A New Strategy for Ultrasensitive Detection Based on Target microRNA-Triggered Rolling Circle Amplification in the Early Diagnosis of Alzheimer’s Disease
Abstract
1. Introduction
2. Results
2.1. Principle of Exo III-GO-RCA Assay
2.2. Assay Feasibility
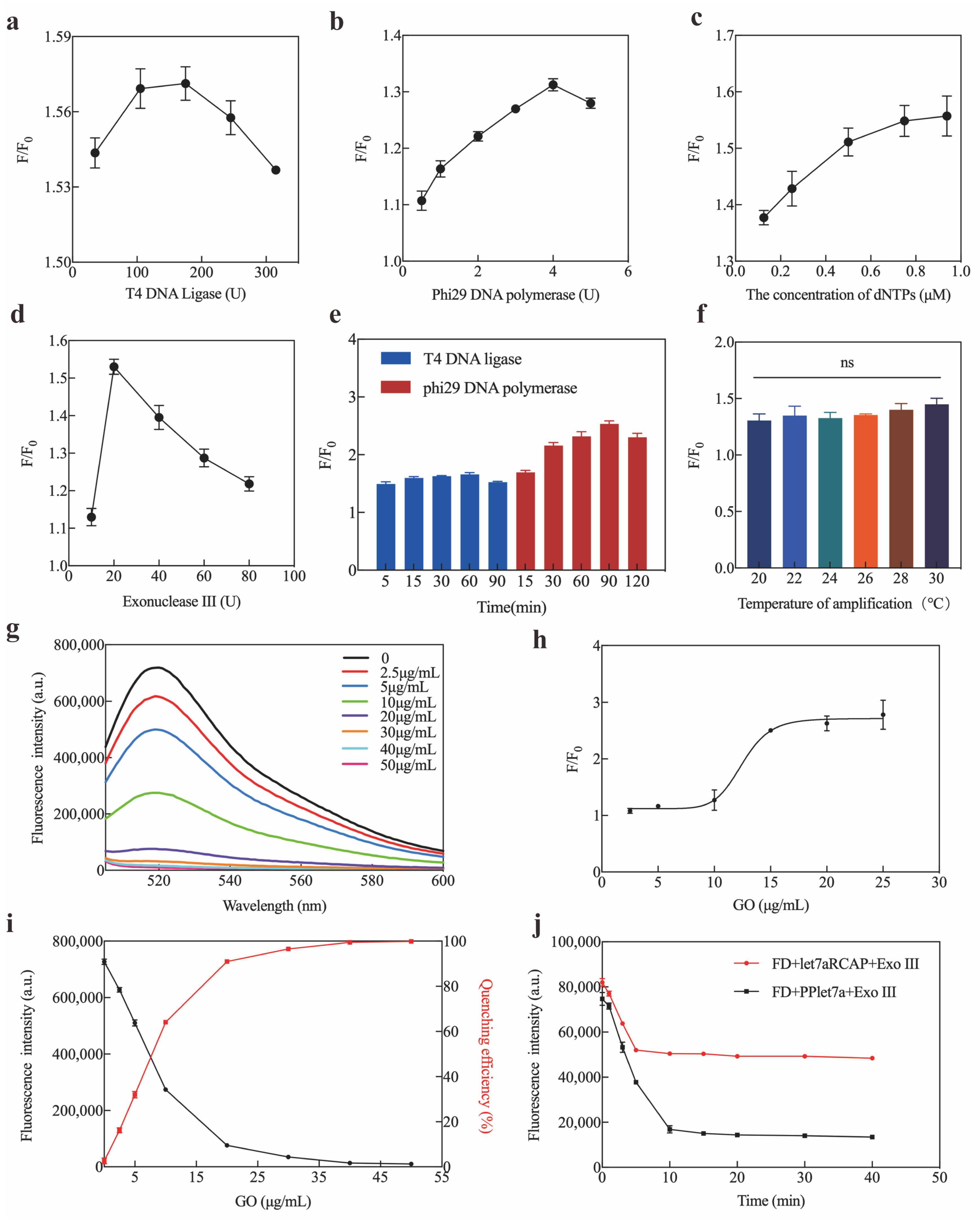
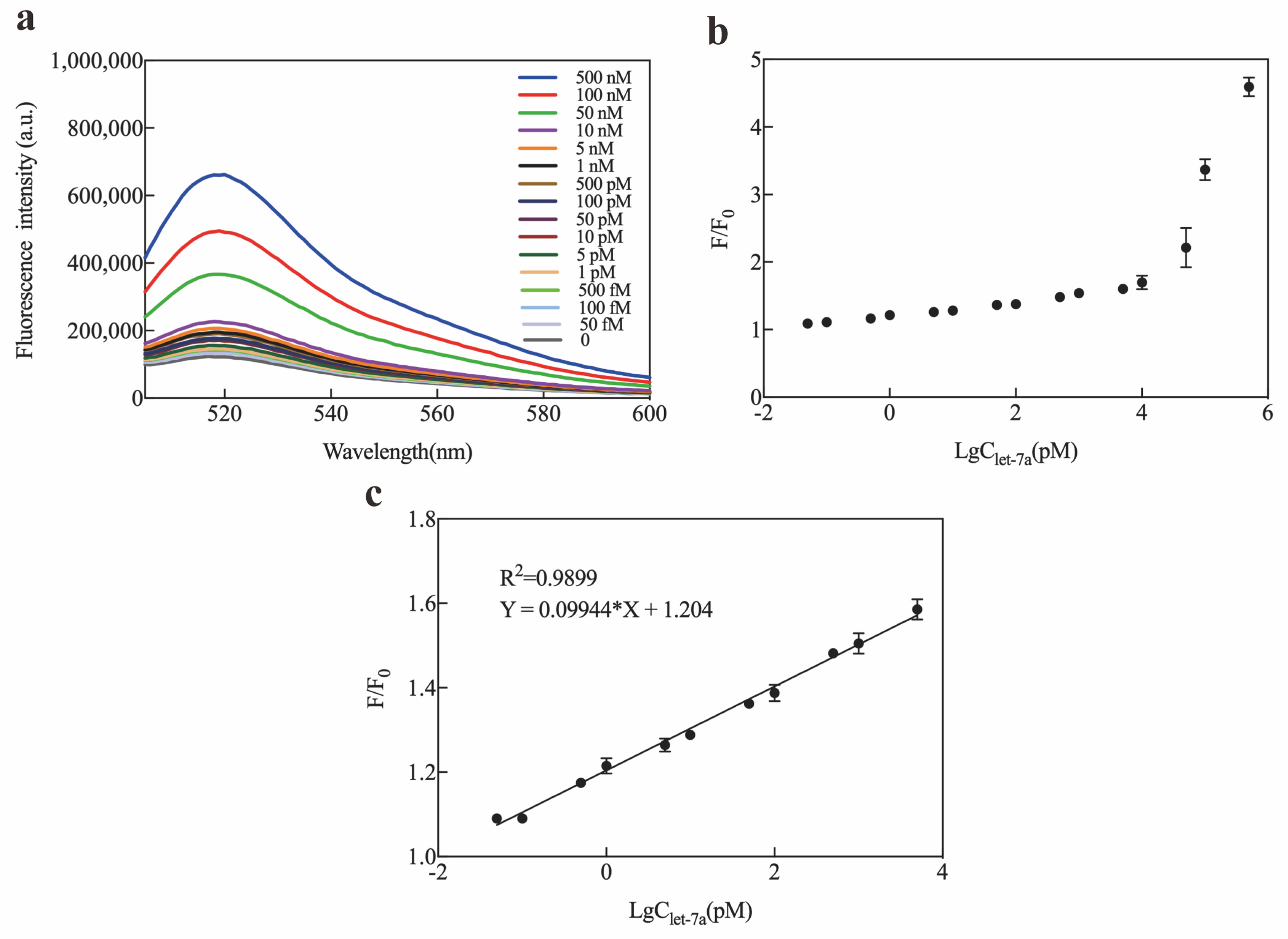
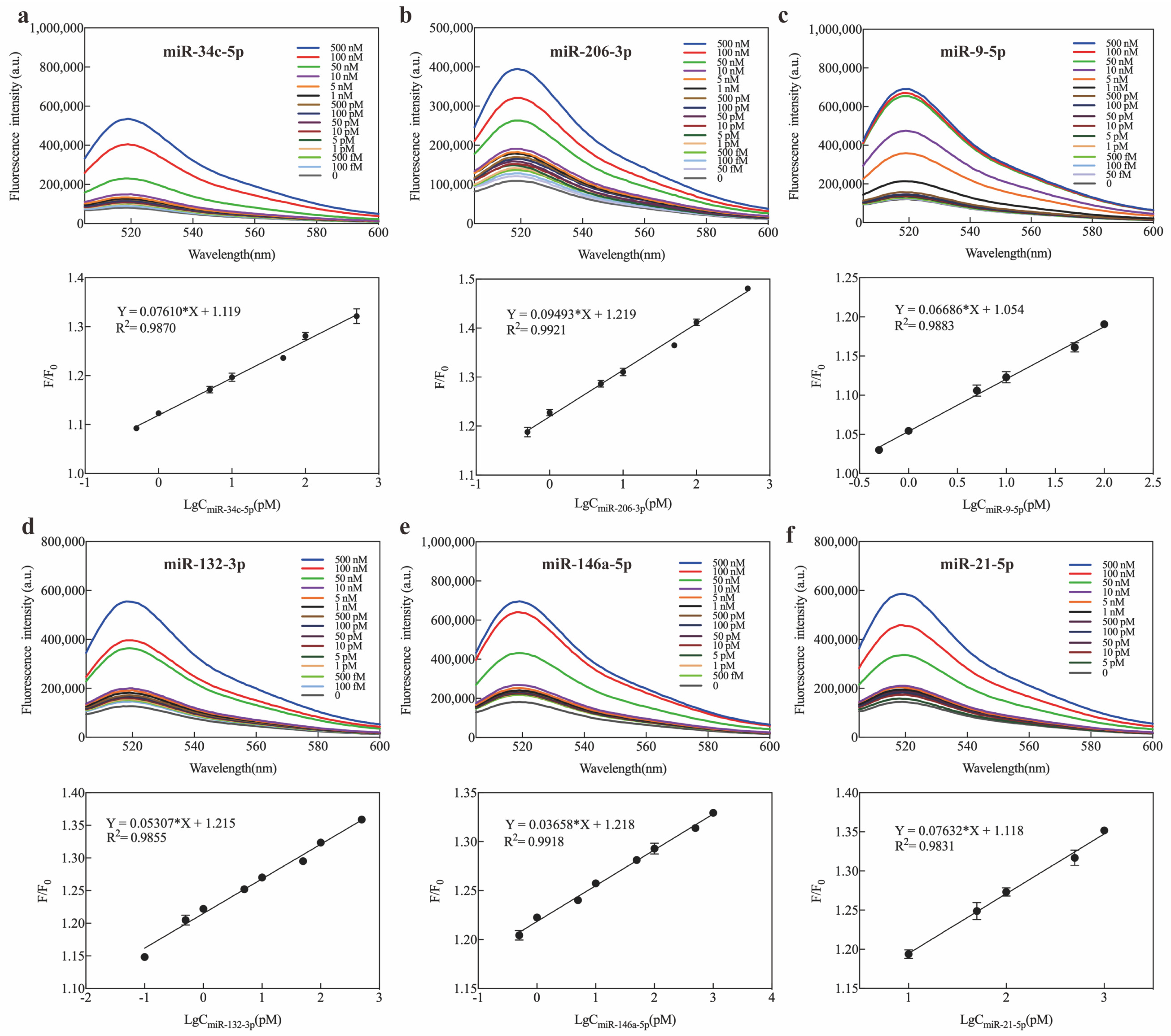
2.3. Optimization and Sensitivity of Exo III-GO-RCA Strategy
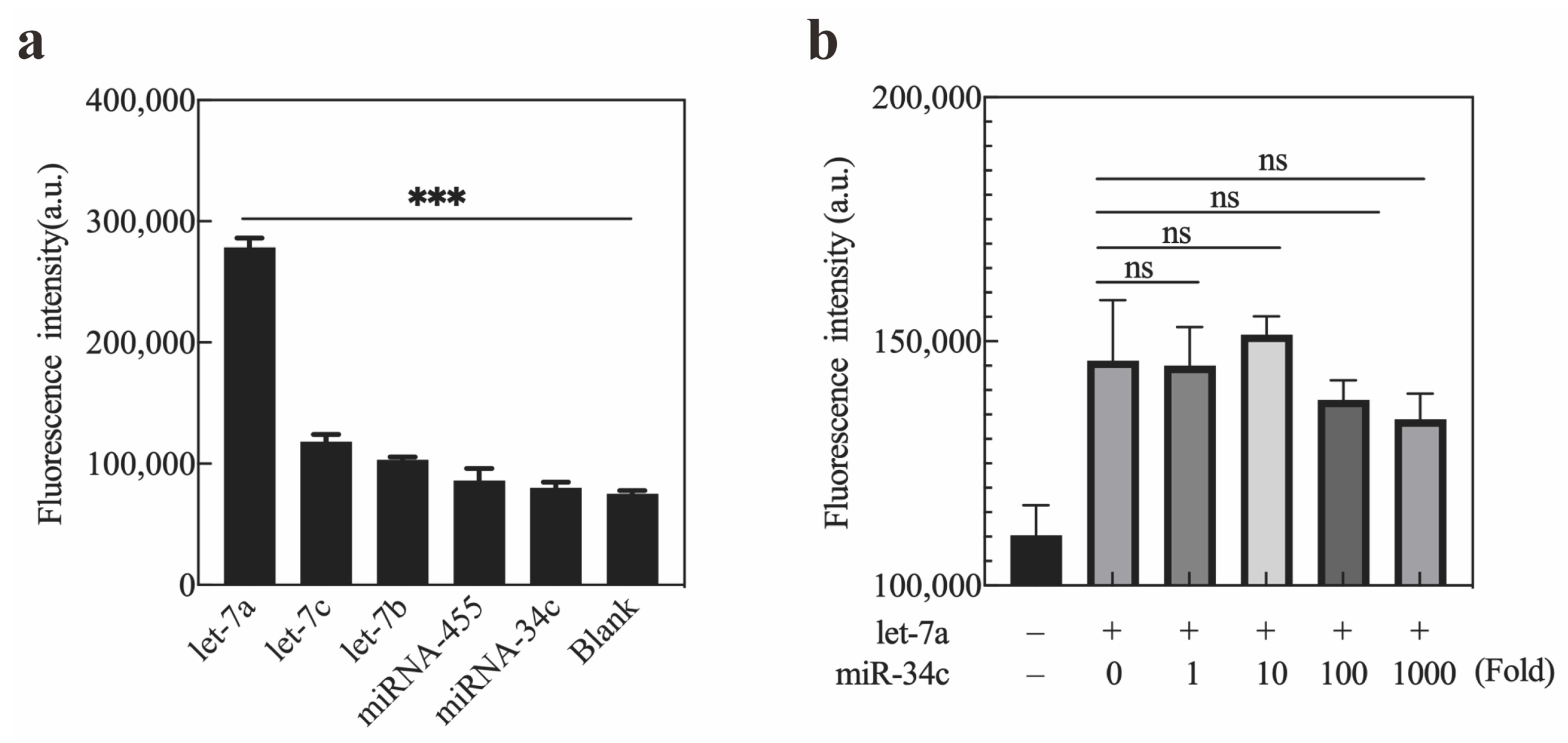
2.4. Specificity of Exo III-GO-RCA
2.5. Application of Exo III-GO-RCA in Real Sample Analysis
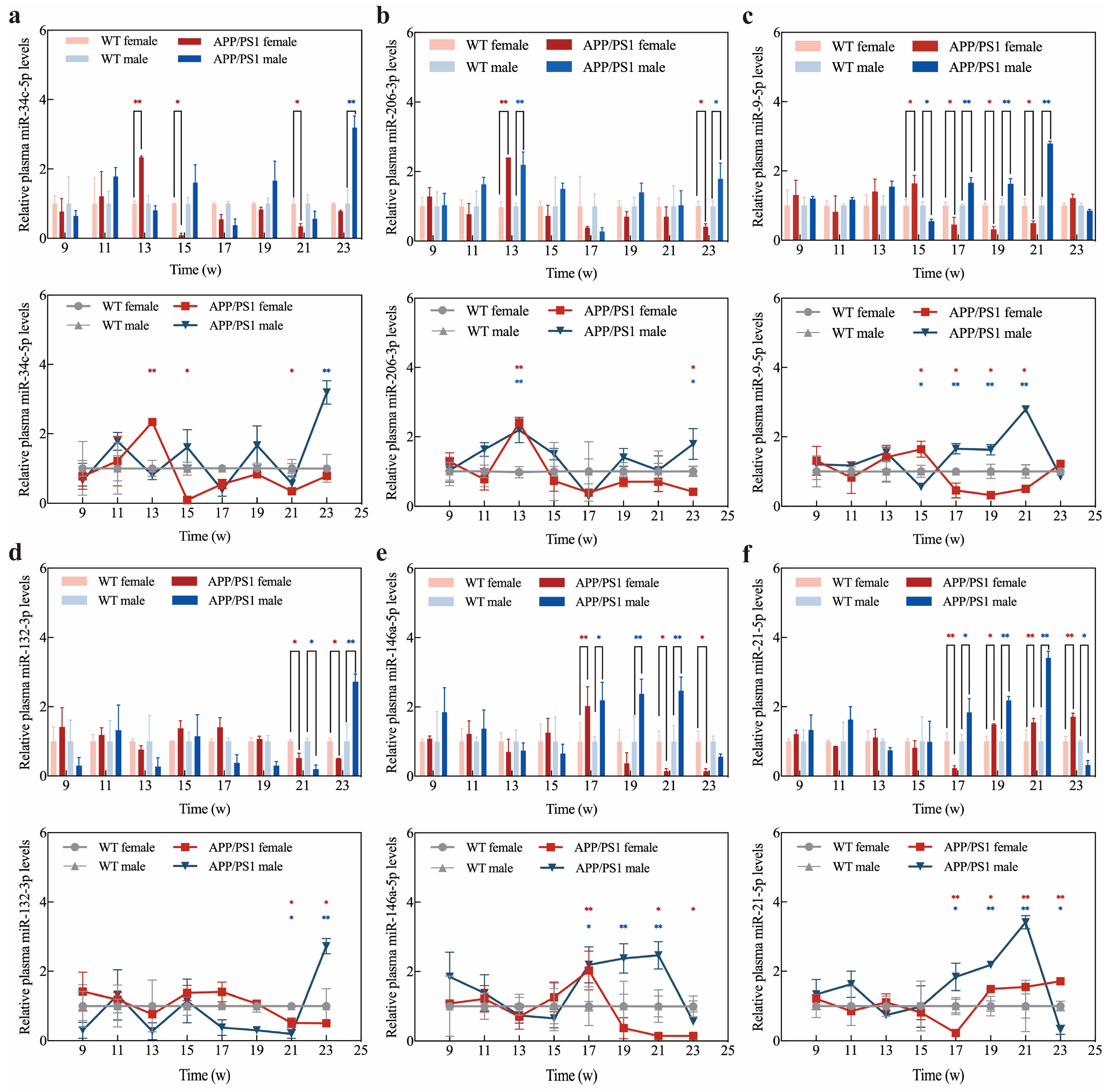
2.6. Application of Exo III-GO-RCA Detection Platform in Early Diagnosis of APP/PS1 Mice
3. Discussion
4. Materials and Methods
4.1. Materials and Apparatus
4.2. Exo III-GO-RCA Reactions
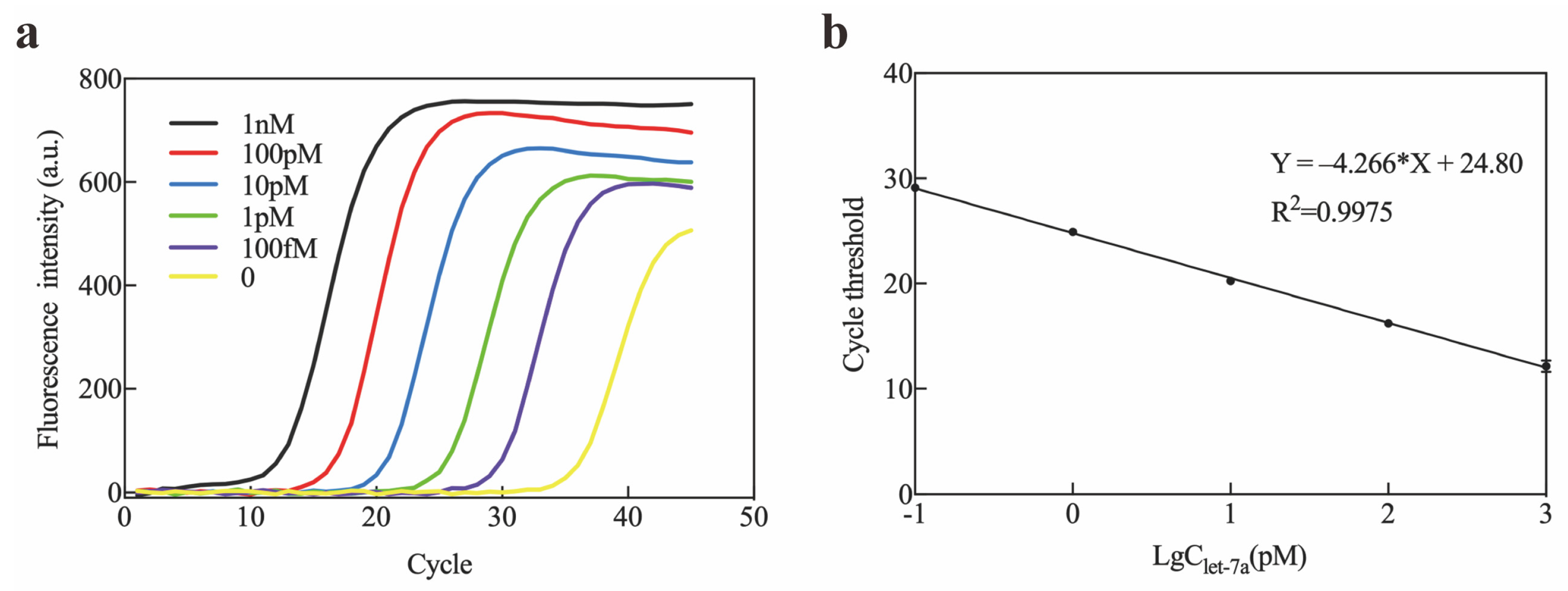
4.3. Mice Plasma miRNA Extraction and qRT-PCR Detection
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Scheltens, P.; De Strooper, B.; Kivipelto, M.; Holstege, H.; Chételat, G.; Teunissen, C.E.; Cummings, J.; van der Flier, W.M. Alzheimer’s disease. Lancet 2021, 397, 1577–1590. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Jin, H.; Xue, Y.-H.; Chen, Q.; Yao, S.-Y.; Du, M.-Q.; Liu, S. Current and future therapeutic strategies for Alzheimer’s disease: An overview of drug development bottlenecks. Front. Aging Neurosci. 2023, 15, 1206572. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chen, J.; Sen, S. MicroRNA as Biomarkers and Diagnostics. J. Cell. Physiol. 2016, 231, 25–30. [Google Scholar] [CrossRef]
- Burgos, K.; Malenica, I.; Metpally, R.; Courtright, A.; Rakela, B.; Beach, T.; Shill, H.; Adler, C.; Sabbagh, M.; Villa, S.; et al. Profiles of extracellular miRNA in cerebrospinal fluid and serum from patients with Alzheimer’s and Parkinson’s diseases correlate with disease status and features of pathology. PLoS ONE 2014, 9, e94839. [Google Scholar] [CrossRef]
- Lukiw, W.J.; Vergallo, A.; Lista, S.; Hampel, H.; Zhao, Y. Biomarkers for Alzheimer’s Disease (AD) and the Application of Precision Medicine. J. Pers. Med. 2020, 10, 138. [Google Scholar] [CrossRef] [PubMed]
- Hou, L.; Li, Q.; Jiang, L.; Qiu, H.; Geng, C.; Hong, J.-S.; Li, H.; Wang, Q. Hypertension and Diagnosis of Parkinson’s Disease: A Meta-Analysis of Cohort Studies. Front. Neurol. 2018, 9, 162. [Google Scholar] [CrossRef]
- Dobricic, V.; Schilling, M.; Schulz, J.; Zhu, L.-S.; Zhou, C.-W.; Fuß, J.; Franzenburg, S.; Zhu, L.-Q.; Parkkinen, L.; Lill, C.M.; et al. Differential microRNA expression analyses across two brain regions in Alzheimer’s disease. Transl. Psychiatry 2022, 12, 352. [Google Scholar] [CrossRef]
- Yoon, S.; Kim, S.E.; Ko, Y.; Jeong, G.H.; Lee, K.H.; Lee, J.; Solmi, M.; Jacob, L.; Smith, L.; Stickley, A.; et al. Differential expression of MicroRNAs in Alzheimer’s disease: A systematic review and meta-analysis. Mol. Psychiatry 2022, 27, 2405–2413. [Google Scholar] [CrossRef]
- Jain, G.; Stuendl, A.; Rao, P.; Berulava, T.; Pena Centeno, T.; Kaurani, L.; Burkhardt, S.; Delalle, I.; Kornhuber, J.; Hüll, M.; et al. A combined miRNA-piRNA signature to detect Alzheimer’s disease. Transl. Psychiatry 2019, 9, 250. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.; Gu, H.; Guo, Q.; Liang, S.; Xue, J.; Yao, F.; Liu, X.; Li, F.; Liu, H.; Sun, L.; et al. Profiling of Serum Exosome MiRNA Reveals the Potential of a MiRNA Panel as Diagnostic Biomarker for Alzheimer’s Disease. Mol. Neurobiol. 2021, 58, 3084–3094. [Google Scholar] [CrossRef]
- Souza, V.C.; Morais, G.S.; Henriques, A.D.; Machado-Silva, W.; Perez, D.I.V.; Brito, C.J.; Camargos, E.F.; Moraes, C.F.; Nóbrega, O.T. Whole-Blood Levels of MicroRNA-9 Are Decreased in Patients With Late-Onset Alzheimer Disease. Am. J. Alzheimer’s Dis. Other Demen. 2020, 35, 1533317520911573. [Google Scholar] [CrossRef] [PubMed]
- Swarbrick, S.; Wragg, N.; Ghosh, S.; Stolzing, A. Systematic Review of miRNA as Biomarkers in Alzheimer’s Disease. Mol. Neurobiol. 2019, 56, 6156–6167. [Google Scholar] [CrossRef] [PubMed]
- Mallick, B.; Ghosh, Z. A complex crosstalk between polymorphic microRNA target sites and AD prognosis. RNA Biol. 2011, 8, 665–673. [Google Scholar] [CrossRef] [PubMed]
- Leidinger, P.; Backes, C.; Deutscher, S.; Schmitt, K.; Mueller, S.C.; Frese, K.; Haas, J.; Ruprecht, K.; Paul, F.; Stähler, C.; et al. A blood based 12-miRNA signature of Alzheimer disease patients. Genome Biol. 2013, 14, R78. [Google Scholar] [CrossRef]
- Fan, D.-Y.; Jian, J.-M.; Huang, S.; Li, W.-W.; Shen, Y.-Y.; Wang, Z.; Zeng, G.-H.; Yi, X.; Jin, W.-S.; Liu, Y.-H.; et al. Establishment of combined diagnostic models of Alzheimer’s disease in a Chinese cohort: The Chongqing Ageing & Dementia Study (CADS). Transl. Psychiatry 2022, 12, 252. [Google Scholar] [CrossRef]
- Liu, K.; Tong, H.; Li, T.; Wang, X.; Chen, Y. Research progress in molecular biology related quantitative methods of MicroRNA. Am. J. Transl. Res. 2020, 12, 3198–3211. [Google Scholar]
- Dong, H.; Lei, J.; Ding, L.; Wen, Y.; Ju, H.; Zhang, X. MicroRNA: Function, detection, and bioanalysis. Chem. Rev. 2013, 113, 6207–6233. [Google Scholar] [CrossRef]
- Hunt, E.A.; Broyles, D.; Head, T.; Deo, S.K. MicroRNA Detection: Current Technology and Research Strategies. Annu. Rev. Anal. Chem. 2015, 8, 217–237. [Google Scholar] [CrossRef]
- Chandrasekaran, A.R.; MacIsaac, M.; Dey, P.; Levchenko, O.; Zhou, L.; Andres, M.; Dey, B.K.; Halvorsen, K. Cellular microRNA detection with miRacles: microRNA-activated conditional looping of engineered switches. Sci. Adv. 2019, 5, eaau9443. [Google Scholar] [CrossRef]
- Pitikultham, P.; Putnin, T.; Pimalai, D.; Sathirapongsasuti, N.; Kitiyakara, C.; Jiang, Q.; Ding, B.; Japrung, D. Ultrasensitive Detection of MicroRNA in Human Saliva via Rolling Circle Amplification Using a DNA-Decorated Graphene Oxide Sensor. ACS Omega 2023, 8, 15266–15275. [Google Scholar] [CrossRef]
- Jiang, W.; Chen, Z.; Lu, J.; Ren, X.; Ma, Y. Ultrasensitive visual detection of miRNA-143 using a CRISPR/Cas12a-based platform coupled with hyperbranched rolling circle amplification. Talanta 2022, 251, 123784. [Google Scholar] [CrossRef] [PubMed]
- Ge, J.; Hu, Y.; Deng, R.; Li, Z.; Zhang, K.; Shi, M.; Yang, D.; Cai, R.; Tan, W. Highly Sensitive MicroRNA Detection by Coupling Nicking-Enhanced Rolling Circle Amplification with MoS Quantum Dots. Anal. Chem. 2020, 92, 13588–13594. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.M.; Li, F.; Zhang, Z.; Zhang, K.; Kang, D.-K.; Ankrum, J.A.; Le, X.C.; Zhao, W. Rolling circle amplification: A versatile tool for chemical biology, materials science and medicine. Chem. Soc. Rev. 2014, 43, 3324–3341. [Google Scholar] [CrossRef]
- Sun, X.; Wang, Y.; Zhang, L.; Liu, S.; Zhang, M.; Wang, J.; Ning, B.; Peng, Y.; He, J.; Hu, Y.; et al. CRISPR-Cas9 Triggered Two-Step Isothermal Amplification Method for E. coli O157:H7 Detection Based on a Metal-Organic Framework Platform. Anal. Chem. 2020, 92, 3032–3041. [Google Scholar] [CrossRef] [PubMed]
- Fan, T.; Mao, Y.; Sun, Q.; Liu, F.; Lin, J.-S.; Liu, Y.; Cui, J.; Jiang, Y. Branched rolling circle amplification method for measuring serum circulating microRNA levels for early breast cancer detection. Cancer Sci. 2018, 109, 2897–2906. [Google Scholar] [CrossRef] [PubMed]
- Tian, B.; Fock, J.; Minero, G.A.S.; Garbarino, F.; Hansen, M.F. Ultrasensitive Real-Time Rolling Circle Amplification Detection Enhanced by Nicking-Induced Tandem-Acting Polymerases. Anal. Chem. 2019, 91, 10102–10109. [Google Scholar] [CrossRef]
- Sung, S.-Y.; Su, Y.-L.; Cheng, W.; Hu, P.-F.; Chiang, C.-S.; Chen, W.-T.; Hu, S.-H. Graphene Quantum Dots-Mediated Theranostic Penetrative Delivery of Drug and Photolytics in Deep Tumors by Targeted Biomimetic Nanosponges. Nano. Lett. 2019, 19, 69–81. [Google Scholar] [CrossRef]
- Liu, B.; Sun, Z.; Zhang, X.; Liu, J. Mechanisms of DNA sensing on graphene oxide. Anal. Chem. 2013, 85, 7987–7993. [Google Scholar] [CrossRef]
- Liu, X.; Wu, Y.; Wu, X.; Zhao, J.X. A graphene oxide-based fluorescence assay for the sensitive detection of DNA exonuclease enzymatic activity. Analyst 2019, 144, 6231–6239. [Google Scholar] [CrossRef]
- Iwe, I.; Li, Z.; Huang, J. Graphene oxide and enzyme-assisted dual-cycling amplification method for sensitive fluorometric determination of DNA. Mikrochim. Acta 2019, 186, 716. [Google Scholar] [CrossRef]
- Li, M.; Xu, X.; Cai, Q.; Luo, X.; Zhou, Z.; Xu, G.; Xie, Y. Graphene oxide-based fluorometric determination of microRNA-141 using rolling circle amplification and exonuclease III-aided recycling amplification. Microchim. Acta 2019, 186, 531. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Lou, Z. Ultrasensitive Detection of Nasopharyngeal Carcinoma-Related MiRNA through Garland Rolling Circle Amplification Integrated Catalytic Hairpin Assembly. ACS Omega 2021, 6, 6460–6465. [Google Scholar] [CrossRef] [PubMed]
- Apiwat, C.; Luksirikul, P.; Kankla, P.; Pongprayoon, P.; Treerattrakoon, K.; Paiboonsukwong, K.; Fucharoen, S.; Dharakul, T.; Japrung, D. Graphene based aptasensor for glycated albumin in diabetes mellitus diagnosis and monitoring. Biosens. Bioelectron. 2016, 82, 140–145. [Google Scholar] [CrossRef]
- Putnin, T.; Waiwinya, W.; Pimalai, D.; Chawjiraphan, W.; Sathirapongsasuti, N.; Japrung, D. Dual sensitive and rapid detection of glycated human serum albumin using a versatile lead/graphene nanocomposite probe as a fluorescence-electrochemical aptasensor. Analyst 2021, 146, 4357–4364. [Google Scholar] [CrossRef] [PubMed]
- Xi, Q.; Zhou, D.-M.; Kan, Y.-Y.; Ge, J.; Wu, Z.-K.; Yu, R.-Q.; Jiang, J.-H. Highly sensitive and selective strategy for microRNA detection based on WS2 nanosheet mediated fluorescence quenching and duplex-specific nuclease signal amplification. Anal. Chem. 2014, 86, 1361–1365. [Google Scholar] [CrossRef] [PubMed]
- Mao, S.; Wu, J.; Yan, J.; Zhang, W.; Zhu, F. Dysregulation of miR-146a: A causative factor in epilepsy pathogenesis, diagnosis, and prognosis. Front. Neurol. 2023, 14, 1094709. [Google Scholar] [CrossRef]
- Boccardi, V.; Poli, G.; Cecchetti, R.; Bastiani, P.; Scamosci, M.; Febo, M.; Mazzon, E.; Bruscoli, S.; Brancorsini, S.; Mecocci, P. miRNAs and Alzheimer’s Disease: Exploring the Role of Inflammation and Vitamin E in an Old-Age Population. Nutrients 2023, 15, 634. [Google Scholar] [CrossRef]
- Santos, F.; Cabreira, V.; Rocha, S.; Massano, J. Blood Biomarkers for the Diagnosis of Neurodegenerative Dementia: A Systematic Review. J. Geriatr. Psychiatry Neurol. 2022, 36, 267–281. [Google Scholar] [CrossRef]
- Wang, W.-X.; Huang, Q.; Hu, Y.; Stromberg, A.J.; Nelson, P.T. Patterns of microRNA expression in normal and early Alzheimer’s disease human temporal cortex: White matter versus gray matter. Acta Neuropathol. 2011, 121, 193–205. [Google Scholar] [CrossRef]
- Giuliani, A.; Gaetani, S.; Sorgentoni, G.; Agarbati, S.; Laggetta, M.; Matacchione, G.; Gobbi, M.; Rossi, T.; Galeazzi, R.; Piccinini, G.; et al. Circulating Inflamma-miRs as Potential Biomarkers of Cognitive Impairment in Patients Affected by Alzheimer’s Disease. Front. Aging Neurosci. 2021, 13, 647015. [Google Scholar] [CrossRef]
- Xie, B.; Liu, Z.; Jiang, L.; Liu, W.; Song, M.; Zhang, Q.; Zhang, R.; Cui, D.; Wang, X.; Xu, S. Increased Serum miR-206 Level Predicts Conversion from Amnestic Mild Cognitive Impairment to Alzheimer’s Disease: A 5-Year Follow-up Study. J. Alzheimer’s Dis. 2017, 55, 509–520. [Google Scholar] [CrossRef]
- Shi, Z.; Zhang, K.; Zhou, H.; Jiang, L.; Xie, B.; Wang, R.; Xia, W.; Yin, Y.; Gao, Z.; Cui, D.; et al. Increased miR-34c mediates synaptic deficits by targeting synaptotagmin 1 through ROS-JNK-p53 pathway in Alzheimer’s Disease. Aging Cell 2020, 19, e13125. [Google Scholar] [CrossRef] [PubMed]
- Rafiey, M.; Nosrati, R.; Babaei, P. Protective effect of miR-34c antagomir against STZ-induced memory impairment by targeting mTOR and PSD-95 in the hippocampus of rats. Neurosci. Lett. 2022, 789, 136881. [Google Scholar] [CrossRef] [PubMed]
- Bhatnagar, S.; Chertkow, H.; Schipper, H.M.; Yuan, Z.; Shetty, V.; Jenkins, S.; Jones, T.; Wang, E. Increased microRNA-34c abundance in Alzheimer’s disease circulating blood plasma. Front. Mol. Neurosci. 2014, 7, 2. [Google Scholar] [CrossRef] [PubMed]
- Cha, D.J.; Mengel, D.; Mustapic, M.; Liu, W.; Selkoe, D.J.; Kapogiannis, D.; Galasko, D.; Rissman, R.A.; Bennett, D.A.; Walsh, D.M. miR-212 and miR-132 Are Downregulated in Neurally Derived Plasma Exosomes of Alzheimer’s Patients. Front. Neurosci. 2019, 13, 1208. [Google Scholar] [CrossRef]
- Xie, B.; Zhou, H.; Zhang, R.; Song, M.; Yu, L.; Wang, L.; Liu, Z.; Zhang, Q.; Cui, D.; Wang, X.; et al. Serum miR-206 and miR-132 as Potential Circulating Biomarkers for Mild Cognitive Impairment. J. Alzheimer’s Dis. 2015, 45, 721–731. [Google Scholar] [CrossRef]
- Wang, S.; Jiang, G.; Wang, S. Neuroprotective Role of MiRNA-9 in Neurological Diseases: A Mini Review. Curr. Mol. Med. 2023, 23, 1007–1011. [Google Scholar] [CrossRef]
- Sørensen, S.S.; Nygaard, A.-B.; Christensen, T. miRNA expression profiles in cerebrospinal fluid and blood of patients with Alzheimer’s disease and other types of dementia—An exploratory study. Transl. Neurodegener. 2016, 5, 6. [Google Scholar] [CrossRef]
- Karuppagounder, S.S.; Pinto, J.T.; Xu, H.; Chen, H.-L.; Beal, M.F.; Gibson, G.E. Dietary supplementation with resveratrol reduces plaque pathology in a transgenic model of Alzheimer’s disease. Neurochem. Int. 2009, 54, 111–118. [Google Scholar] [CrossRef]
- Hébert, S.S.; Horré, K.; Nicolaï, L.; Papadopoulou, A.S.; Mandemakers, W.; Silahtaroglu, A.N.; Kauppinen, S.; Delacourte, A.; De Strooper, B. Loss of microRNA cluster miR-29a/b-1 in sporadic Alzheimer’s disease correlates with increased BACE1/beta-secretase expression. Proc. Natl. Acad. Sci. USA 2008, 105, 6415–6420. [Google Scholar] [CrossRef]
- Zia, A.; Sahebdel, F.; Farkhondeh, T.; Ashrafizadeh, M.; Zarrabi, A.; Hushmandi, K.; Samarghandian, S. A review study on the modulation of SIRT1 expression by miRNAs in aging and age-associated diseases. Int. J. Biol. Macromol. 2021, 188, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Zeng, C.; Meng, X.; Mai, D.; Xu, K.; Qu, S. Overexpression of miR-132-3p contributes to neuronal protection in in vitro and in vivo models of Alzheimer’s disease. Behav. Brain Res. 2022, 417, 113584. [Google Scholar] [CrossRef]
- Hernandez-Rapp, J.; Rainone, S.; Goupil, C.; Dorval, V.; Smith, P.Y.; Saint-Pierre, M.; Vallée, M.; Planel, E.; Droit, A.; Calon, F.; et al. microRNA-132/212 deficiency enhances Aβ production and senile plaque deposition in Alzheimer’s disease triple transgenic mice. Sci. Rep. 2016, 6, 30953. [Google Scholar] [CrossRef]
- Sierksma, A.; Lu, A.; Salta, E.; Vanden Eynden, E.; Callaerts-Vegh, Z.; D’Hooge, R.; Blum, D.; Buée, L.; Fiers, M.; De Strooper, B. Deregulation of neuronal miRNAs induced by amyloid-β or TAU pathology. Mol. Neurodegener. 2018, 13, 54. [Google Scholar] [CrossRef]
- Scoyni, F.; Giudice, L.; Väänänen, M.-A.; Downes, N.; Korhonen, P.; Choo, X.Y.; Välimäki, N.-N.; Mäkinen, P.; Korvenlaita, N.; Rozemuller, A.J.; et al. Alzheimer’s disease-induced phagocytic microglia express a specific profile of coding and non-coding RNAs. Alzheimer’s Dement. 2024, 20, 954–974. [Google Scholar] [CrossRef]
- Su, W.; Li, Z.; Jia, Y.; Zhu, Y.; Cai, W.; Wan, P.; Zhang, Y.; Zheng, S.G.; Zhuo, Y. microRNA-21a-5p/PDCD4 axis regulates mesenchymal stem cell-induced neuroprotection in acute glaucoma. J. Mol. Cell Biol. 2017, 9, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Salt, T.E.; Luong, V.; Wood, N.; Cheung, W.; Maass, A.; Ferrari, G.; Russo-Marie, F.; Sillito, A.M.; Cheetham, M.E.; et al. Targeting amyloid-beta in glaucoma treatment. Proc. Natl. Acad. Sci. USA 2007, 104, 13444–13449. [Google Scholar] [CrossRef] [PubMed]
- Yin, Z.; Geng, X.; Zhang, Z.; Wang, Y.; Gao, X. Rhein Relieves Oxidative Stress in an Aβ1-42 Oligomer-Burdened Neuron Model by Activating the SIRT1/PGC-1α-Regulated Mitochondrial Biogenesis. Front. Pharmacol. 2021, 12, 746711. [Google Scholar] [CrossRef]
- Yin, X.; Zhou, Z.; Qiu, Y.; Fan, X.; Zhao, C.; Bao, J.; Liu, C.; Liu, F.; Qian, W. SIRT1 Regulates Tau Expression and Tau Synaptic Pathology. J. Alzheimer’s Dis. 2021, 84, 895–904. [Google Scholar] [CrossRef]
- Kumar, R.; Chaterjee, P.; Sharma, P.K.; Singh, A.K.; Gupta, A.; Gill, K.; Tripathi, M.; Dey, A.B.; Dey, S. Sirtuin1: A promising serum protein marker for early detection of Alzheimer’s disease. PLoS ONE 2013, 8, e61560. [Google Scholar] [CrossRef]



| Added (pM) | Found (pM) | Recovery (%) | RSD (±%) |
|---|---|---|---|
| 100 | 95.5 | 95.5 | 4.28376963 |
| 1 | 1 | 100 | 2.939 |
| 0.1 | 0.1024 | 102.4 | 4.7109375 |
| Method | Target | Samples | LOD | Analysis Time | Ref. |
|---|---|---|---|---|---|
| Nicking-Enhanced RCA with MoS2 QDs | Let-7a | Serum | 4.6 fM | 5.3 h | [22] |
| Branched RCA | miR-21 | Serum | 1 pM | 5 h | [25] |
| AuNP with CRISPR/Cas12a-assisted HRCA | miR-143 | Cell, urine | 100 aM | 2 h | [21] |
| DSNSA with WS2 nanosheet | miR-21 | Cell | 300 fM | 5 h | [35] |
| RCA integrated catalytic hairpin assembly | Let-7b | Serum | 10 pM | 4.3 h | [32] |
| Nicking-Enhanced RCA with GO | miR-21 | Saliva | 1.4 fM | 2.4 h | [20] |
| Exo III-GO-RCA | Let-7a | Serum | 19.35 fM | 4 h | This work |
| Name | Sequence (5′–3′) | Length |
|---|---|---|
| let-7a-5p | UGAGGUAGUAGGUUGUAUAGUU | 22 bp |
| PPlet7a | p-CTACTACTCACCCTAACCCTAACCCTAACCCTAACC CTAACCCTAAAACTATACAAC | 58 bp |
| F-DNA | FAM-CCCTAACCCTAA | 12 bp |
| let-7b | UGAGGUAGUAGGUUGUGUGGUU | 22 bp |
| let-7c | UGAGGUAGUAGGUUGUAUGGUU | 22 bp |
| miR-455-3p | AUGCAGUCCAUGGGCAUAUACAC | 23 bp |
| miR-34c-5p | AGGCAGUGUAGUUAGCUGAUUGC | 23 bp |
| miR-206-3p | UGGAAUGUAAGGAAGUGUGUGG | 22 bp |
| miR-9-5p | UCUUUGGUUAUCUAGCUGUAUGA | 23 bp |
| miR-132-3p | UAACAGUCUACAGCCAUGGUCG | 22 bp |
| miR-146a-5p | UGAGAACUGAAUUCCAUGGGUU | 22 bp |
| miR-21-5p | UAGCUUAUCAGACUGAUGUUGA | 22 bp |
| PP34c | p-ACTACACTGCCTCCCTAACCCTAACCCTAACCCTA ACCCTAACCCTAAGCAATCAGCTA | 59 bp |
| PP206 | p-CTTACATTCCACCCTAACCCTAACCCTAACCCTAA CCCTAACCCTAACCACACACTTC | 58 bp |
| PP9 | p-GATAACCAAAGACCCTAACCCTAACCCTAACCCT AACCCTAACCCTAATCATACAGCTA | 59 bp |
| PP132 | p-GTAGACTGTTACCCTAACCCTAACCCTAACCCTA ACCCTAACCCTAACGACCATGGCT | 58 bp |
| PP146a | p-TTCAGTTCTCACCCTAACCCTAACCCTAACCCTA ACCCTAACCCTAAAACCCATGGAA | 58 bp |
| PP21 | p-TGATAAGCTACCCTAACCCTAACCCTAACCCTAA CCCTAACCCTAATCAACATCAGTC | 58 bp |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, F.; Zhang, N.; Zhang, Y. A New Strategy for Ultrasensitive Detection Based on Target microRNA-Triggered Rolling Circle Amplification in the Early Diagnosis of Alzheimer’s Disease. Int. J. Mol. Sci. 2024, 25, 9490. https://doi.org/10.3390/ijms25179490
Zhao F, Zhang N, Zhang Y. A New Strategy for Ultrasensitive Detection Based on Target microRNA-Triggered Rolling Circle Amplification in the Early Diagnosis of Alzheimer’s Disease. International Journal of Molecular Sciences. 2024; 25(17):9490. https://doi.org/10.3390/ijms25179490
Chicago/Turabian StyleZhao, Fei, Na Zhang, and Yi Zhang. 2024. "A New Strategy for Ultrasensitive Detection Based on Target microRNA-Triggered Rolling Circle Amplification in the Early Diagnosis of Alzheimer’s Disease" International Journal of Molecular Sciences 25, no. 17: 9490. https://doi.org/10.3390/ijms25179490
APA StyleZhao, F., Zhang, N., & Zhang, Y. (2024). A New Strategy for Ultrasensitive Detection Based on Target microRNA-Triggered Rolling Circle Amplification in the Early Diagnosis of Alzheimer’s Disease. International Journal of Molecular Sciences, 25(17), 9490. https://doi.org/10.3390/ijms25179490







