miR156-SPL and miR169-NF-YA Modules Regulate the Induction of Somatic Embryogenesis in Arabidopsis via LEC- and Auxin-Related Pathways
Abstract
:1. Introduction
2. Results
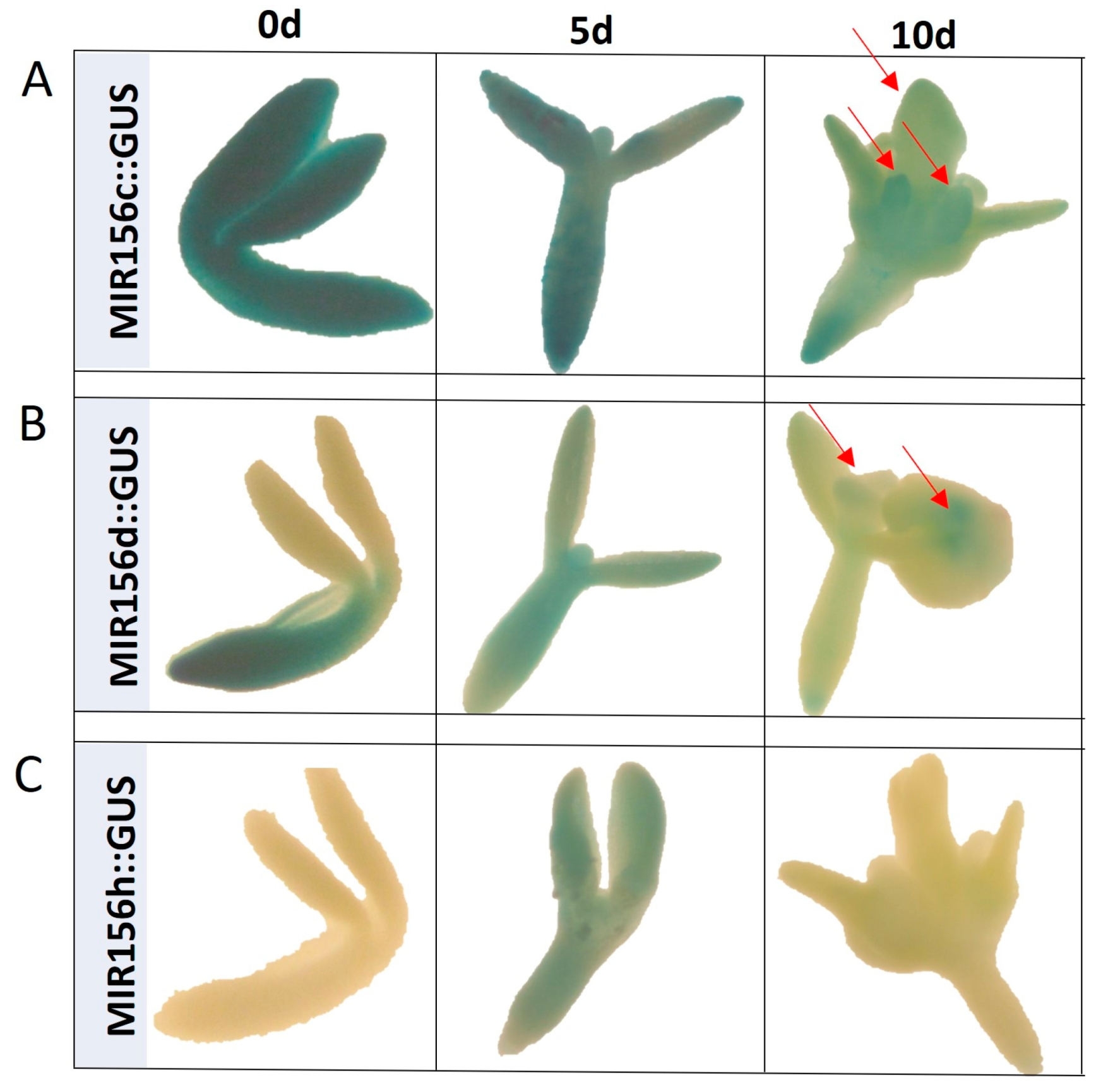
2.1. Different MIRNA156 Genes Regulate SE Induction
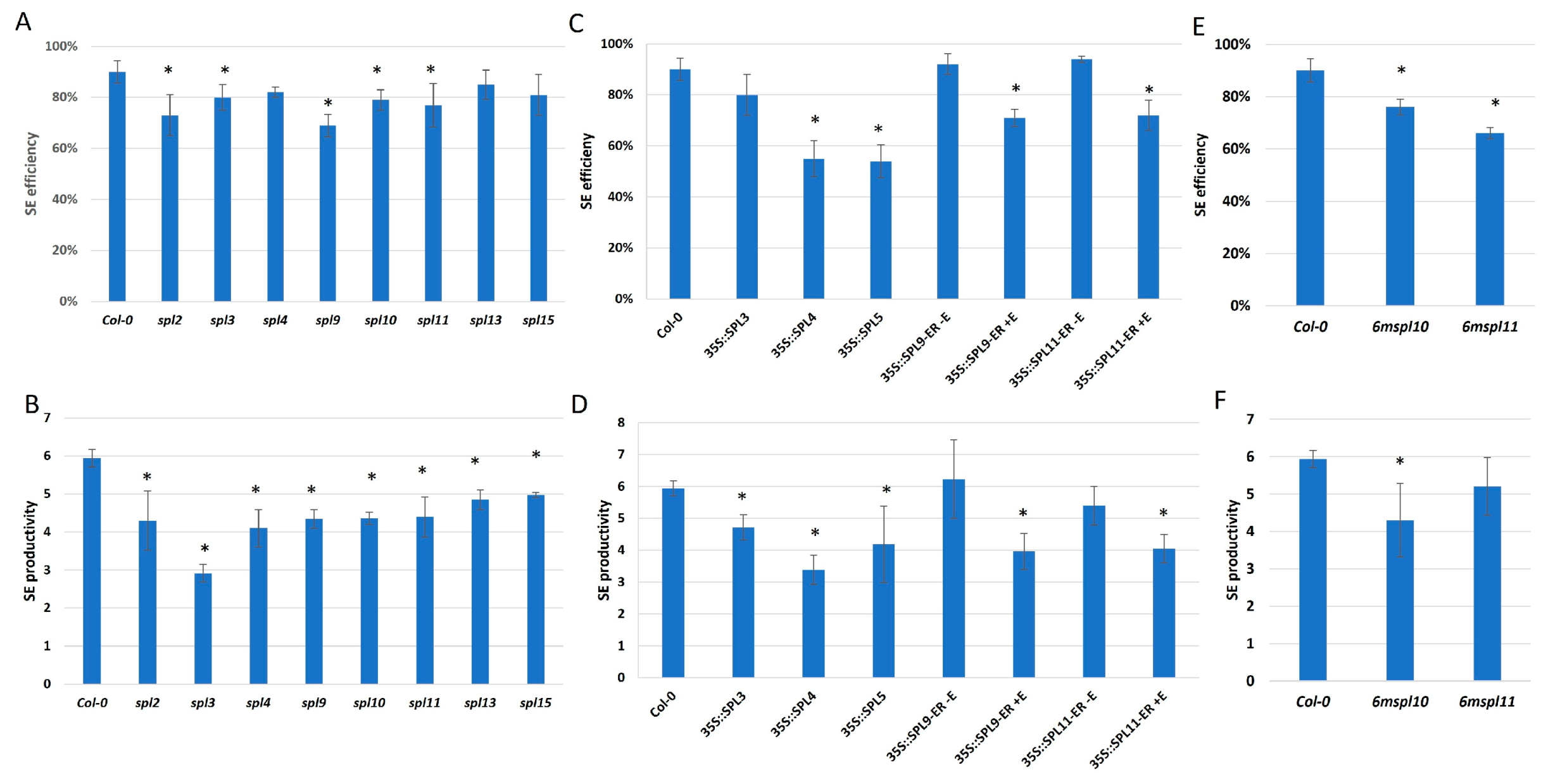
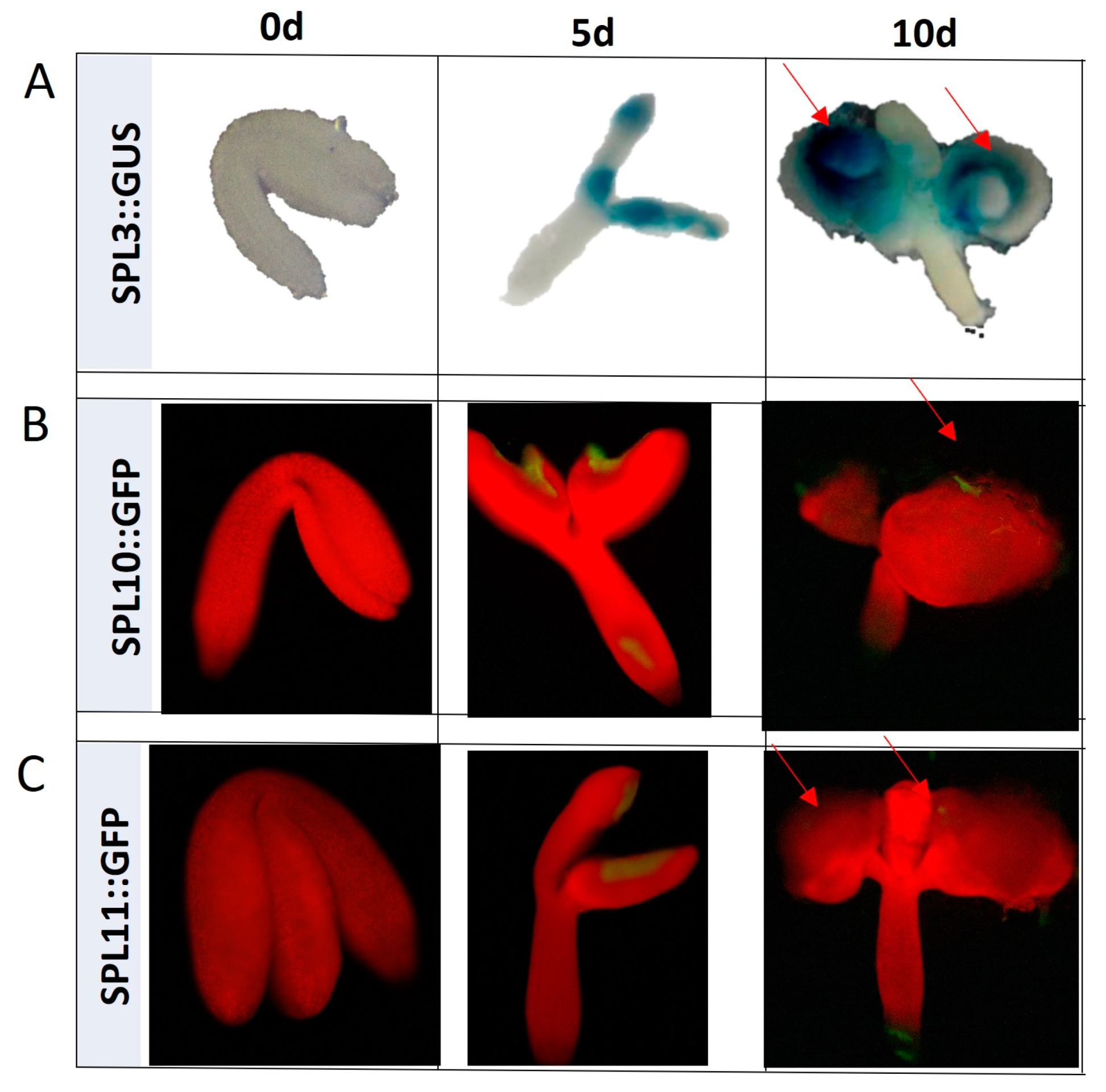
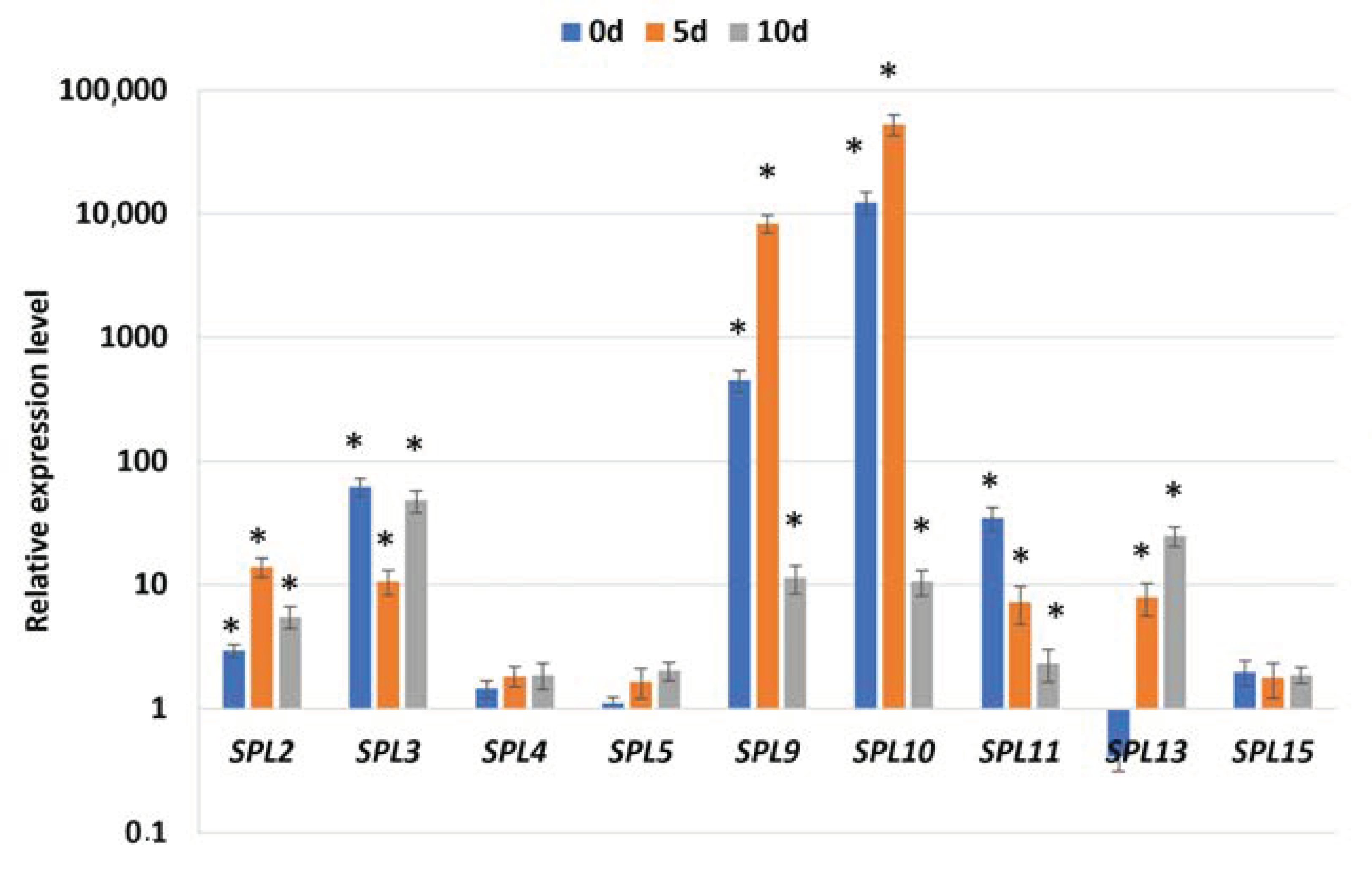
2.2. miR156 Regulates SE Induction via Controlling SPLs
2.3. miR156 via SPL11 Might Regulate LEC Genes in SE
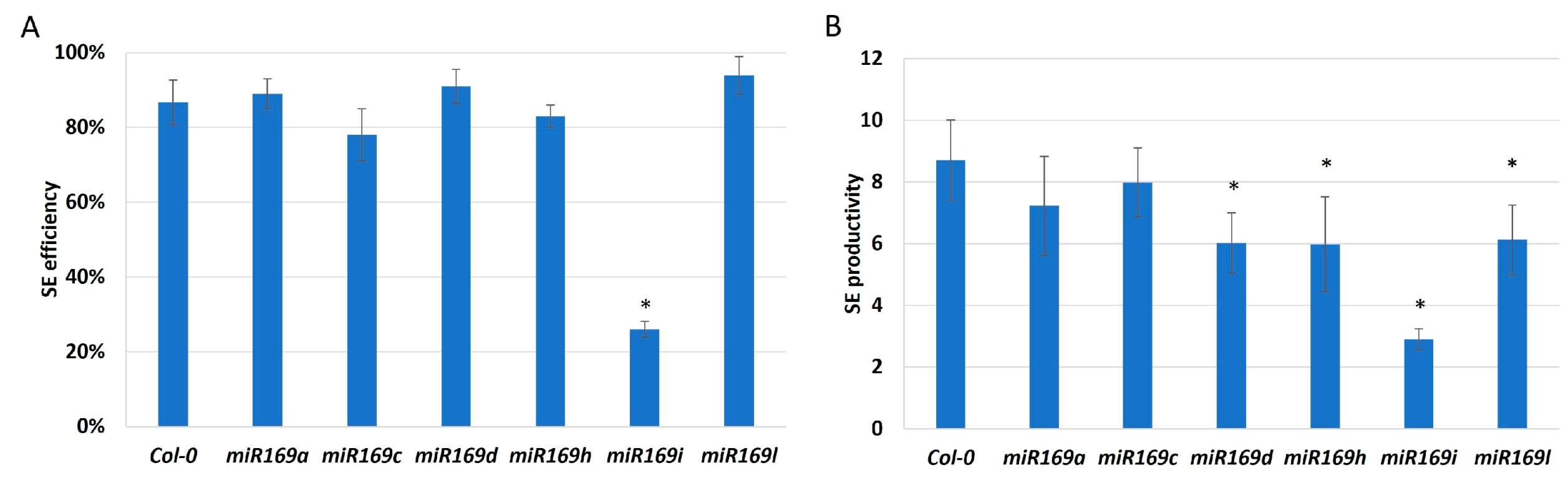
2.4. Different MIRNA169 Genes Contribute to miR169 Regulating SE Induction
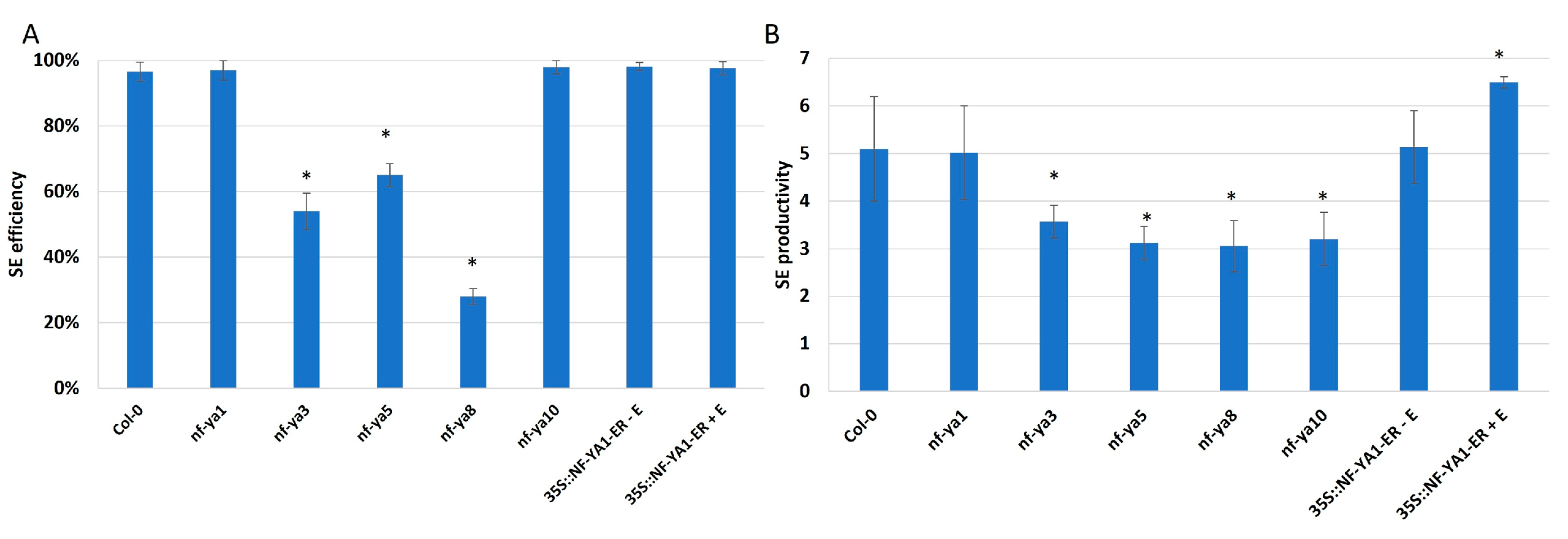
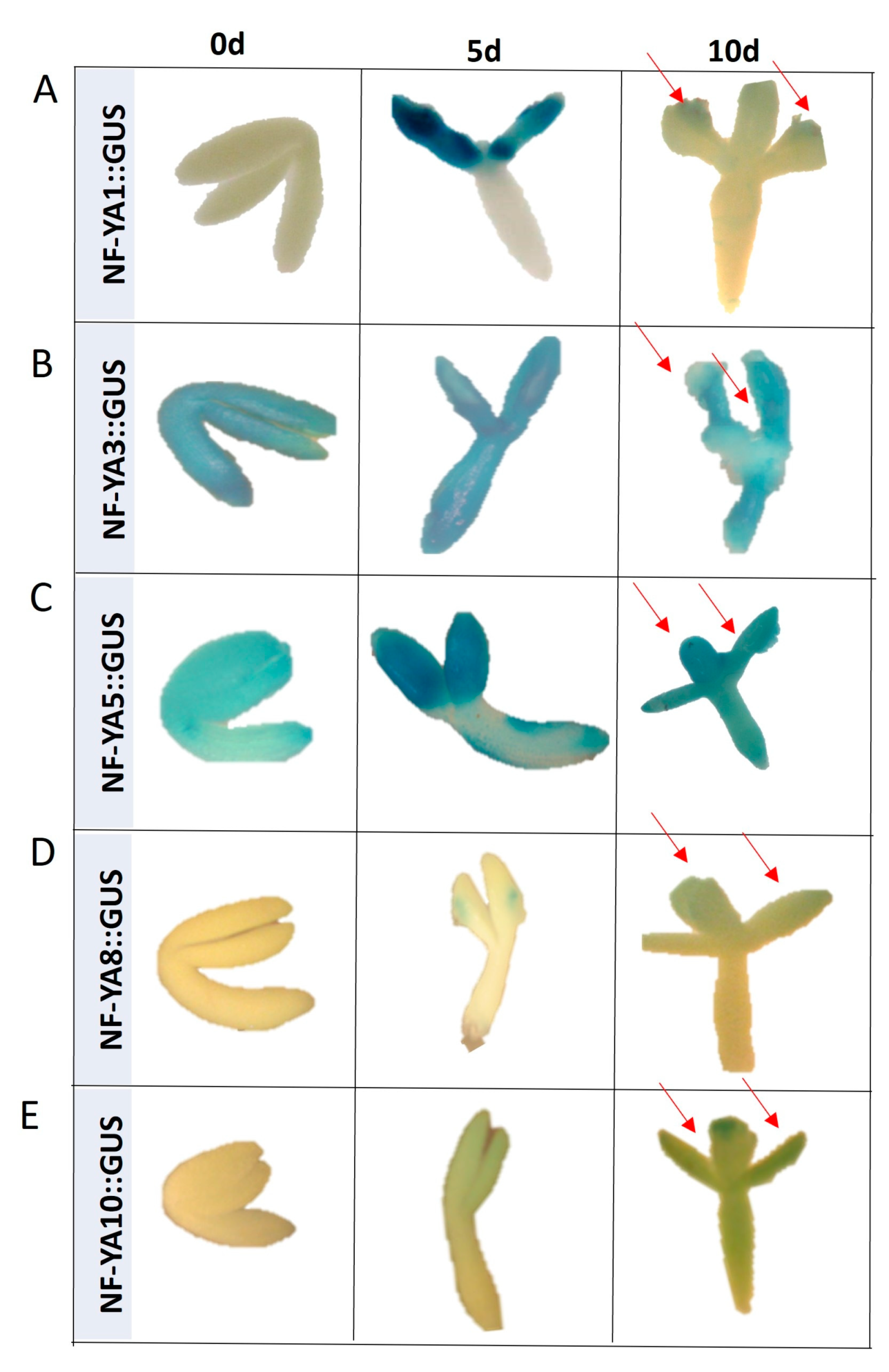
2.5. miR169 Regulates Embryogenic Induction via Controlling NF-YA
NF-YA Control SE Induction
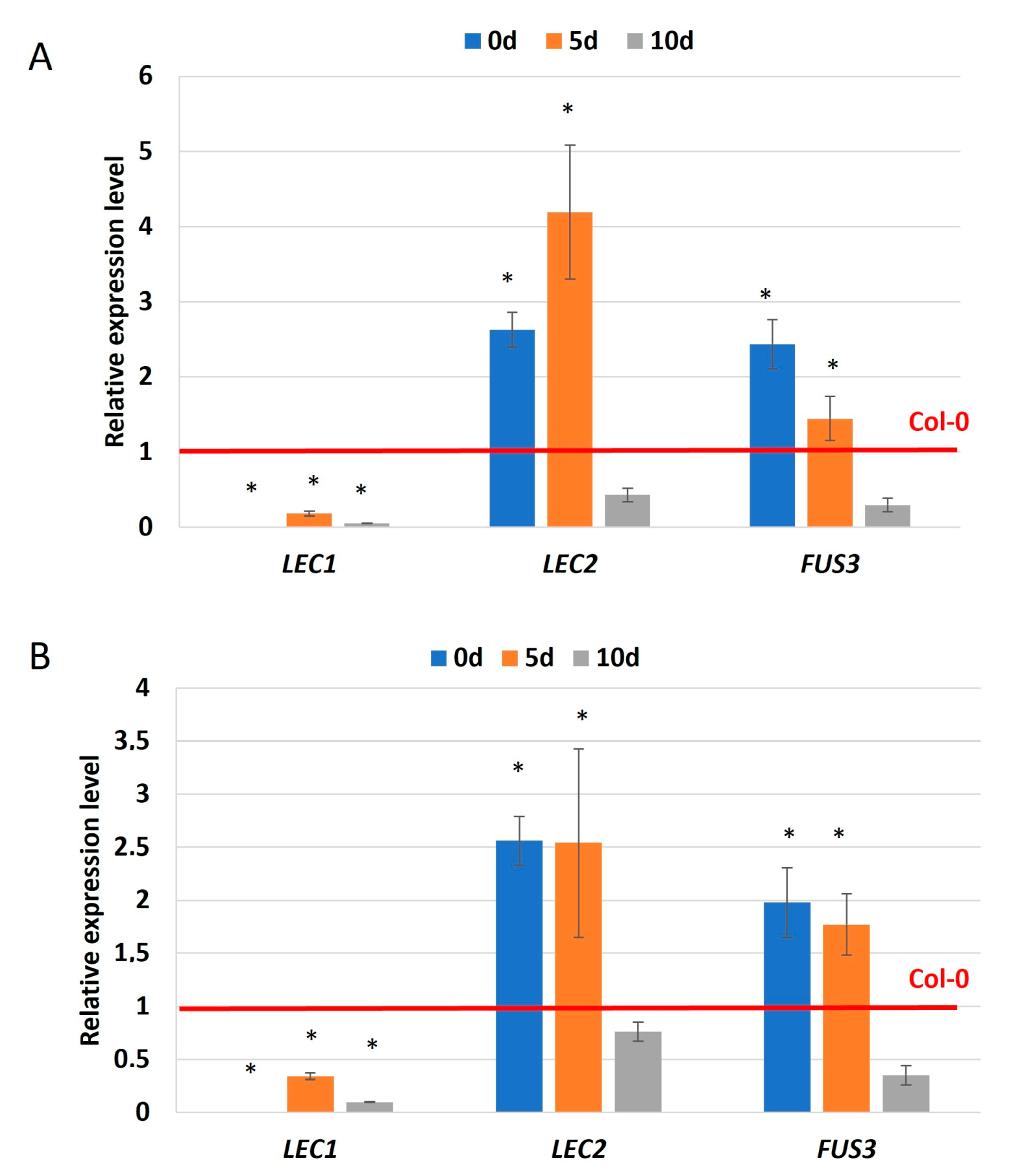
2.6. miR169 via NF-YA5 Regulates the Expression of Genes of Critical Role in SE
3. Discussion
3.1. miR156 through SPLs Controls SE Induction
3.2. Phytohormone and LEC Genes Related Mechanism of miR156-SPL Regulation in SE
3.3. Two miR169 Isoforms via NF-YA TFs Control SE Induction
3.4. miR169-NF-YA5 Module—A New Player in LEC1 and Auxin-Related Mechanisms Controlling SE Induction
4. Materials and Methods
4.1. Plant Material and Growth Conditions
4.2. Somatic Embryogenesis Induction In Vitro
4.3. Analysis of Target Gene Expression
4.4. GUS/GFP Signal Detection
4.5. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Costa, S.; Shaw, P. “Open Minded” Cells: How Cells Can Change Fate. Trends Cell Biol. 2007, 17, 101–106. [Google Scholar] [CrossRef] [PubMed]
- Ochoa-Alejo, N. The Uses of Somatic Embryogenesis for Genetic Transformation. In Somatic Embryogenesis: Fundamental Aspects and Applications; Loyola-Vargas, V.M., Ochoa-Alejo, N., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 415–434. ISBN 978-3-319-33705-0. [Google Scholar]
- Wójcikowska, B.; Wójcik, A.M.; Gaj, M.D. Epigenetic Regulation of Auxin-Induced Somatic Embryogenesis in Plants. Int. J. Mol. Sci. 2020, 21, 2307. [Google Scholar] [CrossRef] [PubMed]
- Salaün, C.; Lepiniec, L.; Dubreucq, B. Genetic and Molecular Control of Somatic Embryogenesis. Plants 2021, 10, 1467. [Google Scholar] [CrossRef]
- Su, Y.H.; Tang, L.P.; Zhao, X.Y.; Zhang, X.S. Plant Cell Totipotency: Insights into Cellular Reprogramming. J. Integr. Plant Biol. 2021, 63, 228–243. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Luo, K.; Li, Z.; Yang, Y. A Novel Method for Induction of Plant Regeneration via Somatic Embryogenesis. Plant Sci. 2009, 177, 43–48. [Google Scholar] [CrossRef]
- Heidmann, I.; de Lange, B.; Lambalk, J.; Angenent, G.C.; Boutilier, K. Efficient Sweet Pepper Transformation Mediated by the BABY BOOM Transcription Factor. Plant Cell Rep. 2011, 30, 1107–1115. [Google Scholar] [CrossRef]
- Belide, S.; Zhou, X.R.; Kennedy, Y.; Lester, G.; Shrestha, P.; Petrie, J.R.; Singh, S.P. Rapid Expression and Validation of Seed-Specific Constructs in Transgenic LEC2 Induced Somatic Embryos of Brassica Napus. Plant Cell Tissue Organ Cult. 2013, 113, 543–553. [Google Scholar] [CrossRef]
- Lowe, K.; Wu, E.; Wang, N.; Hoerster, G.; Hastings, C.; Cho, M.J.; Scelonge, C.; Lenderts, B.; Chamberlin, M.; Cushatt, J.; et al. Morphogenic Regulators Baby Boom and Wuschel Improve Monocot Transformation. Plant Cell 2016, 28, 1998–2015. [Google Scholar] [CrossRef]
- Gliwicka, M.; Nowak, K.; Balazadeh, S.; Mueller-Roeber, B.; Gaj, M.D. Extensive Modulation of the Transcription Factor Transcriptome during Somatic Embryogenesis in Arabidopsis Thaliana. PLoS ONE 2013, 8, e69261. [Google Scholar] [CrossRef]
- Gaj, M.D.; Zhang, S.; Harada, J.J.; Lemaux, P.G. Leafy Cotyledon Genes Are Essential for Induction of Somatic Embryogenesis of Arabidopsis. Planta 2005, 222, 977–988. [Google Scholar] [CrossRef]
- Su, Y.H.; Liu, Y.B.; Bai, B.; Zhang, X.S. Establishment of Embryonic Shoot-Root Axis Is Involved in Auxin and Cytokinin Response during Arabidopsis Somatic Embryogenesis. Front. Plant Sci. 2015, 5, 792. [Google Scholar] [CrossRef] [PubMed]
- Horstman, A.; Bemer, M.; Boutilier, K. A Transcriptional View on Somatic Embryogenesis. Regeneration 2017, 4, 201–216. [Google Scholar] [CrossRef]
- Siddiqui, Z.H.; Abbas, Z.K.; Ansari, M.W.; Khan, M.N. The Role of MiRNA in Somatic Embryogenesis. Genomics 2019, 111, 1026–1033. [Google Scholar] [CrossRef] [PubMed]
- Szyrajew, K.; Bielewicz, D.; Dolata, J.; Wójcik, A.M.; Nowak, K.; Szczygieł-Sommer, A.; Szweykowska-Kulinska, Z.; Jarmolowski, A.; Gaj, M.D. MicroRNAs Are Intensively Regulated during Induction of Somatic Embryogenesis in Arabidopsis. Front. Plant Sci. 2017, 8, 18. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, S.; Han, S.; Wu, T.; Li, X.; Li, W.; Qi, L. Genome-Wide Identification of MicroRNAs in Larch and Stage-Specific Modulation of 11 Conserved MicroRNAs and Their Targets during Somatic Embryogenesis. Planta 2012, 236, 647–657. [Google Scholar] [CrossRef]
- Yang, X.; Wang, L.; Yuan, D.; Lindsey, K.; Zhang, X. Small RNA and Degradome Sequencing Reveal Complex MiRNA Regulation during Cotton Somatic Embryogenesis. J. Exp. Bot. 2013, 64, 1521–1536. [Google Scholar] [CrossRef] [PubMed]
- Chávez-Hernández, E.C.; Alejandri-Ramírez, N.D.; Juárez-González, V.T.; Dinkova, T.D. Maize MiRNA and Target Regulation in Response to Hormone Depletion and Light Exposure during Somatic Embryogenesis. Front. Plant Sci. 2015, 6, 555. [Google Scholar] [CrossRef]
- Wójcik, A.M.; Nodine, M.D.; Gaj, M.D. MiR160 and MiR166/165 Contribute to the LEC2-Mediated Auxin Response Involved in the Somatic Embryogenesis Induction in Arabidopsis. Front. Plant Sci. 2017, 8, 2024. [Google Scholar] [CrossRef]
- Wójcik, A.M.; Mosiolek, M.; Karcz, J.; Nodine, M.D.; Gaj, M.D. Whole Mount in Situ Localization of Mirnas and Mrnas during Somatic Embryogenesis in Arabidopsis. Front. Plant Sci. 2018, 9, 1277. [Google Scholar] [CrossRef]
- Wójcik, A.M.; Gaj, M.D. MiR393 Contributes to the Embryogenic Transition Induced in Vitro in Arabidopsis via the Modification of the Tissue Sensitivity to Auxin Treatment. Planta 2016, 244, 231–243. [Google Scholar] [CrossRef]
- Szczygieł-Sommer, A.; Gaj, M.D. The MiR396–GRF Regulatory Module Controls the Embryogenic Response in Arabidopsis via an Auxin-Related Pathway. Int. J. Mol. Sci. 2019, 20, 5221. [Google Scholar] [CrossRef] [PubMed]
- Luján-Soto, E.; Juárez-González, V.T.; Reyes, J.L.; Dinkova, T.D. Microrna Zma-Mir528 Versatile Regulation on Target Mrnas during Maize Somatic Embryogenesis. Int. J. Mol. Sci. 2021, 22, 5310. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.P.; Zhang, X.S.; Su, Y.H. Regulation of Cell Reprogramming by Auxin during Somatic Embryogenesis. aBIOTECH 2020, 1, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Elhiti, M.; Stasolla, C. Transduction of Signals during Somatic Embryogenesis. Plants 2022, 11, 178. [Google Scholar] [CrossRef]
- Jones-Rhoades, M.W.; Bartel, D.P. Computational Identification of Plant MicroRNAs and Their Targets, Including a Stress-Induced MiRNA. Mol. Cell 2004, 14, 789–799. [Google Scholar] [CrossRef]
- Wu, G.; Park, M.Y.; Conway, S.R.; Wang, J.W.; Weigel, D.; Poethig, R.S. The Sequential Action of MiR156 and MiR172 Regulates Developmental Timing in Arabidopsis. Cell 2009, 138, 750–759. [Google Scholar] [CrossRef]
- Zhang, M.; An, P.; Li, H.; Wang, X.; Zhou, J.; Dong, P.; Zhao, Y.; Wang, Q.; Li, C. The MiRNA-Mediated Post-Transcriptional Regulation of Maize in Response to High Temperature. Int. J. Mol. Sci. 2019, 20, 1754. [Google Scholar] [CrossRef]
- Birkenbihl, R.P.; Jach, G.; Saedler, H.; Huijser, P. Functional Dissection of the Plant-Specific SBP-Domain: Overlap of the DNA-Binding and Nuclear Localization Domains. J. Mol. Biol. 2005, 352, 585–596. [Google Scholar] [CrossRef]
- Yamasaki, K.; Kigawa, T.; Inoue, M.; Tateno, M.; Yamasaki, T.; Yabuki, T.; Aoki, M.; Seki, E.; Matsuda, T.; Nunokawa, E.; et al. A Novel Zinc-Binding Motif Revealed by Solution Structures of DNA-Binding Domains of Arabidopsis SBP-Family Transcription Factors. J. Mol. Biol. 2004, 337, 49–63. [Google Scholar] [CrossRef]
- Xu, M.; Hu, T.; Zhao, J.; Park, M.Y.; Earley, K.W.; Wu, G.; Yang, L.; Poethig, R.S. Developmental Functions of MiR156-Regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) Genes in Arabidopsis Thaliana. PLoS Genet. 2016, 12, e1006263. [Google Scholar] [CrossRef]
- Shikata, M.; Koyama, T.; Mitsuda, N.; Ohme-Takagi, M. Arabidopsis SBP-Box Genes SPL10, SPL11 and SPL2 Control Morphological Change in Association with Shoot Maturation in the Reproductive Phase. Plant Cell Physiol. 2009, 50, 2133–2145. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; He, H.; Yu, D. Identification of Nitrogen Starvation-Responsive MicroRNAs in Arabidopsis Thaliana. PLoS ONE 2012, 7, e48951. [Google Scholar] [CrossRef]
- Gonzalez-Ibeas, D.; Blanca, J.; Donaire, L.; Saladié, M.; Mascarell-Creus, A.; Cano-Delgado, A.; Garcia-Mas, J.; Llave, C.; Aranda, M.A. Analysis of the Melon (Cucumis Melo) Small RNAome by High-Throughput Pyrosequencing. BMC Genom. 2011, 12, 393. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.; Talla, A.; Qiu, W. Small RNA Profiling of Virus-Infected Grapevines: Evidences for Virus Infection-Associated and Variety-Specific MiRNAs. Funct. Integr. Genom. 2012, 12, 659–669. [Google Scholar] [CrossRef]
- Licausi, F.; Weits, D.A.; Pant, B.D.; Scheible, W.R.; Geigenberger, P.; van Dongen, J.T. Hypoxia Responsive Gene Expression Is Mediated by Various Subsets of Transcription Factors and MiRNAs That Are Determined by the Actual Oxygen Availability. New Phytol. 2011, 190, 442–456. [Google Scholar] [CrossRef]
- Zhao, M.; Ding, H.; Zhu, J.K.; Zhang, F.; Li, W.X. Involvement of MiR169 in the Nitrogen-Starvation Responses in Arabidopsis. New Phytol. 2011, 190, 906–915. [Google Scholar] [CrossRef]
- Rhoades, M.W.; Reinhart, B.J.; Lim, L.P.; Burge, C.B.; Bartel, B.; Bartel, D.P. Prediction of Plant MicroRNA Targets. Cells. 2002, 110, 513–520. [Google Scholar] [CrossRef]
- Mantovani, R. The Molecular Biology of the CCAAT-Binding Factor NF-Y. Gene 1999, 239, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Baxevanis, A.D.; Arents, G.; Moudrianakis, E.N.; Landsman, D. A Variety of DNA-Binding and Multimeric Proteins Contain the Histone Fold Motif. Nucleic Acids Res. 1995, 23, 2685–2691. [Google Scholar] [CrossRef]
- Zemzoumi, K.; Frontini, M.; Bellorini, M.; Mantovani, R. NF-Y Histone Fold A1 Helices Help Impart CCAAT Specificity. J. Mol. Biol. 1999, 286, 327–337. [Google Scholar] [CrossRef]
- Zhou, X.; Wang, G.; Sutoh, K.; Zhu, J.K.; Zhang, W. Identification of Cold-Inducible MicroRNAs in Plants by Transcriptome Analysis. Biochim. Biophys. Acta Gene Regul. Mech. 2008, 1779, 780–788. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Yoo, S.J.; Lee, J.H.; Kim, W.; Yoo, S.K.; Fitzgerald, H.; Carrington, J.C.; Ahn, J.H. Genetic Framework for Flowering-Time Regulation by Ambient Temperature-Responsive MiRNAs in Arabidopsis. Nucleic Acids Res. 2010, 38, 3081–3093. [Google Scholar] [CrossRef]
- Li, Y.; Zhao, S.L.; Li, J.L.; Hu, X.H.; Wang, H.; Cao, X.L.; Xu, Y.J.; Zhao, Z.X.; Xiao, Z.Y.; Yang, N.; et al. Osa-MiR169 Negatively Regulates Rice Immunity against the Blast Fungus Magnaporthe Oryzae. Front. Plant Sci. 2017, 8, 2. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Wang, Y.; Gu, Y.; Yan, P.; Zhao, W.; Jiang, T. Genome-Wide Identification of MiR169 Family in Response to ABA and Salt Stress in Poplar. Forests 2023, 14, 961. [Google Scholar] [CrossRef]
- Sorin, C.; Declerck, M.; Christ, A.; Blein, T.; Ma, L.; Lelandais-Brière, C.; Njo, M.F.; Beeckman, T.; Crespi, M.; Hartmann, C. A MiR169 Isoform Regulates Specific NF-YA Targets and Root Architecture in Arabidopsis. New Phytol. 2014, 202, 1197–1211. [Google Scholar] [CrossRef]
- Long, J.M.; Liu, C.Y.; Feng, M.Q.; Liu, Y.; Wu, X.M.; Guo, W.W. MiR156-SPL Modules Regulate Induction of Somatic Embryogenesis in Citrus Callus. J. Exp. Bot. 2018, 69, 2979–2993. [Google Scholar] [CrossRef]
- Nodine, M.D.; Bartel, D.P. MicroRNAs Prevent Precocious Gene Expression and Enable Pattern Formation during Plant Embryogenesis. Genes Dev. 2010, 24, 2678–2692. [Google Scholar] [CrossRef] [PubMed]
- Petroni, K.; Kumimoto, R.W.; Gnesutta, N.; Calvenzani, V.; Fornari, M.; Tonelli, C.; Holt, B.F.; Mantovani, R. The Promiscuous Life of Plant NUCLEAR FACTOR Y Transcription Factors. Plant Cell 2013, 24, 4777–4792. [Google Scholar] [CrossRef]
- Calvenzani, V.; Testoni, B.; Gusmaroli, G.; Lorenzo, M.; Gnesutta, N.; Petroni, K.; Mantovani, R.; Tonelli, C. Interactions and CCAAT-Binding of Arabidopsis Thaliana NF-Y Subunits. PLoS ONE 2012, 7, e42902. [Google Scholar] [CrossRef]
- Su, Y.H.; Zhao, X.Y.; Liu, Y.B.; Zhang, C.L.; O’Neill, S.D.; Zhang, X.S. Auxin-Induced WUS Expression Is Essential for Embryonic Stem Cell Renewal during Somatic Embryogenesis in Arabidopsis. Plant J. 2009, 59, 448–460. [Google Scholar] [CrossRef]
- Mu, J.; Tan, H.; Hong, S.; Liang, Y.; Zuo, J. Arabidopsis Transcription Factor Genes NF-YA1, 5, 6, and 9 Play Redundant Roles in Male Gametogenesis, Embryogenesis, and Seed Development. Mol. Plant 2013, 6, 188–201. [Google Scholar] [CrossRef] [PubMed]
- Fornari, M.; Calvenzani, V.; Masiero, S.; Tonelli, C.; Petroni, K. The Arabidopsis NF-YA3 and NF-YA8 Genes Are Functionally Redundant and Are Required in Early Embryogenesis. PLoS ONE 2013, 8, e82043. [Google Scholar] [CrossRef] [PubMed]
- Sabana, A.A.; Antony, G.; Rahul, C.U.; Rajesh, M.K. In Silico Identification of MicroRNAs and Their Targets Associated with Coconut Embryogenic Calli. Agri Gene 2018, 7, 59–65. [Google Scholar] [CrossRef]
- Zhang, J.; Xue, B.; Gai, M.; Song, S.; Jia, N.; Sun, H. Small RNA and Transcriptome Sequencing Reveal a Potential MiRNA-Mediated Interaction Network That Functions during Somatic Embryogenesis in Lilium Pumilum DC. Fisch. Front. Plant Sci. 2017, 8, 566. [Google Scholar] [CrossRef] [PubMed]
- Qin, Z.; Li, J.; Zhang, Y.; Xiao, Y.; Zhang, X.; Zhong, L.; Liu, H.; Chen, B. Genome-Wide Identification of MicroRNAs Involved in the Somatic Embryogenesis of Eucalyptus. G3 Genes Genomes Genet. 2021, 11, jkab070. [Google Scholar] [CrossRef]
- Wu, X.M.; Kou, S.J.; Liu, Y.L.; Fang, Y.N.; Xu, Q.; Guo, W.W. Genomewide Analysis of Small RNAs in Nonembryogenic and Embryogenic Tissues of Citrus: MicroRNA- and SiRNA-Mediated Transcript Cleavage Involved in Somatic Embryogenesis. Plant Biotechnol. J. 2015, 13, 383–394. [Google Scholar] [CrossRef]
- Armenta-Medina, A.; Lepe-Soltero, D.; Xiang, D.; Datla, R.; Abreu-Goodger, C.; Gillmor, C.S. Arabidopsis Thaliana MiRNAs Promote Embryo Pattern Formation Beginning in the Zygote. Dev. Biol. 2017, 431, 145–151. [Google Scholar] [CrossRef]
- Plotnikova, A.; Kellner, M.J.; Schon, M.A.; Mosiolek, M.; Nodine, M.D. MicroRNA Dynamics and Functions During Arabidopsis Embryogenesis. Plant Cell 2019, 31, 2929–2946. [Google Scholar] [CrossRef]
- Nowak, K.; Morończyk, J.; Wójcik, A.; Gaj, M.D. AGL15 Controls the Embryogenic Reprogramming of Somatic Cells in Arabidopsis through the Histone Acetylation-Mediated Repression of the Mirna Biogenesis Genes. Int. J. Mol. Sci. 2020, 21, 6733. [Google Scholar] [CrossRef]
- Serivichyaswat, P.; Ryu, H.S.; Kim, W.; Kim, S.; Chung, K.S.; Kim, J.J.; Ahn, J.H. Expression of the Floral Repressor MiRNA156 Is Positively Regulated by the AGAMOUS-like Proteins AGL15 and AGL18. Mol. Cells 2015, 38, 259–266. [Google Scholar] [CrossRef]
- Jeyakumar, J.M.J.; Ali, A.; Wang, W.M.; Thiruvengadam, M. Characterizing the Role of the MiR156-SPL Network in Plant Development and Stress Response. Plants 2020, 9, 1206. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.Q.; Lian, H.; Tang, H.; Dolezal, K.; Zhou, C.M.; Yu, S.; Chen, J.H.; Chen, Q.; Liu, H.; Ljung, K.; et al. An IntrinsicmicroRNA Timer Regulates Progressive Decline in Shoot Regenerative Capacity in Plants. Plant Cell 2015, 27, 349–360. [Google Scholar] [CrossRef]
- Wang, L.; Liu, N.; Wang, T.; Li, J.; Wen, T.; Yang, X.; Lindsey, K.; Zhang, X. The GhmiR157a- GhSPL10 Regulatory Module Controls Initial Cellular Dedifferentiation and Callus Proliferation in Cotton by Modulating Ethylene-Mediated Flavonoid Biosynthesis. J. Exp. Bot. 2018, 69, 1081–1093. [Google Scholar] [CrossRef] [PubMed]
- Willmann, M.R.; Mehalick, A.J.; Packer, R.L.; Jenik, P.D. MicroRNAs Regulate the Timing of Embryo Maturation in Arabidopsis. Plant Physiol. 2011, 155, 1871–1884. [Google Scholar] [CrossRef] [PubMed]
- Nowak, K.; Morończyk, J.; Grzyb, M.; Szczygieł-Sommer, A.; Gaj, M.D. MiR172 Regulates WUS during Somatic Embryogenesis in Arabidopsis via AP2. Cells 2022, 11, 718. [Google Scholar] [CrossRef] [PubMed]
- Luo, P.; Di, D.W.; Wu, L.; Yang, J.; Lu, Y.; Shi, W. MicroRNAs Are Involved in Regulating Plant Development and Stress Response through Fine-Tuning of TIR1/AFB-Dependent Auxin Signaling. Int. J. Mol. Sci. 2022, 23, 510. [Google Scholar] [CrossRef]
- Curaba, J.; Singh, M.B.; Bhalla, P.L. MiRNAs in the Crosstalk between Phytohormone Signalling Pathways. J. Exp. Bot. 2014, 65, 1425–1438. [Google Scholar] [CrossRef]
- Yu, S.; Galvão, V.C.; Zhang, Y.C.; Horrer, D.; Zhang, T.Q.; Hao, Y.H.; Feng, Y.Q.; Wang, S.; Schmid, M.; Wang, J.W. Gibberellin Regulates the Arabidopsis Floral Transition through MiR156-Targeted SQUAMOSA PROMOTER BINDING-LIKE Transcription Factors. Plant Cell 2012, 24, 3320–3332. [Google Scholar] [CrossRef]
- Sablowski, R. Cytokinin and WUSCHEL Tie the Knot around Plant Stem Cells. Proc. Natl. Acad. Sci. USA 2009, 106, 16016–16017. [Google Scholar] [CrossRef]
- Wójcik, A.M.; Wójcikowska, B.; Gaj, M.D. Current Perspectives on the Auxin-Mediated Genetic Network That Controls the Induction of Somatic Embryogenesis in Plants. Int. J. Mol. Sci. 2020, 21, 1333. [Google Scholar] [CrossRef]
- Dai, Z.; Wang, J.; Yang, X.; Lu, H.; Miao, X.; Shi, Z. Modulation of Plant Architecture by the MiR156f-OsSPL7-OsGH3.8 Pathway in Rice. J. Exp. Bot. 2018, 69, 5117–5130. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.; Niu, Q.W.; Ng, K.H.; Chua, N.H. The Role of MiR156/SPLs Modules in Arabidopsis Lateral Root Development. Plant J. 2015, 83, 673–685. [Google Scholar] [CrossRef]
- Braybrook, S.A.; Harada, J.J. LECs Go Crazy in Embryo Development. Trends Plant Sci. 2008, 13, 624–630. [Google Scholar] [CrossRef]
- Zhang, M.; Hu, X.; Zhu, M.; Xu, M.; Wang, L. Transcription Factors NF-YA2 and NF-YA10 Regulate Leaf Growth via Auxin Signaling in Arabidopsis. Sci. Rep. 2017, 7, 1395. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Liu, S.; Ren, T.; Niu, M.; Liu, X.; Liu, C.; Wang, H.; Yin, W.; Xia, X. Crucial Abiotic Stress Regulatory Network of NF-Y Transcription Factor in Plants. Int. J. Mol. Sci. 2023, 24, 4426. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, S.; Han, S.; Li, X.; Qi, L. Transcriptome Profiling and in Silico Analysis of Somatic Embryos in Japanese Larch (Larix Leptolepis). Plant Cell Rep. 2012, 31, 1637–1657. [Google Scholar] [CrossRef] [PubMed]
- Jin, F.; Hu, L.; Yuan, D.; Xu, J.; Gao, W.; He, L.; Yang, X.; Zhang, X. Comparative Transcriptome Analysis between Somatic Embryos (SEs) and Zygotic Embryos in Cotton: Evidence for Stress Response Functions in SE Development. Plant Biotechnol. J. 2014, 12, 161–173. [Google Scholar] [CrossRef] [PubMed]
- Li, W.X.; Oono, Y.; Zhu, J.; He, X.J.; Wu, J.M.; Iida, K.; Lu, X.Y.; Cui, X.; Jin, H.; Zhu, J.K. The Arabidopsis NFYA5 Transcription Factor Is Regulated Transcriptionally and Posttranscriptionally to Promote Drought Resistance. Plant Cell 2008, 20, 2238–2251. [Google Scholar] [CrossRef]
- Lotan, T.; Ohto, M.; Yee, K.M.; West, M.A.L.; Lo, R.; Kwong, R.W.; Yamagishi, K.; Fischer, R.L.; Goldberg, R.B.; Harada, J.J. Arabidopsis LEAFY COTYLEDON1 Is Sufficient to Induce Embryo Development in Vegetative Cells. Cell 1998, 93, 1195–1205. [Google Scholar] [CrossRef]
- Gyula, P.; Baksa, I.; Tóth, T.; Mohorianu, I.; Dalmay, T.; Szittya, G. Ambient Temperature Regulates the Expression of a Small Set of SRNAs Influencing Plant Development through NF-YA2 and YUC2. Plant Cell Environ. 2018, 41, 2404–2417. [Google Scholar] [CrossRef]
- Aslam, M.; Sugita, K.; Qin, Y.; Rahman, A. Aux/Iaa14 Regulates Microrna-mediated Cold Stress Response in Arabidopsis Roots. Int. J. Mol. Sci. 2020, 21, 8441. [Google Scholar] [CrossRef]
- Diletta Dolfini, R.G.; Mantovani, R. NF-Y and the Transcriptional Activation of CCAAT Promoters. Crit. Rev. Biochem. Mol. Biol. 2012, 47, 29–49. [Google Scholar] [CrossRef]
- Oldfield, A.J.; Yang, P.; Conway, A.E.; Cinghu, S.; Freudenberg, J.M.; Yellaboina, S.; Jothi, R. Histone-Fold Domain Protein NF-Y Promotes Chromatin Accessibility for Cell Type-Specific Master Transcription Factors. Mol. Cell 2014, 55, 708–722. [Google Scholar] [CrossRef] [PubMed]
- Gaj, M.D. Direct Somatic Embryogenesis as a Rapid and Efficient System for in Vitro Regeneration of Arabidopsis Thaliana. Plant Cell Tissue Organ Cult. 2001, 64, 39–46. [Google Scholar] [CrossRef]
- Heidstra, R.; Sabatini, S. Plant and Animal Stem Cells: Similar yet Different. Nat. Rev. Mol. Cell Biol. 2014, 15, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Tao, Z.; Shen, L.; Gu, X.; Wang, Y.; Yu, H.; He, Y. Embryonic Epigenetic Reprogramming by a Pioneer Transcription Factor in Plants. Nature 2017, 551, 124–128. [Google Scholar] [CrossRef]
- Gamborg, O.L.; Miller, R.A.; Ojima, K. Nutrient Requirements of Suspension Cultures of Soybean Root Cells. Exp. Cell Res. 1968, 50, 151–158. [Google Scholar] [CrossRef]
- Speth, C.; Laubinger, S. Rapid and Parallel Quantification of Small and Large RNA Species. In Plant Circadian Networks: Methods and Protocols; Staiger, D., Ed.; Springer New York: New York, NY, USA, 2014; pp. 93–106. ISBN 978-1-4939-0700-7. [Google Scholar]
- Kraut, M.; Wójcikowska, B.; Ledwoń, A.; Gaj, M.D. Immature Zygotic Embryo Cultures of Arabidopsis. Amodel System for Molecular Studies on Morphogenic Pathways Induced in Vitro. Acta Biol. Cracoviensia Ser. Bot. 2011, 53, 59–67. [Google Scholar] [CrossRef]
- Wójcikowska, B.; Jaskóła, K.; Gąsiorek, P.; Meus, M.; Nowak, K.; Gaj, M.D. LEAFY COTYLEDON2 (LEC2) Promotes Embryogenic Induction in Somatic Tissues of Arabidopsis, via YUCCA-Mediated Auxin Biosynthesis. Planta 2013, 238, 425–440. [Google Scholar] [CrossRef]
- Jefferson, R.A.; Kavanagh, T.A.; Bevan, M.W. GUS Fusions: Beta-glucuronidase as a Sensitive and Versatile Gene Fusion Marker in Higher Plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar] [CrossRef]


| Gene/miRNA | Primer Sequence |
|---|---|
| SPL2 | [15] |
| SPL3 | [15] |
| SPL4 | F-GCGCTTAGCTGGACACAATG R-GTCTGGATCAGTTGACCGCT |
| SPL5 | F-GCGGTCAACTGATCCAGACT R-AGAAGAGAGAGAGCGGGAGG |
| SPL9 | [15] |
| SPL10 | [15] |
| SPL11 | F-GCAGGTTCCATGCTGTCTCT R-ACGACGCCTCGCATTATGAT |
| SPL13 | [15] |
| SPL15 | F-TAATGTGTTCGGGTCAGGCC R -TCCGGATCCATCCTCGAAGT |
| LEC1 | [91] |
| LEC2 | [91] |
| FUS3 | [91] |
| NF-YA5 | [15] |
| YUC4 | [64] |
| YUC10 | [64] |
| PIN1 | F-AACCGTTTCGTCGCTCTCTT R-ACGGAGGTTCATGGCGTAAG |
| miR156 | [15] |
| miR169 | [15] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nowak, K.; Wójcik, A.M.; Konopka, K.; Jarosz, A.; Dombert, K.; Gaj, M.D. miR156-SPL and miR169-NF-YA Modules Regulate the Induction of Somatic Embryogenesis in Arabidopsis via LEC- and Auxin-Related Pathways. Int. J. Mol. Sci. 2024, 25, 9217. https://doi.org/10.3390/ijms25179217
Nowak K, Wójcik AM, Konopka K, Jarosz A, Dombert K, Gaj MD. miR156-SPL and miR169-NF-YA Modules Regulate the Induction of Somatic Embryogenesis in Arabidopsis via LEC- and Auxin-Related Pathways. International Journal of Molecular Sciences. 2024; 25(17):9217. https://doi.org/10.3390/ijms25179217
Chicago/Turabian StyleNowak, Katarzyna, Anna M. Wójcik, Katarzyna Konopka, Alicja Jarosz, Katarzyna Dombert, and Małgorzata D. Gaj. 2024. "miR156-SPL and miR169-NF-YA Modules Regulate the Induction of Somatic Embryogenesis in Arabidopsis via LEC- and Auxin-Related Pathways" International Journal of Molecular Sciences 25, no. 17: 9217. https://doi.org/10.3390/ijms25179217






