CryoEM Workflow Acceleration with Feret Signatures
Abstract
:1. Introduction
2. Results
2.1. Assessment of Preferred Orientation
2.1.1. Data Displaying Strong Preferred Orientations
2.1.2. Data with Good Coverage of Orientations
2.1.3. Feret Signature Robustness: Analysis of Randomly Split Particle Datasets
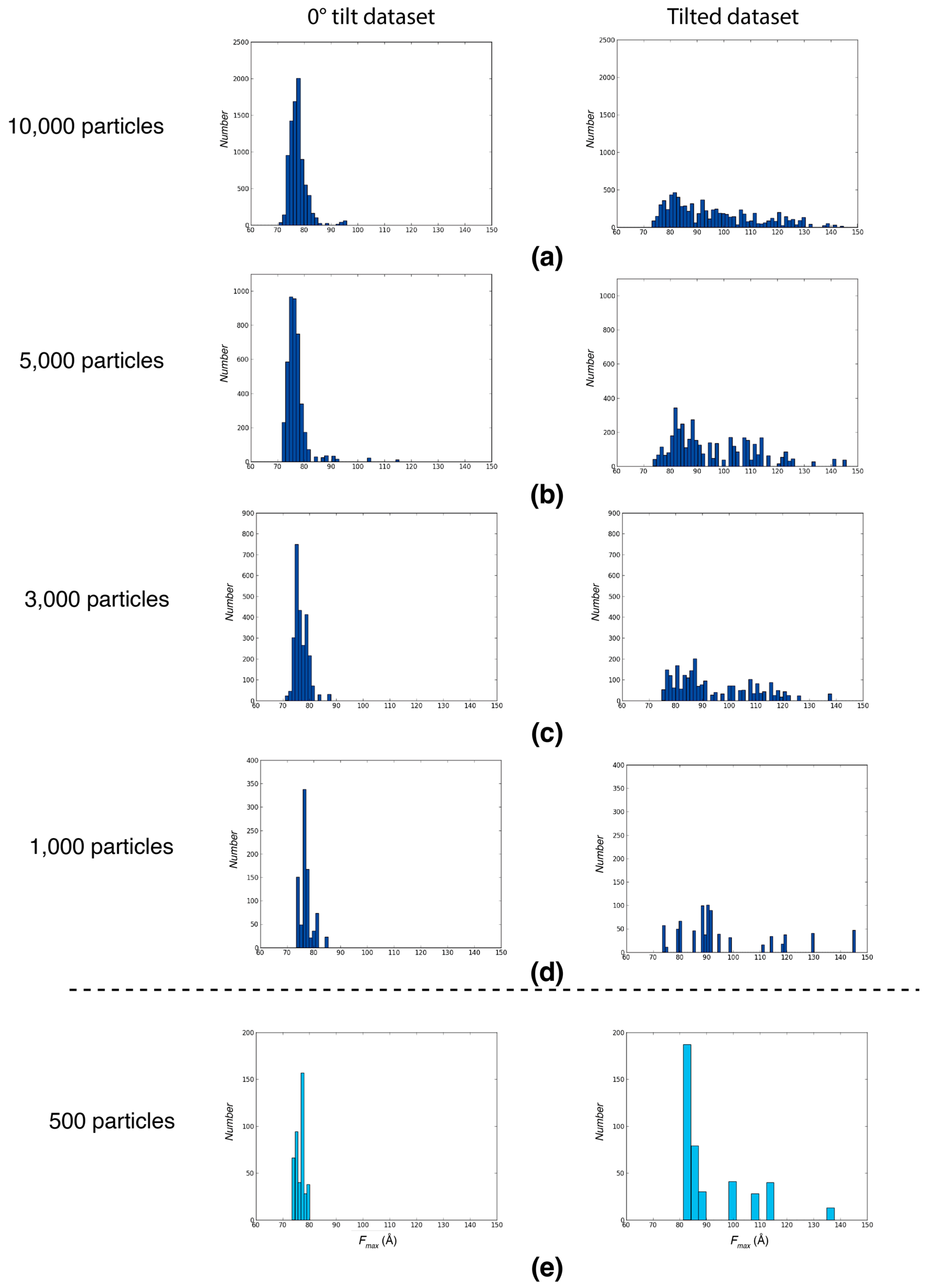
2.1.4. Minimal Number of Particles Required for Preferred Orientation Detection
2.2. Detection of Classification Inconsistencies with Feret Signatures
3. Discussion
4. Materials and Methods
4.1. Materials
4.1.1. CryoEM Data
4.1.2. Feret Signatures: Simulated Feret Signatures from 3D Maps
4.1.3. Feret Analysis
Characterization of Four Integrin Conformation Distributions
Characterization of Influenza Hemagglutinin Trimer Distributions
Characterization of Rabbit Muscle Aldolase Distributions from Tilt Series
Characterization of Beta-Galactosidase
4.2. Methods
4.2.1. Data Processing and Particle Selection
4.2.2. Feret Signature Extraction and Analysis
Loading Image Stacks
Running Feret Signature Evaluation
Classification and Averaging
Filtering and Noise Reduction
Comprehensive Analysis and Visualization
Data Management and Reproducibility
Parameter Optimization and Continuous Refinement
4.2.3. Simulation of Feret Signatures
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 2D | two-dimensional |
| 3D | three-dimensional |
| Å | Ångström |
| cryoEM | cryogenic electron microscopy |
| CTF | contrast transfer function |
| e− | electron |
| eV | electronvolt |
| Fmax | maximum Feret diameter |
| kV | kilovolt |
| SPA | single particle analysis |
Appendix A. Figures
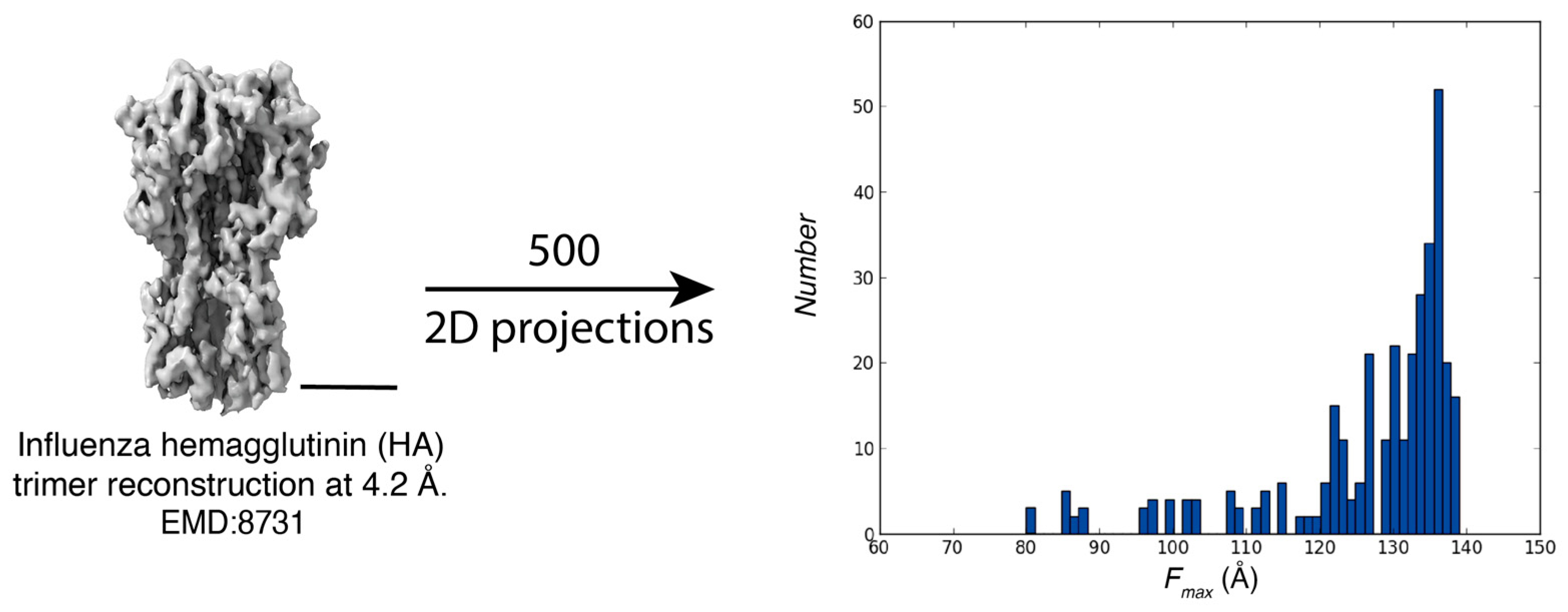

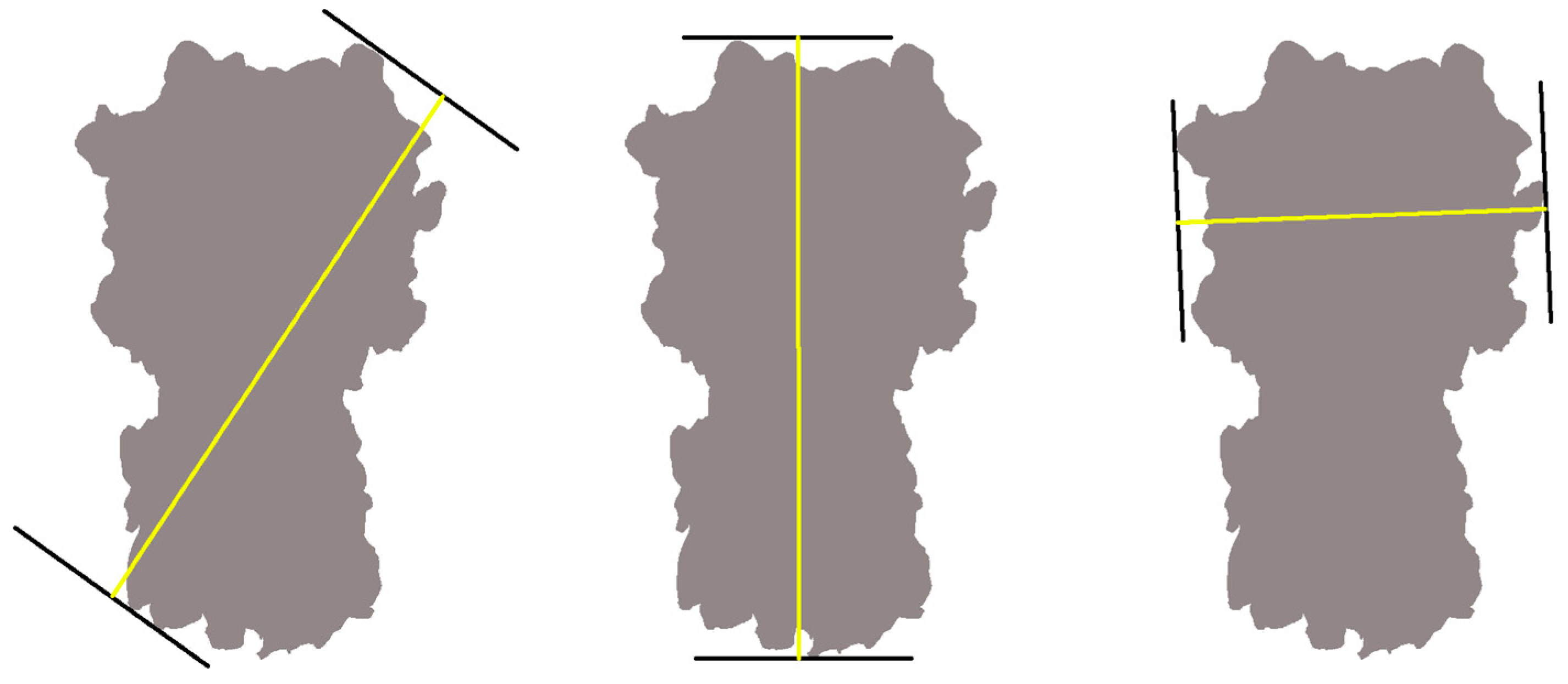
References
- Kühlbrandt, W. The Resolution Revolution. Science 2014, 343, 1443–1444. [Google Scholar] [CrossRef] [PubMed]
- Subramaniam, S. The Cryo-EM Revolution: Fueling the next Phase. IUCrJ 2019, 6, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Callaway, E. The Protein-Imaging Technique Taking over Structural Biology. Nature 2020, 578, 201. [Google Scholar] [CrossRef] [PubMed]
- Danev, R.; Yanagisawa, H.; Kikkawa, M. Cryo-Electron Microscopy Methodology: Current Aspects and Future Directions. Trends Biochem. Sci. 2019, 44, 837–848. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Dang, S. Recent Technical Advances in Sample Preparation for Single-Particle Cryo-EM. Front. Mol. Biosci. 2022, 9, 892459. [Google Scholar] [CrossRef] [PubMed]
- Patel, A.; Toso, D.; Litvak, A.; Nogales, E. Efficient Graphene Oxide Coating Improves Cryo-EM Sample Preparation and Data Collection from Tilted Grids. bioRxiv 2021. [Google Scholar] [CrossRef]
- Glaeser, R.M.; Han, B.-G. Opinion: Hazards Faced by Macromolecules When Confined to Thin Aqueous Films. Biophys. Rep. 2017, 3, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.Z.; Baldwin, P.R.; Davis, J.H.; Williamson, J.R.; Potter, C.S.; Carragher, B.; Lyumkis, D. Addressing Preferred Specimen Orientation in Single-Particle Cryo-EM through Tilting. Nat. Methods 2017, 14, 793–796. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.-T.; Fan, H.; Hu, J.J.; Zhou, Z.H. Overcoming the Preferred Orientation Problem in cryoEM with Self-Supervised Deep-Learning. bioRxiv 2024. [CrossRef]
- Noble, A.J.; Dandey, V.P.; Wei, H.; Brasch, J.; Chase, J.; Acharya, P.; Tan, Y.Z.; Zhang, Z.; Kim, L.Y.; Scapin, G.; et al. Routine Single Particle CryoEM Sample and Grid Characterization by Tomography. eLife 2018, 7, e34257. [Google Scholar] [CrossRef]
- Li, B.; Zhu, D.; Shi, H.; Zhang, X. Effect of Charge on Protein Preferred Orientation at the Air–Water Interface in Cryo-Electron Microscopy. J. Struct. Biol. 2021, 213, 107783. [Google Scholar] [CrossRef] [PubMed]
- Vilela, F.; Bezault, A.; Rodriguez De Francisco, B.; Sauvanet, C.; Xu, X.-P.; Swift, M.F.; Yao, Y.; Marrasi, F.M.; Hanein, D.; Volkmann, N. Characterization of Heterogeneity in Nanodisc Samples Using Feret Signatures. J. Struct. Biol. 2022, 214, 107916. [Google Scholar] [CrossRef] [PubMed]
- Radermacher, M.; Wagenknecht, T.; Verschoor, A.; Frank, J. Three-dimensional Reconstruction from a Single-exposure, Random Conical Tilt Series Applied to the 50S Ribosomal Subunit of Escherichia coli. J. Microsc. 1987, 146, 113–136. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Lander, G.C.; Herzik, M.A. Sub-2 Angstrom Resolution Structure Determination Using Single-Particle Cryo-EM at 200 keV. J. Struct. Biol. X 2020, 4, 100020. [Google Scholar] [CrossRef]
- Xu, X.-P.; Kim, E.; Swift, M.; Smith, J.W.; Volkmann, N.; Hanein, D. Three-Dimensional Structures of Full-Length, Membrane-Embedded Human αIIbβ3 Integrin Complexes. Biophys. J. 2016, 110, 798–809. [Google Scholar] [CrossRef] [PubMed]
- Mastronarde, D.N. Automated Electron Microscope Tomography Using Robust Prediction of Specimen Movements. J. Struct. Biol. 2005, 152, 36–51. [Google Scholar] [CrossRef] [PubMed]
- Punjani, A.; Rubinstein, J.L.; Fleet, D.J.; Brubaker, M.A. cryoSPARC: Algorithms for Rapid Unsupervised Cryo-EM Structure Determination. Nat. Methods 2017, 14, 290–296. [Google Scholar] [CrossRef]
- Volkmann, N.; Hanein, D. Quantitative Fitting of Atomic Models into Observed Densities Derived by Electron Microscopy. J. Struct. Biol. 1999, 125, 176–184. [Google Scholar] [CrossRef] [PubMed]
- Schatz, M.; Van Heel, M. Invariant Classification of Molecular Views in Electron Micrographs. Ultramicroscopy 1990, 32, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Buades, A.; Coll, B.; Morel, J.-M. A Non-Local Algorithm for Image Denoising. In Proceedings of the 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’05), San Diego, CA, USA, 20–26 June 2005; IEEE: San Diego, CA, USA, 2005; Volume 2, pp. 60–65. [Google Scholar]
- Sahoo, P.; Wilkins, C.; Yeager, J. Threshold Selection Using Renyi’s Entropy. Pattern Recogn. 1997, 30, 71–84. [Google Scholar] [CrossRef]
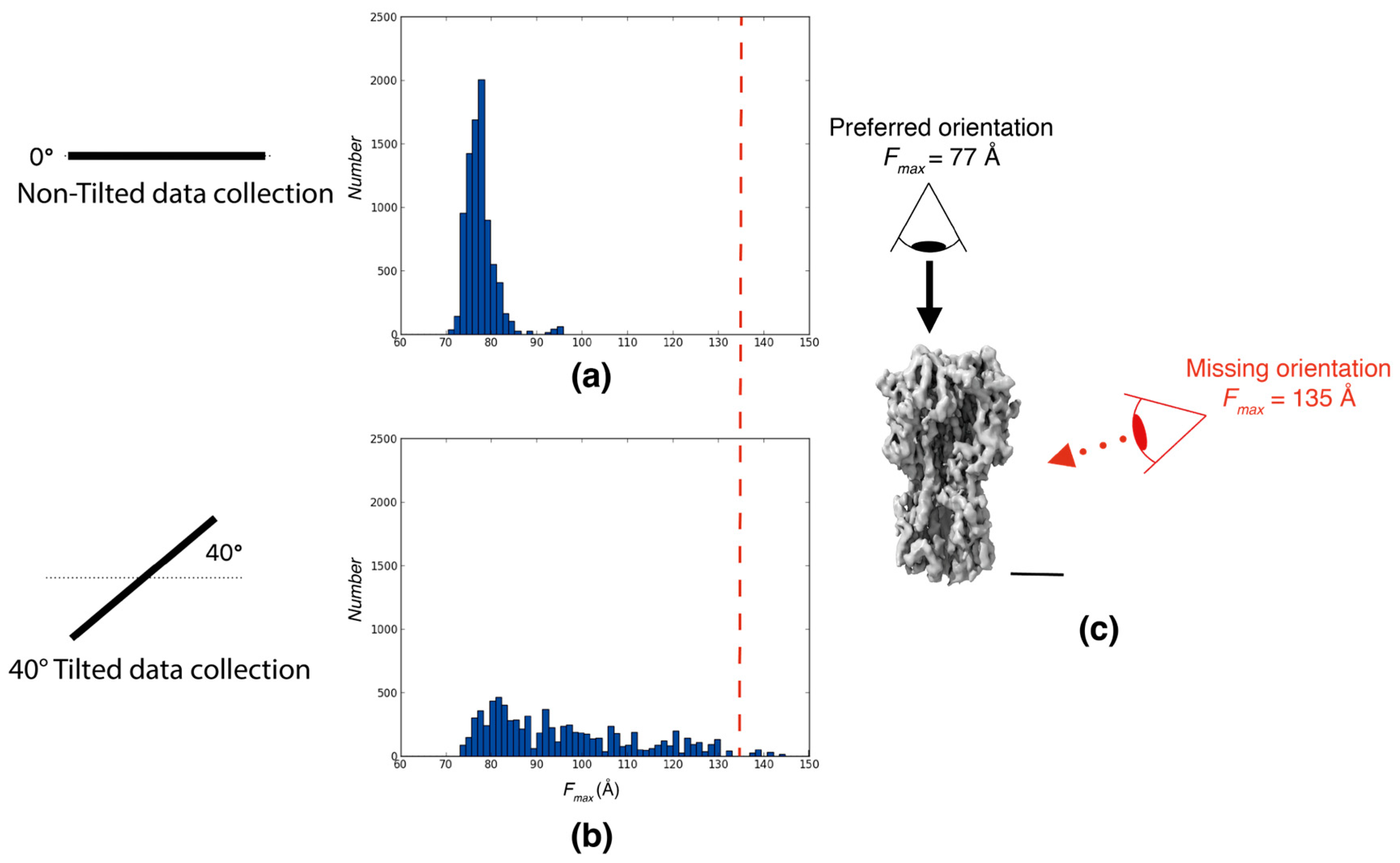
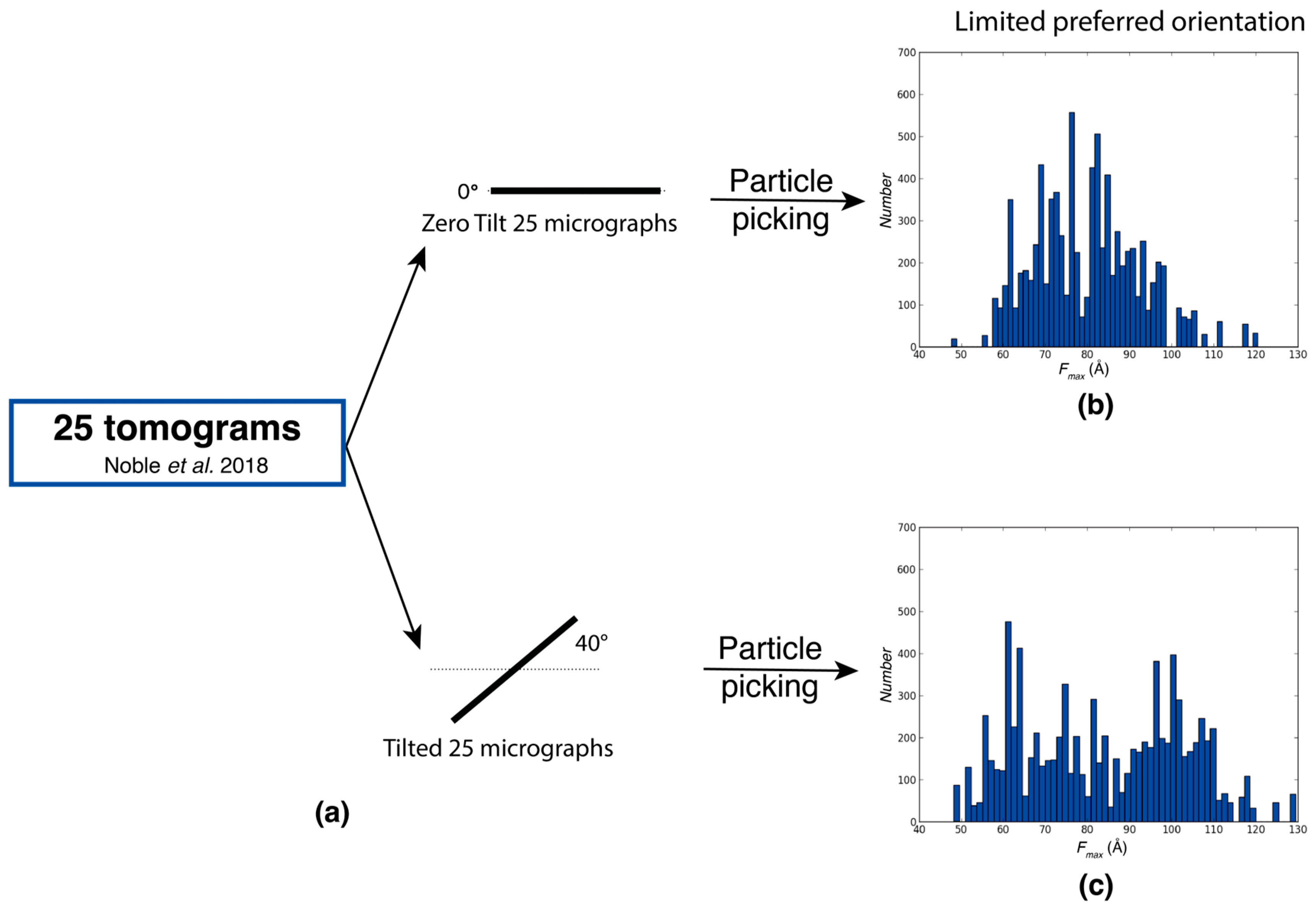

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nottelet, P.; Van Blerkom, P.; Xu, X.-P.; Hanein, D.; Volkmann, N. CryoEM Workflow Acceleration with Feret Signatures. Int. J. Mol. Sci. 2024, 25, 7593. https://doi.org/10.3390/ijms25147593
Nottelet P, Van Blerkom P, Xu X-P, Hanein D, Volkmann N. CryoEM Workflow Acceleration with Feret Signatures. International Journal of Molecular Sciences. 2024; 25(14):7593. https://doi.org/10.3390/ijms25147593
Chicago/Turabian StyleNottelet, Pierre, Peter Van Blerkom, Xiao-Ping Xu, Dorit Hanein, and Niels Volkmann. 2024. "CryoEM Workflow Acceleration with Feret Signatures" International Journal of Molecular Sciences 25, no. 14: 7593. https://doi.org/10.3390/ijms25147593





