Genomic Landscape of Branchio-Oto-Renal Syndrome through Whole-Genome Sequencing: A Single Rare Disease Center Experience in South Korea
Abstract
:1. Introduction
2. Results
2.1. Cohort Description and Clinical Phenotypes
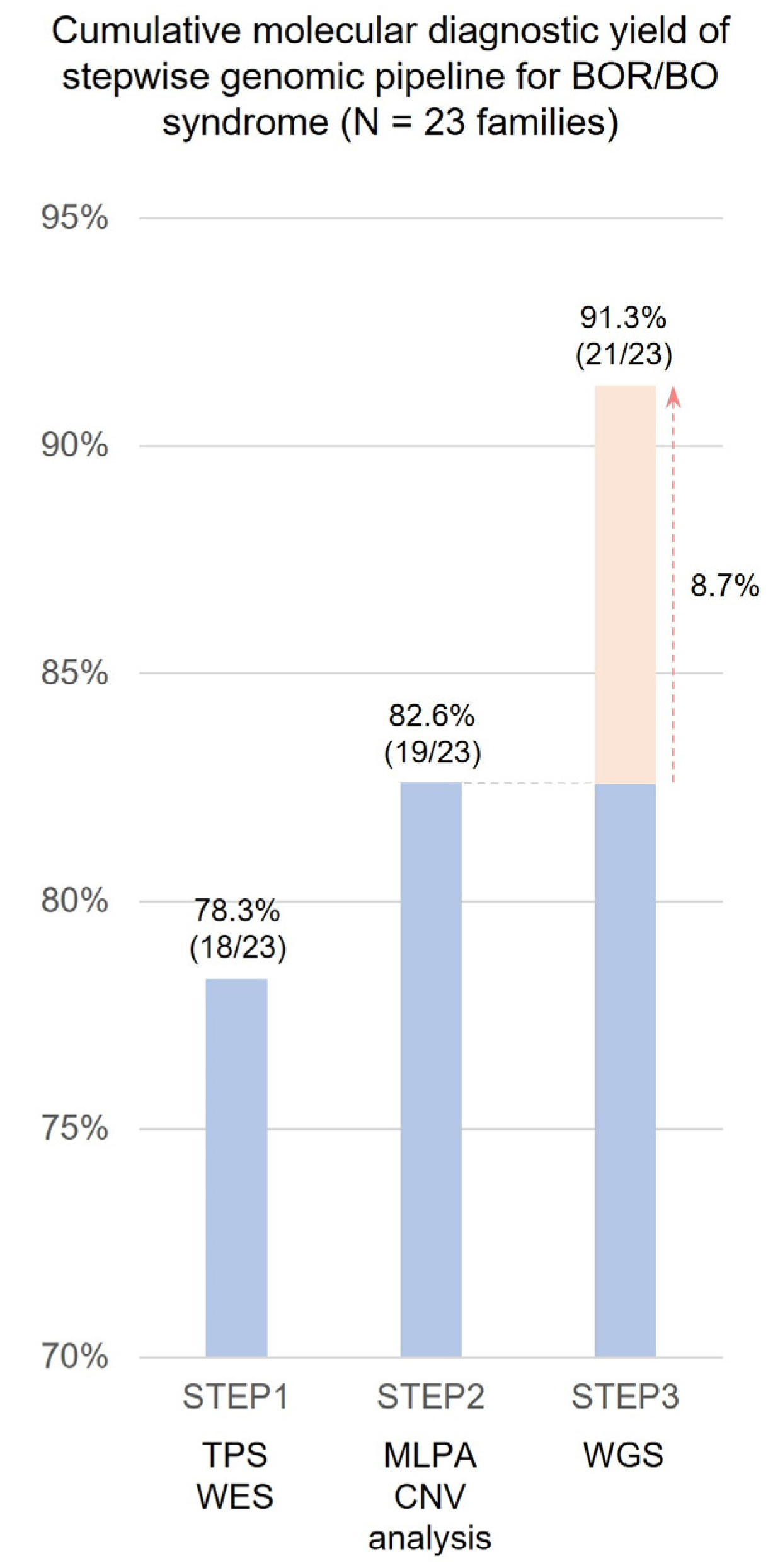
2.2. Stepwise Molecular Diagnostic Yield
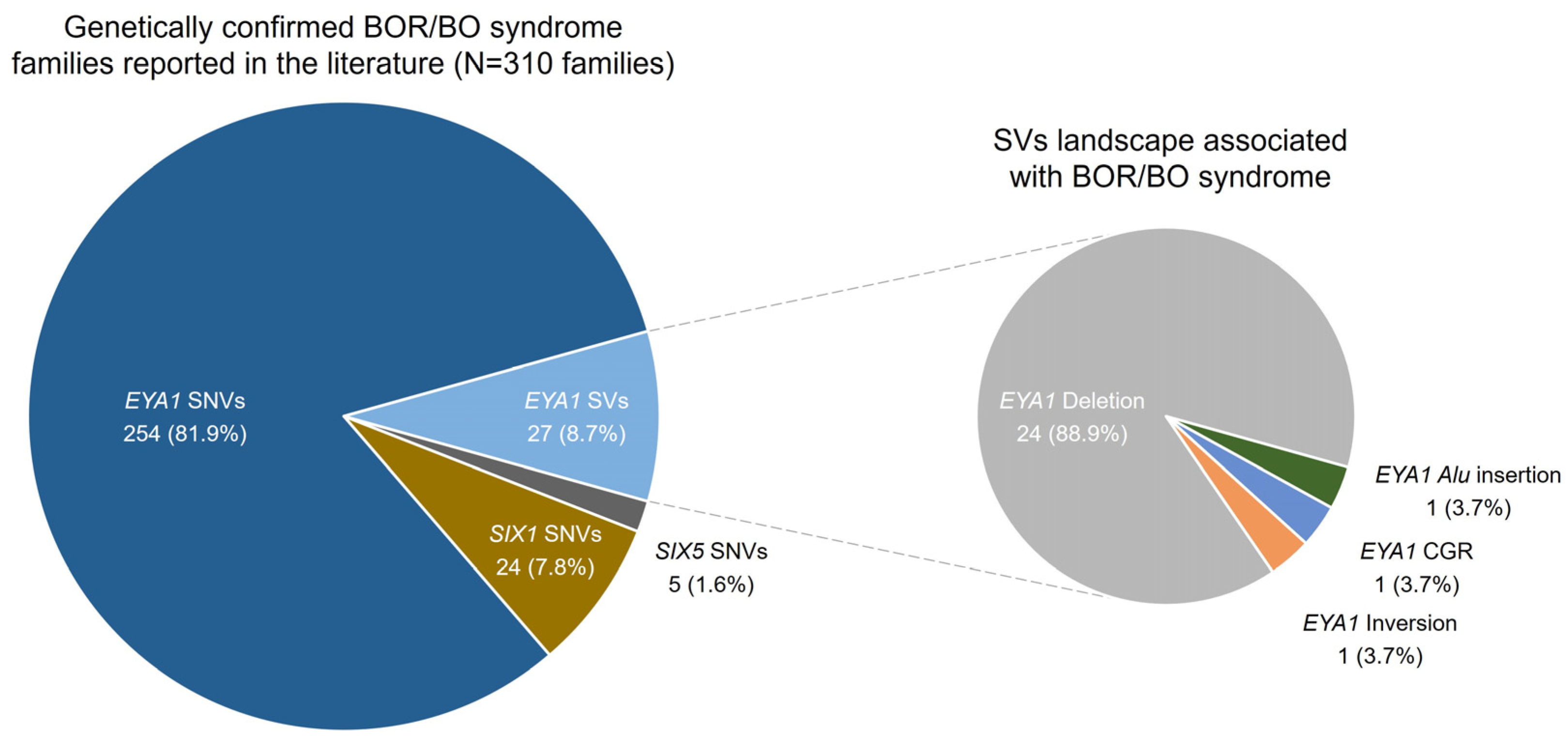
2.3. Genomic Landscape
3. Discussion
| No | PMID | Writer and References | Year | Molecular Diagnostic Yield | Cohort Information | Method | EYA1 | SIX1 | SIX5 | ANKRD11 | SV Annotation |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 9361030 | Sonia Abdelhak et al. [39] | 1997 | 44% (16/36 families) | 36 families clinically diagnosed with BOR syndrome | PCR direct exon sequencing Southern blot | 16 | - | - | - | 1 EYA1 deletion from intron X to exon 16 1 EYA1 deletion from exon 9 to intron IX 1 EYA1 Alu insertion |
| 2 | 10991693 | Sarah Rickard et al. [32] | 2000 | 61% (11/18 families) | 18 families with probable BOR syndrome | PCR direct exon sequencing SSCP analysis | 11 | - | - | - | - |
| 3 | 15146463 | Eugene H. Chang et al. [10] | 2004 | 17% (19/106 families) | 106 families with two or more BOR features | SSCP analysis Bidirectional exon sequencing Semi-quantitative fluorescent multiplex PCR | 19 | - | - | - | 2 entire EYA1 deletions 1 EYA1 deletion from exon 10 to exon 12 |
| 4 | 16491411 | Michiyo Okada et al. [40] | 2006 | 33% (5/15 families) | 15 families with BOR syndrome or BOR-related conditions | PCR direct sequencing RT-PCR | 5 | - | - | - | - |
| 5 | 17637804 | Kirsten Marie Sanggaard et al. [37] | 2007 | 83% (5/6 families) | 6 families clinically diagnosed with BOR syndrome | Marker analysis Linkage analysis MLPA PCR direct exon sequencing | 4 | 1 | - | - | - |
| 6 | 17357085 | Bethan E. Hoskins et al. [13] | 2007 | 5% (5/95 families) | 95 families who met BOR criteria but without EYA1 or SIX1 mutations | PCR direct exon sequencing | - | - | 5 | - | - |
| 7 | 18330911 | Amit Kochhar et al. [33] | 2008 | 4% (10/247 families) | 247 families with BOR syndrome | DHPLC Bidirectional exon sequencing PCR direct exon sequencing | - | 10 | - | - | - |
| 8 | 18220287 | Dana J. Orten et al. [27] | 2008 | 30% (76/248 families) | 248 families with at least one of the major BOR criteria | PCR direct exon sequencing DHPLC | 76 | - | - | - | - |
| 9 | 19206155 | Tracy L. Stockley et al. [41] | 2009 | 82% (14/17 families) | 17 families with a clinical suspicion of BOR syndrome | Bidirectional exon sequencing MLPA | 14 | - | - | - | 1 entire EYA1 deletion 1 EYA1 deletion of exon 9 1 EYA1 deletion from exon 9 to exon 10 |
| 10 | 21280147 | Krug et al. [17] | 2011 | 46% (45/124 families) | 124 families with BOR syndrome | Whole-exome sequencing Multiplex PCR MLPA | 42 | 3 | - | - | - |
| 11 | 22447252 | Shin-Hao Wang et al. [42] | 2012 | 16% (2/12 families) | 12 families who fulfilled the criteria for BOR syndrome | Direct sequencing of EYA1/SIX1 Quantitative PCR | 2 | - | - | - | - |
| 12 | 23840632 | Mee Hyun Song et al. [38] | 2013 | 71% (5/7 families) | 7 families with hearing loss and one or more typical features of BOR syndrome | PCR direct exon sequencing MLPA | 5 | - | - | - | 1 entire EYA1 deletion |
| 13 | 23851940 | Patrick D. Brophy et al. [43] | 2013 | 14% (5/32 families) | 32 BOR probands negative for coding sequence and splice site mutations in known BOR-causing genes | Array-based CGH Long-range PCR | 5 | - | - | - | 1 EYA1 deletion from intron 17 to exon 18 and entire 3′ UTR 4 entire EYA1 deletions |
| 14 | 28583505 | Kyle D. Klingbeil et al. [44] | 2017 | 60% (6/10 families) | 10 families clinically diagnosed with BOR syndrome | Whole-exome sequencing Sanger sequencing | 6 | - | - | - | 2 entire EYA1 deletions |
| 15 | 29500469 | Ai Unzaki et al. [45] | 2018 | 72% (26/36 families) | 36 families clinically diagnosed with BOR syndrome | Direct exon sequencing MLPA Array-based CGH NGS | 22 | 1 | - | - | 1 EYA1 deletion from exon 10 to exon 18 1 EYA1 deletion from exon 2 to exon 3 1 EYA1 deletion from exon 2 to exon 12 1 EYA1 exon 12 deletion 2 EYA1 exon 17 deletions |
| 16 | 31427586 | Michie Ideura et al. [26] | 2019 | 32% (19/59 families) | 59 families clinically diagnosed with BOR/BO syndrome | NGS Array-based CGH | 18 | 1 | - | - | 1 entire EYA1 deletion |
| This study | S. H. Cho et al. | 2024 | 91% (21/23 families) | 23 families with a clinical suspicion of BOR syndrome | MLPA Whole-exome sequencing Whole-genome sequencing | 12 | 8 | - | 1 | 1 EYA1 complex genomic rearrangement 1 EYA1 cryptic inversion 1 entire EYA1 deletion |
4. Materials and Methods
4.1. Participants and Clinical Assessment
4.2. Conventional Genetic Pipeline
4.3. Whole-Genome Sequencing and Bioinformatic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, A.; Francis, M.; Ni, L.; Cremers, C.W.; Kimberling, W.J.; Sato, Y.; Phelps, P.D.; Bellman, S.C.; Wagner, M.J.; Pembrey, M.; et al. Phenotypic manifestations of branchio-oto-renal syndrome. Am. J. Med. Genet. 1995, 58, 365–370. [Google Scholar] [CrossRef]
- Kochhar, A.; Fischer, S.M.; Kimberling, W.J.; Smith, R.J. Branchio-oto-renal syndrome. Am. J. Med. Genet. A 2007, 143A, 1671–1678. [Google Scholar] [CrossRef]
- Vincent, C.; Kalatzis, V.; Abdelhak, S.; Chaib, H.; Compain, S.; Helias, J.; Vaneecloo, F.M.; Petit, C. BOR and BO syndromes are allelic defects of EYA1. Eur. J. Hum. Genet. 1997, 5, 242–246. [Google Scholar] [CrossRef] [PubMed]
- Ruf, R.G.; Berkman, J.; Wolf, M.T.; Nurnberg, P.; Gattas, M.; Ruf, E.M.; Hyland, V.; Kromberg, J.; Glass, I.; Macmillan, J.; et al. A gene locus for branchio-otic syndrome maps to chromosome 14q21.3-q24.3. J. Med. Genet. 2003, 40, 515–519. [Google Scholar] [CrossRef]
- Masuda, M.; Kanno, A.; Nara, K.; Mutai, H.; Morisada, N.; Iijima, K.; Morimoto, N.; Nakano, A.; Sugiuchi, T.; Okamoto, Y.; et al. Phenotype-genotype correlation in patients with typical and atypical branchio-oto-renal syndrome. Sci. Rep. 2022, 12, 969. [Google Scholar] [CrossRef]
- Abdelhak, S.; Kalatzis, V.; Heilig, R.; Compain, S.; Samson, D.; Vincent, C.; Weil, D.; Cruaud, C.; Sahly, I.; Leibovici, M.; et al. A human homologue of the Drosophila eyes absent gene underlies branchio-oto-renal (BOR) syndrome and identifies a novel gene family. Nat. Genet. 1997, 15, 157–164. [Google Scholar] [CrossRef] [PubMed]
- Fraser, F.C.; Sproule, J.R.; Halal, F. Frequency of the branchio-oto-renal (BOR) syndrome in children with profound hearing loss. Am. J. Med. Genet. 1980, 7, 341–349. [Google Scholar] [CrossRef] [PubMed]
- Fraser, F.C.; Ling, D.; Clogg, D.; Nogrady, B. Genetic aspects of the BOR syndrome--branchial fistulas, ear pits, hearing loss, and renal anomalies. Am. J. Med. Genet. 1978, 2, 241–252. [Google Scholar] [CrossRef]
- Fukuda, S.; Kuroda, T.; Chida, E.; Shimizu, R.; Usami, S.; Koda, E.; Abe, S.; Namba, A.; 0 Kitamura, K.; Inuyama, Y. A family affected by branchio-oto syndrome with EYA1 mutations. Auris Nasus Larynx 2001, 28, S7–S11. [Google Scholar] [CrossRef]
- Chang, E.H.; Menezes, M.; Meyer, N.C.; Cucci, R.A.; Vervoort, V.S.; Schwartz, C.E.; Smith, R.J. Branchio-oto-renal syndrome: The mutation spectrum in EYA1 and its phenotypic consequences. Hum. Mutat. 2004, 23, 582–589. [Google Scholar] [CrossRef]
- Feng, H.; Xu, H.; Chen, B.; Sun, S.; Zhai, R.; Zeng, B.; Tang, W.; Lu, W. Genetic and Phenotypic Variability in Chinese Patients With Branchio-Oto-Renal or Branchio-Oto Syndrome. Front. Genet. 2021, 12, 765433. [Google Scholar] [CrossRef] [PubMed]
- Ruf, R.G.; Xu, P.X.; Silvius, D.; Otto, E.A.; Beekmann, F.; Muerb, U.T.; Kumar, S.; Neuhaus, T.J.; Kemper, M.J.; Raymond, R.M., Jr.; et al. SIX1 mutations cause branchio-oto-renal syndrome by disruption of EYA1-SIX1-DNA complexes. Proc. Natl. Acad. Sci. USA 2004, 101, 8090–8095. [Google Scholar] [CrossRef]
- Hoskins, B.E.; Cramer, C.H.; Silvius, D.; Zou, D.; Raymond, R.M.; Orten, D.J.; Kimberling, W.J.; Smith, R.J.; Weil, D.; Petit, C.; et al. Transcription factor SIX5 is mutated in patients with branchio-oto-renal syndrome. Am. J. Hum. Genet. 2007, 80, 800–804. [Google Scholar] [CrossRef] [PubMed]
- Ohto, H.; Kamada, S.; Tago, K.; Tominaga, S.I.; Ozaki, H.; Sato, S.; Kawakami, K. Cooperation of six and eya in activation of their target genes through nuclear translocation of Eya. Mol. Cell Biol. 1999, 19, 6815–6824. [Google Scholar] [CrossRef]
- Lee, S.; Yun, Y.; Cha, J.H.; Han, J.H.; Lee, D.H.; Song, J.J.; Park, M.K.; Lee, J.H.; Oh, S.H.; Choi, B.Y.; et al. Phenotypic and molecular basis of SIX1 variants linked to non-syndromic deafness and atypical branchio-otic syndrome in South Korea. Sci. Rep. 2023, 13, 11776. [Google Scholar] [CrossRef]
- Wang, Y.G.; Sun, S.P.; Qiu, Y.L.; Xing, Q.H.; Lu, W. A novel mutation in EYA1 in a Chinese family with Branchio-oto-renal syndrome. BMC Med. Genet. 2018, 19, 139. [Google Scholar] [CrossRef]
- Krug, P.; Moriniere, V.; Marlin, S.; Koubi, V.; Gabriel, H.D.; Colin, E.; Bonneau, D.; Salomon, R.; Antignac, C.; Heidet, L. Mutation screening of the EYA1, SIX1, and SIX5 genes in a large cohort of patients harboring branchio-oto-renal syndrome calls into question the pathogenic role of SIX5 mutations. Hum. Mutat. 2011, 32, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Jamuar, S.S.; Tan, E.C. Clinical application of next-generation sequencing for Mendelian diseases. Hum. Genom. 2015, 9, 10. [Google Scholar] [CrossRef]
- Bick, D.; Jones, M.; Taylor, S.L.; Taft, R.J.; Belmont, J. Case for genome sequencing in infants and children with rare, undiagnosed or genetic diseases. J. Med. Genet. 2019, 56, 783–791. [Google Scholar] [CrossRef]
- Choy, K.W. Next-Generation Sequencing to Diagnose Suspected Genetic Disorders. N. Engl. J. Med. 2019, 380, 200–201. [Google Scholar] [CrossRef]
- Pajusalu, S.; Kahre, T.; Roomere, H.; Murumets, U.; Roht, L.; Simenson, K.; Reimand, T.; Ounap, K. Large gene panel sequencing in clinical diagnostics-results from 501 consecutive cases. Clin. Genet. 2018, 93, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Aspromonte, M.C.; Bellini, M.; Gasparini, A.; Carraro, M.; Bettella, E.; Polli, R.; Cesca, F.; Bigoni, S.; Boni, S.; Carlet, O.; et al. Characterization of intellectual disability and autism comorbidity through gene panel sequencing. Hum. Mutat. 2020, 41, 1183. [Google Scholar] [CrossRef] [PubMed]
- Pennisi, E. Upstart DNA sequencers could be a ‘game changer’. Science 2022, 376, 1257–1258. [Google Scholar] [CrossRef] [PubMed]
- Brlek, P.; Bulic, L.; Bracic, M.; Projic, P.; Skaro, V.; Shah, N.; Shah, P.; Primorac, D. Implementing Whole Genome Sequencing (WGS) in Clinical Practice: Advantages, Challenges, and Future Perspectives. Cells 2024, 13, 504. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.J.H. Branchiootorenal Spectrum Disorder. In GeneReviews((R)); Adam, M.P., Feldman, J., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Ideura, M.; Nishio, S.Y.; Moteki, H.; Takumi, Y.; Miyagawa, M.; Sato, T.; Kobayashi, Y.; Ohyama, K.; Oda, K.; Matsui, T.; et al. Comprehensive analysis of syndromic hearing loss patients in Japan. Sci. Rep. 2019, 9, 11976. [Google Scholar] [CrossRef]
- Orten, D.J.; Fischer, S.M.; Sorensen, J.L.; Radhakrishna, U.; Cremers, C.W.; Marres, H.A.; Van Camp, G.; Welch, K.O.; Smith, R.J.; Kimberling, W.J. Branchio-oto-renal syndrome (BOR): Novel mutations in the EYA1 gene, and a review of the mutational genetics of BOR. Hum. Mutat. 2008, 29, 537–544. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef]
- Sirmaci, A.; Spiliopoulos, M.; Brancati, F.; Powell, E.; Duman, D.; Abrams, A.; Bademci, G.; Agolini, E.; Guo, S.; Konuk, B.; et al. Mutations in ANKRD11 cause KBG syndrome, characterized by intellectual disability, skeletal malformations, and macrodontia. Am. J. Hum. Genet. 2011, 89, 289–294. [Google Scholar] [CrossRef] [PubMed]
- Wojcik, M.H.; Lemire, G.; Berger, E.; Zaki, M.S.; Wissmann, M.; Win, W.; White, S.M.; Weisburd, B.; Wieczorek, D.; Waddell, L.B.; et al. Genome Sequencing for Diagnosing Rare Diseases. N. Engl. J. Med. 2024, 390, 1985–1997. [Google Scholar] [CrossRef]
- Schobers, G.; Derks, R.; den Ouden, A.; Swinkels, H.; van Reeuwijk, J.; Bosgoed, E.; Lugtenberg, D.; Sun, S.M.; Corominas Galbany, J.; Weiss, M.; et al. Genome sequencing as a generic diagnostic strategy for rare disease. Genome Med. 2024, 16, 32. [Google Scholar] [CrossRef]
- Rickard, S.; Boxer, M.; Trompeter, R.; Bitner-Glindzicz, M. Importance of clinical evaluation and molecular testing in the branchio-oto-renal (BOR) syndrome and overlapping phenotypes. J. Med. Genet. 2000, 37, 623–627. [Google Scholar] [CrossRef] [PubMed]
- Kochhar, A.; Orten, D.J.; Sorensen, J.L.; Fischer, S.M.; Cremers, C.W.; Kimberling, W.J.; Smith, R.J. SIX1 mutation screening in 247 branchio-oto-renal syndrome families: A recurrent missense mutation associated with BOR. Hum. Mutat. 2008, 29, 565. [Google Scholar] [CrossRef] [PubMed]
- Vervoort, V.S.; Smith, R.J.; O’Brien, J.; Schroer, R.; Abbott, A.; Stevenson, R.E.; Schwartz, C.E. Genomic rearrangements of EYA1 account for a large fraction of families with BOR syndrome. Eur. J. Hum. Genet. 2002, 10, 757–766. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Wang, J.; Mitchell, E.; Guo, J.; Wang, L.; Zhang, Y.; Hodge, J.C.; Shen, Y. Recurrent 8q13.2-13.3 microdeletions associated with branchio-oto-renal syndrome are mediated by human endogenous retroviral (HERV) sequence blocks. BMC Med. Genet. 2014, 15, 90. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Valle, A.; Wang, X.; Potocki, L.; Xia, Z.; Kang, S.H.; Carlin, M.E.; Michel, D.; Williams, P.; Cabrera-Meza, G.; Brundage, E.K.; et al. HERV-mediated genomic rearrangement of EYA1 in an individual with branchio-oto-renal syndrome. Am. J. Med. Genet. A 2010, 152A, 2854–2860. [Google Scholar] [CrossRef] [PubMed]
- Sanggaard, K.M.; Rendtorff, N.D.; Kjaer, K.W.; Eiberg, H.; Johnsen, T.; Gimsing, S.; Dyrmose, J.; Nielsen, K.O.; Lage, K.; Tranebjaerg, L. Branchio-oto-renal syndrome: Detection of EYA1 and SIX1 mutations in five out of six Danish families by combining linkage, MLPA and sequencing analyses. Eur. J. Hum. Genet. 2007, 15, 1121–1131. [Google Scholar] [CrossRef] [PubMed]
- Song, M.H.; Kwon, T.J.; Kim, H.R.; Jeon, J.H.; Baek, J.I.; Lee, W.S.; Kim, U.K.; Choi, J.Y. Mutational analysis of EYA1, SIX1 and SIX5 genes and strategies for management of hearing loss in patients with BOR/BO syndrome. PLoS ONE 2013, 8, e67236. [Google Scholar] [CrossRef] [PubMed]
- Abdelhak, S.; Kalatzis, V.; Heilig, R.; Compain, S.; Samson, D.; Vincent, C.; Levi-Acobas, F.; Cruaud, C.; Le Merrer, M.; Mathieu, M.; et al. Clustering of mutations responsible for branchio-oto-renal (BOR) syndrome in the eyes absent homologous region (eyaHR) of EYA1. Hum. Mol. Genet. 1997, 6, 2247–2255. [Google Scholar] [CrossRef]
- Okada, M.; Fujimaru, R.; Morimoto, N.; Satomura, K.; Kaku, Y.; Tsuzuki, K.; Nozu, K.; Okuyama, T.; Iijima, K. EYA1 and SIX1 gene mutations in Japanese patients with branchio-oto-renal (BOR) syndrome and related conditions. Pediatr. Nephrol. 2006, 21, 475–481. [Google Scholar] [CrossRef]
- Stockley, T.L.; Mendoza-Londono, R.; Propst, E.J.; Sodhi, S.; Dupuis, L.; Papsin, B.C. A recurrent EYA1 mutation causing alternative RNA splicing in branchio-oto-renal syndrome: Implications for molecular diagnostics and disease mechanism. Am. J. Med. Genet. A 2009, 149A, 322–327. [Google Scholar] [CrossRef]
- Wang, S.H.; Wu, C.C.; Lu, Y.C.; Lin, Y.H.; Su, Y.N.; Hwu, W.L.; Yu, I.S.; Hsu, C.J. Mutation screening of the EYA1, SIX1, and SIX5 genes in an East Asian cohort with branchio-oto-renal syndrome. Laryngoscope 2012, 122, 1130–1136. [Google Scholar] [CrossRef]
- Brophy, P.D.; Alasti, F.; Darbro, B.W.; Clarke, J.; Nishimura, C.; Cobb, B.; Smith, R.J.; Manak, J.R. Genome-wide copy number variation analysis of a Branchio-oto-renal syndrome cohort identifies a recombination hotspot and implicates new candidate genes. Hum. Genet. 2013, 132, 1339–1350. [Google Scholar] [CrossRef] [PubMed]
- Klingbeil, K.D.; Greenland, C.M.; Arslan, S.; Llamos Paneque, A.; Gurkan, H.; Demir Ulusal, S.; Maroofian, R.; Carrera-Gonzalez, A.; Montufar-Armendariz, S.; Paredes, R.; et al. Novel EYA1 variants causing Branchio-oto-renal syndrome. Int. J. Pediatr. Otorhinolaryngol. 2017, 98, 59–63. [Google Scholar] [CrossRef]
- Unzaki, A.; Morisada, N.; Nozu, K.; Ye, M.J.; Ito, S.; Matsunaga, T.; Ishikura, K.; Ina, S.; Nagatani, K.; Okamoto, T.; et al. Clinically diverse phenotypes and genotypes of patients with branchio-oto-renal syndrome. J. Hum. Genet. 2018, 63, 647–656. [Google Scholar] [CrossRef] [PubMed]
- Bi, Q.; Huang, S.; Wang, H.; Gao, X.; Ma, M.; Han, M.; Lu, S.; Kang, D.; Nourbakhsh, A.; Yan, D.; et al. Preimplantation genetic testing for hereditary hearing loss in Chinese population. J. Assist. Reprod. Genet. 2023, 40, 1721–1732. [Google Scholar] [CrossRef]
- Yi, H.; Yun, Y.; Choi, W.H.; Hwang, H.Y.; Cha, J.H.; Seok, H.; Song, J.J.; Lee, J.H.; Lee, S.Y.; Kim, D. CRISPR-based editing strategies to rectify EYA1 complex genomic rearrangement linked to haploinsufficiency. Mol. Ther. Nucleic Acids 2024, 35, 102199. [Google Scholar] [CrossRef]
- Yun, Y.; Lee, S.Y. Updates on Genetic Hearing Loss: From Diagnosis to Targeted Therapies. J. Audiol. Otol. 2024, 28, 88–92. [Google Scholar] [CrossRef]
- Engels, S.; Kohlhase, J.; McGaughran, J. A SALL1 mutation causes a branchio-oto-renal syndrome-like phenotype. J. Med. Genet. 2000, 37, 458–460. [Google Scholar] [CrossRef]
- Correa-Cerro, L.S.; Kennerknecht, I.; Just, W.; Vogel, W.; Muller, D. The gene for branchio-oculo-facial syndrome does not colocalize to the EYA1-4 genes. J. Med. Genet. 2000, 37, 620–623. [Google Scholar] [CrossRef]
- Trummer, T.; Muller, D.; Schulze, A.; Vogel, W.; Just, W. Branchio-oculo-facial syndrome and branchio-otic/branchio-oto-renal syndromes are distinct entities. J. Med. Genet. 2002, 39, 71–73. [Google Scholar] [CrossRef]
- Lin, A.E.; Semina, E.V.; Daack-Hirsch, S.; Roeder, E.R.; Curry, C.J.; Rosenbaum, K.; Weaver, D.D.; Murray, J.C. Exclusion of the branchio-oto-renal syndrome locus (EYA1) from patients with branchio-oculo-facial syndrome. Am. J. Med. Genet. 2000, 91, 387–390. [Google Scholar] [CrossRef]
- Butcher, D.T.; Cytrynbaum, C.; Turinsky, A.L.; Siu, M.T.; Inbar-Feigenberg, M.; Mendoza-Londono, R.; Chitayat, D.; Walker, S.; Machado, J.; Caluseriu, O.; et al. CHARGE and Kabuki Syndromes: Gene-Specific DNA Methylation Signatures Identify Epigenetic Mechanisms Linking These Clinically Overlapping Conditions. Am. J. Hum. Genet. 2017, 100, 773–788. [Google Scholar] [CrossRef]
- Van der Auwera, G.A.; Carneiro, M.O.; Hartl, C.; Poplin, R.; Del Angel, G.; Levy-Moonshine, A.; Jordan, T.; Shakir, K.; Roazen, D.; Thibault, J.; et al. From FastQ data to high confidence variant calls: The Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 2013, 43, 11.10.1–11.10.33. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Joo, K.; Oh, J.; Han, J.H.; Park, H.R.; Lee, S.; Oh, D.Y.; Woo, S.J.; Choi, B.Y. Severe or Profound Sensorineural Hearing Loss Caused by Novel USH2A Variants in Korea: Potential Genotype-Phenotype Correlation. Clin. Exp. Otorhinolaryngol. 2020, 13, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Kim, M.Y.; Han, J.H.; Park, S.S.; Yun, Y.; Jee, S.C.; Han, J.J.; Lee, J.H.; Seok, H.; Choi, B.Y. Ramifications of POU4F3 variants associated with autosomal dominant hearing loss in various molecular aspects. Sci. Rep. 2023, 13, 12584. [Google Scholar] [CrossRef] [PubMed]
- Jo, H.D.; Han, J.H.; Lee, S.M.; Choi, D.H.; Lee, S.Y.; Choi, B.Y. Genetic Load of Alternations of Transcription Factor Genes in Non-Syndromic Deafness and the Associated Clinical Phenotypes: Experience from Two Tertiary Referral Centers. Biomedicines 2022, 10, 2125. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.H.; Yun, Y.; Lee, D.H.; Cha, J.H.; Lee, S.M.; Lee, J.; Suh, M.H.; Lee, J.H.; Oh, S.H.; Park, M.K.; et al. Novel autosomal dominant TMC1 variants linked to hearing loss: Insight into protein-lipid interactions. BMC Med. Genom. 2023, 16, 320. [Google Scholar] [CrossRef] [PubMed]
- Oza, A.M.; DiStefano, M.T.; Hemphill, S.E.; Cushman, B.J.; Grant, A.R.; Siegert, R.K.; Shen, J.; Chapin, A.; Boczek, N.J.; Schimmenti, L.A.; et al. Expert specification of the ACMG/AMP variant interpretation guidelines for genetic hearing loss. Hum. Mutat. 2018, 39, 1593–1613. [Google Scholar] [CrossRef]
- Talevich, E.; Shain, A.H.; Botton, T.; Bastian, B.C. CNVkit: Genome-Wide Copy Number Detection and Visualization from Targeted DNA Sequencing. PLoS Comput. Biol. 2016, 12, e1004873. [Google Scholar] [CrossRef]
- Krumm, N.; Sudmant, P.H.; Ko, A.; O’Roak, B.J.; Malig, M.; Coe, B.P.; Project, N.E.S.; Quinlan, A.R.; Nickerson, D.A.; Eichler, E.E. Copy number variation detection and genotyping from exome sequence data. Genome Res. 2012, 22, 1525–1532. [Google Scholar] [CrossRef]
- Faust, G.G.; Hall, I.M. SAMBLASTER: Fast duplicate marking and structural variant read extraction. Bioinformatics 2014, 30, 2503–2505. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Scheffler, K.; Halpern, A.L.; Bekritsky, M.A.; Noh, E.; Kallberg, M.; Chen, X.; Kim, Y.; Beyter, D.; Krusche, P.; et al. Strelka2: Fast and accurate calling of germline and somatic variants. Nat. Methods 2018, 15, 591–594. [Google Scholar] [CrossRef] [PubMed]
- Rausch, T.; Zichner, T.; Schlattl, A.; Stutz, A.M.; Benes, V.; Korbel, J.O. DELLY: Structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 2012, 28, i333–i339. [Google Scholar] [CrossRef]
| Family | Sex/Age | Diagnostic Approach * | Gene | Variant [NM/NP No.] | Zygosity/ Inheritance | ACMG Classification # | Affected Domain | Branchial Anomalies | Preauricular Pits | Hearing Loss | Renal Anomalies | EAC Anomalies | Middle Ear Anomalies | Inner Ear Anomalies | Typical/ Atypical |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BOR01 | F/15 | WES | EYA1 | c.1319G>A;p.Arg440Gln [NM_000503.6/NP_000494.2] | Het/ de novo | Pathogenic | ED | O | O | MHL | O | X | O | O | Typical |
| BOR02 | F/22 | WGS | EYA1 | Complex genomic rearrangement g.[71211857_712282326inv;712111857_71215145del] | Het/AD | Pathogenic | N/D | O | O | MHL | X | X | O | O | Typical |
| F/56 | WGS | EYA1 | Complex genomic rearrangement g.[71211857_712282326inv;712111857_71215145del] | Het/AD | Pathogenic | N/D | X | O | MHL | X | O | O | X | Atypical | |
| M/32 | WGS | EYA1 | Complex genomic rearrangement g.[71211857_712282326inv;712111857_71215145del] | Het/AD | Pathogenic | N/D | O | O | SNHL | O | X | X | O | Typical | |
| F/29 | WGS | EYA1 | Complex genomic rearrangement g.[71211857_712282326inv;712111857_71215145del] | Het/AD | Pathogenic | N/D | O | O | SNHL | X | X | X | O | Typical | |
| BOR03 | M/31 | WES | EYA1 | c.1623_1626dup:p.Gln543AsnfsTer90 [NM_000503.6/NP_000494.2] | Het/ de novo | Pathogenic | ED | O | O | SNHL | X | X | X | O | Typical |
| BOR04 | F/0 | WES | EYA1 | c.1598-2A>C:p.? [NM_000503.6/NP_000494.2] | Het/AD | Pathogenic | N/D | O | O | MHL | X | O | O | X | Typical |
| F/30 | WES | EYA1 | c.1598-2A>C:p.? [NM_000503.6/NP_000494.2] | Het/AD | Pathogenic | N/D | O | O | MHL | X | X | O | X | Typical | |
| BOR05 | F/4 | WGS | EYA1 | Cryptic inversion c.49-7047[NC_000008.11:g.71448124]inv | Het/AD | Pathogenic | N/D | O | O | SNHL | X | O | X | O | Typical |
| F/33 | WGS | EYA1 | Cryptic inversion c.49-7047[NC_000008.11:g.71448124]inv | Het/AD | Pathogenic | N/D | O | X | MHL | X | X | O | X | Typical | |
| BOR06 | F/8 | WES | EYA1 | c.1081C>T:p.Arg361Ter [NM_000503.6/NP_000494.2] | Het/AD | Pathogenic | ED | O | O | MHL | X | O | O | O | Typical |
| M/8 | WES | EYA1 | c.1081C>T:p.Arg361Ter [NM_000503.6/NP_000494.2] | Het/AD | Pathogenic | ED | O | O | MHL | X | X | O | O | Typical | |
| BOR07 | M/17 | WES | EYA1 | c.1220G>A:p.Arg407Gln [NM_000503.6/NP_000494.2] | Het / de novo | Pathogenic | ED | O | O | MHL | O | O | O | O | Typical |
| BOR08 | M/6 | WES | EYA1 | c.1276G>A:p.Gly426Ser [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | MHL | O | O | O | X | Typical |
| M/9 | WES | EYA1 | c.1276G>A:p.Gly426Ser [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | X | X | X | X | X | Atypical | |
| BOR09 | F/12 | MLPA | EYA1 | Deletion | Het/AD | Pathogenic | N/D | O | O | MHL | X | O | O | O | Typical |
| BOR10 | M/7 | WES | EYA1 | c.1081C>T:p.Arg361Ter [NM_000503.6/NP_000494.2] | Het/AD | Pathogenic | ED | X | O | MHL | O | O | O | O | Typical |
| BOR11 | F/7 | WES | EYA1 | c.1715G>A:p.Trp572Ter [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | MHL | X | X | O | O | Typical |
| F/12 | WES | EYA1 | c.1715G>A:p.Trp572Ter [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | MHL | X | X | O | X | Typical | |
| BOR12 | M/31 | WES | EYA1 | c.802C>T:p.Gln268Ter [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | MHL | X | X | O | X | Typical |
| M/28 | WES | EYA1 | c.802C>T:p.Gln268Ter [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | MHL | X | X | O | X | Typical | |
| M/64 | WES | EYA1 | c.802C>T:p.Gln268Ter [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | SNHL | X | X | X | O | Typical | |
| F/56 | WES | EYA1 | c.802C>T:p.Gln268Ter [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | MHL | X | X | O | O | Typical | |
| F/49 | WES | EYA1 | c.802C>T:p.Gln268Ter [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | MHL | X | X | O | X | Typical | |
| F/20 | WES | EYA1 | c.802C>T:p.Gln268Ter [NM_000503.6/NP_000494.2] | Het/AD | Likely Pathogenic | ED | O | O | MHL | X | X | O | X | Typical | |
| BOR13 | F/31 | WES | SIX1 | c.501G>C:p.Gln167His [NM_005982.4/NP_005973.1] | Het/AD | Likely Pathogenic | HD | O | X | SNHL | X | X | X | X | Atypical |
| F/60 | WES | SIX1 | c.501G>C:p.Gln167His [NM_005982.4/NP_005973.1] | Het/AD | Likely Pathogenic | HD | X | X | SNHL | X | X | X | X | Atypical | |
| F/30 | WES | SIX1 | c.501G>C:p.Gln167His [NM_005982.4/NP_005973.1] | Het/AD | Likely Pathogenic | HD | X | X | SNHL | X | X | X | X | Atypical | |
| BOR14 | F/10 | WES | SIX1 | c.386_391del:p.Tyr129_Cys130del [NM_005982.4/NP_005973.1] | Het/AD | Pathogenic | HD | X | O | SNHL | X | X | X | X | Atypical |
| F/40 | WES | SIX1 | c.386_391del:p.Tyr129_Cys130del [NM_005982.4/NP_005973.1] | Het/AD | Pathogenic | HD | X | O | SNHL | X | X | X | X | Atypical | |
| BOR15 | F/11 | WES | SIX1 | c.397_399del:p.Glu133del [NM_005982.4/NP_005973.1] | Het/ de novo | Likely Pathogenic | HD | X | O | SNHL | X | X | X | O | Atypical |
| BOR16 | M/76 | WES | SIX1 | c.21del:p.Phe7LeufsTer82 [NM_005982.4/NP_005973.1] | Het/AD | Pathogenic | SD | X | O | SNHL | X | X | X | X | Atypical |
| BOR17 | M/10 | WES | SIX1 | c.386A>C:p.Tyr129Ser [NM_005982.4/NP_005973.1] | Het/AD | Likely Pathogenic | HD | X | X | SNHL | X | X | X | X | Atypical |
| M/44 | WES | SIX1 | c.386A>C:p.Tyr129Ser [NM_005982.4/NP_005973.1] | Het/AD | Likely Pathogenic | HD | X | X | MHL | X | X | O | X | Atypical | |
| BOR18 | M/22 | WES | SIX1 | c.176A>C:p.His59Pro [NM_005982.4/NP_005973.1] | Het/AD | Likely Pathogenic | SD | X | O | SNHL | X | X | X | X | Atypical |
| BOR19 | M/16 | WES | SIX1 | c.513G>T:p.Trp171Cys [NM_005982.4/NP_005973.1] | Het/ de novo | Pathogenic | HD | O | O | CHL | X | X | X | X | Typical |
| BOR20 | M/1 | WES | SIX1 | c.376_378del:p.Glu126del [NM_005982.4/NP_005973.1] | Het/AD | Likely Pathogenic | HD | X | O | SNHL | X | X | X | X | Atypical |
| M/1 | WES | SIX1 | c.376_378del:p.Glu126del [NM_005982.4/NP_005973.1] | Het/AD | Likely Pathogenic | HD | X | X | SNHL | X | X | X | X | Atypical | |
| BOR21 | F/1 | WES | ANKRD11 | c.2409_2412del:p.Glu805ArgfsTer57 [NM_013275.6/NP_037407.4] | Het/AD | Pathogenic | Linker region | O | O | SNHL | X | X | X | X | Typical |
| BOR22 | M/13 | WGS | Negative | - | - | - | - | O | O | MHL | O | X | O | O | Typical |
| BOR23 | F/51 | WES | Negative | - | - | - | - | X | O | MHL | X | X | O | X | N/D |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cho, S.H.; Jeong, S.H.; Choi, W.H.; Lee, S.-Y. Genomic Landscape of Branchio-Oto-Renal Syndrome through Whole-Genome Sequencing: A Single Rare Disease Center Experience in South Korea. Int. J. Mol. Sci. 2024, 25, 8149. https://doi.org/10.3390/ijms25158149
Cho SH, Jeong SH, Choi WH, Lee S-Y. Genomic Landscape of Branchio-Oto-Renal Syndrome through Whole-Genome Sequencing: A Single Rare Disease Center Experience in South Korea. International Journal of Molecular Sciences. 2024; 25(15):8149. https://doi.org/10.3390/ijms25158149
Chicago/Turabian StyleCho, Sung Ho, Sung Ho Jeong, Won Hoon Choi, and Sang-Yeon Lee. 2024. "Genomic Landscape of Branchio-Oto-Renal Syndrome through Whole-Genome Sequencing: A Single Rare Disease Center Experience in South Korea" International Journal of Molecular Sciences 25, no. 15: 8149. https://doi.org/10.3390/ijms25158149






