum. Genome Biol. Evol. 2015, 7, 1179–1191. [Google Scholar] [CrossRef]
- Kim, Y.K.; Jo, S.; Cheon, S.H.; Joo, M.J.; Hong, J.R.; Kwak, M.; Kim, K.J. Plastome Evolution and Phylogeny of Orchidaceae, with 24 New Sequences. Front. Plant Sci. 2020, 11, 22. [Google Scholar] [CrossRef]
- Zavala-Páez, M.; Vieira, L.D.N.; Baura, V.A.D.; Balsanelli, E.; Souza, E.M.D.; Cevallos, M.C.; Smidt, E.D.C. Comparative plastid genomics of neotropical Bulbophyllum (Orchidaceae; Epidendroideae). Front. Plant Sci. 2020, 11, 799. [Google Scholar] [CrossRef]
- Feng, Y.L.; Wicke, S.; Li, J.W.; Han, Y.; Lin, C.S.; Li, D.Z.; Zhou, T.T.; Huang, W.C.; Huang, L.Q.; Jin, X.H. Lineage-specific reductions of plastid genomes in an orchid tribe with partially and fully mycoheterotrophic species. Genome Biol. Evol. 2016, 8, 2164–2175. [Google Scholar] [CrossRef] [PubMed]
- Sanderson, M.J.; Copetti, D.; Burquez, A.; Bustamante, E.; Charboneau, J.L.M.; Eguiarte, L.E.; Kumar, S.; Lee, H.O.; Lee, J.; McMahon, M.; et al. Exceptional reduction of the plastid genome of saguaro cactus (Carnegiea gigantea): Loss of the ndh gene suite and inverted repeat. Am. J. Bot. 2015, 102, 1115–1127. [Google Scholar] [CrossRef] [PubMed]
- Thode, V.A.; Lohmann, L.G. Comparative Chloroplast Genomics at Low Taxonomic Levels: A Case Study Using Amphilophium (Bignonieae, Bignoniaceae). Front. Plant Sci. 2019, 10, 796. [Google Scholar] [CrossRef] [PubMed]
- Dugas, D.V.; Hernandez, D.; Koenen, E.J.; Schwarz, E.; Straub, S.; Hughes, C.E.; Jansen, R.K.; Nageswara-Rao, M.; Staats, M.; Trujillo, J.T.; et al. Mimosoid legume plastome evolution: IR expansion, tandem repeat expansions, and accelerated rate of evolution in clpP. Sci. Rep. 2015, 5, 16958. [Google Scholar] [CrossRef] [PubMed]
- Cavalier-Smith, T. Chloroplast evolution: Secondary symbiogenesis and multiple losses. Curr. Biol. 2002, 12, R62–R64. [Google Scholar] [CrossRef] [PubMed]
- Gu, C.; Ma, L.; Wu, Z.; Chen, K.; Wang, Y. Comparative analyses of chloroplast genomes from 22 Lythraceae species: Inferences for phylogenetic relationships and genome evolution within Myrtales. BMC Plant Biol. 2019, 19, 281. [Google Scholar] [CrossRef] [PubMed]
- Iriarte, A.; Lamolle, G.; Musto, H. Codon Usage Bias: An endless tale. J. Mol. Evol. 2021, 89, 589–593. [Google Scholar] [CrossRef]
- Fay, J.C.; Wu, C.-I. Sequence Divergence, Functional Constraint, and Selection in Protein Evolution. Annu. Rev. Genom. Hum. Genet. 2003, 4, 213–235. [Google Scholar] [CrossRef]
- Zhang, J. Rates of conservative and radical nonsynonymous nucleotide substitutions in mammalian nuclear genes. J. Mol. Evol. 2000, 50, 56–68. [Google Scholar] [CrossRef]
- Chase, M.W.; Cameron, K.M.; Freudestein, J.V.; Pridgeon, A.M.; Salazar, G.; Van den Berg, C.; Schuiteman, A. An update classification of Orchidaceae. Bot. J. Linn. Soc. 2015, 177, 151–174. [Google Scholar] [CrossRef]
- Freudenstein, J.V.; Chase, M.W. Phylogenetic relationships in Epidendroideae (Orchidaceae), one of the great flowering plant radiations: Progressive specialization and diversification. Ann. Bot. 2015, 115, 665–681. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, N.; Lewis, P.O.; Lemmon, E.M.; Lemmon, A.R.; Holsinger, K.E. Anchored phylogenomics improves the resolution of evolutionary relationships in the rapid radiation of Protea L. Am. J. Bot. 2017, 104, 102–115. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.-J.; Yu, W.-B.; Yang, J.-B.; Song, Y.; DePamphilis, C.W.; Yi, T.S.; Li, D.Z. GetOrganelle: A fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 2020, 21, 241. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Wick, R.R.; Schultz, M.B.; Zobel, J.; Holt, K.E. Bandage: Interactive visualization of de novo genome assemblies. Bioinformatics 2015, 31, 3350–3352. [Google Scholar] [CrossRef] [PubMed]
- Qu, X.J.; Moore, M.J.; Li, D.Z.; Yi, T.S. PGA: A software package for rapid, accurate, and flexible batch annotation of plastomes. Plant Methods. 2019, 15, 50. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Greiner, S.; Lehwark, P.; Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: Expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 2019, 47, W59–W64. [Google Scholar] [CrossRef]
- Rissman, A.I.; Mau, B.; Biehl, B.S.; Darling, A.E.; Glasner, J.D.; Perna, N.T. Reordering contigs of draft genomes using the Mauve aligner. Bioinformatics 2009, 25, 2071–2073. [Google Scholar] [CrossRef]
- Amiryousefi, A.; Hyvonen, J.; Poczai, P. Irscope: An online program to visualize the junction sites of chloroplast genomes. Bioinformatics 2018, 34, 3030–3031. [Google Scholar] [CrossRef]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef] [PubMed]
- Villanueva, R.A.M.; Chen, Z.J. ggplot2: Elegant Graphics for Data Analysis (2nd Ed.). Meas. Interdiscip. Res. Perspect. 2019, 17, 160–167. [Google Scholar] [CrossRef]
- Zhang, D.; Gao, F.; Jakovlić, I.; Zou, H.; Zhang, J.; Li, W.X.; Wang, G.T. PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Mol. Ecol. Resour. 2020, 20, 348–355. [Google Scholar] [CrossRef] [PubMed]
- Xia, X. Dambe7: New and improved tools for data analysis in molecular biology and evolution. Mol. Biol. Evol. 2018, 35, 1550–1552. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. Tbtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhang, Z.; Vang, S.; Yu, J.; Wong, G.K.; Wang, J. Correlation between Ka/Ks and Ks is related to substitution model and evolutionary lineage. J. Mol. Evol. 2009, 68, 414–423. [Google Scholar] [CrossRef] [PubMed]
- Brudno, M.; Malde, S.; Poliakov, A.; Do, C.B.; Couronne, O.; Dubchak, I.; Batzoglou, S. Glocal alignment: Finding rearrangements during alignment. Bioinformatics 2003, 19, i54–i62. [Google Scholar] [CrossRef]
- Rozas, J.; Sánchez-DelBarrio, J.C.; Messeguer, X.; Rozas, R. Dnasp, dna polymorphism analyses by the coalescent and other methods. Bioinformatics 2003, 19, 2496–2497. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Capella-Gutierrez, S.; Silla-Martinez, J.M.; Gabaldon, T. Trimal: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the 2010 Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar]
- Stamatakis, A.; Hoover, P.; Rougemont, J. A rapid bootstrap algorithm for the RAxML Web servers. Syst. Biol. 2008, 57, 758–771. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evol. Int. J. Org. Evol. 1985, 39, 783–791. [Google Scholar] [CrossRef]
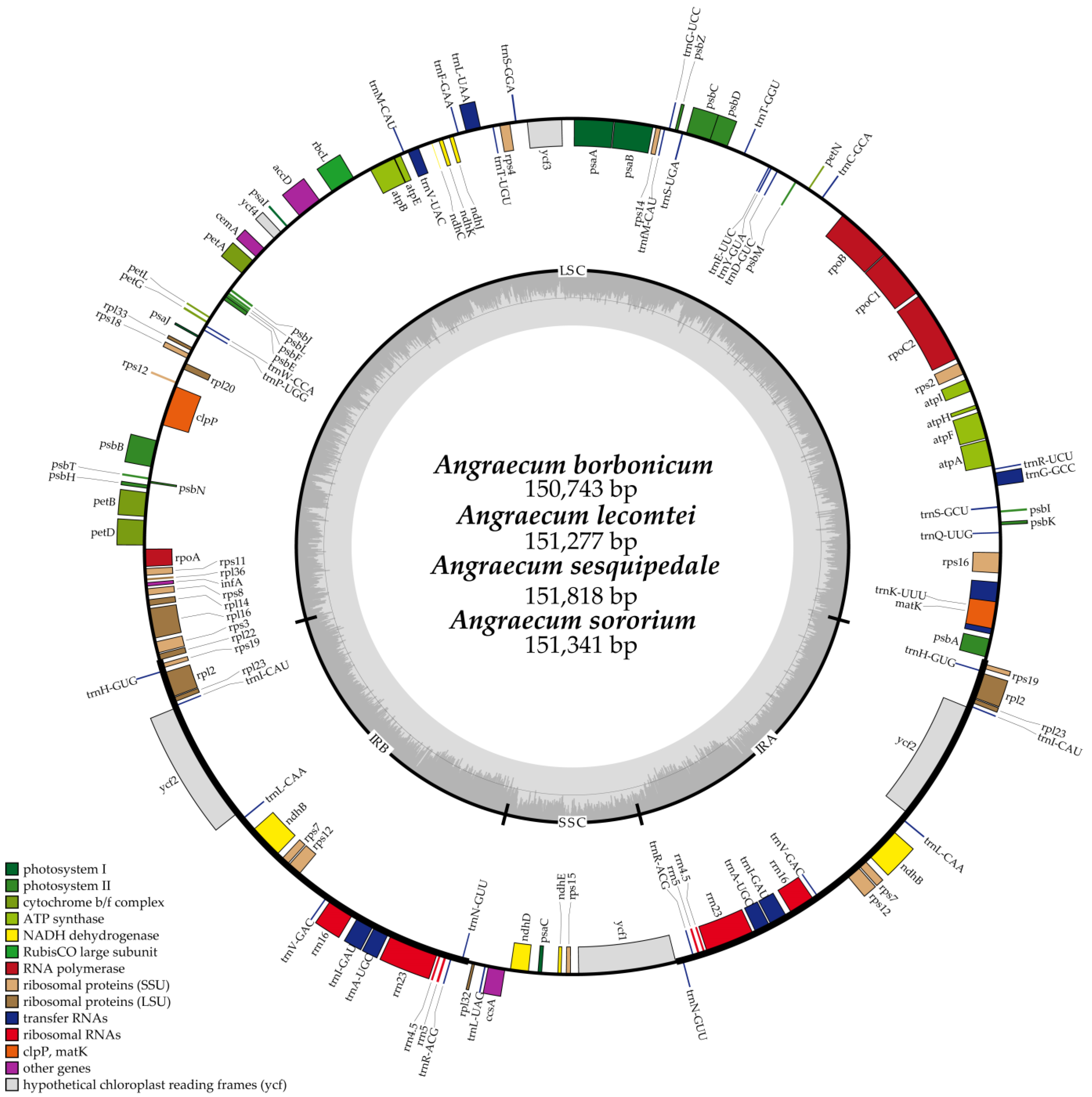


| Taxa | Size (bp) | GC Content (%) | LSC Size in bp (%) | IR Size in bp (%) | SSC Size in bp (%) | Total Number of Gene | CDS | tRNA Gene | rRNA Gene | Number of ndh Fragment |
|---|---|---|---|---|---|---|---|---|---|---|
| A. borbonicum | 150,743 | 36.7 | 88,206 (58.5) | 25,387 (33.7) | 11,763 (7.8) | 120 | 74 | 38 | 8 | 7 |
| A. lecomtei | 151,277 | 36.8 | 87,889 (58.1) | 25,733 (34.0) | 11,922 (7.9) | 120 | 74 | 38 | 8 | 7 |
| A. sesquipedale | 151,818 | 36.8 | 88,904 (58.6) | 25,982 (34.2) | 11,760 (7.7) | 120 | 74 | 38 | 8 | 7 |
| A. sororium | 151,341 | 36.9 | 87,992 (58.1) | 25,875 (34.2) | 11,599 (7.7) | 120 | 74 | 38 | 8 | 7 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, C.-Y.; Lin, W.-J.; Li, R.; Wu, Y.; Liu, Z.-J.; Li, M.-H. Characterization of Angraecum (Angraecinae, Orchidaceae) Plastomes and Utility of Sequence Variability Hotspots. Int. J. Mol. Sci. 2024, 25, 184. https://doi.org/10.3390/ijms25010184
Zhou C-Y, Lin W-J, Li R, Wu Y, Liu Z-J, Li M-H. Characterization of Angraecum (Angraecinae, Orchidaceae) Plastomes and Utility of Sequence Variability Hotspots. International Journal of Molecular Sciences. 2024; 25(1):184. https://doi.org/10.3390/ijms25010184
Chicago/Turabian StyleZhou, Cheng-Yuan, Wen-Jun Lin, Ruyi Li, Yuhan Wu, Zhong-Jian Liu, and Ming-He Li. 2024. "Characterization of Angraecum (Angraecinae, Orchidaceae) Plastomes and Utility of Sequence Variability Hotspots" International Journal of Molecular Sciences 25, no. 1: 184. https://doi.org/10.3390/ijms25010184





