Identification of Apple Flower Development-Related Gene Families and Analysis of Transcriptional Regulation
Abstract
:1. Introduction
2. Results
2.1. Identification of C2H2, DELLA, and FKF1 Gene Families in the Apple Genome
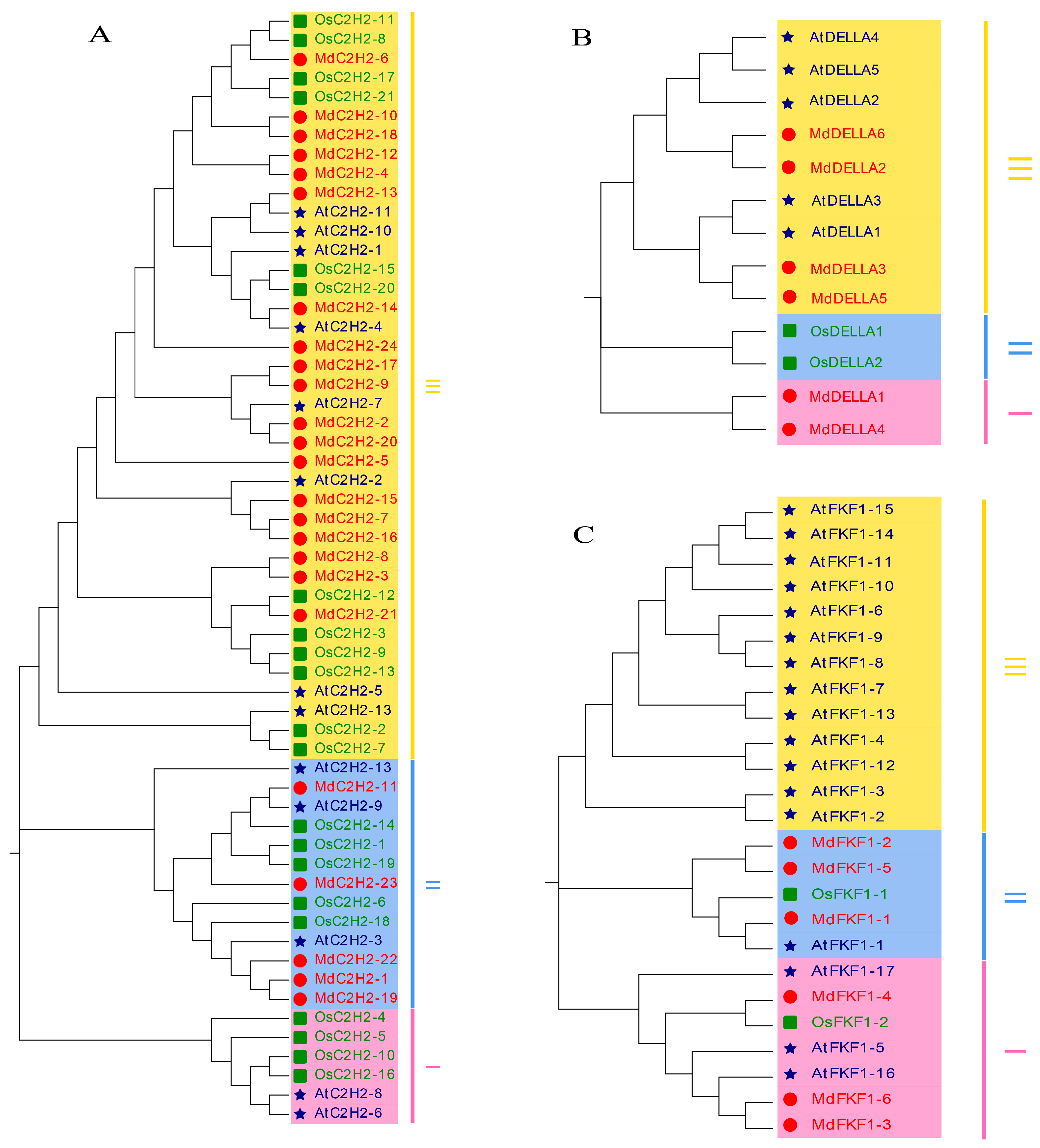
2.2. Phylogenetic Analysis of the C2H2, DELLA, and FKF1 Gene Families
2.3. Gene Structure and Conserved Protein Motif Analysis of the C2H2, DELLA, and FKF1 Gene Families
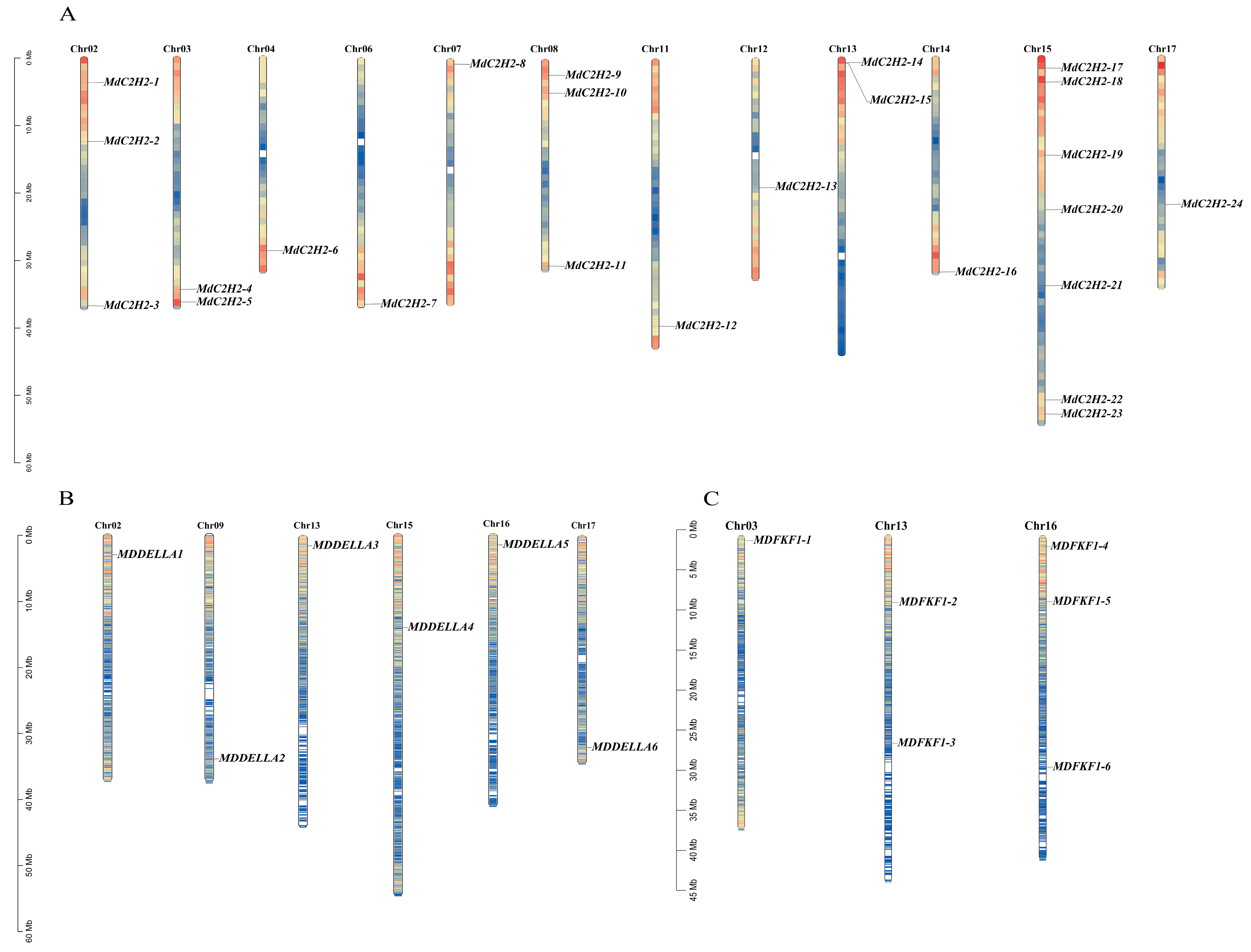
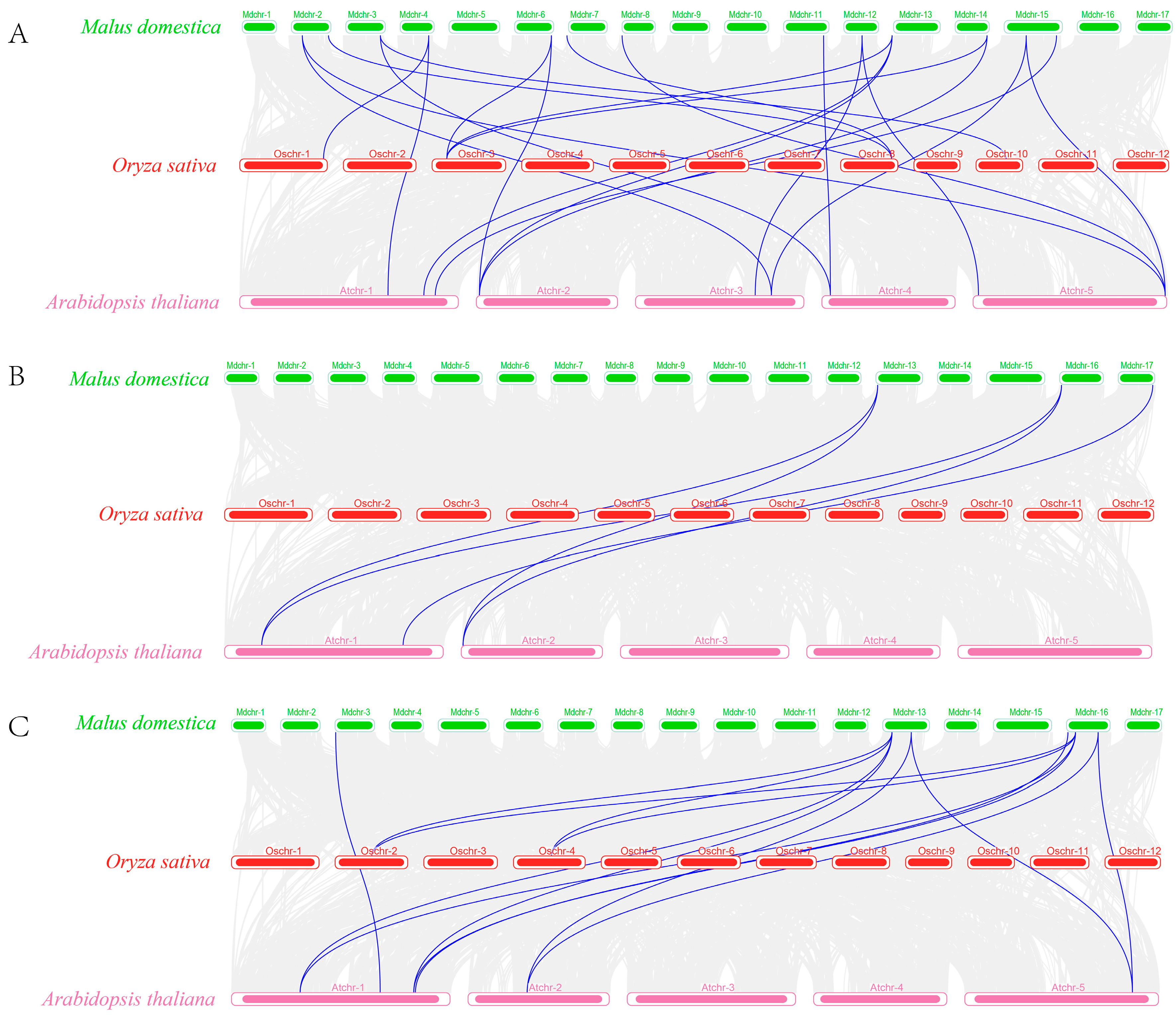
2.4. Chromosomal Distribution and Covariance Analysis of C2H2, DELLA, and FKF1 Genes in Apples
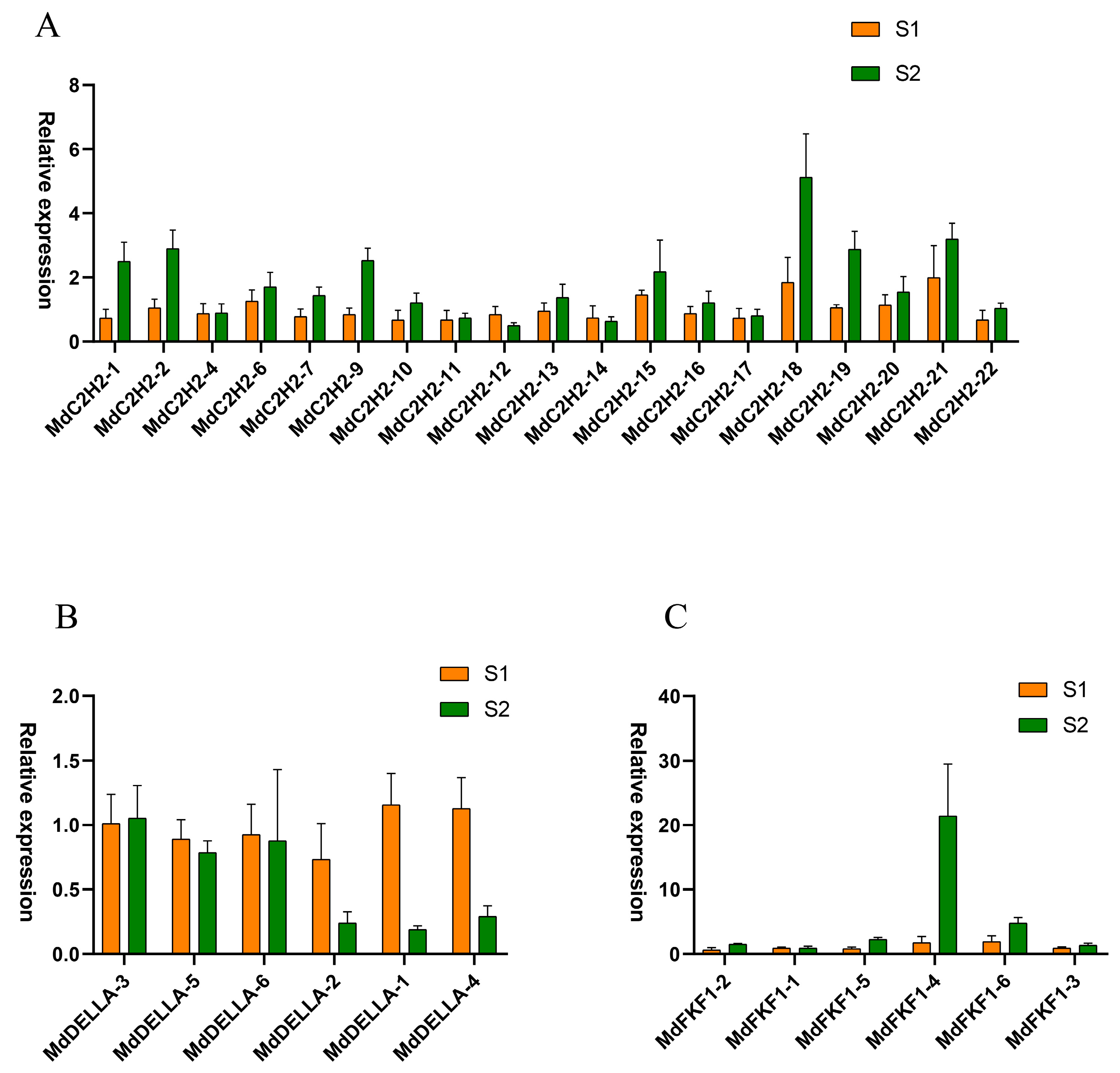
2.5. Expression Profiles of C2H2, DELLA, and FKF1 Genes in Apple Flower Buds
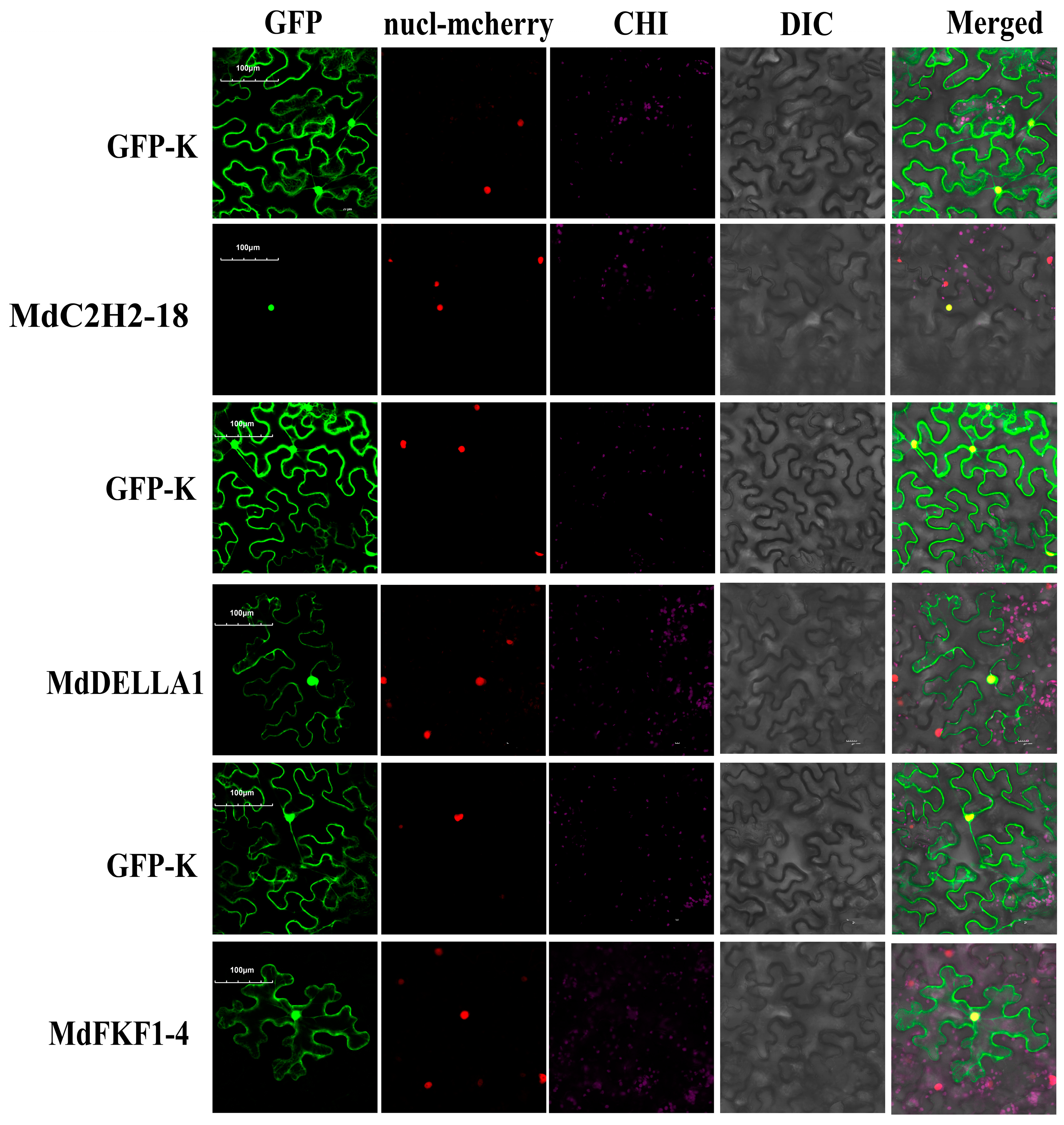
2.6. Subcellular Localization of Apple C2H2, DELLA, and FKF1 Proteins
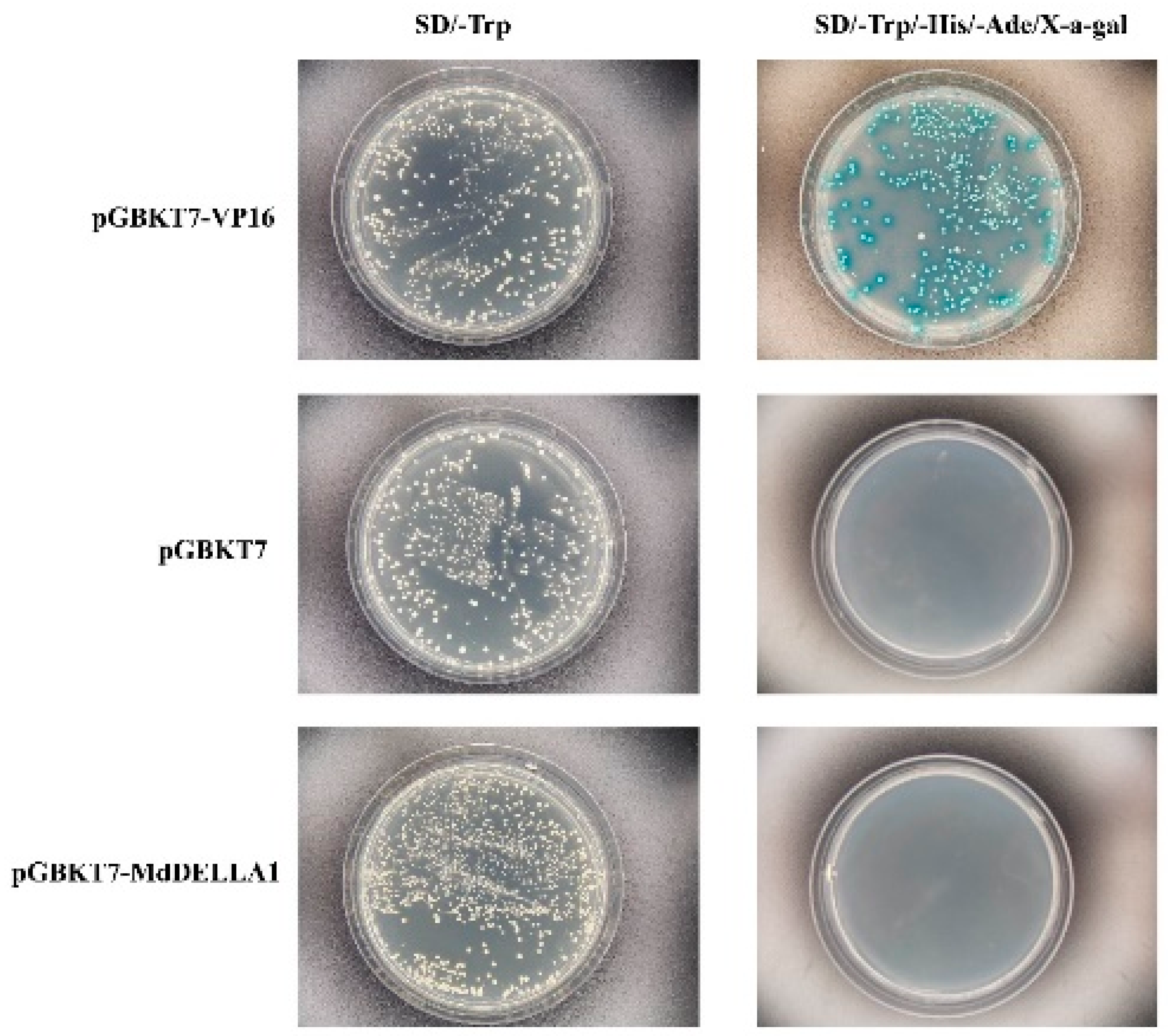
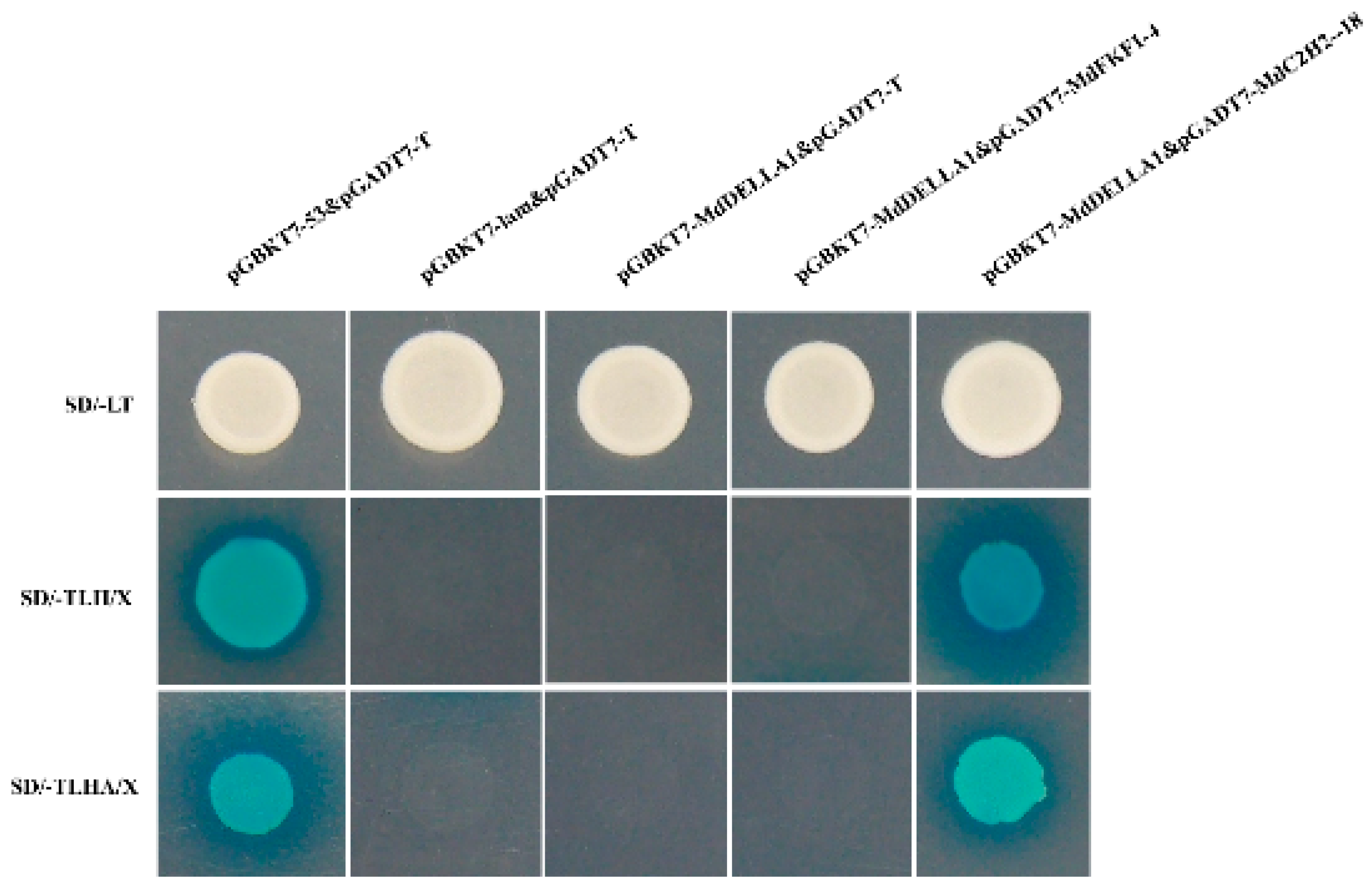
2.7. Yeast Two-Hybrid Validation
3. Materials and Methods
3.1. Test Materials
3.2. Identification and Prediction of Physicochemical Properties of C2H2, DELLA, and FKF1 Genes in Apples
3.3. Gene Structure, Analysis, and Phylogenetic Tree Construction
3.4. Chromosome Distribution and Co-Linearity Analysis
3.5. Quantitative Fluorescence Analysis
3.6. Subcellular Localization
3.7. Yeast Hybridization
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Osnato, M.; Cota, I.; Nebhnani, P.; Cereijo, U.; Pelaz, S. Photoperiod control of plant growth: Flowering time genes beyond flowering. Front. Plant Sci. 2022, 12, 805635. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.Z.; Zhou, Y.P.; Lv, T.X.; Xie, C.P. Research progress on the autonomous flowering time pathway in Arabidopsis. Physiol. Mol. Biol. Plants 2017, 23, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Dennis, E.S.; Finnegan, E.J.; Bilodeau, P.; Chhaudhury, A.; Genger, R.; Helliwell, C.A.; Sheldon, C.C.; Bagnall, D.J.; Peacock, W.J. Vernalization and the initiation of flowering. Semin. Cell Dev. Biol. 1996, 7, 441–448. [Google Scholar] [CrossRef]
- Bao, S.; Hua, C.; Shen, L.; Yu, H. New insights into gibberellin signaling in regulating flowering in Arabidopsis. J. Integr. Plant Biol. 2020, 62, 118–131. [Google Scholar] [CrossRef] [PubMed]
- Susila, H.; Nasim, Z.; Ahn, J.H. Ambient temperature-responsive mechanisms coordinate regulation of flowering time. Int. J. Mol. Sci. 2018, 19, 3196. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W. Regulation of flowering time by the miR156-mediated age pathway. J. Exp. Bot. 2014, 65, 4723–4730. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Liu, H.; Wang, F.; Ma, Z.; Mei, C.; Ma, F.; Mao, K. Apple MdbHLH4 promotes the flowering transition through interactions with FLOWERING LOCUS C and transcriptional activation of FLOWERING LOCUS T. Sci. Hortic. 2023, 322, 112444. [Google Scholar] [CrossRef]
- Lyu, T.; Cao, J. Cys2/His2 zinc-finger proteins in transcriptional regulation of flower development. Int. J. Mol. Sci. 2018, 19, 2589. [Google Scholar] [CrossRef]
- Zhang, H.; Zhao, T.; Zhuang, P.; Song, Z.; Du, H. NbCZF1, a novel C2H2-type zinc finger protein, as a new regulator of SsCut-induced plant immunity in Nicotiana benthamiana. Plant Cell Physiol. 2016, 57, 2472–2484. [Google Scholar] [CrossRef]
- Ming, N.; Ma, N.; Jiao, B.; Lv, W.; Meng, Q. Genome wide identification of C2H2-type zinc finger proteins of tomato and expression analysis under different abiotic stresses. Plant Mol. Biol. Rep. 2020, 38, 75–94. [Google Scholar] [CrossRef]
- Li, S.; Li, Y.; Cai, Q.; Li, X.; Sun, Y.; Yu, T.; Yang, J.; Zhang, J. Analysis of the C2H2 gene family in maize (Zea mays L.) under cold stress: Identification and expression. Life Sci. 2022, 13, 122. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, H.; Wang, H.L.; Zhang, T.; Zeng, X.Q.; Chen, H.; Wang, Z.W.; Zhang, J.; Zheng, H.; Tang, J.; Ling, Y.H. NONSTOP GLUMES1 encodes a C2H2 zinc finger protein that regulates spikelet development in rice. Plant Cell 2020, 32, 392–413. [Google Scholar] [CrossRef] [PubMed]
- Prunet, N.; Yang, W.; Das, P.; Meyerowitz, E.M.; Jack, T.P. SUPERMAN prevents class B gene expression and promotes stem cell termination in the fourth whorl of Arabidopsis thaliana flowers. Proc. Natl. Acad. Sci. USA 2017, 114, 7166–7171. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Yu, H.; Xia, S.; Cui, Y.; Yu, X.; Liu, H.; Zeng, D.; Hu, J.; Zhang, Q.; Gao, Z. The C2H2 zinc-finger protein LACKING RUDIMENTARY GLUME 1 regulates spikelet development in rice. Sci. Bull. 2020, 65, 753–764. [Google Scholar] [CrossRef] [PubMed]
- Shuai, Y.; Feng, G.; Yang, Z.; Liu, Q.; Han, J.; Xu, X.; Nie, G.; Huang, L.; Zhang, X. Genome-wide identification of C2H2-type zinc finger gene family members and their expression during abiotic stress responses in orchardgrass (Dactylis glomerata). Genome 2022, 64, 189–203. [Google Scholar] [CrossRef] [PubMed]
- Duan, S.F.; Zhao, Y.; Xiang, G.S.; Baldwin, T.C.; Liang, Y.L. Genome-wide identification and expression analysis of the C2H2-zinc finger transcription factor gene family and screening of candidate genes involved in floral development in Coptis teeta Wall (Ranunculaceae). Front. Genet. 2024, 15, 1349673. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Coulter, J.A.; Li, Y.; Zhang, X.; Meng, J.; Zhang, J.; Liu, Y. Genome-wide identification and analysis of the Q-type C2H2 gene family in potato (Solanum tuberosum L.). Int. J. Biol. Macromol. 2020, 153, 327–340. [Google Scholar] [CrossRef] [PubMed]
- Sun, T. The molecular mechanism and evolution of the GA–GID1–DELLA signaling module in plants. Curr. Biol. 2011, 21, 338–345. [Google Scholar] [CrossRef]
- Ikeda, A.; Ueguchi-Tanaka, M.; Sonoda, Y.; Kitano, H.; Koshioka, M.; Futsuhara, Y.; Matsuoka, M.; Yamaguchi, J. Slender rice, aconstitutive gibberellin response mutant, is caused by a nullmutation of the SLRl gene, an ortholog of the height regulating gene GAI/RGA/RHT/D8. Plant Cell 2001, 13, 999–1010. [Google Scholar] [CrossRef]
- Shohat, H.; Illouz-Eliaz, N.; Kanno, Y.; Seo, M.; Weiss, D. The tomato DELLA protein PROCERA promotes abscisic acid responses in guard cells by upregulating an abscisic acid transporter. Plant Physiol. 2020, 184, 518–528. [Google Scholar] [CrossRef]
- Guan, H.; Huang, X.; Zhu, Y.; Xie, B.; Liu, H.; Song, S.; Hao, Y.; Chen, R. Identification of DELLA genes and key stage for GA sensitivity in bolting and flowering of flowering Chinese cabbage. Int. J. Mol. Sci. 2021, 22, 12092. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Hu, W.W.; Shen, L.; Lee, L.Y.C.; Tao, Z.; Han, J.H.; Yu, H. Global identification of DELLA target genes during arabidopsis flower development. Plant Physiol. 2008, 147, 1126–1142. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Cheng, T.; Wang, P.; Luo, Y.; Wang, J.; Yang, L. Genome-wide bioinformatics analysis of DELLA-family proteins from plants. Plant Omics 2013, 6, 201–207. [Google Scholar]
- Yoshida, H.; Hirano, K.; Sato, T.; Mitsuda, N.; Nomoto, M.; Maeo, K.; Koketsu, E.; Mitani, R. DELLA protein functions as a transcriptional activator through the DNA binding of the indeterminate domain family proteins. Proc. Natl. Acad. Sci. USA 2014, 111, 7861–7866. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, S.; Böhlenius, H.; Moritz, T.; Nilsson, O. GA4 is the active gibberellin in the regulation of LEAFY transcription and Arabidopsis floral initiation. Plant Cell 2006, 18, 2172–2181. [Google Scholar] [CrossRef] [PubMed]
- Xue, H.; Gao, X.; He, P.; Xiao, G. Origin, evolution, and molecular function of DELLA proteins in plants. Crop J. 2022, 10, 287–299. [Google Scholar] [CrossRef]
- Jia, Q.; Xiao, Z.X.; Wong, F.L.; Sun, S.; Liang, K.J.; Lam, H.M. Genome-wide analyses of the soybean F-box gene family in response to salt stress. Int. J. Mol. Sci. 2017, 18, 818. [Google Scholar] [CrossRef] [PubMed]
- Han, S.H.; Yoo, S.C.; Lee, B.D.; An, G.; Paek, N.C. Rice FLAVIN-BINDING, KELCH REPEAT, F-BOX 1 (OsFKF1) promotes flowering independent of photoperiod. Plant Cell Environ. 2015, 38, 2527–2540. [Google Scholar] [CrossRef] [PubMed]
- Jia, F.; Wu, B.; Li, H.; Huang, J.; Zheng, C. Genome-wide identification and characterisation of F-box family in maize. Mol. Genet. Genom. 2013, 288, 559–577. [Google Scholar] [CrossRef]
- Li, F.; Zhang, X.; Hu, R.; Wu, F.; Ma, J.; Meng, Y.; Fu, Y.F. Identification and molecular characterization of FKF1 and GI homologous genes in soybean. PLoS ONE 2013, 8, e79036. [Google Scholar] [CrossRef]
- Song, Y.H.; Smith, R.W.; To, B.J.; Millar, A.J.; Imaizumi, T. FKF1 conveys timing information for CONSTANS stabilization in photoperiodic flowering. Science 2012, 336, 1045–1049. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Li, X.; Zeng, B.; Zhong, M.; Yang, J.; Yang, P.; Li, X.; He, C.; Lin, J.; Liu, X.; et al. FKF1 F-box protein promotes flowering in part by negatively regulating DELLA protein stability under long-day photoperiod in Arabidopsis. J. Integr. Plant Biol. 2020, 62, 1717–1740. [Google Scholar] [CrossRef]
- Becker, A.; Theißen, G. The major clades of MADS-box genes and their role in the development and evolution of flowering plants. Mol. Phylogenet. Evol. 2003, 29, 464–489. [Google Scholar] [CrossRef] [PubMed]
- Söding, J. Protein homology detection by HMM–HMM comparison. Bioinformatics 2005, 21, 951–960. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, Z.; Li, X.; Chen, Z.; Liu, Y.; Zhang, F.; Dai, Q.; Yu, Q.; Li, N. Identification and analysis of the expression of the PIP5K gene family in tomatoes. Int. J. Mol. Sci. 2023, 25, 159. [Google Scholar] [CrossRef]
- Sarwar, R.; Jiang, T.; Ding, P.; Gao, Y.; Tan, X.; Zhu, K. Genome-wide analysis and functional characterization of the DELLA gene family associated with stress tolerance in B. napus. BMC Plant Biol. 2021, 21, 286. [Google Scholar] [CrossRef]
- Jouffrey, V.; Leonard, A.S.; Ahnert, S.E. Gene duplication and subsequent diversifification strongly affect phenotypic evolvability and robustness. R. Soc. Open Sci. 2021, 8, 201636. [Google Scholar] [CrossRef]
- Neugebauer, K.M. On the importance of being co-transcriptional. J. Cell Sci. 2002, 115, 3865–3871. [Google Scholar] [CrossRef]
- Chung, B.Y.W.; Simons, C.; Firth, A.E.; Brown, C.M.; Hellens, R.P. Effect of 5’UTR introns on gene expression in Arabidopsis thaliana. BMC Genom. 2006, 7, 120. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Zhao, Z.; Chen, H.; Ao, W.; Lu, L.; Liu, J.; Li, X.; Sun, Y. Genome-wide characterization and expression of DELLA genes in Cucurbita moschata reveal their potential roles under development and abiotic stress. Front. Plant Sci. 2023, 14, 1137126. [Google Scholar] [CrossRef]
- Zhong, G.Y.; Yang, Y. Characterization of grape Gibberellin Insensitive1 mutant alleles in transgenic Arabidopsis. Transgenic Res. 2012, 21, 725–741. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Li, Y.; Huang, J.; Jiao, R. DoDELLA1, a DELLA protein from Dioscorea opposite, regulates the growth and development in transgenic tobacco by controlling gibberellin level. Plant Growth Regul. 2022, 97, 571–583. [Google Scholar] [CrossRef]
- Wen, L.H.; Zhong, W.J.; Huo, X.M.; Zhuang, W.B.; Ni, Z.J.; Gao, Z.H. Expression analysis of ABA- and GA-related genes during four stages of bud dormancy in Japanese apricot (Prunus mume Sieb. et Zucc). J. Hortic. Sci. Biotechnol. 2016, 91, 362–369. [Google Scholar] [CrossRef]
- Gagne, J.M.; Downes, B.P.; Shiu, S.-H.; Durski, A.M.; Vierstra, R.D. The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc. Natl. Acad. Sci. USA 2002, 99, 11519–11524. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.; Nijhawan, A.; Arora, R.; Agarwal, P.; Ray, S.; Sharma, P.; Kapoor, S.; Tyagi, A.K.; Khurana, J.P. F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol. 2007, 143, 1467–1483. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Zhang, A.; Huang, L.; Yu, X.; Yang, K.; Fan, S.; Cao, J. BcMF20, a putative pollen-specific transcription factor from Brassica campestris ssp. Chinensis. Mol. Biol. Rep. 2011, 38, 5321–5325. [Google Scholar] [CrossRef] [PubMed]
- Shan, Z.; Luo, X.; Wu, M.; Wei, L.; Fan, Z.; Zhu, Y. Genome-wide identification and expression of GRAS gene family members in cassava. BMC Plant Biol. 2020, 20, 46. [Google Scholar] [CrossRef]
- Nelson, D.C.; Lasswell, J.; Rogg, L.E.; Cohen, M.A.; Bartel, B. FKF1, a clock-controlled gene that regulates the transition to flowering in Arabidopsis. Cell 2000, 101, 331–340. [Google Scholar] [CrossRef]
- Jorge, H.G.; Asier, B.M.; Renaud, D.; Blázquez, M.A. Origin of Gibberellin-Dependent transcriptional regulation by molecular exploitation of a transactivation domain in DELLA proteins. Mol. Biol. Evol. 2019, 36, 908–918. [Google Scholar]

| Gene Name | Gene ID | Chromosome Location | Length of CDS (bp) | Physicochemical Properties | Subcellular Localization | |||
|---|---|---|---|---|---|---|---|---|
| No. | Mw | pI | GRAVY | |||||
| MdC2H2-1 | MD02G1048300 | Chr2 | 2716 | 473 | 53,935.18 | 8.58 | −0.785 | nucl |
| MdC2H2-3 | MD02G1313400 | Chr2 | 2983 | 365 | 40,796.93 | 8.8 | −0.608 | nucl |
| MdC2H2-2 | MD02G1151700 | Chr2 | 3781 | 520 | 54,260.67 | 8.83 | −0.495 | nucl |
| MdC2H2-4 | MD03G1258200 | Chr3 | 3533 | 464 | 50,955.33 | 9.16 | −0.637 | nucl |
| MdC2H2-5 | MD03G1283900 | Chr3 | 2591 | 534 | 58,928.67 | 8.82 | −0.818 | nucl |
| MdC2H2-6 | MD04G1203000 | Chr4 | 3703 | 522 | 57,092.7 | 8.99 | −0.746 | nucl |
| MdC2H2 7 | MD06G1234600 | Chr6 | 3271 | 503 | 56,383.07 | 8.94 | −0.884 | nucl |
| MdC2H2 8 | MD07G1008700 | Chr7 | 3170 | 389 | 43,693.48 | 7.04 | −0.797 | nucl |
| MdC2H2-11 | MD08G1239600 | Chr8 | 5069 | 373 | 42,921.8 | 6.24 | −1.145 | cyto |
| MdC2H2-10 | MD08G1063600 | Chr8 | 3377 | 488 | 54,409.49 | 8.86 | −0.18 | nucl |
| MdC2H2-9 | MD08G1033100 | Chr8 | 4082 | 535 | 55,755.13 | 8.81 | −0.437 | nucl |
| MdC2H2-12 | MD11G1278800 | Chr11 | 3430 | 466 | 51,228.26 | 9.2 | −0.691 | nucl |
| MdC2H2-13 | MD12G1120300 | Chr12 | 2650 | 530 | 58,194.12 | 9.23 | −0.666 | nucl |
| MdC2H2-14 | MD13G1013100 | Chr13 | 6347 | 600 | 64,123.49 | 9.29 | −0.717 | nucl |
| MdC2H2-15 | MD13G1015000 | Chr13 | 3018 | 483 | 52,950.02 | 8.94 | −0.784 | nucl |
| MdC2H2-16 | MD14G1241600 | Chr14 | 3364 | 503 | 56,507.32 | 9.02 | −0.881 | nucl |
| MdC2H2-23 | MD15G1431100 | Chr15 | 4846 | 294 | 34,436.24 | 7.92 | −0.773 | nucl |
| MdC2H2-17 | MD15G1029700 | Chr15 | 3481 | 541 | 56,410.92 | 8.62 | −0.445 | nucl |
| MdC2H2-18 | MD15G1058000 | Chr15 | 3342 | 533 | 58,976.45 | 8.54 | −0.798 | cyto |
| MdC2H2-20 | MD15G1266200 | Chr15 | 3521 | 528 | 55,077.62 | 9.04 | −0.481 | nucl |
| MdC2H2-21 | MD15G1323400 | Chr15 | 1076 | 358 | 40,054.06 | 5.73 | −0.636 | nucl |
| MdC2H2-19 | MD15G1186600 | Chr15 | 1835 | 363 | 41,314.08 | 9.16 | −0.684 | nucl |
| MdC2H2-22 | MD15G1411000 | Chr15 | 2708 | 369 | 42,755.28 | 8.34 | −0.932 | nucl |
| MdC2H2-24 | MD17G1186000 | Chr17 | 2471 | 521 | 56,980.48 | 8.34 | −0.668 | nucl |
| MdDELLA3 | MD13G1022100 | Chr13 | 1947 | 635 | 69,734.67 | 5.3 | −0.28 | nucl |
| MdDELLA5 | MD16G1023300 | Chr16 | 1919 | 639 | 70,235.28 | 5.25 | −0.287 | nucl |
| MdDELLA6 | MD17G1260700 | Chr17 | 1754 | 584 | 63,697.47 | 4.95 | −0.289 | nucl |
| MdDELLA2 | MD09G1264800 | Chr09 | 1742 | 580 | 63,166.16 | 5.09 | −0.215 | chlo |
| MdDELLA1 | MD02G1039600 | Chr02 | 1640 | 546 | 59,826.8 | 5.63 | −0.114 | chlo |
| MdDELLA4 | MD15G1180500 | Chr15 | 1643 | 547 | 59,896.09 | 5.53 | −0.091 | nucl |
| MdFKF1-2 | MD13G1117000 | Chr13 | 3505 | 371 | 41,108.05 | 5.75 | −0.138 | nucl |
| MdFKF1-1 | MD03G1007500 | Chr3 | 2972 | 437 | 49,203.71 | 9.47 | −0.243 | cyto |
| MdFKF1-5 | MD16G1117600 | Chr16 | 3235 | 386 | 43,284.64 | 6.2 | −0.21 | cyto |
| MdFKF1-4 | MD16G1018000 | Chr16 | 2898 | 632 | 70,550.83 | 5.53 | −0.351 | nucl |
| MdFKF1-6 | MD16G1255700 | Chr16 | 6558 | 623 | 67,868.46 | 5.29 | −0.112 | nucl |
| MdFKF1-3 | MD13G1249400 | Chr13 | 5468 | 623 | 68,016.72 | 5.34 | −0.107 | nucl |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mei, C.; Li, X.; Yan, P.; Feng, B.; Mamat, A.; Wang, J.; Li, N. Identification of Apple Flower Development-Related Gene Families and Analysis of Transcriptional Regulation. Int. J. Mol. Sci. 2024, 25, 7510. https://doi.org/10.3390/ijms25147510
Mei C, Li X, Yan P, Feng B, Mamat A, Wang J, Li N. Identification of Apple Flower Development-Related Gene Families and Analysis of Transcriptional Regulation. International Journal of Molecular Sciences. 2024; 25(14):7510. https://doi.org/10.3390/ijms25147510
Chicago/Turabian StyleMei, Chuang, Xianguo Li, Peng Yan, Beibei Feng, Aisajan Mamat, Jixun Wang, and Ning Li. 2024. "Identification of Apple Flower Development-Related Gene Families and Analysis of Transcriptional Regulation" International Journal of Molecular Sciences 25, no. 14: 7510. https://doi.org/10.3390/ijms25147510






