Crosstalk among Alternative Polyadenylation, Genetic Variants and Ubiquitin Modification Contribute to Lung Adenocarcinoma Risk
Abstract
:1. Introduction
2. Results
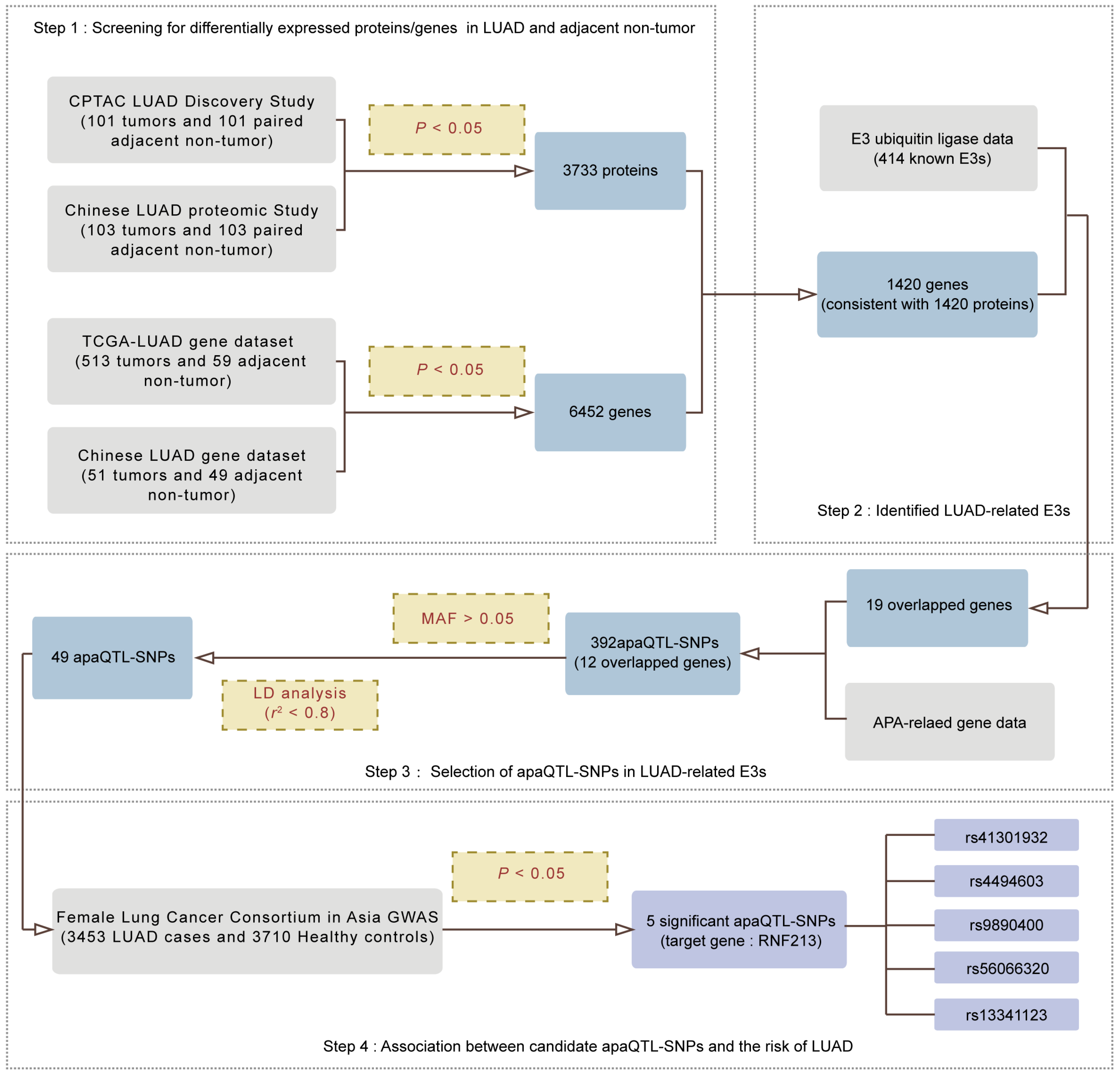
2.1. Screening for APA-Related E3s in LUAD
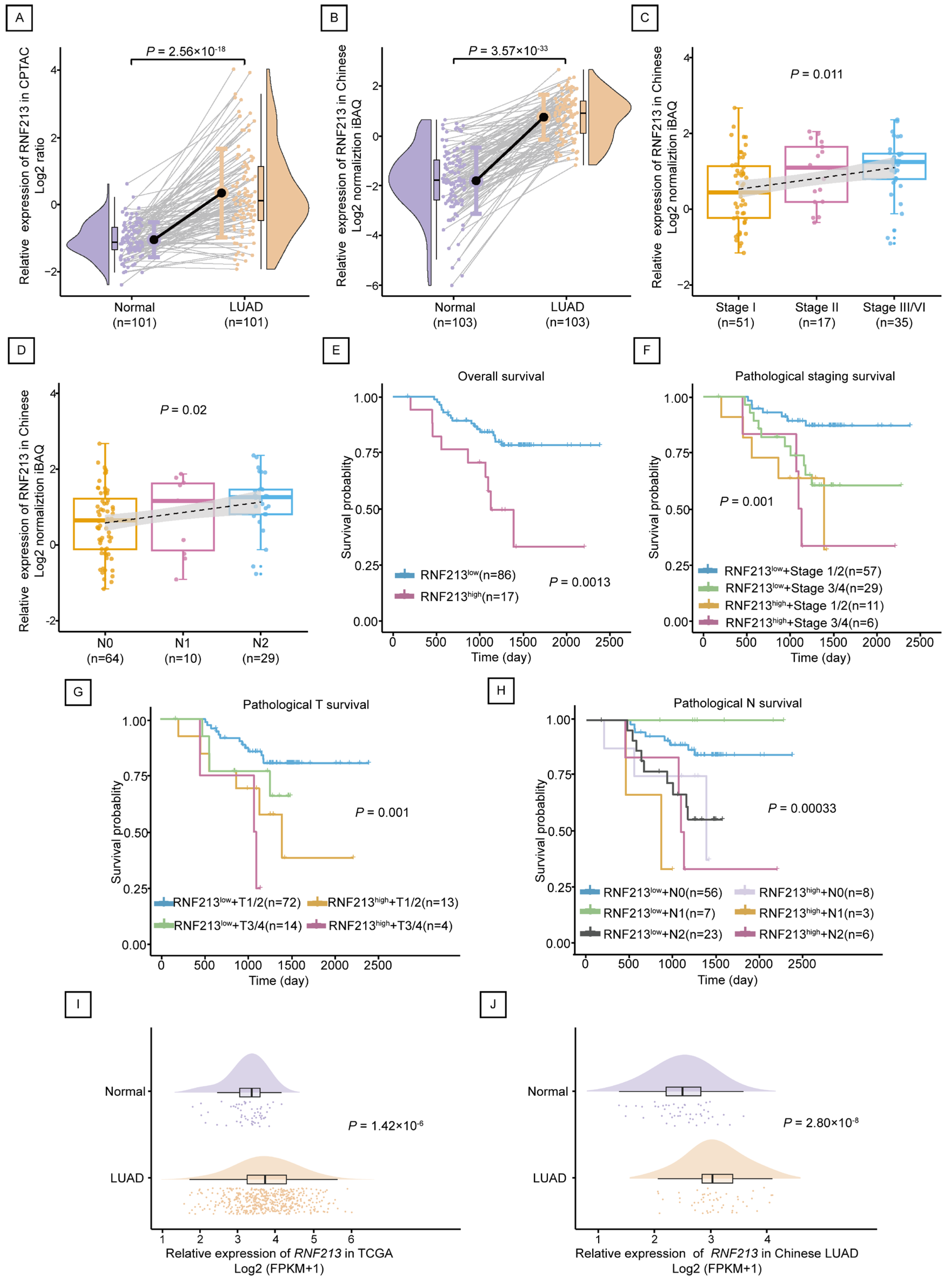
2.2. RNF213 Was Positively Correlated with the Poor Prognosis of LUAD
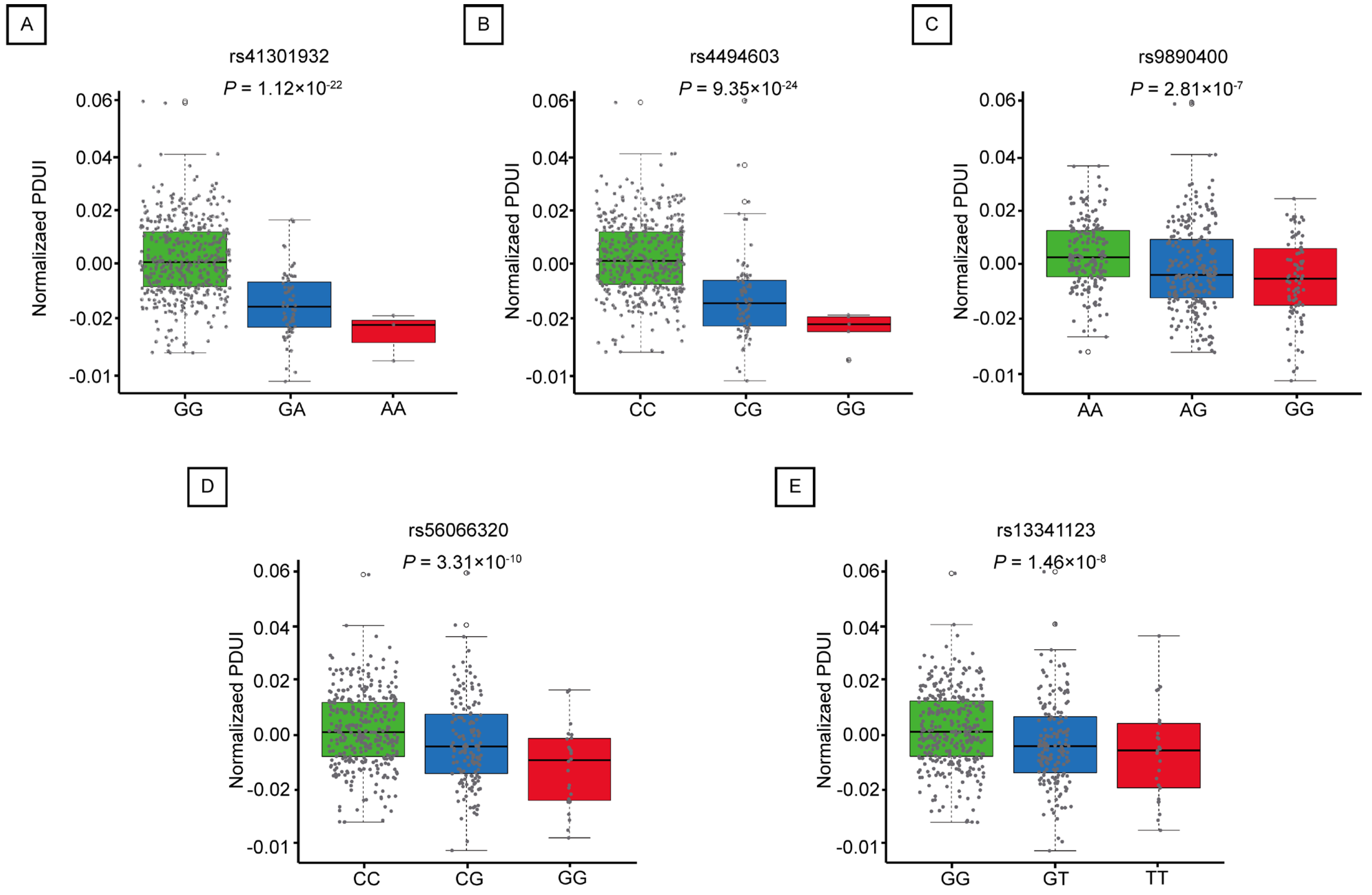
2.3. Mutations in apaQTL-SNPs Regulating RNF213 Are Associated with an Increased Risk of LUAD
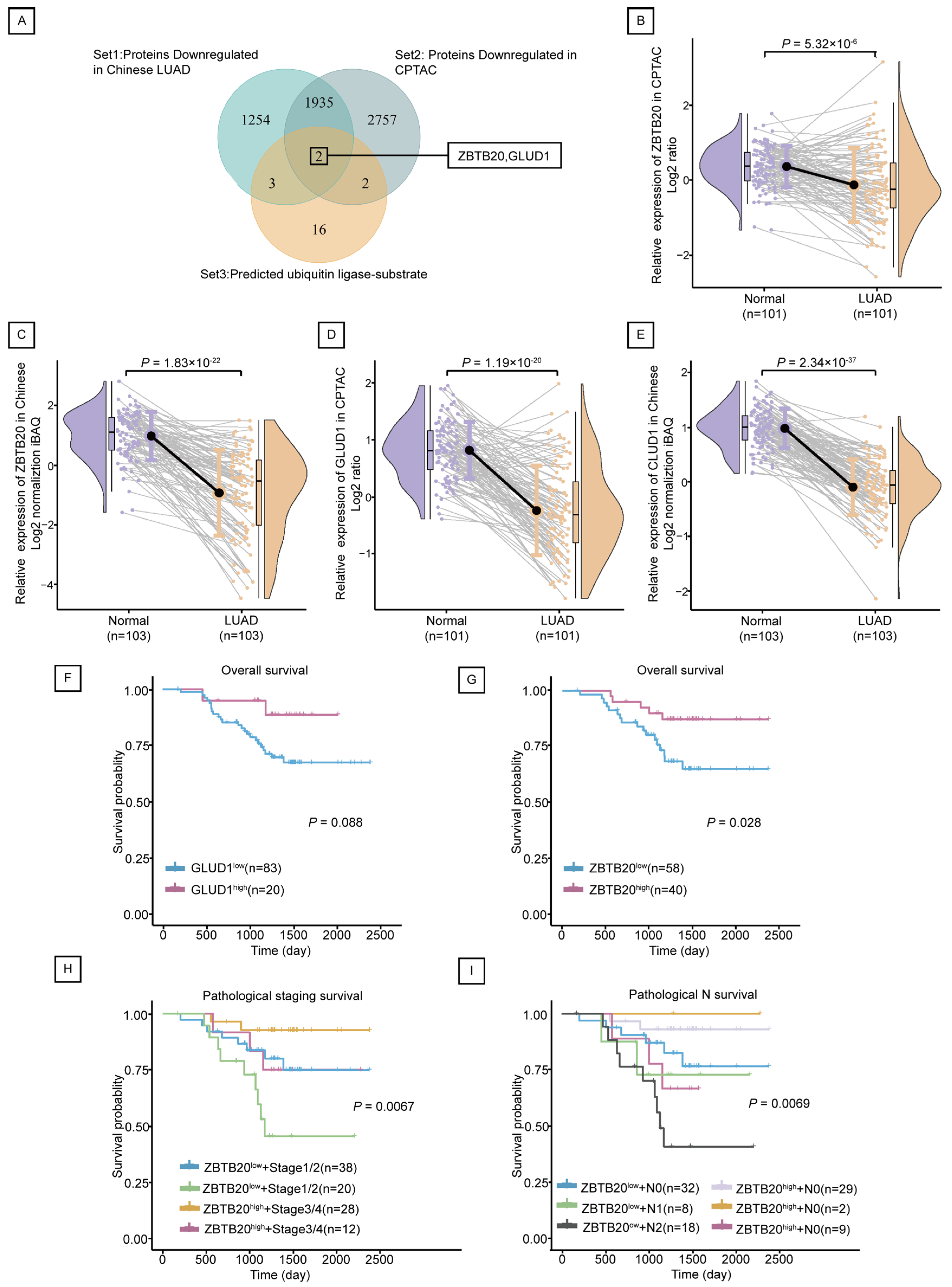
2.4. ZBTB20 Is a Potential Substrate for Ubiquitination by RNF213 in LUAD
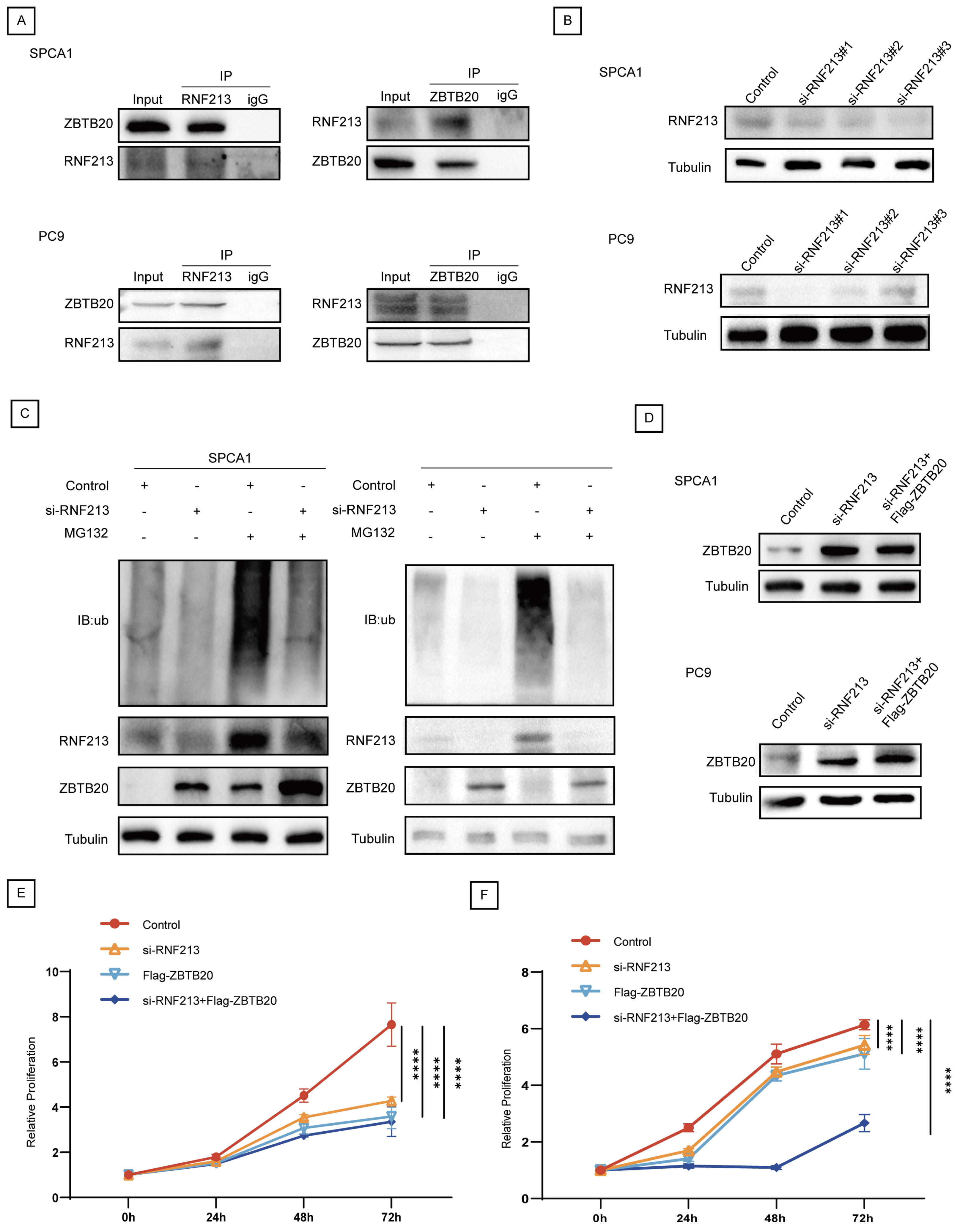
2.5. RNF213 Mediates ZBTB20 Ubiquitination and Degradation to Suppress LUAD Progression
3. Discussion
4. Materials and Methods
4.1. Acquisition and Processing of Proteomics and Genomics Data
4.2. Prediction of Substrate Proteins
4.3. Protein-Protein Docking Assay
4.4. Cell Culture and Transfection
4.5. Western Blot Assay
4.6. Co-Immunoprecipitation (Co-IP) and Ubiquitination Assay
4.7. Cell Counting Kit-8 (CCK8) Assay
4.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Zheng, X.; Tang, H.; Zang, Y.S.; Zeng, C.; Liu, X.; Shen, Y.; Pang, Y.; Wang, S.; Xie, F.; et al. E3 ligase MKRN3 is a tumor suppressor regulating PABPC1 ubiquitination in non-small cell lung cancer. J. Exp. Med. 2021, 218, e20210151. [Google Scholar] [CrossRef] [PubMed]
- Relli, V.; Trerotola, M.; Guerra, E.; Alberti, S. Abandoning the Notion of Non-Small Cell Lung Cancer. Trends Mol. Med. 2019, 25, 585–594. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, Z.; Yu, Z. Identification of a novel glycolysis-related gene signature for predicting metastasis and survival in patients with lung adenocarcinoma. J. Transl. Med. 2019, 17, 423. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Wang, Q.; Lei, Z.; Feng, M.; Zhao, Z.; Wang, Y.; Wei, G. DTL promotes cancer progression by PDCD4 ubiquitin-dependent degradation. J. Exp. Clin. Cancer Res. 2019, 38, 350. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Lu, Y.; Wang, M.; Wu, Y.; Wang, X.; Li, Y. Roles of E3 ubiquitin ligases in gastric cancer carcinogenesis and their effects on cisplatin resistance. J. Mol. Med. 2021, 99, 193–212. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Dai, X.; Jiang, W.; Li, Y.; Wei, W. RBR E3 ubiquitin ligases in tumorigenesis. Semin. Cancer Biol. 2020, 67 Pt 2, 131–144. [Google Scholar] [CrossRef]
- Senft, D.; Qi, J.; Ronai, Z.A. Ubiquitin ligases in oncogenic transformation and cancer therapy. Nat. Rev. Cancer 2018, 18, 69–88. [Google Scholar] [CrossRef] [PubMed]
- Tantai, J.; Pan, X.; Chen, Y.; Shen, Y.; Ji, C. TRIM46 activates AKT/HK2 signaling by modifying PHLPP2 ubiquitylation to promote glycolysis and chemoresistance of lung cancer cells. Cell Death Dis. 2022, 13, 285. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Guo, H.; Li, Q.; Wu, W.; Yu, M.; Zhang, L.; Li, C.; Song, J.; Wang, Z.; Zhang, J.; et al. The AMPK-HOXB9-KRAS axis regulates lung adenocarcinoma growth in response to cellular energy alterations. Cell Rep. 2022, 40, 111210. [Google Scholar] [CrossRef] [PubMed]
- Banh, R.S.; Iorio, C.; Marcotte, R.; Xu, Y.; Cojocari, D.; Rahman, A.A.; Pawling, J.; Zhang, W.; Sinha, A.; Rose, C.M.; et al. PTP1B controls non-mitochondrial oxygen consumption by regulating RNF213 to promote tumour survival during hypoxia. Nat. Cell Biol. 2016, 18, 803–813. [Google Scholar] [CrossRef] [PubMed]
- Otten, E.G.; Werner, E.; Crespillo-Casado, A.; Boyle, K.B.; Dharamdasani, V.; Pathe, C.; Santhanam, B.; Randow, F. Ubiquitylation of lipopolysaccharide by RNF213 during bacterial infection. Nature 2021, 594, 111–116. [Google Scholar] [CrossRef] [PubMed]
- Dai, Z.T.; Xiang, Y.; Wang, Y.; Bao, L.Y.; Wang, J.; Li, J.P.; Zhang, H.M.; Lu, Z.; Ponnambalam, S.; Liao, X.H. Prognostic value of members of NFAT family for pan-cancer and a prediction model based on NFAT2 in bladder cancer. Aging 2021, 13, 13876–13897. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.T.; Wang, Y.H.; Cai, X.Y.; Dong, Y.; Cui, Q.; Zhou, Y.N.; Yang, X.W.; Lu, W.F.; Zhang, M. RNF115 promotes lung adenocarcinoma through Wnt/beta-catenin pathway activation by mediating APC ubiquitination. Cancer Metab. 2021, 9, 7. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Liang, J.; Wen, J.; Luo, M.; Li, J.; Sun, X.; Xu, X.; Li, J.; Wang, D.; Wang, J.; et al. Mutations of RNF213 are responsible for sporadic cerebral cavernous malformation and lead to a mulberry-like cluster in zebrafish. J. Cereb. Blood Flow Metab. 2021, 41, 1251–1263. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xu, W.; Kang, W.; Wong, S.H.; Wang, M.; Zhou, Y.; Fang, X.; Zhang, X.; Yang, H.; Wong, C.H.; et al. Genomic analysis of liver cancer unveils novel driver genes and distinct prognostic features. Theranostics 2018, 8, 1740–1751. [Google Scholar] [CrossRef] [PubMed]
- Tian, H.; Yu, K.; He, L.; Xu, H.; Han, C.; Zhang, X.; Wang, X.; Zhang, X.; Zhang, L.; Gao, G.; et al. RNF213 modulates gamma-herpesvirus infection and reactivation via targeting the viral Replication and Transcription Activator. Proc. Natl. Acad. Sci. USA 2023, 120, e2218825120. [Google Scholar] [CrossRef] [PubMed]
- Shah, A.; Mittleman, B.E.; Gilad, Y.; Li, Y.I. Benchmarking sequencing methods and tools that facilitate the study of alternative polyadenylation. Genome Biol. 2021, 22, 291. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhang, Q.; Miao, Y.R.; Yang, J.; Yang, W.; Yu, F.; Wang, D.; Guo, A.Y.; Gong, J. SNP2APA: A database for evaluating effects of genetic variants on alternative polyadenylation in human cancers. Nucleic Acids Res. 2020, 48, D226–D232. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Xie, Y.; Li, Q.; Zhang, Z.; Chen, J.; Zhang, K.; Xia, X.; Yu, D.; Chen, D.; Yu, Z.; et al. CPSF6-mediated XBP1 3′UTR shortening attenuates cisplatin-induced ER stress and elevates chemo-resistance in lung adenocarcinoma. Drug Resist. Updates 2023, 68, 100933. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Peng, F.; Wang, D.; Li, Y.; Li, J.S.; Li, L.; Li, W. 3′aQTL-atlas: An atlas of 3′UTR alternative polyadenylation quantitative trait loci across human normal tissues. Nucleic Acids Res. 2022, 50, D39–D45. [Google Scholar] [CrossRef]
- Lan, Q.; Hsiung, C.A.; Matsuo, K.; Hong, Y.C.; Seow, A.; Wang, Z.; Hosgood, H.D., 3rd; Chen, K.; Wang, J.C.; Chatterjee, N.; et al. Genome-wide association analysis identifies new lung cancer susceptibility loci in never-smoking women in Asia. Nat. Genet. 2012, 44, 1330–1335. [Google Scholar] [CrossRef] [PubMed]
- Stoyanov, D.; Stoyanov, G.S.; Ivanov, M.N.; Spasov, R.H.; Tonchev, A.B. Transcription Factor Zbtb20 as a Regulator of Malignancy and Its Practical Applications. Int. J. Mol. Sci. 2023, 24, 13763. [Google Scholar] [CrossRef]
- Liu, G.; Zhou, L.; Zhang, H.; Chen, R.; Zhang, Y.; Li, L.; Lu, J.Y.; Jiang, H.; Liu, D.; Qi, S.; et al. Regulation of hepatic lipogenesis by the zinc finger protein Zbtb20. Nat. Commun. 2017, 8, 14824. [Google Scholar] [CrossRef] [PubMed]
- To, J.C.; Chiu, A.P.; Tschida, B.R.; Lo, L.H.; Chiu, C.H.; Li, X.X.; Kuka, T.P.; Linden, M.A.; Amin, K.; Chan, W.C.; et al. ZBTB20 regulates WNT/CTNNB1 signalling pathway by suppressing PPARG during hepatocellular carcinoma tumourigenesis. JHEP Rep. 2021, 3, 100223. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhou, X.; Zhang, M.; Cheng, L.; Zhang, Y.; Wang, X. ZBTB20 promotes cell migration and invasion of gastric cancer by inhibiting IkappaBalpha to induce NF-kappaB activation. Artif. Cells Nanomed. Biotechnol. 2019, 47, 3862–3872. [Google Scholar] [CrossRef] [PubMed]
- Okazaki, S.; Morimoto, T.; Kamatani, Y.; Kamimura, T.; Kobayashi, H.; Harada, K.; Tomita, T.; Higashiyama, A.; Takahashi, J.C.; Nakagawara, J.; et al. Moyamoya Disease Susceptibility Variant RNF213 p.R4810K Increases the Risk of Ischemic Stroke Attributable to Large-Artery Atherosclerosis. Circulation 2019, 139, 295–298. [Google Scholar] [CrossRef] [PubMed]
- Bang, O.Y.; Chung, J.W.; Kim, D.H.; Won, H.H.; Yeon, J.Y.; Ki, C.S.; Shin, H.J.; Kim, J.S.; Hong, S.C.; Kim, D.K.; et al. Moyamoya Disease and Spectrums of RNF213 Vasculopathy. Transl. Stroke Res. 2020, 11, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Pollaci, G.; Gorla, G.; Potenza, A.; Carrozzini, T.; Canavero, I.; Bersano, A.; Gatti, L. Novel Multifaceted Roles for RNF213 Protein. Int. J. Mol. Sci. 2022, 23, 4492. [Google Scholar] [CrossRef] [PubMed]
- Esquela-Kerscher, A.; Slack, F.J. Oncomirs—microRNAs with a role in cancer. Nat. Rev. Cancer 2006, 6, 259–269. [Google Scholar] [CrossRef] [PubMed]
- Mayr, C. What Are 3′ UTRs Doing? Cold Spring Harb. Perspect. Biol. 2019, 11, a034728. [Google Scholar] [CrossRef] [PubMed]
- Gillette, M.A.; Satpathy, S.; Cao, S.; Dhanasekaran, S.M.; Vasaikar, S.V.; Krug, K.; Petralia, F.; Li, Y.; Liang, W.W.; Reva, B.; et al. Proteogenomic Characterization Reveals Therapeutic Vulnerabilities in Lung Adenocarcinoma. Cell 2020, 182, 200–225.e35. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.Y.; Zhang, C.; Wang, X.; Zhai, L.; Ma, Y.; Mao, Y.; Qian, K.; Sun, C.; Liu, Z.; Jiang, S.; et al. Integrative Proteomic Characterization of Human Lung Adenocarcinoma. Cell 2020, 182, 245–261.e17. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Tao, H.; He, J.; Huang, S.Y. The HDOCK server for integrated protein-protein docking. Nat. Protoc. 2020, 15, 1829–1852. [Google Scholar] [CrossRef] [PubMed]


| SNPs | Gene | Location (hg37) | Genotypes | Cases, n (100%) | Controls, n (100%) | Adjusted OR (95%CI) a | p |
|---|---|---|---|---|---|---|---|
| rs41301932 | RNF213 | chr17:78406974 | GG | 3109 (90.04%) | 3413 (92.00%) | 1 (ref) | - |
| GA | 333 (9.64%) | 291 (7.84%) | 1.26 (1.07–1.49) | 0.005 | |||
| AA | 11 (0.32%) | 6 (0.16%) | 2.05 (0.76–5.57) | 0.157 | |||
| Dominant model | 1.28 (1.09–1.51) | 0.003 | |||||
| Additive model | 1.28 (1.09–1.50) | 0.002 | |||||
| rs4494603 | RNF213 | chr17:78408213 | CC | 3056 (88.50%) | 3373 (90.92%) | 1 (ref) | - |
| CG | 385 (11.15%) | 327 (8.81%) | 1.31 (1.12–1.53) | 0.001 | |||
| GG | 12 (0.35%) | 10 (0.27%) | 1.32 (0.57–3.07) | 0.516 | |||
| Dominant model | 1.31 (1.12–1.52) | 0.001 | |||||
| Additive model | 1.29 (1.11–1.49) | 0.001 | |||||
| rs9890400 | RNF213 | chr17:78416493 | AA | 1097 (31.77%) | 1267 (34.15%) | 1 (ref) | - |
| AG | 1641 (47.52%) | 1732 (46.68%) | 1.09 (0.98–1.21) | 0.106 | |||
| GG | 715 (20.71%) | 711 (19.16%) | 1.16 (1.02–1.32) | 0.028 | |||
| Dominant model | 1.11 (1.01–1.23) | 0.037 | |||||
| Additive model | 1.08 (1.01–1.15) | 0.023 | |||||
| rs56066320 | RNF213 | chr17:78418906 | CC | 2942 (85.20%) | 3257 (87.79%) | 1 (ref) | - |
| CG | 493 (14.28%) | 437 (11.78%) | 1.25 (1.09–1.44) | 0.002 | |||
| GG | 18 (0.52%) | 16 (0.43%) | 1.30 (0.66–2.55) | 0.453 | |||
| Dominant model | 1.25 (1.09–1.44) | 0.001 | |||||
| Additive model | 1.23 (1.08–1.40) | 0.001 | |||||
| rs13341123 | RNF213 | chr17:78427033 | GG | 2987 (86.50%) | 3297 (88.87%) | 1 (ref) | - |
| GT | 451 (13.06%) | 400 (10.78%) | 1.24 (1.07–1.43) | 0.003 | |||
| TT | 15 (0.44%) | 13 (0.35%) | 1.14 (0.79–1.66) | 0.486 | |||
| Dominant model | 1.24 (1.08–1.43) | 0.003 | |||||
| Additive model | 1.23 (1.07–1.41) | 0.003 |
| Variables | CPTAC Discovery Study | Chinese LUAD Study | TCGA-LUAD | FLCCA GWAS | ||||
|---|---|---|---|---|---|---|---|---|
| Case (n = 101) | p Value (RNF213) | Case (n = 103) | p Value (RNF213) | Case (n = 515) | p Value (RNF213) | Case (n = 3453) | Control (n = 3710) | |
| Age, n (100%) | 0.357 | 0.632 | 0.958 | |||||
| ≤60 | 41 (40.59%) | 51 (49.51%) | 159 (30.87%) | 1669 (48.33%) | 2013 (54.26%) | |||
| >60 | 60 (59.41%) | 52 (50.49%) | 337 (65.44%) | 1784 (51.67%) | 1697 (45.74%) | |||
| Unknown | 0 | 0 | 19 (3.69%) | 0 | 0 | |||
| Gender | 0.979 | 0.176 | 0.617 | |||||
| Female | 37 (36.63%) | 65 (63.11%) | 276 (53.59%) | |||||
| Male | 64 (63.37%) | 38 (36.89%) | 239 (46.41%) | |||||
| Stage, n (100%) | 0.795 | 0.011 | 0.867 | |||||
| I | 54 (53.47%) | 51 (49.51%) | 276 (53.59%) | |||||
| II | 29 (28.71%) | 17 (16.51%) | 121 (23.50%) | |||||
| III/IV | 18 (17.82%) | 35 (33.98%) | 110 (21.36%) | |||||
| Unknown | 0 | 0 | 8 (1.55%) | |||||
| Pathologic T, n (100%) | 0.306 | 0.672 | 0.581 | |||||
| T1/T2 | 89 (88.12%) | 85 (82.52%) | 446 (86.60%) | |||||
| T3/T4 | 12 (11.88%) | 18 (17.48%) | 66 (12.82%) | |||||
| Unknown | 0 | 0 | 3 (0.58%) | |||||
| Pathologic N, n (100%) | 0.646 | 0.020 | 0.744 | |||||
| N0 | 71 (70.30%) | 64 (62.14%) | 332 (64.47%) | |||||
| N1 | 14 (13.86%) | 10 (9.71%) | 95 (18.45%) | |||||
| N2/N3 | 16 (15.84%) | 29 (28.16%) | 76 (14.76%) | |||||
| Unknown | 0 | 0 | 12 (2.30%) | |||||
| Pathologic M, n (100%) | 0.230 | 0.282 | 0.757 | |||||
| M0 | 83 (82.18) | 101 (98.06%) | 346 (67.18%) | |||||
| M1 | 1 (0.99%) | 2 (1.94%) | 25 (4.85%) | |||||
| Unknown | 17 (16.83%) | 0 (0%) | 144 (27.77%) | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, Y.; Yuan, Y.; Xu, H.; Zhang, W.; Ning, A.; Li, S.; Chen, Q.; Tao, X.; Pan, G.; Tian, T.; et al. Crosstalk among Alternative Polyadenylation, Genetic Variants and Ubiquitin Modification Contribute to Lung Adenocarcinoma Risk. Int. J. Mol. Sci. 2024, 25, 8084. https://doi.org/10.3390/ijms25158084
Wu Y, Yuan Y, Xu H, Zhang W, Ning A, Li S, Chen Q, Tao X, Pan G, Tian T, et al. Crosstalk among Alternative Polyadenylation, Genetic Variants and Ubiquitin Modification Contribute to Lung Adenocarcinoma Risk. International Journal of Molecular Sciences. 2024; 25(15):8084. https://doi.org/10.3390/ijms25158084
Chicago/Turabian StyleWu, Yutong, Yanqiong Yuan, Huiwen Xu, Wendi Zhang, Anhui Ning, Siqi Li, Qiong Chen, Xiaobo Tao, Gongbu Pan, Tian Tian, and et al. 2024. "Crosstalk among Alternative Polyadenylation, Genetic Variants and Ubiquitin Modification Contribute to Lung Adenocarcinoma Risk" International Journal of Molecular Sciences 25, no. 15: 8084. https://doi.org/10.3390/ijms25158084





