Computational Screening for the Dipeptidyl Peptidase-IV Inhibitory Peptides from Putative Hemp Seed Hydrolyzed Peptidome as a Potential Antidiabetic Agent
Abstract
1. Introduction
2. Results and Discussion
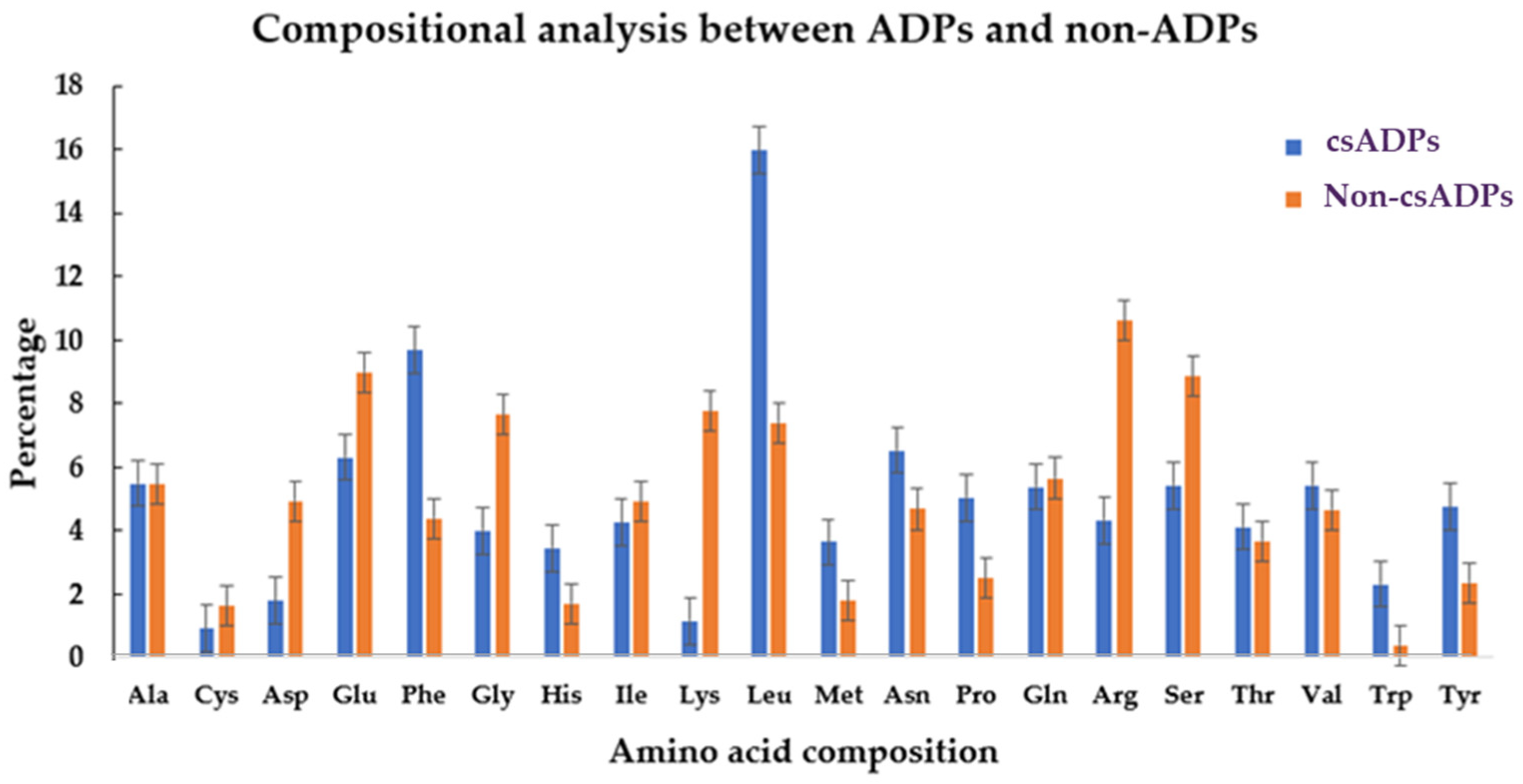
2.1. Hemp Seed Putative DPPIV Inhibitory Peptide Screening Using a Bioinformatics Approach
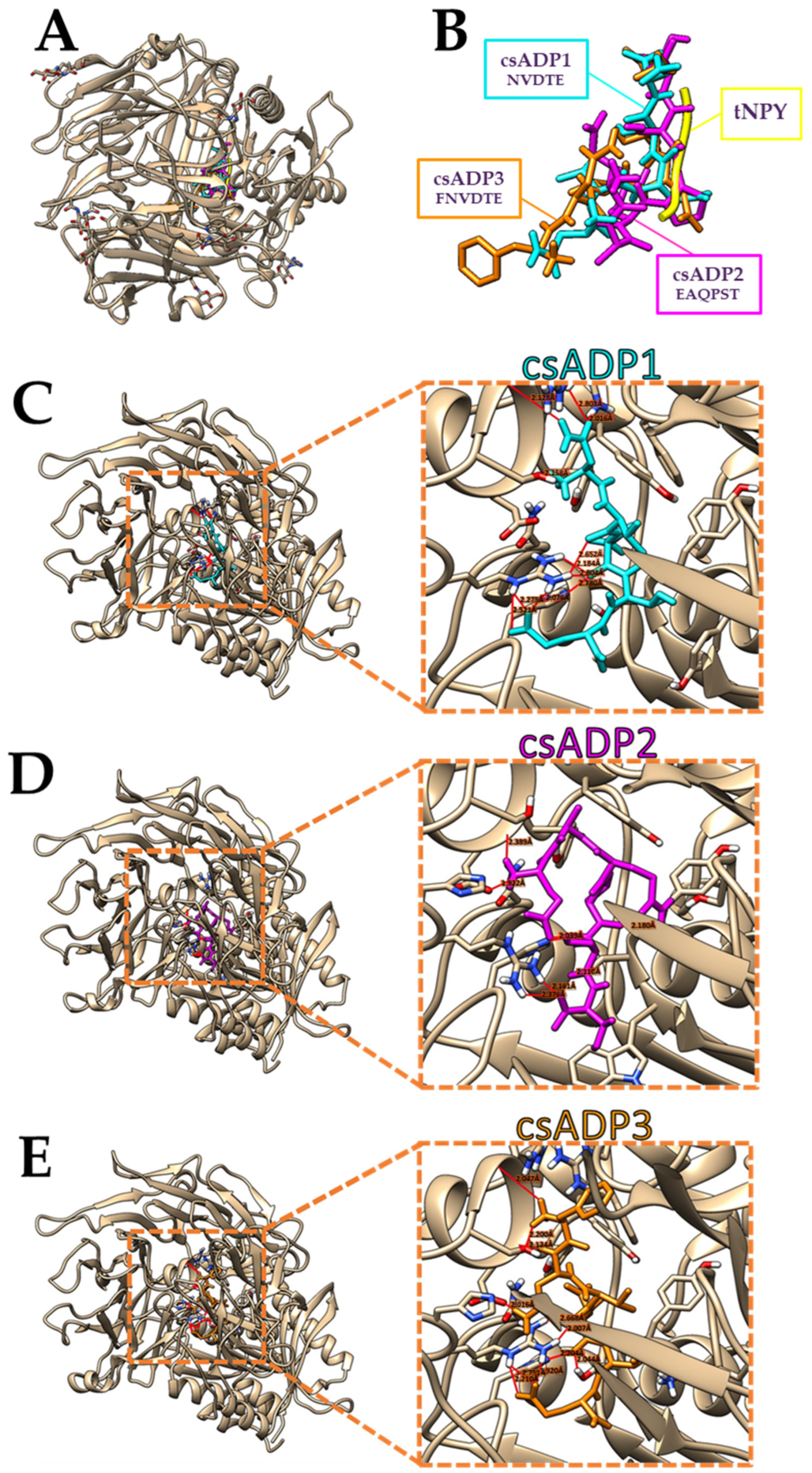
2.2. Protein–Peptide Docking Simulations
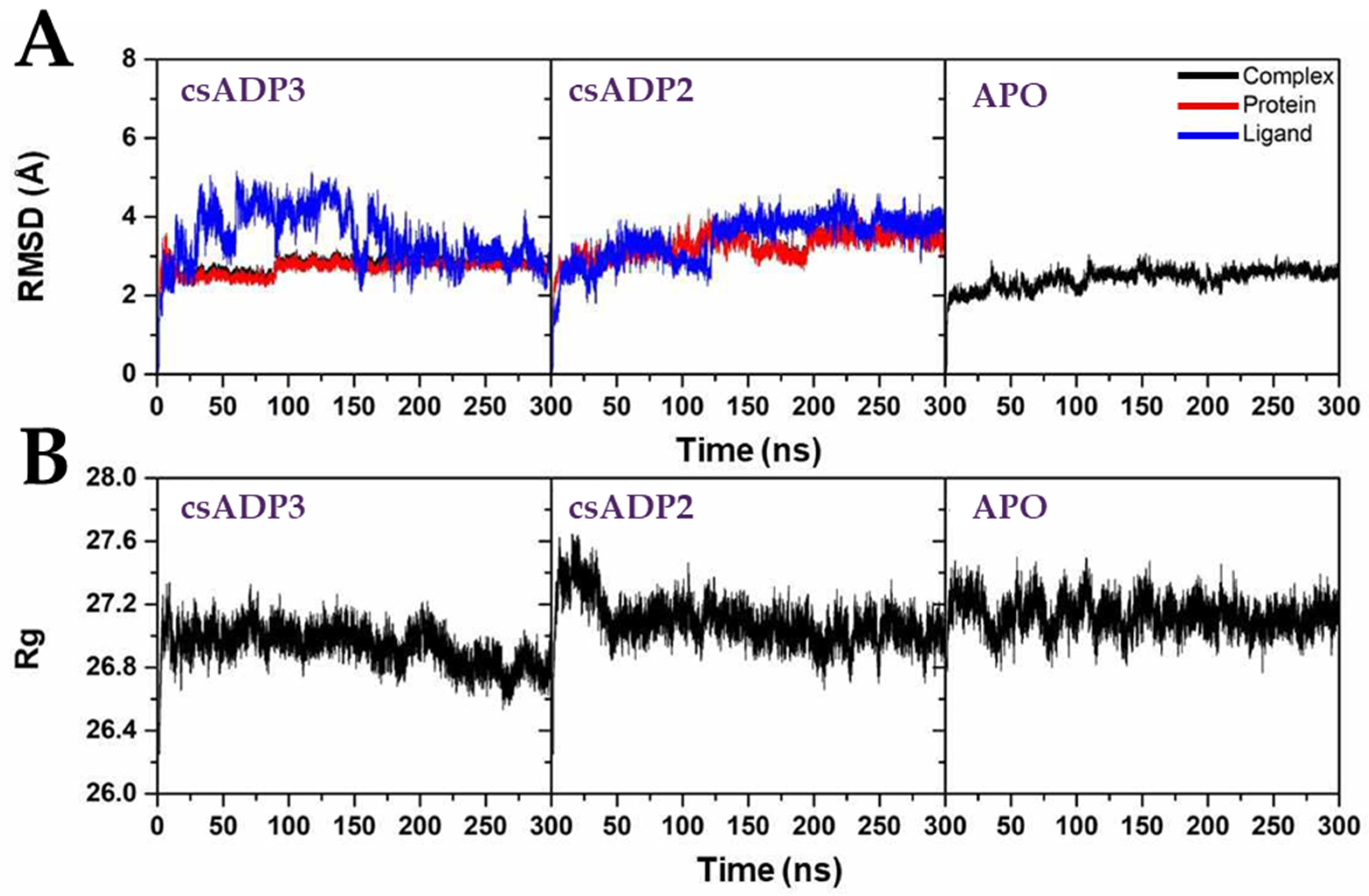
2.3. Molecular Dynamics Stimulation
3. Materials and Methods
3.1. Hemp Seed Putative Hydrolyzed Peptidome Dataset Preparation and Computer-Aided DPPIV Inhibitory Peptide Screening
3.2. Protein–Peptide Docking Simulations
3.3. Molecular Dynamics Simulations
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahmad, E.; Lim, S.; Lamptey, R.; Webb, D.R.; Davies, M.J. Type 2 diabetes. Lancet 2022, 400, 1803–1820. [Google Scholar] [CrossRef]
- World Health Organization. Global Report on Diabetes. Erişim tarihi Volume 22. Available online: http://apps.who.int/iris/bitstream/10665/204871/1/9789241565257_eng.%20pdf?%20ua=%201%20Yay%C4%B1nlanma%20tarihi%202016 (accessed on 5 August 2023).
- Deshpande, A.D.; Harris-Hayes, M.; Schootman, M. Epidemiology of diabetes and diabetes-related complications. Phys. Ther. 2008, 88, 1254–1264. [Google Scholar] [CrossRef]
- Aertgeerts, K.; Ye, S.; Tennant, M.G.; Kraus, M.L.; Rogers, J.; Sang, B.C.; Skene, R.J.; Webb, D.R.; Prasad, G.S. Crystal structure of human dipeptidyl peptidase IV in complex with a decapeptide reveals details on substrate specificity and tetrahedral intermediate formation. Protein Sci. 2004, 13, 412–421. [Google Scholar] [CrossRef] [PubMed]
- Lambeir, A.-M.; Durinx, C.; Scharpé, S.; De Meester, I. Dipeptidyl-peptidase IV from bench to bedside: An update on structural properties, functions, and clinical aspects of the enzyme DPP IV. Crit. Rev. Clin. Lab. Sci. 2003, 40, 209–294. [Google Scholar] [CrossRef] [PubMed]
- Deacon, C.F. Physiology and pharmacology of DPP-4 in glucose homeostasis and the treatment of type 2 diabetes. Front. Endocrinol. 2019, 10, 440649. [Google Scholar]
- Kirino, Y.; Sato, Y.; Kamimoto, T.; Kawazoe, K.; Minakuchi, K. Altered dipeptidyl peptidase-4 activity during the progression of hyperinsulinemic obesity and islet atrophy in spontaneously late-stage type 2 diabetic rats. Am. J. Physiol. Endocrinol. Metab. 2011, 300, E372–E379. [Google Scholar] [CrossRef] [PubMed]
- Verma, N.; Manna, S.K. Advanced glycation end products (AGE) potently induce autophagy through activation of RAF protein kinase and nuclear factor κB (NF-κB). J. Biol. Chem. 2016, 291, 1481–1491. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Zhang, L.; Joo, D.; Sun, S.-C. NF-κB signaling in inflammation. Signal Transduct. Target. Ther. 2017, 2, 17023. [Google Scholar] [CrossRef]
- Trzaskalski, N.A.; Fadzeyeva, E.; Mulvihill, E.E. Dipeptidyl peptidase-4 at the interface between inflammation and metabolism. Clin. Med. Insights Endocrinol. Diabetes 2020, 13, 1179551420912972. [Google Scholar] [CrossRef] [PubMed]
- Popko, K.; Gorska, E.; Stelmaszczyk-Emmel, A.; Plywaczewski, R.; Stoklosa, A.; Gorecka, D.; Pyrzak, B.; Demkow, U. Proinflammatory cytokines Il-6 and TNF-α and the development of inflammation in obese subjects. Eur. J. Med. Res. 2010, 15, 120. [Google Scholar] [CrossRef] [PubMed]
- Abbott, C.A.; McCaughan, G.W.; Baker, E.; Sutherland, G.R. Genomic organization, exact localization, and tissue expression of the human CD26 (dipeptidyl peptidase IV) gene. Immunogenetics 1994, 40, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Röhrborn, D.; Wronkowitz, N.; Eckel, J. DPP4 in diabetes. Front. Immunol. 2015, 6, 386. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Jia, Y.; Sun, S.; Meng, L. Adverse event profiles of dipeptidyl peptidase-4 inhibitors: Data mining of the public version of the FDA adverse event reporting system. BMC Pharmacol. Toxicol. 2020, 21, 68. [Google Scholar] [CrossRef]
- Rossino, G.; Marchese, E.; Galli, G.; Verde, F.; Finizio, M.; Serra, M.; Linciano, P.; Collina, S. Peptides as Therapeutic Agents: Challenges and Opportunities in the Green Transition Era. Molecules 2023, 28, 7165. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Kumar, S.; Kumar, S.; Kumar, D. DPP-IV Inhibitors from natural sources: An alternative approach for treatment and management of diabetes. Indian J. Nat. Prod. Resour. (IJNPR) [Former. Nat. Prod. Radiance (NPR)] 2020, 10, 227–237. [Google Scholar]
- Zhao, L.; Zhang, M.; Pan, F.; Li, J.; Dou, R.; Wang, X.; Wang, Y.; He, Y.; Wang, S.; Cai, S. In silico analysis of novel dipeptidyl peptidase-IV inhibitory peptides released from Macadamia integrifolia antimicrobial protein 2 (MiAMP2) and the possible pathways involved in diabetes protection. Curr. Res. Food Sci. 2021, 4, 603–611. [Google Scholar] [CrossRef] [PubMed]
- Agustia, F.C.; Murdiati, A.; Indrati, R. Production of Dipeptidyl Peptidase-IV Inhibitory Peptides from Germinated Jack Bean [Canavalia ensiformis (L.) DC.] Flour. Prev. Nutr. Food Sci. 2023, 28, 149. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Sun, Y.; Li, Y.; Wu, Q.; Wang, L. Identification and Characterization of the Seed Storage Proteins and Related Genes of Cannabis sativa L. Front. Nutr. 2021, 8, 678421. [Google Scholar] [CrossRef]
- House, J.D.; Neufeld, J.; Leson, G. Evaluating the quality of protein from hemp seed (Cannabis sativa L.) products through the use of the protein digestibility-corrected amino acid score method. J. Agric. Food Chem. 2010, 58, 11801–11807. [Google Scholar] [CrossRef] [PubMed]
- King, J.; Raymond, B. Hempseed (Cannabis sativa L.) Proteins: Composition, Structure, Enzymatic Modification, and Functional or Bioactive Properties. In Sustainable Protein Sources; Elsevier: Amsterdam, The Netherlands, 2024; pp. 323–338. [Google Scholar]
- Do, D.T.; Ye, A.; Singh, H.; Acevedo-Fani, A. Protein bodies from hemp seeds: Isolation, microstructure and physicochemical characterisation. Food Hydrocoll. 2024, 149, 109597. [Google Scholar] [CrossRef]
- Santos-Sánchez, G.; Álvarez-López, A.I.; Ponce-Espana, E.; Carrillo-Vico, A.; Bollati, C.; Bartolomei, M.; Lammi, C.; Cruz-Chamorro, I. Hempseed (Cannabis sativa) protein hydrolysates: A valuable source of bioactive peptides with pleiotropic health-promoting effects. Trends Food Sci. Technol. 2022, 127, 303–318. [Google Scholar] [CrossRef]
- Capcanari, T.; Boaghe, E.; Negoiţa, C. Hemp (Cannabis sativa L.) seeds nutritional aspects and food production perspectives: A review. Food Syst. 2024, 7, 52–58. [Google Scholar] [CrossRef]
- Prasertsuk, K.; Prongfa, K.; Suttiwanich, P.; Harnkit, N.; Sangkhawasi, M.; Promta, P.; Chumnanpuen, P. Computer-aided screening for potential coronavirus 3-chymotrypsin-like protease (3CLpro) inhibitory peptides from putative hemp seed trypsinized peptidome. Molecules 2022, 28, 50. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Pangloli, P.; Dia, V.P. Comparative Physicochemical and Functional Analyses of Protein Ingredients and Their Enzymatic Hydrolysates from Industrial Hempseed (Cannabis sativa L.) Hearts. ACS Food Sci. Technol. 2024; Online ahead of print. [Google Scholar] [CrossRef]
- Li, J.; Bollati, C.; Bartolomei, M.; Mazzolari, A.; Arnoldi, A.; Vistoli, G.; Lammi, C. Hempseed (Cannabis sativa) peptide H3 (IGFLIIWV) exerts cholesterol-lowering effects in human hepatic cell line. Nutrients 2022, 14, 1804. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.-R.; Qian, P.; Sun, Z.; Zhou, X.-H.; Chen, T.-P.; He, J.-F.; Zhang, H.; Wu, J. Hempseed protein derived antioxidative peptides: Purification, identification and protection from hydrogen peroxide-induced apoptosis in PC12 cells. Food Chem. 2010, 123, 1210–1218. [Google Scholar] [CrossRef]
- Agustia, F.C.; Supriyadi; Murdiati, A.; Indrati, R. Formation of dipeptidyl peptidase-IV (DPP-IV) inhibitory peptides from Jack Bean (Canavalia ensiformis (L.) DC.) sprout in simulated digestion. Food Sci. Biotechnol. 2024, 33, 645–655. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.; Jang, J.; Seo, H.H.; Lee, J.H.; Moh, S.H. Cannabis sativa (Hemp) seed-derived peptides WVYY and PSLPA modulate the Nrf2 signaling pathway in human keratinocytes. bioRxiv 2024. [Google Scholar] [CrossRef]
- Wenyu, M.; Yunfei, L.; Lulu, S.; Jingjing, W.; Jianjun, C.; Nan, Q. Anti-fatigue effect of hemp seed oligopeptide in mice and its sex difference. Food Mach. 2024, 39, 139–144. [Google Scholar]
- Hwangbo, Y.; Pan, J.H.; Lee, J.J.; Kim, T.; Kim, J.H. Production of protein hydrolysates from hemp (Cannabis sativa L.) seed and its protective effects against dexamethasone-induced muscle atrophy. Food Biosci. 2024, 59, 104046. [Google Scholar] [CrossRef]
- Girgih, A.T.; He, R.; Malomo, S.; Offengenden, M.; Wu, J.; Aluko, R.E. Structural and functional characterization of hemp seed (Cannabis sativa L.) protein-derived antioxidant and antihypertensive peptides. J. Funct. Foods 2014, 6, 384–394. [Google Scholar] [CrossRef]
- Samsamikor, M.; Mackay, D.S.; Mollard, R.C.; Alashi, A.M.; Aluko, R.E. Hemp seed protein and its hydrolysate vs. casein protein consumption in adults with hypertension: A double-blind crossover study. Am. J. Clin. Nutr. 2024; in press. [Google Scholar]
- Cruz-Chamorro, I.; Santos-Sánchez, G.; Bollati, C.; Bartolomei, M.; Li, J.; Arnoldi, A.; Lammi, C. Hempseed (Cannabis sativa) peptides WVSPLAGRT and IGFLIIWV exert anti-inflammatory activity in the LPS-stimulated human hepatic cell line. J. Agric. Food Chem. 2022, 70, 577–583. [Google Scholar] [CrossRef] [PubMed]
- Millan-Linares, M.C.; Rivero-Pino, F.; Gonzalez-de la Rosa, T.; Villanueva, A.; Montserrat-de la Paz, S. Identification, characterization, and molecular docking of immunomodulatory oligopeptides from bioavailable hempseed protein hydrolysates. Food Res. Int. 2024, 176, 113712. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Liang, K.; Jin, Y.; Zhang, M.; Chen, Y.; Wu, H.; Lai, F. Identification and characterization of two novel α-glucosidase inhibitory oligopeptides from hemp (Cannabis sativa L.) seed protein. J. Funct. Foods 2016, 26, 439–450. [Google Scholar] [CrossRef]
- Nayak, A.P.; Green, B.J.; Sussman, G.; Berlin, N.; Lata, H.; Chandra, S.; ElSohly, M.A.; Hettick, J.M.; Beezhold, D.H. Characterization of Cannabis sativa allergens. Ann. Allergy Asthma Immunol. 2013, 111, 32–37.e34. [Google Scholar] [CrossRef] [PubMed]
- Charoenkwan, P.; Nantasenamat, C.; Hasan, M.M.; Moni, M.A.; Manavalan, B.; Shoombuatong, W. StackDPPIV: A novel computational approach for accurate prediction of dipeptidyl peptidase IV (DPP-IV) inhibitory peptides. Methods 2022, 204, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Das, D.; Kabir, M.E.; Sarkar, S.; Wann, S.B.; Kalita, J.; Manna, P. Antidiabetic potential of soy protein/peptide: A therapeutic insight. Int. J. Biol. Macromol. 2022, 194, 276–288. [Google Scholar] [CrossRef] [PubMed]
- Mojica, L.; De Mejía, E.G. Optimization of enzymatic production of anti-diabetic peptides from black bean (Phaseolus vulgaris L.) proteins, their characterization and biological potential. Food Funct. 2016, 7, 713–727. [Google Scholar] [CrossRef] [PubMed]
- Vainerman, E.; Golovina, T.; Rogozhin, S. Low-molecular-weight thiol in macromolecules of Canabis sativa 11S globulin–Edestin. Food/Nahrung 1986, 30, 398–400. [Google Scholar] [CrossRef]
- Malomo, S.A.; He, R.; Aluko, R.E. Structural and functional properties of hemp seed protein products. J. Food Sci. 2014, 79, C1512–C1521. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Esteban, J.I.; Pinela, J.; Ćirić, A.; Calhelha, R.C.; Soković, M.; Ferreira, I.C.; Barros, L.; Torija-Isasa, E.; de Cortes Sánchez-Mata, M. Chemical composition and biological activities of whole and dehulled hemp (Cannabis sativa L.) seeds. Food Chem. 2022, 374, 131754. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Gong, H.; Mao, X. Dipeptidyl peptidase-IV inhibitory activity and related molecular mechanism of bovine α-lactalbumin-derived peptides. Molecules 2020, 25, 3009. [Google Scholar] [CrossRef]
- Nongonierma, A.B.; FitzGerald, R.J. Features of dipeptidyl peptidase IV (DPP-IV) inhibitory peptides from dietary proteins. J. Food Biochem. 2019, 43, e12451. [Google Scholar] [CrossRef]
- Charoenkwan, P.; Kanthawong, S.; Nantasenamat, C.; Hasan, M.M.; Shoombuatong, W. iDPPIV-SCM: A sequence-based predictor for identifying and analyzing dipeptidyl peptidase IV (DPP-IV) inhibitory peptides using a scoring card method. J. Proteome Res. 2020, 19, 4125–4136. [Google Scholar] [CrossRef] [PubMed]
- Timmons, P.B.; Hewage, C.M. HAPPENN is a novel tool for hemolytic activity prediction for therapeutic peptides which employs neural networks. Sci. Rep. 2020, 10, 10869. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Cheng, J.; Wu, H. Discovery of food-derived dipeptidyl peptidase IV inhibitory peptides: A review. Int. J. Mol. Sci. 2019, 20, 463. [Google Scholar] [CrossRef]
- Ogata, S.; Misumi, Y.; Tsuji, E.; Takami, N.; Oda, K.; Ikehara, Y. Identification of the active site residues in dipeptidyl peptidase IV by affinity labeling and site-directed mutagenesis. Biochemistry 1992, 31, 2582–2587. [Google Scholar] [CrossRef] [PubMed]
- Mathur, V.; Alam, O.; Siddiqui, N.; Jha, M.; Manaithiya, A.; Bawa, S.; Sharma, N.; Alshehri, S.; Alam, P.; Shakeel, F. Insight into structure activity relationship of DPP-4 inhibitors for development of antidiabetic agents. Molecules 2023, 28, 5860. [Google Scholar] [CrossRef] [PubMed]
- Jeffrey, G.A.; Jeffrey, G.A. An Introduction to Hydrogen Bonding; Oxford University Press: New York, NY, USA, 1997; Volume 12. [Google Scholar]
- Schön, A.; Madani, N.; Smith, A.B.; Lalonde, J.M.; Freire, E. Some Binding-Related Drug Properties are Dependent on Thermodynamic Signature. Chem. Biol. Drug Des. 2011, 77, 161–165. [Google Scholar] [CrossRef] [PubMed]
- Kastritis, P.L.; Bonvin, A.M. On the binding affinity of macromolecular interactions: Daring to ask why proteins interact. J. R. Soc. Interface 2013, 10, 20120835. [Google Scholar] [CrossRef] [PubMed]
- Abbad, A.; Anga, L.; Faouzi, A.; Iounes, N.; Nourlil, J. Effect of identified non-synonymous mutations in DPP4 receptor binding residues among highly exposed human population in Morocco to MERS-CoV through computational approach. PLoS ONE 2021, 16, e0258750. [Google Scholar] [CrossRef]
- Naik, S.; Deora, N.; Pal, S.K.; Ahmed, M.Z.; Alqahtani, A.S.; Shukla, P.K.; Venkatraman, K.; Kumar, S. Purification, biochemical characterization, and DPP-IV and α-amylase inhibitory activity of Berberine from Cardiospermum halicacabum. J. Mol. Recognit. 2022, 35, e2983. [Google Scholar] [CrossRef] [PubMed]
- Ryskjær, J.; Deacon, C.F.; Carr, R.D.; Krarup, T.; Madsbad, S.; Holst, J.; Vilsbøll, T. Plasma dipeptidyl peptidase-IV activity in patients with type-2 diabetes mellitus correlates positively with HbAlc levels, but is not acutely affected by food intake. Eur. J. Endocrinol. 2006, 155, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Nauck, M.A.; Meier, J.J. Incretin hormones: Their role in health and disease. Diabetes Obes. Metab. 2018, 20, 5–21. [Google Scholar] [CrossRef] [PubMed]
- Han, S.J.; Kim, H.J.; Choi, S.-E.; Kang, Y.; Lee, K.W.; Kim, D.J. Incretin secretion and serum DPP-IV activity in Korean patients with type 2 diabetes. Diabetes Res. Clin. Pract. 2010, 89, e49–e52. [Google Scholar] [CrossRef] [PubMed]
- Mune, M.A.M.; Minka, S.R.; Henle, T. Investigation on antioxidant, angiotensin converting enzyme and dipeptidyl peptidase IV inhibitory activity of Bambara bean protein hydrolysates. Food Chem. 2018, 250, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Rivero-Pino, F.; Espejo-Carpio, F.J.; Guadix, E.M. Identification of dipeptidyl peptidase IV (DPP-IV) inhibitory peptides from vegetable protein sources. Food Chem. 2021, 354, 129473. [Google Scholar] [CrossRef]
- Luo, Z.; Su, K.; Zhang, X. Potential of plant proteins digested in silico by gastrointestinal enzymes as nutritional supplement for COVID-19 patients. Plant Foods Hum. Nutr. 2020, 75, 583–591. [Google Scholar] [CrossRef] [PubMed]
- Vermeirssen, V.; van der Bent, A.; Van Camp, J.; van Amerongen, A.; Verstraete, W. A quantitative in silico analysis calculates the angiotensin I converting enzyme (ACE) inhibitory activity in pea and whey protein digests. Biochimie 2004, 86, 231–239. [Google Scholar] [CrossRef] [PubMed]
- Villanueva, A.; Rivero-Pino, F.; Martin, M.E.; Gonzalez-de la Rosa, T.; Montserrat-de la Paz, S.; Millan-Linares, M.C. Identification of the Bioavailable Peptidome of Chia Protein Hydrolysate and the In Silico Evaluation of Its Antioxidant and ACE Inhibitory Potential. J. Agric. Food Chem. 2024, 72, 3189–3199. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Heo, L.; Lee, M.S.; Seok, C. GalaxyPepDock: A protein–peptide docking tool based on interaction similarity and energy optimization. Nucleic Acids Res. 2015, 43, W431–W435. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Jin, B.; Li, H.; Huang, S.-Y. HPEPDOCK: A web server for blind peptide–protein docking based on a hierarchical algorithm. Nucleic Acids Res. 2018, 46, W443–W450. [Google Scholar] [CrossRef] [PubMed]
- Meng, E.C.; Pettersen, E.F.; Couch, G.S.; Huang, C.C.; Ferrin, T.E. Tools for integrated sequence-structure analysis with UCSF Chimera. BMC Bioinform. 2006, 7, 339. [Google Scholar] [CrossRef] [PubMed]
- Xue, L.C.; Rodrigues, J.P.; Kastritis, P.L.; Bonvin, A.M.; Vangone, A. PRODIGY: A web server for predicting the binding affinity of protein–protein complexes. Bioinformatics 2016, 32, 3676–3678. [Google Scholar] [CrossRef] [PubMed]
- Clementel, D.; Del Conte, A.; Monzon, A.M.; Camagni, G.F.; Minervini, G.; Piovesan, D.; Tosatto, S.C. RING 3.0: Fast generation of probabilistic residue interaction networks from structural ensembles. Nucleic Acids Res. 2022, 50, W651–W656. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Huang, J.; He, B. AntiDMPpred: A web service for identifying anti-diabetic peptides. PeerJ 2022, 10, e13581. [Google Scholar] [CrossRef]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [PubMed]
- Ryckaert, J.-P.; Ciccotti, G.; Berendsen, H.J.C. Numerical integration of the cartesian equations of motion of a system with constraints: Molecular dynamics of n-alkanes. J. Comput. Phys. 1977, 23, 327–341. [Google Scholar] [CrossRef]
- Uberuaga, B.P.; Anghel, M.; Voter, A.F. Synchronization of trajectories in canonical molecular-dynamics simulations: Observation, explanation, and exploitation. J. Chem. Phys. 2004, 120, 6363–6374. [Google Scholar] [CrossRef] [PubMed]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]



| Peptide ID | Sequence (Protein/Protease) | Length | iDPPIV-SCM Prediction Scores | AllerCatPro | HemoPred | ToxinPred |
|---|---|---|---|---|---|---|
| csADP1 | NVDTE (Edestin2/Pepsin) | 5 | 335.75 | No evidence | Non-hemolytic | 0.59 |
| csADP2 | EAQPST (Vicilin/Pepsin) | 6 | 325.6 | No evidence | Non-hemolytic | 0.49 |
| csADP3 | FNVDTE (Edestin2/Pepsin) | 6 | 322.6 | No evidence | Non-hemolytic | 0.57 |
| Peptide ID | Secondary Structures | csADP Residues | DPPIV Residues | Distance (Å) |
|---|---|---|---|---|
| csADP1 |  | Glu5 | Arg125 | 2.273 |
| Glu5 | Arg125 | 2.521 | ||
| Asp3 | Arg125 | 2.184 | ||
| Val2 | Arg125 | 2.652 | ||
| Asp3 | Arg125 | 2.604 | ||
| Glu5 | Arg125 | 1.976 | ||
| Asn1 | Arg669 | 2.803 | ||
| Asn1 | Arg669 | 2.016 | ||
| Asn1 | Ser209 | 2.158 | ||
| Asn1 | Val207 | 2.128 | ||
| csADP2 |  | Thr6 | Arg125 | 2.376 |
| Thr6 | Arg125 | 2.161 | ||
| Ser5 | Ser630 | 2.001 | ||
| Pro4 | Tyr631 | 2.18 | ||
| Glu1 | Glu205 | 2.389 | ||
| Glu1 | Glu205 | 1.922 | ||
| Ser5 | His740 | 2.152 | ||
| csADP3 |  | Glu6 | Arg125 | 2.757 |
| Glu6 | Arg125 | 2.24 | ||
| Val3 | Arg125 | 2.668 | ||
| Val3 | Arg125 | 2.007 | ||
| Glu6 | Arg125 | 1.92 | ||
| Asp4 | Ser630 | 2.044 | ||
| Phe1 | Val207 | 2.047 | ||
| Phe1 | Ser630 | 2.2 | ||
| Asn2 | Glu205 | 2.124 | ||
| Asn2 | Glu205 | 2.016 |
| Peptide ID | PRODIGY | Molecular Docking Scores | AntiDMPred | ||
|---|---|---|---|---|---|
| ΔG (kcal mol−1) | Kd (M) at 25 °C | GalaxyPepDock | HPEPDOCK | ||
| csADP1 | −10.2 | 3.50 × 10−7 | 0.872 | −114.142 | 0.51 |
| csADP2 | −11.7 | 2.50 × 10−9 | 0.954 | −143.62 | 0.50 |
| csADP3 | −10.6 | 1.80 × 10−8 | 0.872 | −169.78 | 0.47 |
| Bond | csADP1 | csADP2 | csADP3 | |||
|---|---|---|---|---|---|---|
| Intra | Inter | Intra | Inter | Intra | Inter | |
| H-Bond | 573 | 7 | 567 | 7 | 561 | 9 |
| π-π Stacking | 83 | 0 | 80 | 0 | 82 | 1 |
| π-Cation | 4 | 0 | 3 | 0 | 5 | 0 |
| Ionic | 15 | 1 | 15 | 0 | 11 | 1 |
| Disulfide | 0 | 0 | 0 | 0 | 0 | 0 |
| π-H-Bond | 2 | 0 | 4 | 1 | 4 | 1 |
| Van der Waals | 623 | 7 | 606 | 8 | 622 | 14 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thongtak, A.; Yutisayanuwat, K.; Harnkit, N.; Noikaew, T.; Chumnanpuen, P. Computational Screening for the Dipeptidyl Peptidase-IV Inhibitory Peptides from Putative Hemp Seed Hydrolyzed Peptidome as a Potential Antidiabetic Agent. Int. J. Mol. Sci. 2024, 25, 5730. https://doi.org/10.3390/ijms25115730
Thongtak A, Yutisayanuwat K, Harnkit N, Noikaew T, Chumnanpuen P. Computational Screening for the Dipeptidyl Peptidase-IV Inhibitory Peptides from Putative Hemp Seed Hydrolyzed Peptidome as a Potential Antidiabetic Agent. International Journal of Molecular Sciences. 2024; 25(11):5730. https://doi.org/10.3390/ijms25115730
Chicago/Turabian StyleThongtak, Arisa, Kulpariya Yutisayanuwat, Nathaphat Harnkit, Tipanart Noikaew, and Pramote Chumnanpuen. 2024. "Computational Screening for the Dipeptidyl Peptidase-IV Inhibitory Peptides from Putative Hemp Seed Hydrolyzed Peptidome as a Potential Antidiabetic Agent" International Journal of Molecular Sciences 25, no. 11: 5730. https://doi.org/10.3390/ijms25115730
APA StyleThongtak, A., Yutisayanuwat, K., Harnkit, N., Noikaew, T., & Chumnanpuen, P. (2024). Computational Screening for the Dipeptidyl Peptidase-IV Inhibitory Peptides from Putative Hemp Seed Hydrolyzed Peptidome as a Potential Antidiabetic Agent. International Journal of Molecular Sciences, 25(11), 5730. https://doi.org/10.3390/ijms25115730







