Comparative Bioinformatic Analysis Reveals Conserved Regions in SARS-CoV-2 Genome for RAPID Pandemic Response
Abstract
:1. Introduction
2. Results
2.1. Comparative Analysis of Whole-Genome Data Using Bioinformatic Tools Revealed Conserved Regions of SARS-CoV-2
2.2. Data Availability
2.3. Code Availability
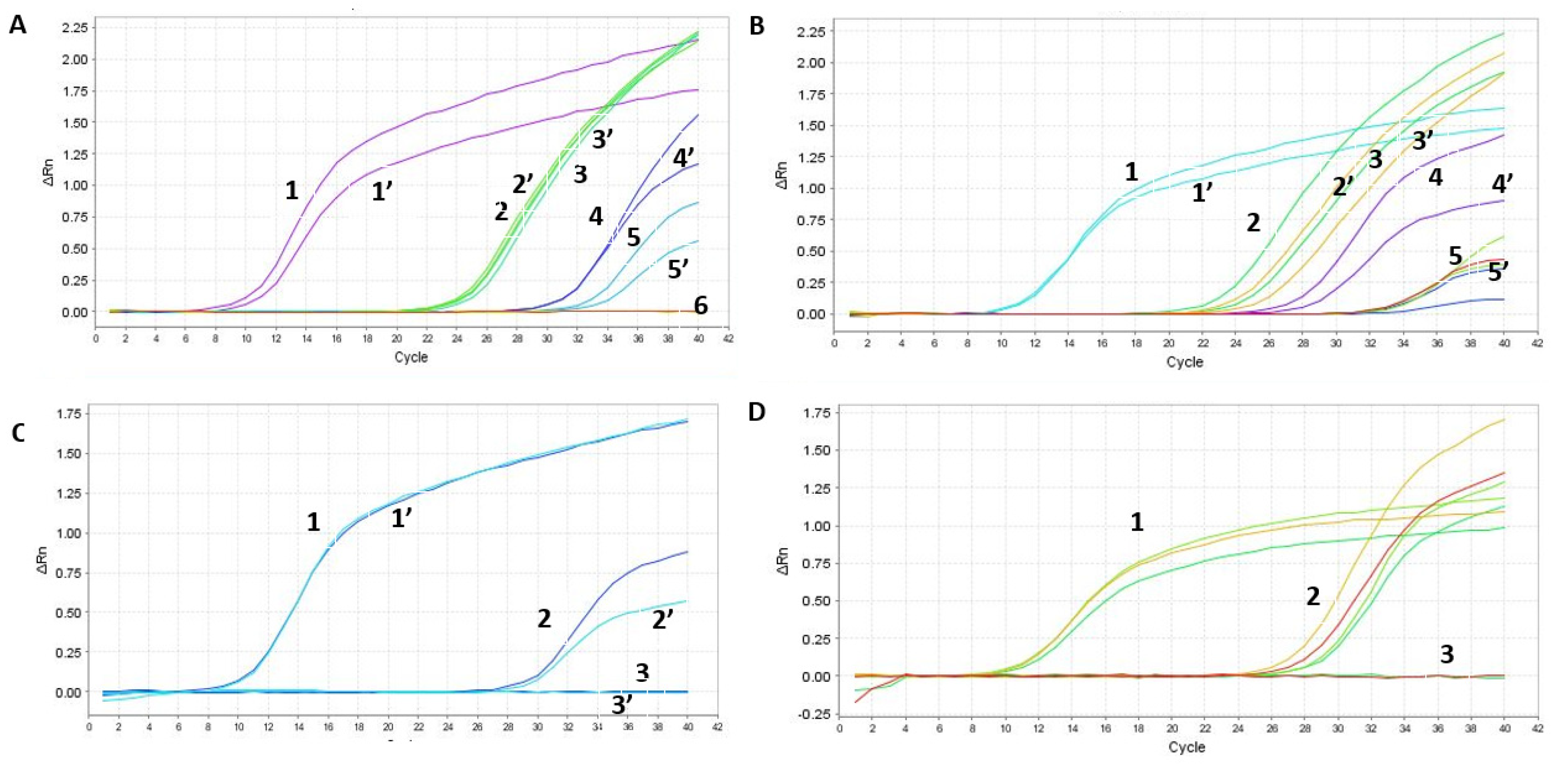
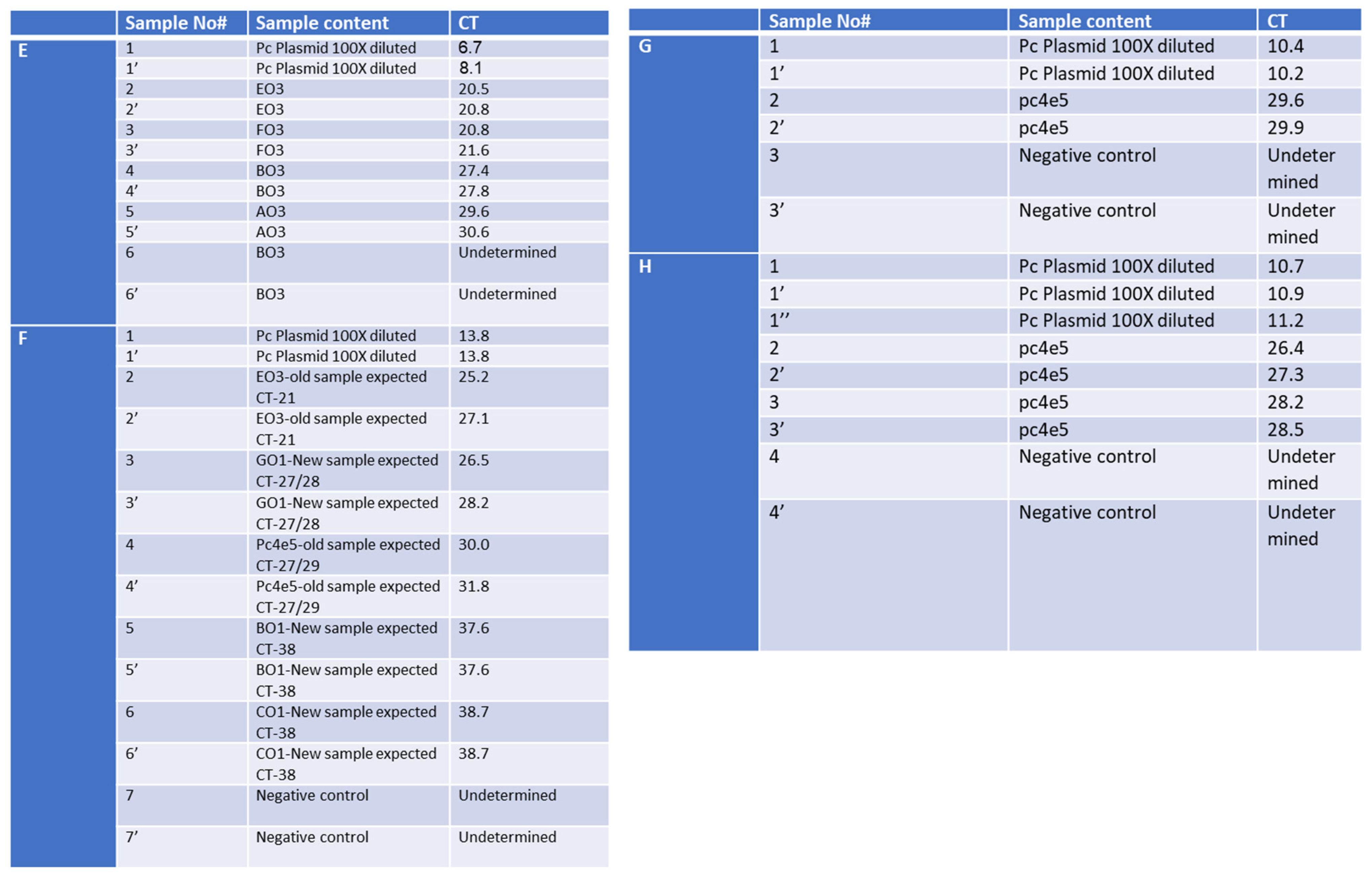
2.4. Evaluation of SET1 Primer Selectivity and Optimization of One-Step RT-qPCR Assay Using Synthetic DNA and RNA
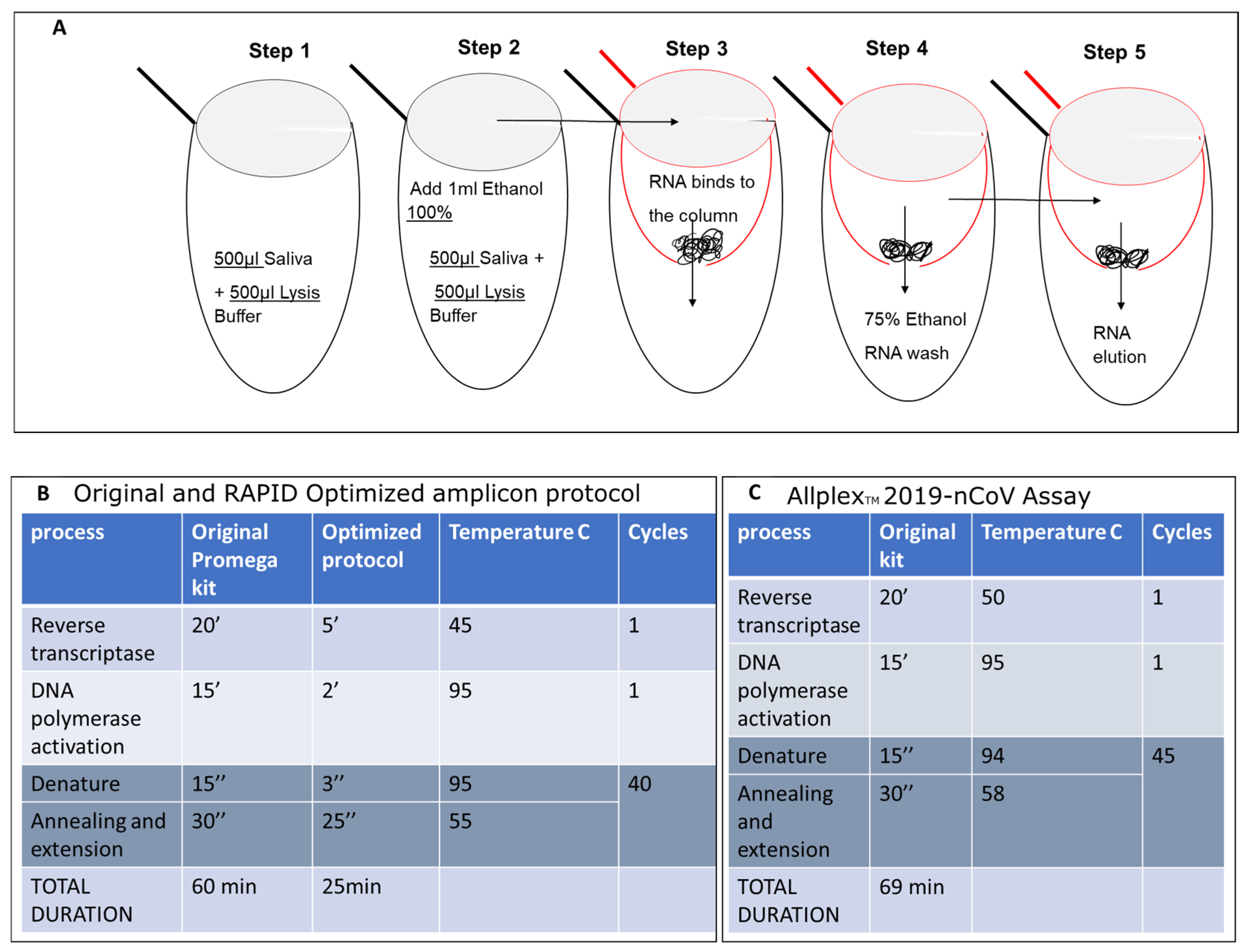
2.5. Evaluation of SET1 Primers Using the Shorter RAPID One-Step RT-qPCR Protocol on Nucleic Acid Isolated from Saliva
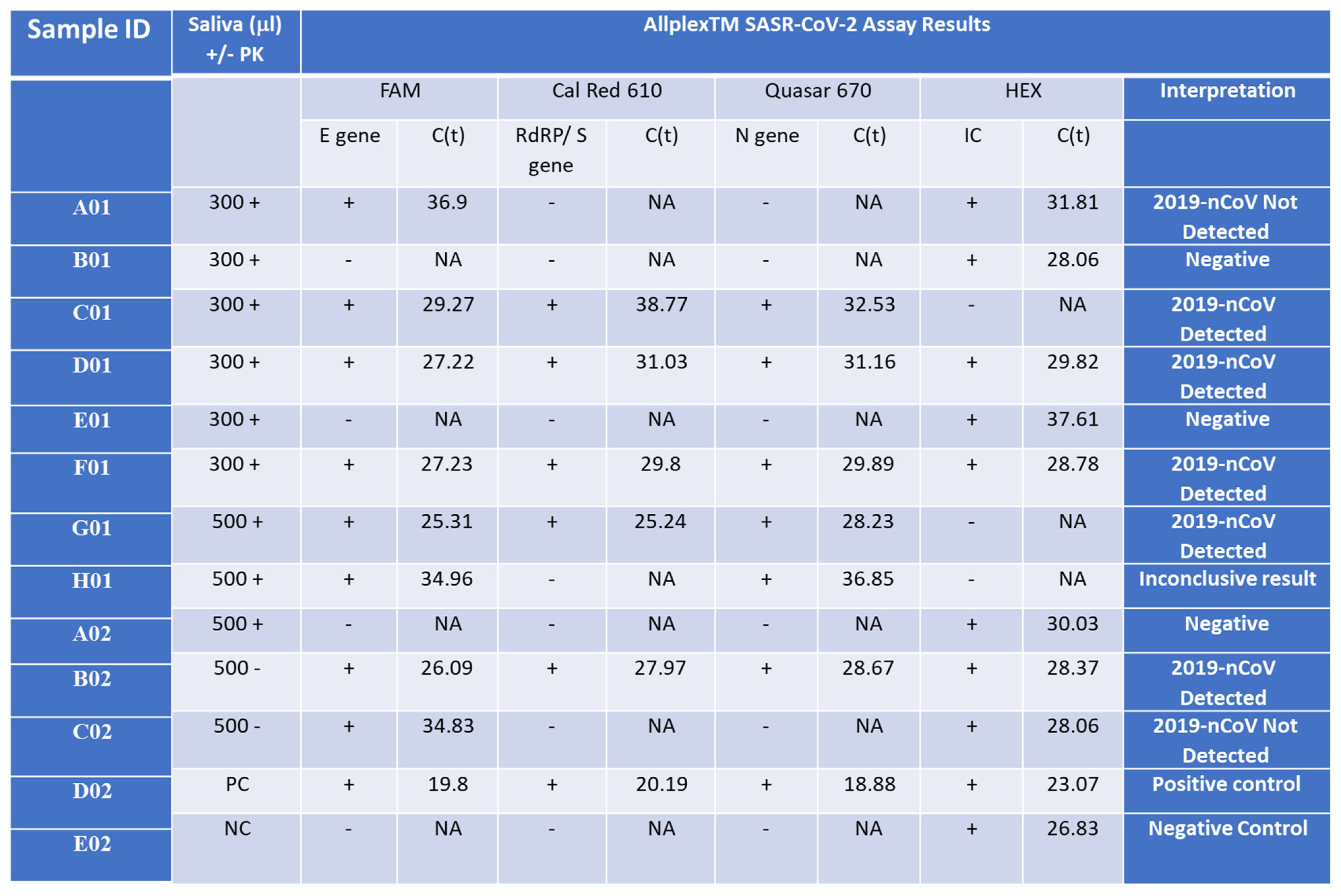
2.6. Validation of the Shortened RAPID One-Step qPCR Assay Using Various Positive and Negative Samples
2.7. Optimization and Significant Reduction in Time of the One-Step qPCR Method, Enhancing Sensitivity for Future SARS-CoV-2 Variants
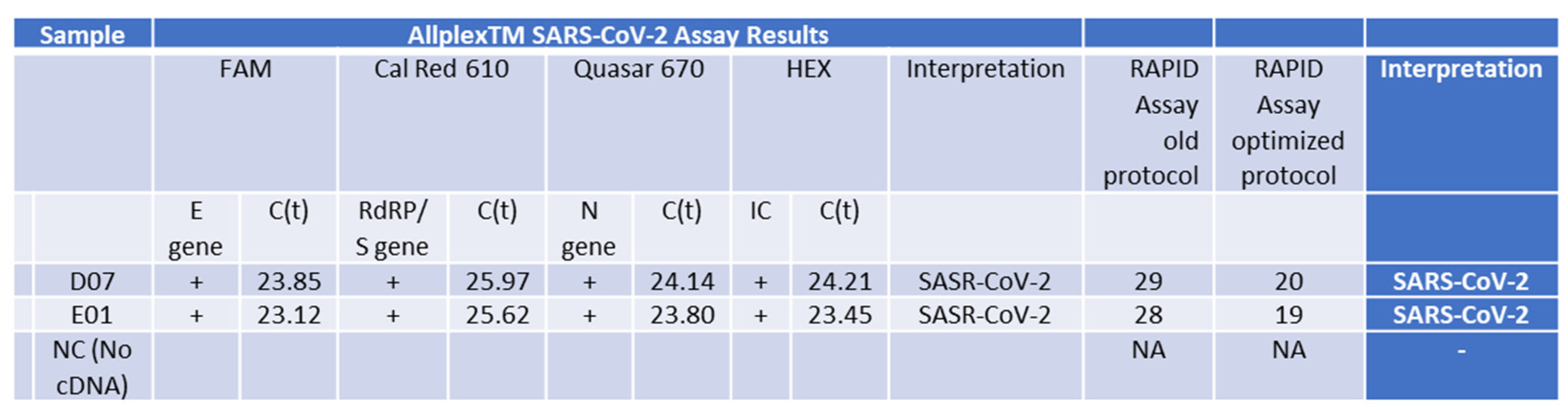
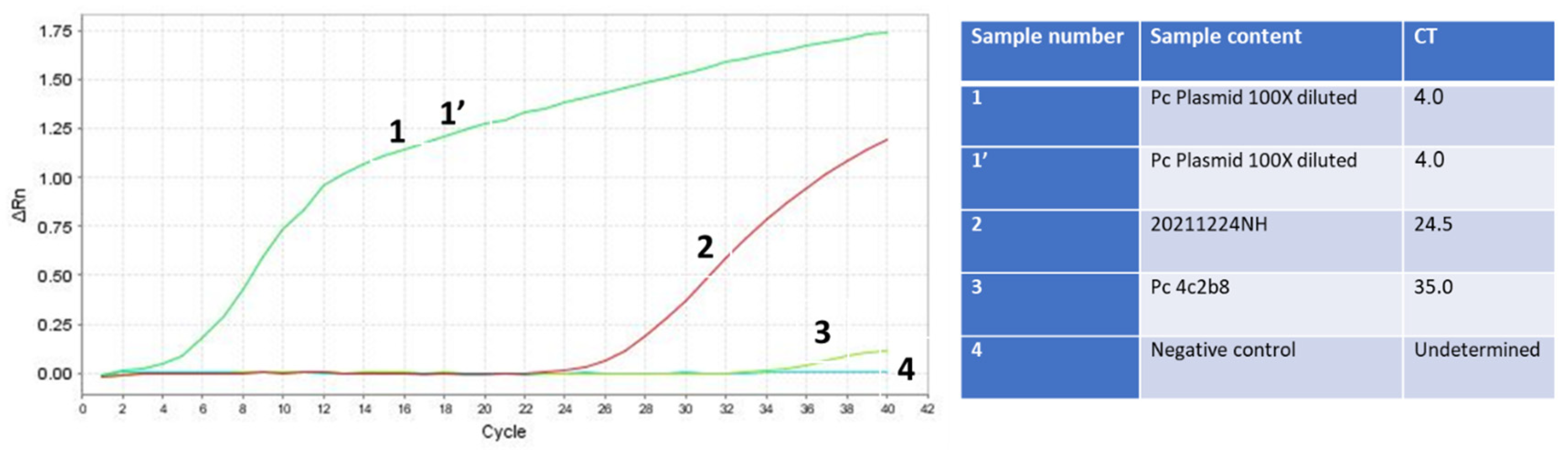
2.8. Validation of the Optimized Assay Using Positive Samples, including a Critical Case Study
3. Discussion
4. Materials and Methods
- -
- Nucleic acid was extracted using the Seegene STARlet equipment following the manufacturer’s protocol.
- -
- Allplex™ 2019-nCoV Assay was ordered from Seegene (Cat. No. RP10243X).
- Ordered reagents:
- -
- GoTaq® 1-Step RT-qPCR System (Promega, Cat. No. A6020) was used for the one-step qPCR.
- -
- Primer sets (Prime time® Std qPCR Assay (500rXn), as well as synthetic RNA and DNA were custom ordered from Integrated DNA Technologies (IDT).
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mercer, T.; Almond, N.; Crone, M.A.; Chain, P.S.G.; Deshpande, A.; Eveleigh, D.; Freemont, P.; Fuchs, S.; Garlick, R.; Huggett, J.; et al. The Coronavirus Standards Working Group’s roadmap for improved population testing. Nat. Biotechnol. 2022, 40, 1563–1568. [Google Scholar] [CrossRef]
- Torge, D.; Bernardi, S.; Arcangeli, M.; Bianchi, S. Histopathological Features of SARS-CoV-2 in Extrapulmonary Organ Infection: A Systematic Review of Literature. Pathogens 2022, 11, 867. [Google Scholar] [CrossRef] [PubMed]
- Chu, V.T.; Schwartz, N.G.; Donnelly, M.A.; Chuey, M.R.; Soto, R.; Yousaf, A.R.; Schmitt-Matzen, E.N.; Sleweon, S.; Ruffin, J.; Thornburg, N.; et al. Comparison of Home Antigen Testing With RT-PCR and Viral Culture During the Course of SARS-CoV-2 Infection. JAMA Intern. Med. 2022, 182, 701–709. [Google Scholar] [CrossRef] [PubMed]
- SelectScience. Seegene Introduces New SARS-CoV-2 Variants Detection Test. Available online: http://www.selectscience.net/product-news/seegene-introduces-new-sars-cov-2-variants-detection-test/?artID=55308 (accessed on 12 March 2024).
- Beniac, D.R.; Andonov, A.; Grudeski, E.; Booth, T.F. Architecture of the SARS coronavirus prefusion spike. Nat. Struct. Mol. Biol. 2006, 13, 751–752. [Google Scholar] [CrossRef] [PubMed]
- Delmas, B.; Laude, H. Assembly of coronavirus spike protein into trimers and its role in epitope expression. J. Virol. 1990, 64, 5367–5375. [Google Scholar] [CrossRef] [PubMed]
- Masters, P.S. The molecular biology of coronaviruses. Adv. Virus Res. 2006, 66, 193–292. [Google Scholar] [CrossRef] [PubMed]
- Mortola, E.; Roy, P. Efficient assembly and release of SARS coronavirus-like particles by a heterologous expression system. FEBS Lett. 2004, 576, 174–178. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Zheng, X.; Gai, W.; Zhao, Y.; Wang, H.; Wang, H.; Feng, N.; Chi, H.; Qiu, B.; Li, N.; et al. MERS-CoV virus-like particles produced in insect cells induce specific humoural and cellular immunity in rhesus macaques. Oncotarget 2017, 8, 12686–12694. [Google Scholar] [CrossRef] [PubMed]
- Tracking SARS-CoV-2 Variants. Available online: https://www.who.int/activities/tracking-SARS-CoV-2-variants (accessed on 12 March 2024).
- Schmidt, C. New Coronavirus Variants Are Urgently Being Tracked around the World. Scientific American. Available online: https://www.scientificamerican.com/article/new-coronavirus-variants-are-urgently-being-tracked-around-the-world/ (accessed on 12 March 2024).
- Amicone, M.; Borges, V.; Alves, M.J.; Isidro, J.; Zé-Zé, L.; Duarte, S.; Vieira, L.; Guiomar, R.; Gomes, J.P.; Gordo, I. Mutation rate of SARS-CoV-2 and emergence of mutators during experimental evolution. Evol. Med. Public Health 2022, 10, 142–155. [Google Scholar] [CrossRef]
- Cosar, B.; Karagulleoglu, Z.Y.; Unal, S.; Ince, A.T.; Uncuoglu, D.B.; Tuncer, G.; Kilinc, B.R.; Ozkan, Y.E.; Ozkoc, H.C.; Demir, I.N.; et al. SARS-CoV-2 Mutations and their Viral Variants. Cytokine Growth Factor. Rev. 2022, 63, 10–22. [Google Scholar] [CrossRef]
- Domnich, A.; De Pace, V.; Pennati, B.M.; Caligiuri, P.; Varesano, S.; Bruzzone, B.; Orsi, A. Evaluation of extraction-free RT-qPCR methods for SARS-CoV-2 diagnostics. Arch. Virol. 2021, 166, 2825–2828. [Google Scholar] [CrossRef] [PubMed]
- Alshahrani, M.M. Inhibition of SARS–CoV–2 NSP–15 by Uridine–5′–Monophosphate Analogues Using QSAR Modelling, Molecular Dynamics Simulations, and Free Energy Landscape. Saudi Pharm. J. 2024, 32, 101914. [Google Scholar] [CrossRef] [PubMed]
- Deshmukh, V.; Motwani, R.; Kumar, A.; Kumari, C.; Raza, K. Histopathological observations in COVID-19: A systematic review. J. Clin. Pathol. 2020, 74, 76–83. [Google Scholar] [CrossRef] [PubMed]
- Belizário, J.E. Immunity, virus evolution, and effectiveness of SARS-CoV-2 vaccines. Braz. J. Med. Biol. Res. 2021, 54, e10725. [Google Scholar] [CrossRef]
- Chedid, M.; Waked, R.; Haddad, E.; Chetata, N.; Saliba, G.; Choucair, J. Antibiotics in treatment of COVID-19 complications: A review of frequency, indications, and efficacy. J. Infect. Public. Health 2021, 14, 570–576. [Google Scholar] [CrossRef] [PubMed]
- Gagliotti, C.; Buttazzi, R.; Ricchizzi, E.; Di Mario, S.; Tedeschi, S.; Moro, M.L. Community use of antibiotics during the COVID-19 lockdown. Infect. Dis. 2021, 53, 142–144. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.; Liu, Y.; Wang, J.; Fallowfield, J.A.; Wang, J.; Li, X.; Shi, J.; Pan, H.; Zou, S.; Zhang, H.; et al. Clinical course and risk factors for mortality of COVID-19 patients with pre-existing cirrhosis: A multicentre cohort study. Gut 2021, 70, 433–436. [Google Scholar] [CrossRef] [PubMed]
- Taylor, L. COVID-19: Antimicrobial misuse in Americas sees drug resistant infections surge, says WHO. BMJ 2021, 375, n2845. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Li, C.; Yuan, J.; Lu, T.; Okamoto, H.; Murphy, D.G. Full-length genome sequences of five hepatitis C virus isolates representing subtypes 3g, 3h, 3i and 3k, and a unique genotype 3 variant. J. Gen. Virol. 2013, 94, 543–548. [Google Scholar] [CrossRef]
- Hepatitis C Virus Subtype 1a Polyprotein Gene, Complete cds. 2009. Available online: https://www.ncbi.nlm.nih.gov/nuccore/AF009606 (accessed on 12 March 2024).
- Gall, A.; Morris, C.; Kellam, P.; Berry, N. Complete Genome Sequence of the WHO International Standard for HIV-1 RNA Determined by Deep Sequencing. Genome Announc. 2014, 2, e01254-13. [Google Scholar] [CrossRef]
- HIV-1 Isolate NIBSC-1 from United Kingdom Gag Protein (Gag), Pol Protein (pol), Vif Protein (Vif), Vpr Protein (Vpr), tat Protein (Tat), rev Protein (Rev), vpu Protein (Vpu), Envelope Glycoprotein (Env), and Nef Protein (Nef) Genes, Complete cds. 2014. Available online: https://www.ncbi.nlm.nih.gov/nuccore/KJ019215 (accessed on 12 March 2024).
- Influenza A Virus (A/Kerala/RGCB140815/2014(H1N1)) Segment 1 Polymerase PB2 (PB2) Gene, Complete cds. 2016. Available online: https://www.ncbi.nlm.nih.gov/nuccore/KU922529 (accessed on 12 March 2024).
- Jones, S.; Prasad, R.; Nair, A.S.; Dharmaseelan, S.; Usha, R.; Nair, R.R.; Pillai, R.M. Whole-Genome Sequences of Influenza A(H1N1)pdm09 Virus Isolates from Kerala, India. Genome Announc. 2017, 5, e00598-17. [Google Scholar] [CrossRef] [PubMed]
- Oong, X.Y.; Ng, K.T.; Tan, J.L.; Chan, K.G.; Kamarulzaman, A.; Chan, Y.F.; Sam, I.C.; Tee, K.K. Whole-Genome Phylogenetic Analysis of Influenza B/Phuket/3073/2013-Like Viruses and Unique Reassortants Detected in Malaysia between 2012 and 2014. PLoS ONE 2017, 12, e0170610. [Google Scholar] [CrossRef] [PubMed]
- Influenza B Virus (B/Malaysia/9012150273/2009) Segment 4 Hemagglutinin (HA) Gene, Complete cds. 2017. Available online: https://www.ncbi.nlm.nih.gov/nuccore/KX269900.1 (accessed on 12 March 2024).
- Human Poliovirus 1 Isolate 558/BRA-PE/88, Complete Genome. 2013. Available online: https://www.ncbi.nlm.nih.gov/nuccore/KF537633 (accessed on 12 March 2024).
- Tavares, F.N.; da Costa, E.V.; Kew, O.M.; da Silva, E.E. Complete Genome Sequence of the Last Representative Genotype of Wild Indigenous Poliovirus Type 1, Which Circulated in Brazil. Genome Announc. 2013, 1, e00811-13. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Wang, Y.; Wang, W.; Nie, K.; Zhao, Y.; Su, J.; Deng, Y.; Zhou, W.; Li, Y.; Wang, H.; et al. Complete Genome Sequence of Middle East Respiratory Syndrome Coronavirus (MERS-CoV) from the First Imported MERS-CoV Case in China. Genome Announc. 2015, 3, e00818-15. [Google Scholar] [CrossRef]
- Middle East Respiratory Syndrome Coronavirus Strain ChinaGD01, Complete Genome. 2015. Available online: https://www.ncbi.nlm.nih.gov/nuccore/KT006149 (accessed on 12 March 2024).
- SARS Coronavirus Sin2500, Complete Genome-Nucleotide-NCBI. Available online: https://www.ncbi.nlm.nih.gov/nuccore/30468042 (accessed on 12 March 2024).
- Shental, N.; Levy, S.; Wuvshet, V.; Skorniakov, S.; Shalem, B.; Ottolenghi, A.; Greenshpan, Y.; Steinberg, R.; Edri, A.; Gillis, R.; et al. Efficient high-throughput SARS-CoV-2 testing to detect asymptomatic carriers. Sci. Adv. 2020, 6, eabc5961. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Karpuj, M.V.; Shaytov, D.R.; Shemer-Avni, Y.; Gideon, M.; Frenkel, Z.M.; Sivan, S. Comparative Bioinformatic Analysis Reveals Conserved Regions in SARS-CoV-2 Genome for RAPID Pandemic Response. Int. J. Mol. Sci. 2024, 25, 5764. https://doi.org/10.3390/ijms25115764
Karpuj MV, Shaytov DR, Shemer-Avni Y, Gideon M, Frenkel ZM, Sivan S. Comparative Bioinformatic Analysis Reveals Conserved Regions in SARS-CoV-2 Genome for RAPID Pandemic Response. International Journal of Molecular Sciences. 2024; 25(11):5764. https://doi.org/10.3390/ijms25115764
Chicago/Turabian StyleKarpuj, Marcela Viviana, D. R. Shaytov, Yonat Shemer-Avni, Michael Gideon, Zakharia M. Frenkel, and Sarit Sivan. 2024. "Comparative Bioinformatic Analysis Reveals Conserved Regions in SARS-CoV-2 Genome for RAPID Pandemic Response" International Journal of Molecular Sciences 25, no. 11: 5764. https://doi.org/10.3390/ijms25115764




