Discovery of Pathogenic Variants Associated with Idiopathic Recurrent Pregnancy Loss Using Whole-Exome Sequencing
Abstract
:1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Subjects
4.2. Whole-Exome Sequencing
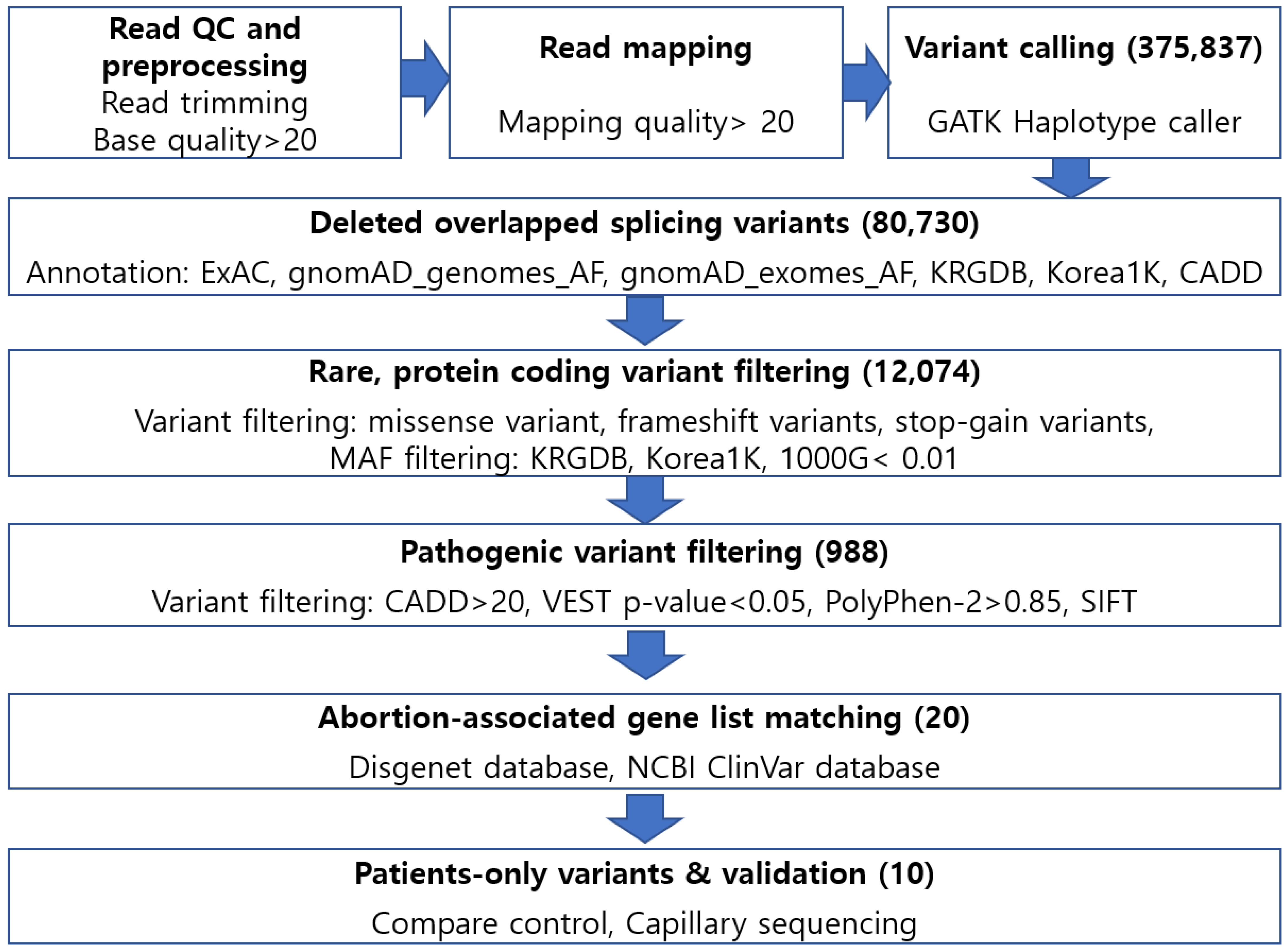
4.3. Annotation and Variant Filtering
4.4. Additional Annotation for Potential Variants
4.5. Validation
4.6. Replication of Patient and Control Samples
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Practice Committee of the American Society for Reproductive Medicine. Definitions of infertility and recurrent pregnancy loss: A committee opinion. Fertil. Steril. 2013, 99, 63. [Google Scholar] [CrossRef] [PubMed]
- El Hachem, H.; Crepaux, V.; May-Panloup, P.; Descamps, P.; Legendre, G.; Bouet, P.E. Recurrent pregnancy loss: Current perspectives. Int. J. Womens Health 2017, 9, 331–345. [Google Scholar] [CrossRef]
- Stirrat, G.M. Recurrent miscarriage I: Definition and epidemiology. Lancet 1990, 336, 673–675. [Google Scholar] [CrossRef] [PubMed]
- Stirrat, G.M. Recurrent miscarriage II: Clinical associations, causes, and management. Lancet 1990, 336, 728–733. [Google Scholar] [CrossRef]
- Lee, J.Y.; Ahn, E.H.; Kim, J.O.; Park, H.S.; Ryu, C.S.; Kim, J.H.; Kim, Y.R.; Lee, W.S.; Kim, N.K. Associations between microRNA (miR-25, miR-32, miR-125, and miR-222) polymorphisms and recurrent implantation failure in Korean women. Hum. Genom. 2019, 13, 68. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.H.; Kim, J.H.; Park, H.W.; Park, H.S.; An, H.J.; Kim, Y.R.; Ahn, E.H.; Lee, W.S.; Kim, N.K. Associations between HOTAIR polymorphisms rs4759314, rs920778, rs1899663, and rs7958904 and risk of primary ovarian insufficiency in Korean women. Maturitas 2021, 144, 74–80. [Google Scholar] [CrossRef]
- Hu, Y.; Liu, C.M.; Qi, L.; He, T.Z.; Shi-Guo, L.; Hao, C.J.; Cui, Y.; Zhang, N.; Xia, H.F.; Ma, X. Two common SNPs in pri-miR-125a alter the mature miRNA expression and associate with recurrent pregnancy loss in a Han-Chinese population. RNA Biol 2011, 8, 861–872. [Google Scholar] [CrossRef] [PubMed]
- Qiao, Y.; Wen, J.; Tang, F.; Martell, S.; Shomer, N.; Leung, P.C.K.; Stephenson, M.D.; Rajcan-Separovic, E. Whole exome sequencing in recurrent early pregnancy loss. MHR Basic Sci. Reprod. Med. 2016, 22, 364–372. [Google Scholar] [CrossRef] [PubMed]
- Quintero-Ronderos, P.; Laissue, P. Genetic Variants Contributing to Early Recurrent Pregnancy Loss Etiology Identified by Sequencing Approaches. Reprod. Sci. 2020, 27, 1541–1552. [Google Scholar] [CrossRef]
- Sawyer, S.L.; Hartley, T.; Dyment, D.A.; Beaulieu, C.L.; Schwartzentruber, J.; Smith, A.; Bedford, H.M.; Bernard, G.; Bernier, F.P.; Brais, B.; et al. Utility of whole-exome sequencing for those near the end of the diagnostic odyssey: Time to address gaps in care. Clin. Genet. 2016, 89, 275–284. [Google Scholar] [CrossRef]
- Rentzsch, P.; Schubach, M.; Shendure, J.; Kircher, M. CADD-Splice—Improving genome-wide variant effect prediction using deep learning-derived splice scores. Genome Med. 2021, 13, 31. [Google Scholar] [CrossRef] [PubMed]
- Schubach, M.; Maass, T.; Nazaretyan, L.; Röner, S.; Kircher, M. CADD v1.7: Using protein language models, regulatory CNNs and other nucleotide-level scores to improve genome-wide variant predictions. Nucleic Acids Res. 2024, 52, D1143–D1154. [Google Scholar] [CrossRef]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A method and server for predicting damaging missense mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef]
- Ng, P.C.; Henikoff, S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003, 31, 3812–3814. [Google Scholar] [CrossRef]
- Douville, C.; Masica, D.L.; Stenson, P.D.; Cooper, D.N.; Gygax, D.M.; Kim, R.; Ryan, M.; Karchin, R. Assessing the Pathogenicity of Insertion and Deletion Variants with the Variant Effect Scoring Tool (VEST-Indel). Hum. Mutat. 2016, 37, 28–35. [Google Scholar] [CrossRef]
- Carter, H.; Douville, C.; Stenson, P.D.; Cooper, D.N.; Karchin, R. Identifying Mendelian disease genes with the Variant Effect Scoring Tool. BMC Genom. 2013, 14, S3. [Google Scholar] [CrossRef] [PubMed]
- Combrinck, M.; Gilbert, J.D.; Byard, R.W. Pseudoxanthoma Elasticum and Sudden Death. J. Forensic Sci. 2011, 56, 418–422. [Google Scholar] [CrossRef] [PubMed]
- Camacho, M.; Rengel, C.; López-Herrero, E.; Carrillo, J.L.; Eslava, A.J.; Valdivielso, P. Approach to the management of pregnancy in patients with pseudoxanthoma elasticum: A review. J. Obstet. Gynaecol. 2016, 36, 1061–1066. [Google Scholar] [CrossRef]
- Bercovitch, L.; Leroux, T.; Terry, S.; Weinstock, M.A. Pregnancy and obstetrical outcomes in pseudoxanthoma elasticum. Br. J. Dermatol. 2004, 151, 1011–1018. [Google Scholar] [CrossRef]
- Koscinski, I.; Viville, S.; Porchet, N.; Bernigaud, A.; Escande, F.; Defossez, A.; Buisine, M.-P. MUC4 gene polymorphism and expression in women with implantation failure. Hum. Reprod. 2006, 21, 2238–2245. [Google Scholar] [CrossRef]
- Kim, J.-H.; Park, H.-S.; Lee, J.-Y.; Ko, E.-J.; Kim, Y.-R.; Cho, H.-Y.; Lee, W.-S.; Ahn, E.-H.; Kim, N.-K. Association Study between Mucin 4 (MUC4) Polymorphisms and Idiopathic Recurrent Pregnancy Loss in a Korean Population. Genes 2022, 13, 937. [Google Scholar] [CrossRef] [PubMed]
- Kikas, T.; Rull, K.; Beaumont, R.N.; Freathy, R.M.; Laan, M. The Effect of Genetic Variation on the Placental Transcriptome in Humans. Front. Genet. 2019, 10, 550. [Google Scholar] [CrossRef]
- Bai, Q.; Assou, S.; Haouzi, D.; Ramirez, J.-M.; Monzo, C.; Becker, F.; Gerbal-Chaloin, S.; Hamamah, S.; De Vos, J. Dissecting the First Transcriptional Divergence During Human Embryonic Development. Stem Cell Rev. Rep. 2012, 8, 150–162. [Google Scholar] [CrossRef]
- Velez Edwards, D.R.; Edwards, T.L.; Bray, M.J.; Torstenson, E.; Jones, S.; Shrubsole, M.J.; Muff, H.J.; Hartmann, K.E. Nonsteroidal Anti-inflammatory Drug Interaction with Prostacyclin Synthase Protects from Miscarriage. Sci. Rep. 2017, 7, 9874. [Google Scholar] [CrossRef]
- Foraker, A.B.; Ray, A.; Da Silva, T.C.; Bareford, L.M.; Hillgren, K.M.; Schmittgen, T.D.; Swaan, P.W. Dynamin 2 Regulates Riboflavin Endocytosis in Human Placental Trophoblasts. Mol. Pharmacol. 2007, 72, 553–562. [Google Scholar] [CrossRef]
- Carpenter, T.O. CYP24A1 loss of function: Clinical phenotype of monoallelic and biallelic mutations. J. Steroid Biochem. Mol. Biol. 2017, 173, 337–340. [Google Scholar] [CrossRef]
- Hou, H.; Zhang, J.Y.; Chen, D.; Deng, F.; Morse, A.N.; Qiu, X.; He, P.; Lash, G.E. Altered decidual and placental catabolism of vitamin D may contribute to the aetiology of spontaneous miscarriage. Placenta 2020, 92, 1–8. [Google Scholar] [CrossRef]
- Pathare, A.D.S.; Hinduja, I. Endometrial Expression of Cell Adhesion Genes in Recurrent Implantation Failure Patients in Ongoing IVF Cycle. Reprod. Sci. 2022, 29, 513–523. [Google Scholar] [CrossRef] [PubMed]
- Suhorutshenko, M.; Kukushkina, V.; Velthut-Meikas, A.; Altmäe, S.; Peters, M.; Mägi, R.; Krjutškov, K.; Koel, M.; Codoñer, F.M.; Martinez-Blanch, J.F.; et al. Endometrial receptivity revisited: Endometrial transcriptome adjusted for tissue cellular heterogeneity. Hum. Reprod. 2018, 33, 2074–2086. [Google Scholar] [CrossRef]
- Bushby, K.M.D.; Collins, J.; Hicks, D. Collagen Type VI Myopathies BT—Progress in Heritable Soft Connective Tissue Diseases; Halper, J., Ed.; Springer: Dordrecht, The Netherlands, 2014; pp. 185–199. ISBN 978-94-007-7893-1. [Google Scholar]
- Diao, H.; Aplin, J.D.; Xiao, S.; Chun, J.; Li, Z.; Chen, S.; Ye, X. Altered Spatiotemporal Expression of Collagen Types I, III, IV, and VI in Lpar3-Deficient Peri-Implantation Mouse Uterus1. Biol. Reprod. 2011, 84, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Bustamante-Marin, X.M.; Yin, W.-N.; Sears, P.R.; Werner, M.E.; Brotslaw, E.J.; Mitchell, B.J.; Jania, C.M.; Zeman, K.L.; Rogers, T.D.; Herring, L.E.; et al. Lack of GAS2L2 Causes PCD by Impairing Cilia Orientation and Mucociliary Clearance. Am. J. Hum. Genet. 2019, 104, 229–245. [Google Scholar] [CrossRef] [PubMed]
- Vanaken, G.J.; Bassinet, L.; Boon, M.; Mani, R.; Honoré, I.; Papon, J.-F.; Cuppens, H.; Jaspers, M.; Lorent, N.; Coste, A.; et al. Infertility in an adult cohort with primary ciliary dyskinesia: Phenotype–gene association. Eur. Respir. J. 2017, 50, 1700314. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Francioli, L.C.; Goodrich, J.K.; Collins, R.L.; Kanai, M.; Wang, Q.; Alföldi, J.; Watts, N.A.; Vittal, C.; Gauthier, L.D.; et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 2024, 625, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Auton, A.; Abecasis, G.R.; Altshuler, D.M.; Durbin, R.M.; Abecasis, G.R.; Bentley, D.R.; Chakravarti, A.; Clark, A.G.; Donnelly, P.; Eichler, E.E.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Jung, K.S.; Hong, K.-W.; Jo, H.Y.; Choi, J.; Ban, H.-J.; Cho, S.B.; Chung, M. KRGDB: The large-scale variant database of 1722 Koreans based on whole genome sequencing. Database 2020, 2020, baz146. [Google Scholar] [CrossRef]
- Jeon, S.; Bhak, Y.; Choi, Y.; Jeon, Y.; Kim, S.; Jang, J.; Jang, J.; Blazyte, A.; Kim, C.; Kim, Y.; et al. Korean Genome Project: 1094 Korean personal genomes with clinical information. Sci. Adv. 2023, 6, eaaz7835. [Google Scholar] [CrossRef]
- Kircher, M.; Witten, D.M.; Jain, P.; O’Roak, B.J.; Cooper, G.M.; Shendure, J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 2014, 46, 310–315. [Google Scholar] [CrossRef]
- Sim, N.-L.; Kumar, P.; Hu, J.; Henikoff, S.; Schneider, G.; Ng, P.C. SIFT web server: Predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 2012, 40, W452–W457. [Google Scholar] [CrossRef]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022, 50, W216–W221. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Gable, A.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING database in 2023: Protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 2023, 51, D638–D646. [Google Scholar] [CrossRef]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3--new capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef]
| Position | rsID | Gene | Transcript_ID | HGVS.c | HGVS.p | MAF frequency | Pathogenicity | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Replication Study | 1000 Genomes | KRGDB | KOREA1K | gnomAD Genomes_AF | VEST p-Value | VEST Missense | VEST Stop-Gain | CADD | SIFT | Polyphen2 | Detected Individual Count | ||||||
| chr2 233274367 | rs575918099 | ALPG | NM_031313.3 | c.1384G>A | p.G462S | . | 0.0002 | 0.0038 | 0.0027 | . | 0.0170 | 0.775 | . | 24.4 | 0 | 1 | 1 |
| chr2 238283646 | rs116238578 | COL6A3 | NM_057165.5 | c.2470G>A | p.V824M | 0.01327 | 0.0016 | 0.0013 | 0.0087 | 0.0003 | 0.0207 | 0.737 | . | 23.8 | 0 | 0.995 | 1 |
| chr3 195488449 | rs200737893 | MUC4 | NM_138297.5 | c.1659G>A | p.W553 * | 0.00443 | 0.0004 | 0.0013 | 0.0027 | 0.0000 | 0.0060 | . | 0.634 | 51 | N/A | N/A | 1 |
| chr10 115334164 | rs542838125 | HABP2 | NM_001177660.2 | c.145G>T | p.D49Y | 0.00443 | . | 0.0013 | 0.0005 | 0.0000 | 0.0150 | 0.799 | . | 32 | 0 | 0.987 | 1 |
| chr16 16276686 | rs758166222 | ABCC6 | NM_001171.5 | c.2045A>T | p.K682M | . | . | 0.0010 | 0.0011 | . | 0.0145 | 0.805 | . | 24.5 | 0 | 1 | 2 |
| chr16 16282707 | rs527236047 | ABCC6 | NM_001171.5 | c.1760C>G | p.S587C | 0.01327 | . | 0.0025 | 0.0022 | 0.0000 | 0.0398 | 0.613 | . | 24 | 0.03 | 0.979 | 1 |
| chr17 34074883 | rs140842796 | GAS2L2 | NM_139285.4 | c.817C>T | p.R273C | 0.00885 | 0.0018 | 0.0013 | 0.0044 | 0.0004 | 0.0075 | 0.93 | . | 28.7 | 0 | 1 | 1 |
| chr19 10897280 | rs763894364 | DNM2 | NM_001190716.2 | c.890G>A | p.R297H | . | 0.0002 | 0.0013 | 0.0005 | 0.0000 | 0.0129 | 0.827 | . | 28.6 | 0 | 0.942 | 1 |
| chr20 48130848 | rs13306027 | PTGIS | NM_000961.4 | c.940G>T | p.E314 * | 0.00443 | . | 0.0013 | 0.0011 | 0.0000 | 0.0177 | . | 0.569 | 33 | N/A | N/A | 1 |
| chr20 52789538 | rs114476330 | CYP24A1 | NM_001128915.2 | c.359G>A | p.R120H | . | 0.0002 | 0.0013 | 0.0033 | 0.0000 | 0.0159 | 0.788 | . | 32 | 0 | 1 | 1 |
| Groups | Gene (rsID) (# of Patients/# of Controls) | |||
|---|---|---|---|---|
| Group 1 | ABCC6 (rs758166222) | DNM2 (rs763894364) | ALPG (rs575918099) | CYP24A1 (rs114476330) |
| 0/0 | 0/0 | 0/0 | 0/0 | |
| Group 2 | MUC4 (rs200737893) | HABP2 (rs542838125) | GAS2L2 (rs140842796) | |
| 1/0 | 1/0 | 2/0 | ||
| Group 3 | ABCC6 (rs527236047) | PTGIS (rs13306027) | ||
| 0/3 | 0/1 | |||
| Group 4 | COL6A3 (rs116238578) | |||
| 2/1 | ||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, J.Y.; Moon, J.; Hu, H.-J.; Ryu, C.S.; Ko, E.J.; Ahn, E.H.; Kim, Y.R.; Kim, J.H.; Kim, N.K. Discovery of Pathogenic Variants Associated with Idiopathic Recurrent Pregnancy Loss Using Whole-Exome Sequencing. Int. J. Mol. Sci. 2024, 25, 5447. https://doi.org/10.3390/ijms25105447
Lee JY, Moon J, Hu H-J, Ryu CS, Ko EJ, Ahn EH, Kim YR, Kim JH, Kim NK. Discovery of Pathogenic Variants Associated with Idiopathic Recurrent Pregnancy Loss Using Whole-Exome Sequencing. International Journal of Molecular Sciences. 2024; 25(10):5447. https://doi.org/10.3390/ijms25105447
Chicago/Turabian StyleLee, Jeong Yong, JaeWoo Moon, Hae-Jin Hu, Chang Soo Ryu, Eun Ju Ko, Eun Hee Ahn, Young Ran Kim, Ji Hyang Kim, and Nam Keun Kim. 2024. "Discovery of Pathogenic Variants Associated with Idiopathic Recurrent Pregnancy Loss Using Whole-Exome Sequencing" International Journal of Molecular Sciences 25, no. 10: 5447. https://doi.org/10.3390/ijms25105447






