Circadian Rhythm and Sleep Analyses in a Fruit Fly Model of Fragile X Syndrome Using a Video-Based Automated Behavioral Research System
Abstract
:1. Introduction
2. Results
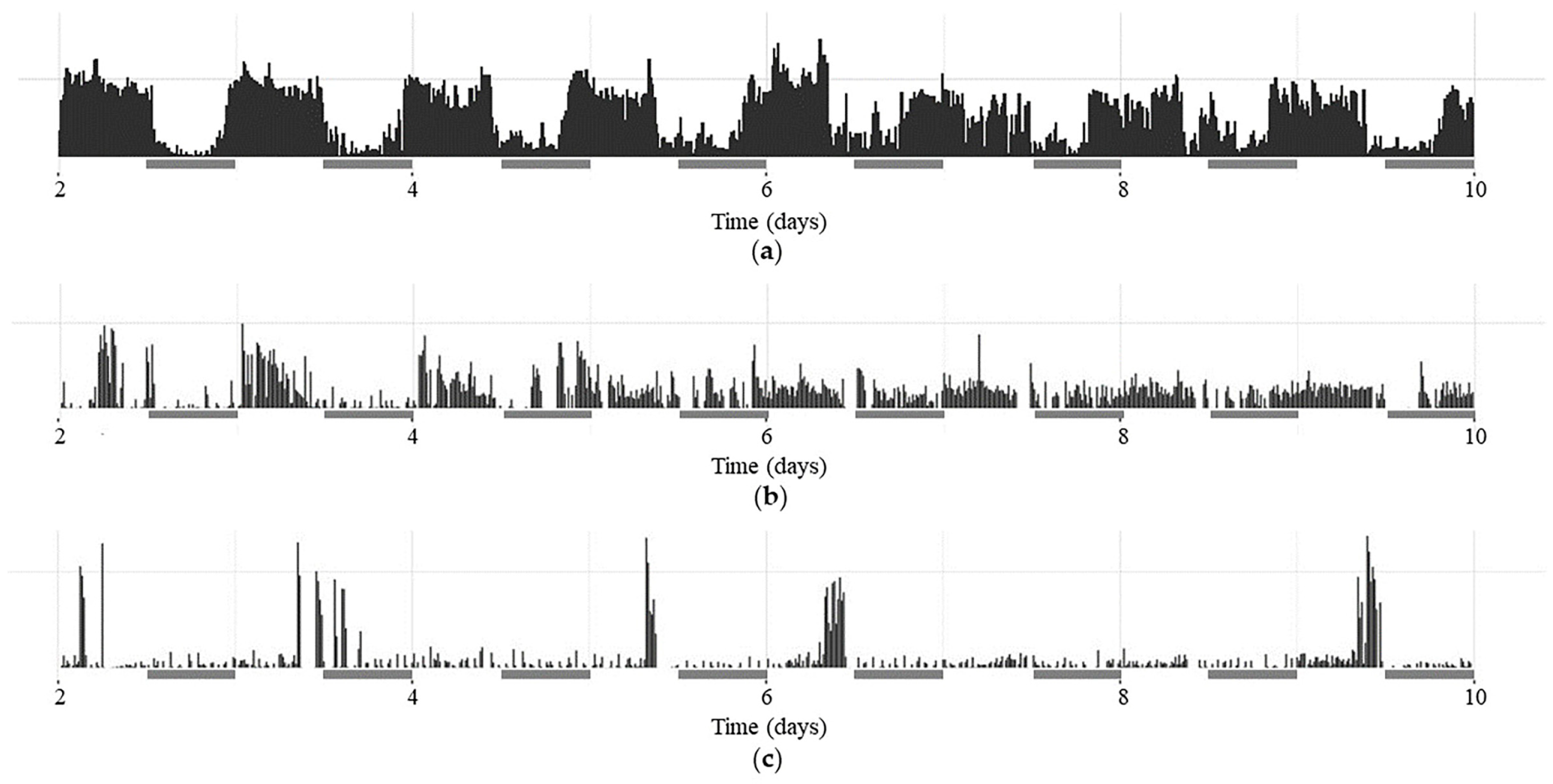
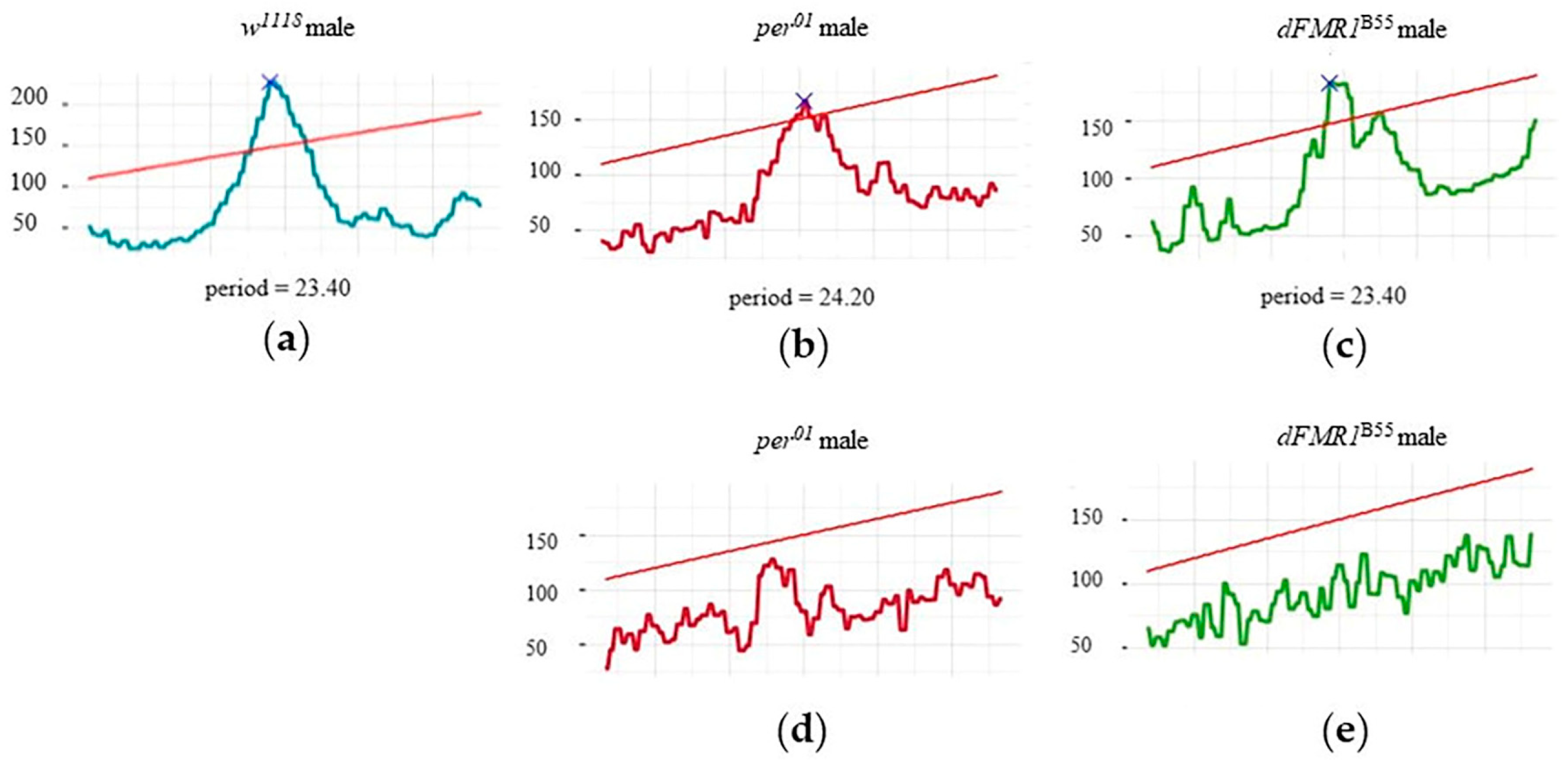
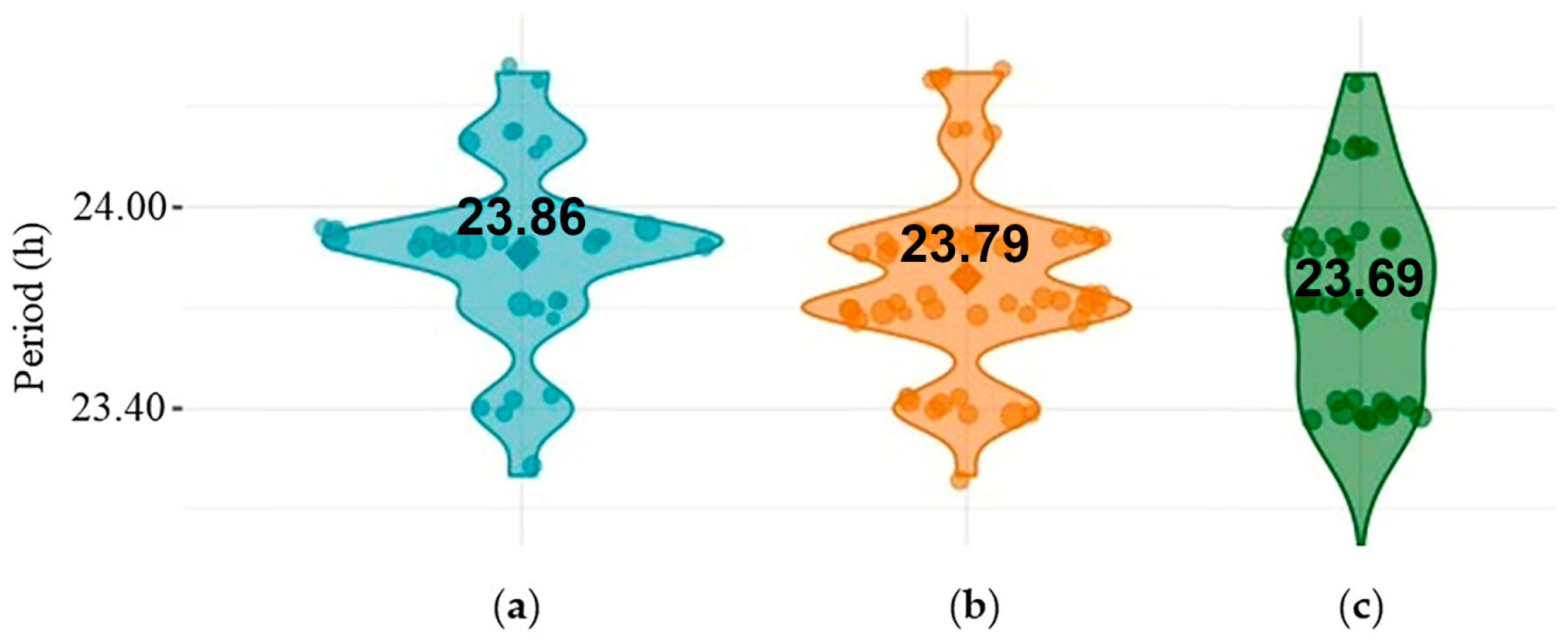
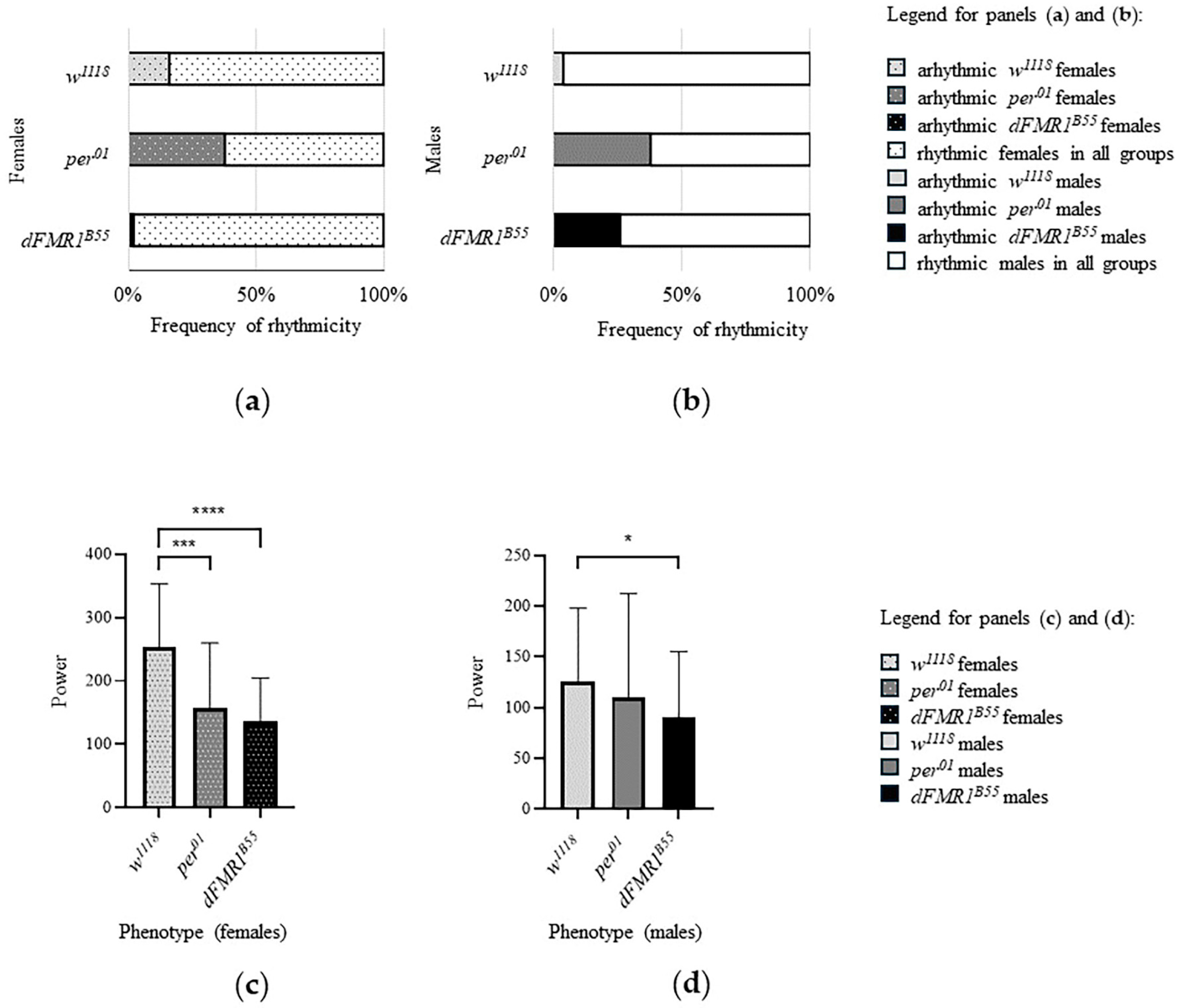
2.1. CR Analysis in the w1118, per01 and dFMR1B55 Groups
2.2. Sleep Analysis in the w1118, per01, and dFMR1B55 Groups
3. Discussion
4. Materials and Methods
4.1. Flies
4.2. Video Monitoring of Locomotor Activity
4.3. CR and Sleep Pattern Analyses
- Number of bouts: number of sleeping episodes;
- Sleep bout length: duration of bout (i.e., sleeping episode);
- Day sleep latency: the duration between ZT00 and first sleeping bout after ZT00;
- Night sleep latency: the duration between ZT12 and first sleeping bout after ZT12;
- Total sleep time (TST): time spent sleeping;
- Wake after sleep onset (WASO: the number of minutes a fly spends awake after having initially fallen asleep and is calculated using the formula: total night time from ZT12 to ZT24 (720 min) − (TST in NT–night sleep latency).
- Sleep fragmentation index (SFI) calculated using the formula: number of bouts/TST.
4.4. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Verkerk, A.J.M.H.; Pieretti, M.; Sutcliffe, J.S.; Fu, Y.-H.; Kuhl, D.P.A.; Pizzuti, A.; Reiner, O.; Richards, S.; Victoria, M.F.; Zhang, F.; et al. Identification of a Gene (FMR-1) Containing a CGG Repeat Coincident with a Breakpoint Cluster Region Exhibiting Length Variation in Fragile X Syndrome. Cell 1991, 65, 905–914. [Google Scholar] [CrossRef]
- Feng, G.; Deák, P.; Chopra, M.; Hall, L.M. Cloning and Functional Analysis of TipE, a Novel Membrane Protein That Enhances Drosophila Para Sodium Channel Function. Cell 1995, 82, 1001–1011. [Google Scholar] [CrossRef]
- Weiler, I.J.; Irwin, S.A.; Klintsova, A.Y.; Spencer, C.M.; Brazelton, A.D.; Miyashiro, K.; Comery, T.A.; Patel, B.; Eberwine, J.; Greenough, W.T. Fragile X Mental Retardation Protein Is Translated near Synapses in Response to Neurotransmitter Activation. Proc. Natl. Acad. Sci. USA 1997, 94, 5395–5400. [Google Scholar] [CrossRef] [PubMed]
- Richter, J.D.; Zhao, X. The Molecular Biology of FMRP: New Insights into Fragile X Syndrome. Nat. Rev. Neurosci. 2021, 22, 209–222. [Google Scholar] [CrossRef] [PubMed]
- Higashimori, H.; Morel, L.; Huth, J.; Lindemann, L.; Dulla, C.; Taylor, A.; Freeman, M.; Yang, Y. Astroglial FMRP-Dependent Translational down-Regulation of mGluR5 Underlies Glutamate Transporter GLT1 Dysregulation in the Fragile X Mouse. Human. Mol. Genet. 2013, 22, 2041–2054. [Google Scholar] [CrossRef]
- Salcedo-Arellano, M.J.; Hagerman, R.J.; Martínez-Cerdeño, V. Fragile X syndrome: Clinical presentation, pathology and treatment. GMM 2023, 156, 3599. [Google Scholar] [CrossRef]
- Wang, H. Developmentally-Programmed FMRP Expression in Oligodendrocytes: A Potential Role of FMRP in Regulating Translation in Oligodendroglia Progenitors. Human. Mol. Genet. 2003, 13, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Tassone, F. Newborn Screening for Fragile X Syndrome. JAMA Neurol. 2014, 71, 355. [Google Scholar] [CrossRef]
- Devitt, N.; Gallagher, L.; Reilly, R. Autism Spectrum Disorder (ASD) and Fragile X Syndrome (FXS): Two Overlapping Disorders Reviewed through Electroencephalography—What Can Be Interpreted from the Available Information? Brain Sci. 2015, 5, 92–117. [Google Scholar] [CrossRef]
- Huddleston, L.B.; Visootsak, J.; Sherman, S.L. Cognitive Aspects of Fragile X Syndrome. WIRES Cogn. Sci. 2014, 5, 501–508. [Google Scholar] [CrossRef]
- Tassone, F.; Hagerman, R.J.; Iklé, D.N.; Dyer, P.N.; Lampe, M.; Willemsen, R.; Oostra, B.A.; Taylor, A.K. FMRP Expression as a Potential Prognostic Indicator in Fragile X Syndrome. Am. J. Med. Genet. 1999, 84, 250–261. [Google Scholar] [CrossRef]
- Kaufmann, W.E.; Kidd, S.A.; Andrews, H.F.; Budimirovic, D.B.; Esler, A.; Haas-Givler, B.; Stackhouse, T.; Riley, C.; Peacock, G.; Sherman, S.L.; et al. Autism Spectrum Disorder in Fragile X Syndrome: Cooccurring Conditions and Current Treatment. Pediatrics 2017, 139, S194–S206. [Google Scholar] [CrossRef] [PubMed]
- Protic, D.D.; Aishworiya, R.; Salcedo-Arellano, M.J.; Tang, S.J.; Milisavljevic, J.; Mitrovic, F.; Hagerman, R.J.; Budimirovic, D.B. Fragile X Syndrome: From Molecular Aspect to Clinical Treatment. IJMS 2022, 23, 1935. [Google Scholar] [CrossRef] [PubMed]
- Thurman, A.J.; McDuffie, A.; Hagerman, R.; Abbeduto, L. Psychiatric Symptoms in Boys with Fragile X Syndrome: A Comparison with Nonsyndromic Autism Spectrum Disorder. Res. Dev. Disabil. 2014, 35, 1072–1086. [Google Scholar] [CrossRef] [PubMed]
- Hagerman, R.J.; Hagerman, P.J. (Eds.) Fragile X Syndrome and Premutation Disorders: New Developments and Treatments; Mac Keith Press: London, UK, 2020; ISBN 978-1-911612-38-4. [Google Scholar]
- Bailey, D.B.; Raspa, M.; Olmsted, M.; Holiday, D.B. Co-occurring Conditions Associated with FMR1 Gene Variations: Findings from a National Parent Survey. Am. J. Med. Genet. Pt. A 2008, 146A, 2060–2069. [Google Scholar] [CrossRef]
- Saldarriaga, W.; Tassone, F.; González-Teshima, L.Y.; Forero-Forero, J.V.; Ayala-Zapata, S.; Hagerman, R. Fragile X Syndrome. Colomb. Med. 2014, 45, 190–198. [Google Scholar] [CrossRef]
- Protic, D.; Salcedo-Arellano, M.J.; Dy, J.B.; Potter, L.A.; Hagerman, R.J. New Targeted Treatments for Fragile X Syndrome. CPR 2019, 15, 251–258. [Google Scholar] [CrossRef]
- Budimirovic, D.B.; Protic, D.D.; Delahunty, C.M.; Andrews, H.F.; Choo, T.; Xu, Q.; Berry-Kravis, E.; Kaufmann, W.E.; FORWARD Consortium. Sleep Problems in Fragile X Syndrome: Cross-sectional Analysis of a Large Clinic-based Cohort. Am. J. Med. Genet. Pt. A 2022, 188, 1029–1039. [Google Scholar] [CrossRef]
- Jenkins, V.K.; Larkin, A.; Thurmond, J. The FlyBase Consortium Using FlyBase: A Database of Drosophila Genes and Genetics. In Drosophila; Dahmann, C., Ed.; Methods in Molecular Biology; Springer: New York, NY, USA, 2022; Volume 2540, pp. 1–34. ISBN 978-1-07-162540-8. [Google Scholar]
- Dahlhaus, R. Of Men and Mice: Modeling the Fragile X Syndrome. Front. Mol. Neurosci. 2018, 11, 41. [Google Scholar] [CrossRef]
- Drozd, M.; Bardoni, B.; Capovilla, M. Modeling Fragile X Syndrome in Drosophila. Front. Mol. Neurosci. 2018, 11, 124. [Google Scholar] [CrossRef] [PubMed]
- Trajković, J.; Makevic, V.; Pesic, M.; Pavković-Lučić, S.; Milojevic, S.; Cvjetkovic, S.; Hagerman, R.; Budimirovic, D.B.; Protic, D. Drosophila Melanogaster as a Model to Study Fragile X-Associated Disorders. Genes 2022, 14, 87. [Google Scholar] [CrossRef] [PubMed]
- Inoue, S.B.; Shimoda, M.; Nishinokubi, I.; Siomi, M.C.; Okamura, M.; Nakamura, A.; Kobayashi, S.; Ishida, N.; Siomi, H. A Role for the Drosophila Fragile X-Related Gene in Circadian Output. Curr. Biol. 2002, 12, 1331–1335. [Google Scholar] [CrossRef] [PubMed]
- Bolduc, F.V.; Valente, D.; Nguyen, A.T.; Mitra, P.P.; Tully, T. An Assay for Social Interaction in Drosophila Fragile X Mutants. Fly 2010, 4, 216–225. [Google Scholar] [CrossRef] [PubMed]
- Dockendorff, T.C.; Su, H.S.; McBride, S.M.J.; Yang, Z.; Choi, C.H.; Siwicki, K.K.; Sehgal, A.; Jongens, T.A. Drosophila Lacking Dfmr1 Activity Show Defects in Circadian Output and Fail to Maintain Courtship Interest. Neuron 2002, 34, 973–984. [Google Scholar] [CrossRef] [PubMed]
- Bushey, D.; Tononi, G.; Cirelli, C. The Drosophila Fragile X Mental Retardation Gene Regulates Sleep Need. J. Neurosci. 2009, 29, 1948–1961. [Google Scholar] [CrossRef] [PubMed]
- Morales, J.; Hiesinger, P.R.; Schroeder, A.J.; Kume, K.; Verstreken, P.; Jackson, F.R.; Nelson, D.L.; Hassan, B.A. Drosophila Fragile X Protein, DFXR, Regulates Neuronal Morphology and Function in the Brain. Neuron 2002, 34, 961–972. [Google Scholar] [CrossRef]
- Sekine, T.; Yamaguchi, T.; Hamano, K.; Siomi, H.; Saez, L.; Ishida, N.; Shimoda, M. Circadian Phenotypes of Drosophila Fragile X Mutants in Alternative Genetic Backgrounds. Zool. Sci. 2008, 25, 561–571. [Google Scholar] [CrossRef] [PubMed]
- Tataroglu, O.; Emery, P. Studying Circadian Rhythms in Drosophila Melanogaster. Methods 2014, 68, 140–150. [Google Scholar] [CrossRef] [PubMed]
- Klarsfeld, A.; Leloup, J.-C.; Rouyer, F. Circadian Rhythms of Locomotor Activity in Drosophila. Behav. Process. 2003, 64, 161–175. [Google Scholar] [CrossRef]
- Shafer, O.T.; Keene, A.C. The Regulation of Drosophila Sleep. Curr. Biol. 2021, 31, R38–R49. [Google Scholar] [CrossRef]
- Dubowy, C.; Sehgal, A. Circadian Rhythms and Sleep in Drosophila Melanogaster. Genetics 2017, 205, 1373–1397. [Google Scholar] [CrossRef] [PubMed]
- Helfrich, C.; Engelmann, W. Evidences for Circadian Rhythmicity in the Per° Mutant of Drosophila Melanogaster. Z. Für Naturforschung C 1987, 42, 1335–1338. [Google Scholar] [CrossRef]
- Helfrich-Förster, C. The Locomotor Activity Rhythm of Drosophila Melanogaster Is Controlled by a Dual Oscillator System. J. Insect Physiol. 2001, 47, 877–887. [Google Scholar] [CrossRef]
- Napolitano, L.M.; Tompkins, L. Neural Control of Homosexual Courtship in Drosophila Melanogaster. J. Neurogenet. 1989, 6, 87–94. [Google Scholar] [CrossRef] [PubMed]
- Takemori, T. Exploring the Genetic Underpinnings of Aggression in Drosophila Melanogaster. J. Undergrad. Neurosci. Educ. 2021, 19, R31–R34. [Google Scholar]
- Li, W.; Wang, Z.; Syed, S.; Lyu, C.; Lincoln, S.; O’Neil, J.; Nguyen, A.D.; Feng, I.; Young, M.W. Chronic Social Isolation Signals Starvation and Reduces Sleep in Drosophila. Nature 2021, 597, 239–244. [Google Scholar] [CrossRef] [PubMed]
- Andrew, D.R.; Moe, M.E.; Chen, D.; Tello, J.A.; Doser, R.L.; Conner, W.E.; Ghuman, J.K.; Restifo, L.L. Spontaneous Motor-Behavior Abnormalities in Two Drosophila Models of Neurodevelopmental Disorders. J. Neurogenet. 2021, 35, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Stojkovic, M.; Petrovic, M.; Capovilla, M.; Milojevic, S.; Makevic, V.; Budimirovic, D.B.; Corscadden, L.; He, S.; Protic, D. Using a Combination of Novel Research Tools to Understand Social Interaction in the Drosophila Melanogaster Model for Fragile X Syndrome. Biology 2024, 13, 432. [Google Scholar] [CrossRef]
- Jagannathan, S.R.; Jeans, T.; Van De Poll, M.N.; Van Swinderen, B. Multivariate Classification of Multichannel Long-Term Electrophysiology Data Identifies Different Sleep Stages in Fruit Flies. Sci. Adv. 2024, 10, eadj4399. [Google Scholar] [CrossRef]
- Keleş, M.F.; Sapci, A.O.B.; Brody, C.; Palmer, I.; Le, C.; Taştan, Ö.; Keleş, S.; Wu, M.N. FlyVISTA, an Integrated Machine Learning Platform for Deep Phenotyping of Sleep in Drosophila. bioRxiv 2023. [Google Scholar] [CrossRef]
- Zimmerman, J.E.; Raizen, D.M.; Maycock, M.H.; Maislin, G.; Pack, A.I. A Video Method to Study Drosophila Sleep. Sleep 2008, 31, 1587–1598. [Google Scholar] [CrossRef] [PubMed]
- Godino-Gimeno, A.; Thörnqvist, P.-O.; Chivite, M.; Míguez, J.M.; Winberg, S.; Cerdá-Reverter, J.M. Obesity Impairs Cognitive Function with No Effects on Anxiety-like Behaviour in Zebrafish. IJMS 2023, 24, 12316. [Google Scholar] [CrossRef] [PubMed]
- Geissmann, Q.; Garcia Rodriguez, L.; Beckwith, E.J.; Gilestro, G.F. Rethomics: An R Framework to Analyse High-Throughput Behavioural Data. PLoS ONE 2019, 14, e0209331. [Google Scholar] [CrossRef]
- Ghosh, A.; Sheeba, V. VANESSA—Shiny Apps for Accelerated Time-Series Analysis and Visualization of Drosophila Circadian Rhythm and Sleep Data. J. Biol. Rhythm. 2022, 37, 222–231. [Google Scholar] [CrossRef] [PubMed]
- Agar, G.; Brown, C.; Sutherland, D.; Coulborn, S.; Oliver, C.; Richards, C. Sleep Disorders in Rare Genetic Syndromes: A Meta-Analysis of Prevalence and Profile. Mol. Autism 2021, 12, 18. [Google Scholar] [CrossRef]
- Kidd, S.A.; Lachiewicz, A.; Barbouth, D.; Blitz, R.K.; Delahunty, C.; McBrien, D.; Visootsak, J.; Berry-Kravis, E. Fragile X Syndrome: A Review of Associated Medical Problems. Pediatrics 2014, 134, 995–1005. [Google Scholar] [CrossRef] [PubMed]
- Kronk, R.; Bishop, E.E.; Raspa, M.; Bickel, J.O.; Mandel, D.A.; Bailey, D.B. Prevalence, Nature, and Correlates of Sleep Problems Among Children with Fragile X Syndrome Based on a Large Scale Parent Survey. Sleep 2010, 33, 679–687. [Google Scholar] [CrossRef] [PubMed]
- Kronk, R.; Dahl, R.; Noll, R. Caregiver Reports of Sleep Problems on a Convenience Sample of Children with Fragile X Syndrome. Am. J. Intellect. Dev. Disabil. 2009, 114, 383–392. [Google Scholar] [CrossRef]
- Richdale, A.L. A Descriptive Analysis of Sleep Behaviour in Children with Fragile X. J. Intellect. Dev. Disabil. 2003, 28, 135–144. [Google Scholar] [CrossRef]
- Symons, F.J.; Byiers, B.J.; Raspa, M.; Bishop, E.; Bailey, D.B. Self-Injurious Behavior and Fragile X Syndrome: Findings from the National Fragile X Survey. Am. J. Intellect. Dev. Disabil. 2010, 115, 473–481. [Google Scholar] [CrossRef]
- Coulson, R.L.; LaSalle, J.M. Epigenetics of Circadian Rhythms in Imprinted Neurodevelopmental Disorders. In Progress. in Molecular Biology and Translational Science; Elsevier: Amsterdam, The Netherlands, 2018; Volume 157, pp. 67–92. ISBN 978-0-12-813565-5. [Google Scholar]
- Allen, C.N.; Nitabach, M.N.; Colwell, C.S. Membrane Currents, Gene Expression, and Circadian Clocks. Cold Spring Harb. Perspect. Biol. 2017, 9, a027714. [Google Scholar] [CrossRef] [PubMed]
- Saré, R.M.; Harkless, L.; Levine, M.; Torossian, A.; Sheeler, C.A.; Smith, C.B. Deficient Sleep in Mouse Models of Fragile X Syndrome. Front. Mol. Neurosci. 2017, 10, 280. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Fang, Z.; Jud, C.; Vansteensel, M.J.; Kaasik, K.; Lee, C.C.; Albrecht, U.; Tamanini, F.; Meijer, J.H.; Oostra, B.A.; et al. Fragile X-Related Proteins Regulate Mammalian Circadian Behavioral Rhythms. Am. J. Human. Genet. 2008, 83, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Rørth, P. A Modular Misexpression Screen in Drosophila Detecting Tissue-Specific Phenotypes. Proc. Natl. Acad. Sci. USA 1996, 93, 12418–12422. [Google Scholar] [CrossRef]
- Simon, A.F.; Chou, M.-T.; Salazar, E.D.; Nicholson, T.; Saini, N.; Metchev, S.; Krantz, D.E. A Simple Assay to Study Social Behavior in Drosophila: Measurement of Social Space within a Group. Genes. Brain Behav. 2012, 11, 243–252. [Google Scholar] [CrossRef]

| Sleep Parameter | Sex | Females | Males | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Phenotype | w1118 | per01 | dFMR1B55 | p Value | w1118 | per01 | dFMR1B55 | p Value | |
| No of bouts | Mean ± SD | 10.18 ± 5.92 | 7.21 ± 7.55 | 16.15 ± 6.83 | 16.11 ± 5.63 | 12.79 ± 8.34 | 20.50 ± 8.68 | ||
| Median [min–max] | 10 [1–29] | 5 [1–32] | 16 [1–31] | <0.0001 | 16 [1–29] | 11 [1–34] | 22 [2–38] | <0.0001 | |
| Sleep bout length (min) | Mean ± SD | 11.97 ± 6.51 | 7.45 ± 2.94 | 16.92 ± 6.63 | 22.97 ± 16.37 | 10.68 ± 6.09 | 18.05 ± 10.40 | ||
| Median [min–max] | 10.0 [5–36.33] | 6.50 [5–18.60] | 16.65 [5–36.18] | <0.0001 | 16.24 [6.63–95.60] | 8.33 [5–44.24] | 16.47 [5.17–60.78] | <0.0001 | |
| Sleep latency (min) | Mean ± SD | 237.45 ± 139.95 | 223.09 ± 226.05 | 147.15 ± 88.87 | 127.64 ± 86.61 | 135,32 ± 104.08 | 89.10 ± 83.37 | ||
| Median [min–max] | 241 [0–666] | 153 [1–1440] | 135.50 [0–435] | <0.0001 | 127.50 [0–489] | 100 [0–509] | 67 [0–453] | <0.0001 | |
| TST (min) | Mean ± SD | 119.25 ± 120.70 | 38.67 ± 70.64 | 270.44 ± 133.37 | 318.26 ± 155.96 | 138.73 ± 150.49 | 357.65 ± 169.94 | ||
| Median [min–max] | 81 [0–647] | 8.50 [0–370] | 300 [0–499] | <0.0001 | 325 [0–617] | 82.50 [0–694] | 414.50 [13–606] | <0.0001 | |
| Sleep Parameter | Sex | Females | Males | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Phenotype | w1118 | per01 | dFMR1B55 | p Value | w1118 | per01 | dFMR1B55 | p Value | |
| No of bouts | Mean ± SD | 21.53 ± 6.91 | 10.18 ± 5.18 | 17.10 ± 8.41 | 23.30 ± 8.76 | 14.34 ± 4.95 | 15.96 ± 10.14 | ||
| Median [min-max] | 22 [6–37] | 9.50 [2–30] | 16 [2–43] | <0.0001 | 23.50 [1–44] | 15 [5–32] | 14 [1–47] | <0.0001 | |
| Sleep bout length (min) | Mean ± SD | 22.37 ± 14.60 | 40.58 ± 27.35 | 37.73 ± 38.03 | 15.63 ± 8.52 | 32.15 ± 16.49 | 73.40 ± 109.02 | ||
| Median [min-max] | 17.19 [5.67–87.71] | 34.03 [6.59–142] | 27.42 [9.67–286.5] | <0.0001 | 12.66 [5–41] | 29.28 [9.09–101.14] | 40.75 [9.26–656] | <0.0001 | |
| Sleep latency (min) | Mean ± SD | 94.20 ± 48.81 | 136.86 ± 51.86 | 109.84 ± 81.30 | 67.98 ± 68.28 | 100.99 ± 49.87 | 78.36 ± 46.62 | ||
| Median [min-max] | 94.50 [4–376] | 134.50 [51–261] | 110 [1–514] | <0.0001 | 54.50 [6–576] | 101.5 [2–230] | 86 [1–179] | <0.0001 | |
| TST (min) | Mean ± SD | 409.81 ± 127.62 | 332.60 ± 146.64 | 447.26 ± 102.53 | 348.66 ± 162.56 | 405.44 ± 136.74 | 520.98 ± 81.72 | ||
| Median [min-max] | 416 [34–618] | 324 [18–630] | 458.50 [57–622] | <0.0001 | 347 [5–638] | 418 [97–699] | 537 [231–672] | <0.0001 | |
| WASO (min) | Mean ± SD | 404.40 ± 161.20 | 524.3 ± 176 | 382.60 ± 152.20 | 439.30 ± 199.10 | 415.5 ± 174.3 | 277.40 ± 83.40 | ||
| Median [min-max] | 392.50 [112–1009] | 541.50 [153–962] | 351 [122–1042] | <0.0001 | 416 [93–1291] | 408 [24–812] | 272.50 [83–538] | <0.0001 | |
| SFI | Mean ± SD | 0.06 ± 0.03 | 0.04 ± 0.03 | 0.04 ± 0.02 | 0.08 ± 0.04 | 0.04 ± 0.02 | 0.03 ± 0.03 | ||
| Median [min-max] | 0.06 [0.01–0.18] | 0.03 [0.01–0.15] | 0.04 [0–0.11] | <0.0001 | 0.08 [0.02–0.2] | 0.04 [0.01–0.11] | 0.02 [0–0.11] | <0.0001 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Milojevic, S.; Ghosh, A.; Makevic, V.; Stojkovic, M.; Capovilla, M.; Tosti, T.; Budimirovic, D.; Protic, D. Circadian Rhythm and Sleep Analyses in a Fruit Fly Model of Fragile X Syndrome Using a Video-Based Automated Behavioral Research System. Int. J. Mol. Sci. 2024, 25, 7949. https://doi.org/10.3390/ijms25147949
Milojevic S, Ghosh A, Makevic V, Stojkovic M, Capovilla M, Tosti T, Budimirovic D, Protic D. Circadian Rhythm and Sleep Analyses in a Fruit Fly Model of Fragile X Syndrome Using a Video-Based Automated Behavioral Research System. International Journal of Molecular Sciences. 2024; 25(14):7949. https://doi.org/10.3390/ijms25147949
Chicago/Turabian StyleMilojevic, Sara, Arijit Ghosh, Vedrana Makevic, Maja Stojkovic, Maria Capovilla, Tomislav Tosti, Dejan Budimirovic, and Dragana Protic. 2024. "Circadian Rhythm and Sleep Analyses in a Fruit Fly Model of Fragile X Syndrome Using a Video-Based Automated Behavioral Research System" International Journal of Molecular Sciences 25, no. 14: 7949. https://doi.org/10.3390/ijms25147949








