Pathological Involvement of Protein Phase Separation and Aggregation in Neurodegenerative Diseases
Abstract
1. Introduction
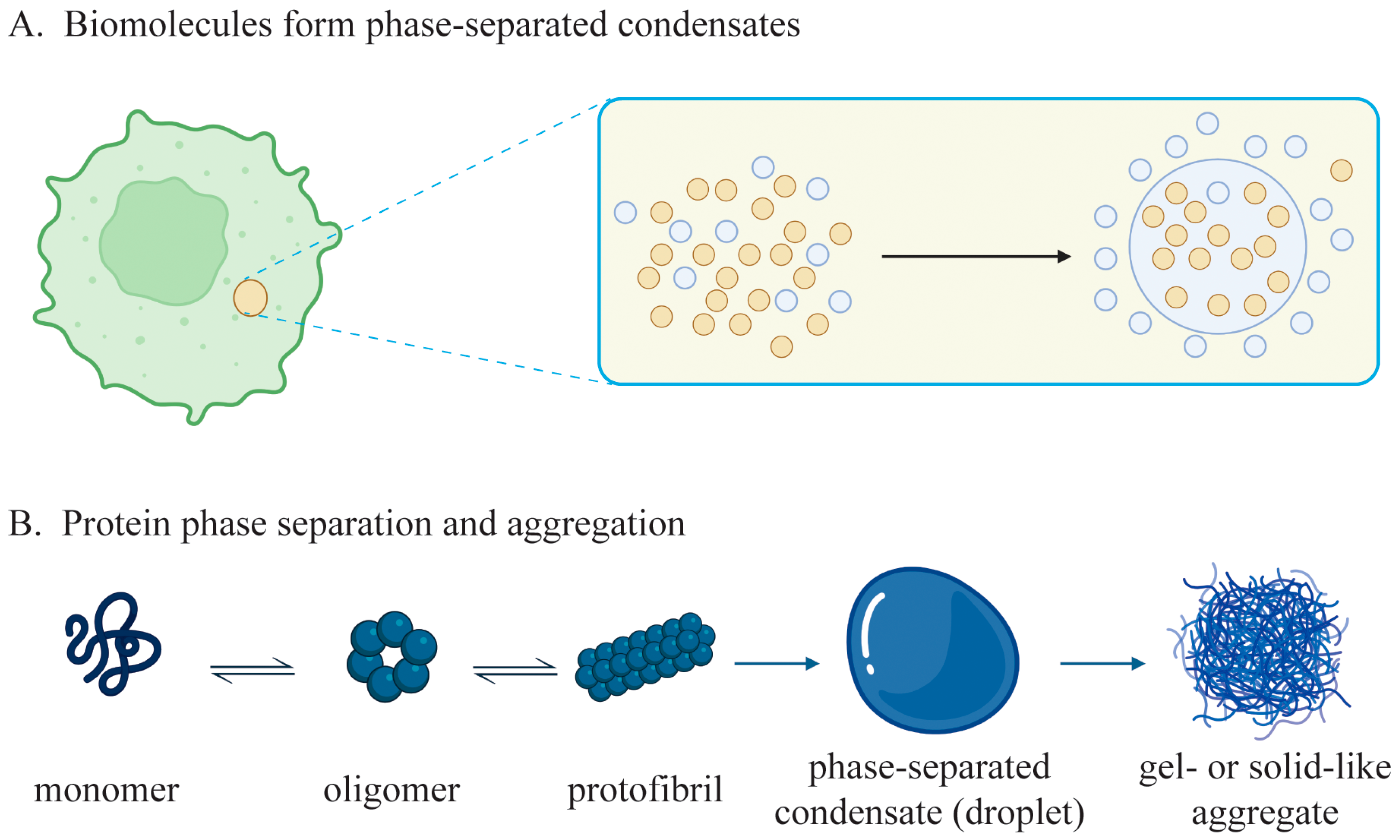
2. Overview of Biomolecular Phase Separation and Aggregation
2.1. Definition and Classification of Phase Separation and Aggregation
2.2. Association between Phase Separation and Neurodegenerative Diseases
3. Types and Characteristics of Common Neurodegenerative Diseases
3.1. Protein Phase Separation and Aggregation in AD

3.1.1. Effects of Gene Mutations on Protein Phase Separation and Aggregation in AD
3.1.2. Effects of PTMs on Protein Phase Separation and Aggregation in AD
| Proteins | PTMs | Modified Sites | Regulators | Effects on Aggregation | References |
|---|---|---|---|---|---|
| Aβ (or APP) | Phosphorylation | Thr668, Thr686, Thr654, Ser655, Ser675, Tyr653, Tyr682, Tyr687 | GSK-3β, CDK5, ERK1, JNK, ROCK1 | ↑ | [59] |
| ROCK2, BACE1, PIN1, heparin | ↓ | ||||
| Acetylation | Lys132, Lys134 | SIRT1 | ↓ | [60,61] | |
| SIRT2 | ↑ | ||||
| Ubiquitylation | Unidentified | UCHL1 | ↓ | [62] | |
| Glycosylation | Thr291, Thr292, Ser295, Thr296, Thr576, Thr577, Thr584 | GalNAc-T2 | ↓ | [63,64] | |
| GalNAc-T3 | ↑ | ||||
| tau | Phosphorylation | Thr181, Thr212, Ser404, Ser202, Thr231, Thr320 | PKA, GSK-3β, Pin1 | ↑ | [65,66] |
| PP1 | ↓ | ||||
| Acetylation | Lys280, Lys174 | EP300 | ↑ | [67,68] | |
| Ubiquitination | Lys 48, Lys63, Lys257, Lys259, Lys274, Lys281, Lys321, Lys375, Lys385 | CHIP | ↑ | [69] | |
| TH006 | ↓ | ||||
| α-syn | Phosphorylation | Tyr39, Tyr125, Ser129, Tyr133, Tyr136 | CK1, CK2, PLK2 | ↑ | [70,71,72,73] |
| Ser87 | CK1 | ↓ | [74] | ||
| Ubiquitylation | Met1, Lys6, Lys10, Lys12, Lys21, Lys23, Lys32, Lys34, Lys43, Lys45, Lys96, Lys102 | SIAH | ↑ | [75,76,77] | |
| CHIP | ↓ | [78] | |||
| Glycosylation | Thr33, Thr59, Thr64, Thr72, Thr75, Thr81, Ser87 | OGT | ↓ | [79] | |
| Sumoylation | Lys96, Lys102 | HPC2, PIAS2, TRIM28 | ↑↓ | [80,81,82] | |
| HTT | Phosphorylation | Ser13, Ser16 | TBK1 | ↓ | [83] |
| Methylation | Arg200, Arg205 | PRMT4 and PRMT6 | ↓ | [84] | |
| Ubiquitylation | Unidentified | Ube3a | ↓ | [85] | |
| SOD1 | Phosphorylation | Thr2 | ATM/CHK2, ATR/CHK1 | ↓ | [86,87] |
| Acetylation | Unidentified | HDAC6 | ↓ | ||
| Sumoylation | Lys75 | SUMO1, SUMO3 | ↑ | ||
| TDP-43 | Phosphorylation | Ser48, Ser379. Ser403, Ser404, Ser409, Ser410 | CK1, CK2, CDC7, TTBK1, TTBK2 | ↑ | [88] |
| Ubiquitination | Unidentified | Parkin | ↑ | [89] | |
| FUS | Phosphorylation | Tyr526 | Abl | ↑ | [90] |
| Acetylation | Lys510 | HDACs | ↓ | [91] |
3.2. Protein Phase Separation and Aggregation in PD
3.2.1. Effects of Gene Mutations on Protein Phase Separation and Aggregation in PD
3.2.2. Effects of PTMs on Protein Phase Separation and Aggregation in PD
3.3. Protein Phase Separation and Aggregation in HD
3.3.1. Effects of Gene Mutations on Protein Phase Separation and Aggregation in HD
3.3.2. Effects of PTMs on Protein Phase Separation and Aggregation in HD
3.4. Protein Phase Separation and Aggregation in ALS
3.4.1. Effects of Gene Mutations on Protein Phase Separation and Aggregation in ALS
3.4.2. Effects of PTMs on Protein Phase Separation and Aggregation in ALS
3.5. Protein Phase Separation and Aggregation in AMD
3.5.1. Effects of Genetic Variations on Protein Phase Separation and Aggregation in AMD
3.5.2. Effects of Dysregulated Metabolism on Protein Phase Separation and Aggregation in AMD
3.6. Protein Phase Separation and Aggregation in DR
3.7. Contributions of RNA Binding Proteins to Neurodegenerative Diseases
4. Techniques to Study Protein Phase Separation and Aggregation
5. Treatments of Neurodegenerative Diseases by Reducing Protein Aggregation
5.1. Targeting the PTMs of Amyloidogenic Proteins
5.2. Targeting the Genes Encoding Amyloidogenic Proteins
5.3. Other Emerging Therapeutic Approaches
6. Conclusions and Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- GBD Nervous System Disorders Collaborators. Global, regional, and national burden of disorders affecting the nervous system, 1990–2021: A systematic analysis for the Global Burden of Disease Study 2021. Lancet Neurol. 2024, 23, 344–381. [Google Scholar] [CrossRef] [PubMed]
- Ding, C.; Wu, Y.; Chen, X.; Chen, Y.; Wu, Z.; Lin, Z.; Kang, D.; Fang, W.; Chen, F. Global, regional, and national burden and attributable risk factors of neurological disorders: The Global Burden of Disease study 1990–2019. Front. Public Health 2022, 10, 952161. [Google Scholar] [CrossRef]
- Huang, Y.; Li, Y.; Pan, H.; Han, L. Global, regional, and national burden of neurological disorders in 204 countries and territories worldwide. J. Glob. Health 2023, 13, 04160. [Google Scholar] [CrossRef] [PubMed]
- Wilson, D.M., 3rd; Cookson, M.R.; Van Den Bosch, L.; Zetterberg, H.; Holtzman, D.M.; Dewachter, I. Hallmarks of neurodegenerative diseases. Cell 2023, 186, 693–714. [Google Scholar] [CrossRef] [PubMed]
- Calabrese, G.; Molzahn, C.; Mayor, T. Protein interaction networks in neurodegenerative diseases: From physiological function to aggregation. J. Biol. Chem. 2022, 298, 102062. [Google Scholar] [CrossRef]
- Das, S.; Zhang, Z.; Ang, L.C. Clinicopathological overlap of neurodegenerative diseases: A comprehensive review. J. Clin. Neurosci. 2020, 78, 30–33. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Wang, S.; Wang, W.; Shi, J.; Stovall, D.B.; Li, D.; Sui, G. Phase-Separated Subcellular Compartmentation and Related Human Diseases. Int. J. Mol. Sci. 2022, 23, 5491. [Google Scholar] [CrossRef]
- Alberti, S.; Hyman, A.A. Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat. Rev. Mol. Cell Biol. 2021, 22, 196–213. [Google Scholar] [CrossRef]
- Vendruscolo, M.; Fuxreiter, M. Protein condensation diseases: Therapeutic opportunities. Nat. Commun. 2022, 13, 5550. [Google Scholar] [CrossRef]
- Nedelsky, N.B.; Taylor, J.P. Bridging biophysics and neurology: Aberrant phase transitions in neurodegenerative disease. Nat. Rev. Neurol. 2019, 15, 272–286. [Google Scholar] [CrossRef]
- Wu, X.; Cai, Q.; Feng, Z.; Zhang, M. Liquid-Liquid Phase Separation in Neuronal Development and Synaptic Signaling. Dev. Cell 2020, 55, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Hurtle, B.T.; Xie, L.; Donnelly, C.J. Disrupting pathologic phase transitions in neurodegeneration. J. Clin. Investig. 2023, 133, e168549. [Google Scholar] [CrossRef]
- Mathieu, C.; Pappu, R.V.; Taylor, J.P. Beyond aggregation: Pathological phase transitions in neurodegenerative disease. Science 2020, 370, 56–60. [Google Scholar] [CrossRef]
- Wen, J.H.; He, X.H.; Feng, Z.S.; Li, D.Y.; Tang, J.X.; Liu, H.F. Cellular Protein Aggregates: Formation, Biological Effects, and Ways of Elimination. Int. J. Mol. Sci. 2023, 24, 8593. [Google Scholar] [CrossRef]
- Chung, C.I.; Yang, J.; Shu, X. Chemogenetic Minitool for Dissecting the Roles of Protein Phase Separation. ACS Cent. Sci. 2023, 9, 1466–1479. [Google Scholar] [CrossRef] [PubMed]
- Bashir, S.; Aiman, A.; Chaudhary, A.A.; Khan, N.; Ahanger, I.A.; Sami, N.; Almugri, E.A.; Ali, M.A.M.; Khan, S.U.; Shahid, M.; et al. Probing protein aggregation through spectroscopic insights and multimodal approaches: A comprehensive review for counteracting neurodegenerative disorders. Heliyon 2024, 10, e27949. [Google Scholar] [CrossRef]
- Lopez-Laguna, H.; Sanchez, J.; Unzueta, U.; Mangues, R.; Vazquez, E.; Villaverde, A. Divalent Cations: A Molecular Glue for Protein Materials. Trends Biochem. Sci. 2020, 45, 992–1003. [Google Scholar] [CrossRef] [PubMed]
- Chaudhary, H.; Iashchishyn, I.A.; Romanova, N.V.; Rambaran, M.A.; Musteikyte, G.; Smirnovas, V.; Holmboe, M.; Ohlin, C.A.; Svedruzic, Z.M.; Morozova-Roche, L.A. Polyoxometalates as Effective Nano-inhibitors of Amyloid Aggregation of Pro-inflammatory S100A9 Protein Involved in Neurodegenerative Diseases. ACS Appl. Mater. Interfaces 2021, 13, 26721–26734. [Google Scholar] [CrossRef]
- Carapeto, A.P.; Marcuello, C.; Faísca, P.F.N.; Rodrigues, M.S. Morphological and Biophysical Study of S100A9 Protein Fibrils by Atomic Force Microscopy Imaging and Nanomechanical Analysis. Biomolecules 2024, 14, 1091. [Google Scholar] [CrossRef]
- Sanders, E.; Csondor, R.; Sulskis, D.; Baronaite, I.; Smirnovas, V.; Maheswaran, L.; Horrocks, J.; Munro, R.; Georgiadou, C.; Horvath, I.; et al. The Stabilization of S100A9 Structure by Calcium Inhibits the Formation of Amyloid Fibrils. Int. J. Mol. Sci. 2023, 24, 13200. [Google Scholar] [CrossRef]
- Han, J.Y.; Choi, T.S.; Kim, H.I. Molecular Role of Ca2+ and Hard Divalent Metal Cations on Accelerated Fibrillation and Interfibrillar Aggregation of alpha-Synuclein. Sci. Rep. 2018, 8, 1895. [Google Scholar] [CrossRef]
- Cuny, G.D. Foreword: Neurodegenerative diseases: Challenges and opportunities. Future Med. Chem. 2012, 4, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Citron, M.; Vigo-Pelfrey, C.; Teplow, D.B.; Miller, C.; Schenk, D.; Johnston, J.; Winblad, B.; Venizelos, N.; Lannfelt, L.; Selkoe, D.J. Excessive production of amyloid beta-protein by peripheral cells of symptomatic and presymptomatic patients carrying the Swedish familial Alzheimer disease mutation. Proc. Natl. Acad. Sci. USA 1994, 91, 11993–11997. [Google Scholar] [CrossRef] [PubMed]
- Hur, J.Y. gamma-Secretase in Alzheimer’s disease. Exp. Mol. Med. 2022, 54, 433–446. [Google Scholar] [CrossRef]
- Thijssen, E.H.; La Joie, R.; Strom, A.; Fonseca, C.; Iaccarino, L.; Wolf, A.; Spina, S.; Allen, I.E.; Cobigo, Y.; Heuer, H.; et al. Plasma phosphorylated tau 217 and phosphorylated tau 181 as biomarkers in Alzheimer’s disease and frontotemporal lobar degeneration: A retrospective diagnostic performance study. Lancet Neurol. 2021, 20, 739–752. [Google Scholar] [CrossRef]
- Vazquez-Velez, G.E.; Zoghbi, H.Y. Parkinson’s Disease Genetics and Pathophysiology. Annu. Rev. Neurosci. 2021, 44, 87–108. [Google Scholar] [CrossRef]
- Zhou, Y.; Wang, Y.; Wan, J.; Zhao, Y.; Pan, H.; Zeng, Q.; Zhou, X.; He, R.; Zhou, X.; Xiang, Y.; et al. Mutational spectrum and clinical features of GBA1 variants in a Chinese cohort with Parkinson’s disease. NPJ Parkinsons Dis. 2023, 9, 129. [Google Scholar] [CrossRef]
- Moreira, A.C.; Silva, T.; Mesquita, G.; Gomes, A.C.; Bento, C.M.; Neves, J.V.; Rodrigues, D.F.; Rodrigues, P.N.; Almeida, A.A.; Santambrogio, P.; et al. H-Ferritin Produced by Myeloid Cells Is Released to the Circulation and Plays a Major Role in Liver Iron Distribution during Infection. Int. J. Mol. Sci. 2021, 23, 269. [Google Scholar] [CrossRef]
- Levi, S.; Rovida, E. Neuroferritinopathy: From ferritin structure modification to pathogenetic mechanism. Neurobiol. Dis. 2015, 81, 134–143. [Google Scholar] [CrossRef]
- Cattaneo, E.; Zuccato, C.; Tartari, M. Normal huntingtin function: An alternative approach to Huntington’s disease. Nat. Rev. Neurosci. 2005, 6, 919–930. [Google Scholar] [CrossRef]
- Folger, A.; Chen, C.; Kabbaj, M.H.; Frey, K.; Wang, Y. Neurodegenerative disease-associated inclusion bodies are cleared by selective autophagy in budding yeast. Autophagy Rep. 2023, 2, 2236407. [Google Scholar] [CrossRef] [PubMed]
- Stamenkovic, S.; Ducic, T.; Stamenkovic, V.; Kranz, A.; Andjus, P.R. Imaging of glial cell morphology, SOD1 distribution and elemental composition in the brainstem and hippocampus of the ALS hSOD1(G93A) rat. Neuroscience 2017, 357, 37–55. [Google Scholar] [CrossRef] [PubMed]
- Brettschneider, J.; Del Tredici, K.; Toledo, J.B.; Robinson, J.L.; Irwin, D.J.; Grossman, M.; Suh, E.; Van Deerlin, V.M.; Wood, E.M.; Baek, Y.; et al. Stages of pTDP-43 pathology in amyotrophic lateral sclerosis. Ann. Neurol. 2013, 74, 20–38. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, I.R.; Bigio, E.H.; Ince, P.G.; Geser, F.; Neumann, M.; Cairns, N.J.; Kwong, L.K.; Forman, M.S.; Ravits, J.; Stewart, H.; et al. Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann. Neurol. 2007, 61, 427–434. [Google Scholar] [CrossRef]
- Assoni, A.F.; Foijer, F.; Zatz, M. Amyotrophic Lateral Sclerosis, FUS and Protein Synthesis Defects. Stem. Cell Rev. Rep. 2023, 19, 625–638. [Google Scholar] [CrossRef]
- Mahley, R.W.; Rall, S.C., Jr. Apolipoprotein E: Far more than a lipid transport protein. Annu. Rev. Genom. Hum. Genet. 2000, 1, 507–537. [Google Scholar] [CrossRef]
- McKay, G.J.; Patterson, C.C.; Chakravarthy, U.; Dasari, S.; Klaver, C.C.; Vingerling, J.R.; Ho, L.; de Jong, P.T.; Fletcher, A.E.; Young, I.S.; et al. Evidence of association of APOE with age-related macular degeneration: A pooled analysis of 15 studies. Hum. Mutat. 2011, 32, 1407–1416. [Google Scholar] [CrossRef]
- Kiser, P.D.; Zhang, J.; Badiee, M.; Li, Q.; Shi, W.; Sui, X.; Golczak, M.; Tochtrop, G.P.; Palczewski, K. Catalytic mechanism of a retinoid isomerase essential for vertebrate vision. Nat. Chem. Biol. 2015, 11, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Choi, E.H.; Suh, S.; Sander, C.L.; Hernandez, C.J.O.; Bulman, E.R.; Khadka, N.; Dong, Z.; Shi, W.; Palczewski, K.; Kiser, P.D. Insights into the pathogenesis of dominant retinitis pigmentosa associated with a D477G mutation in RPE65. Hum. Mol. Genet. 2018, 27, 2225–2243. [Google Scholar] [CrossRef]
- Shea, Y.F.; Pan, Y.; Mak, H.K.; Bao, Y.; Lee, S.C.; Chiu, P.K.; Chan, H.F. A systematic review of atypical Alzheimer’s disease including behavioural and psychological symptoms. Psychogeriatrics 2021, 21, 396–406. [Google Scholar] [CrossRef]
- Schneider, A.; Biernat, J.; von Bergen, M.; Mandelkow, E.; Mandelkow, E.M. Phosphorylation that detaches tau protein from microtubules (Ser262, Ser214) also protects it against aggregation into Alzheimer paired helical filaments. Biochemistry 1999, 38, 3549–3558. [Google Scholar] [CrossRef]
- Kopke, E.; Tung, Y.C.; Shaikh, S.; Alonso, A.C.; Iqbal, K.; Grundke-Iqbal, I. Microtubule-associated protein tau. Abnormal phosphorylation of a non-paired helical filament pool in Alzheimer disease. J. Biol. Chem. 1993, 268, 24374–24384. [Google Scholar] [CrossRef] [PubMed]
- Dosztanyi, Z.; Csizmok, V.; Tompa, P.; Simon, I. IUPred: Web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 2005, 21, 3433–3434. [Google Scholar] [CrossRef] [PubMed]
- Goate, A.M. Molecular genetics of Alzheimer’s disease. Geriatrics 1997, 52 (Suppl. S2), S9–S12. [Google Scholar]
- Ryan, N.S.; Nicholas, J.M.; Weston, P.S.J.; Liang, Y.; Lashley, T.; Guerreiro, R.; Adamson, G.; Kenny, J.; Beck, J.; Chavez-Gutierrez, L.; et al. Clinical phenotype and genetic associations in autosomal dominant familial Alzheimer’s disease: A case series. Lancet Neurol. 2016, 15, 1326–1335. [Google Scholar] [CrossRef] [PubMed]
- Lanoiselee, H.M.; Nicolas, G.; Wallon, D.; Rovelet-Lecrux, A.; Lacour, M.; Rousseau, S.; Richard, A.C.; Pasquier, F.; Rollin-Sillaire, A.; Martinaud, O.; et al. APP, PSEN1, and PSEN2 mutations in early-onset Alzheimer disease: A genetic screening study of familial and sporadic cases. PLoS Med. 2017, 14, e1002270. [Google Scholar] [CrossRef]
- Scheuner, D.; Eckman, C.; Jensen, M.; Song, X.; Citron, M.; Suzuki, N.; Bird, T.D.; Hardy, J.; Hutton, M.; Kukull, W.; et al. Secreted amyloid beta-protein similar to that in the senile plaques of Alzheimer’s disease is increased in vivo by the presenilin 1 and 2 and APP mutations linked to familial Alzheimer’s disease. Nat. Med. 1996, 2, 864–870. [Google Scholar] [CrossRef]
- Goode, B.L.; Chau, M.; Denis, P.E.; Feinstein, S.C. Structural and functional differences between 3-repeat and 4-repeat tau isoforms. Implications for normal tau function and the onset of neurodegenetative disease. J. Biol. Chem. 2000, 275, 38182–38189. [Google Scholar] [CrossRef]
- Lu, M.; Kosik, K.S. Competition for microtubule-binding with dual expression of tau missense and splice isoforms. Mol. Biol. Cell 2001, 12, 171–184. [Google Scholar] [CrossRef]
- Strang, K.H.; Golde, T.E.; Giasson, B.I. MAPT mutations, tauopathy, and mechanisms of neurodegeneration. Lab. Investig. 2019, 99, 912–928. [Google Scholar] [CrossRef]
- Ramesh, M.; Gopinath, P.; Govindaraju, T. Role of Post-translational Modifications in Alzheimer’s Disease. Chembiochem 2020, 21, 1052–1079. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.Y.; Clark, A.W.; Rosales, J.L.; Chapman, K.; Fung, T.; Johnston, R.N. Elevated neuronal Cdc2-like kinase activity in the Alzheimer disease brain. Neurosci. Res. 1999, 34, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Pei, J.J.; Grundke-Iqbal, I.; Iqbal, K.; Bogdanovic, N.; Winblad, B.; Cowburn, R.F. Accumulation of cyclin-dependent kinase 5 (cdk5) in neurons with early stages of Alzheimer’s disease neurofibrillary degeneration. Brain Res. 1998, 797, 267–277. [Google Scholar] [CrossRef] [PubMed]
- Pei, J.J.; Braak, E.; Braak, H.; Grundke-Iqbal, I.; Iqbal, K.; Winblad, B.; Cowburn, R.F. Distribution of active glycogen synthase kinase 3beta (GSK-3beta) in brains staged for Alzheimer disease neurofibrillary changes. J. Neuropathol. Exp. Neurol. 1999, 58, 1010–1019. [Google Scholar] [CrossRef] [PubMed]
- Vieira, S.I.; Rebelo, S.; Esselmann, H.; Wiltfang, J.; Lah, J.; Lane, R.; Small, S.A.; Gandy, S.; da Cruz, E.S.E.F.; da Cruz, E.S.O.A. Retrieval of the Alzheimer’s amyloid precursor protein from the endosome to the TGN is S655 phosphorylation state-dependent and retromer-mediated. Mol. Neurodegener. 2010, 5, 40. [Google Scholar] [CrossRef]
- Wen, W.; Li, P.; Liu, P.; Xu, S.; Wang, F.; Huang, J.H. Post-Translational Modifications of BACE1 in Alzheimer’s Disease. Curr. Neuropharmacol. 2022, 20, 211–222. [Google Scholar] [CrossRef]
- Song, W.J.; Son, M.Y.; Lee, H.W.; Seo, H.; Kim, J.H.; Chung, S.H. Enhancement of BACE1 Activity by p25/Cdk5-Mediated Phosphorylation in Alzheimer’s Disease. PLoS ONE 2015, 10, e0136950. [Google Scholar] [CrossRef]
- Oliveira, J.M.; Henriques, A.G.; Martins, F.; Rebelo, S.; da Cruz e Silva, O.A. Amyloid-beta Modulates Both AbetaPP and Tau Phosphorylation. J. Alzheimers Dis. 2015, 45, 495–507. [Google Scholar] [CrossRef]
- Zhang, T.; Chen, D.; Lee, T.H. Phosphorylation Signaling in APP Processing in Alzheimer’s Disease. Int. J. Mol. Sci. 2019, 21, 209. [Google Scholar] [CrossRef]
- Bai, N.; Li, N.; Cheng, R.; Guan, Y.; Zhao, X.; Song, Z.; Xu, H.; Yi, F.; Jiang, B.; Li, X.; et al. Inhibition of SIRT2 promotes APP acetylation and ameliorates cognitive impairment in APP/PS1 transgenic mice. Cell Rep. 2022, 40, 111062. [Google Scholar] [CrossRef]
- Li, N.; Bai, N.; Zhao, X.; Cheng, R.; Wu, X.; Jiang, B.; Li, X.; Xue, M.; Xu, H.; Guo, Q.; et al. Cooperative effects of SIRT1 and SIRT2 on APP acetylation. Aging Cell 2023, 22, e13967. [Google Scholar] [CrossRef] [PubMed]
- Maniv, I.; Sarji, M.; Bdarneh, A.; Feldman, A.; Ankawa, R.; Koren, E.; Magid-Gold, I.; Reis, N.; Soteriou, D.; Salomon-Zimri, S.; et al. Altered ubiquitin signaling induces Alzheimer’s disease-like hallmarks in a three-dimensional human neural cell culture model. Nat. Commun. 2023, 14, 5922. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Ku, X.; Zou, X.; Hou, J.; Yan, W.; Zhang, Y. Comprehensive analysis of O-glycosylation of amyloid precursor protein (APP) using targeted and multi-fragmentation MS strategy. Biochim. Biophys. Acta Gen. Subj. 2021, 1865, 129954. [Google Scholar] [CrossRef]
- Tachida, Y.; Iijima, J.; Takahashi, K.; Suzuki, H.; Kizuka, Y.; Yamaguchi, Y.; Tanaka, K.; Nakano, M.; Takakura, D.; Kawasaki, N.; et al. O-GalNAc glycosylation determines intracellular trafficking of APP and Abeta production. J. Biol. Chem. 2023, 299, 104905. [Google Scholar] [CrossRef]
- Abasi, L.S.; Elathram, N.; Movva, M.; Deep, A.; Corbett, K.D.; Debelouchina, G.T. Phosphorylation regulates tau’s phase separation behavior and interactions with chromatin. Commun. Biol. 2024, 7, 251. [Google Scholar] [CrossRef]
- Noble, W.; Hanger, D.P.; Miller, C.C.; Lovestone, S. The importance of tau phosphorylation for neurodegenerative diseases. Front. Neurol. 2013, 4, 83. [Google Scholar] [CrossRef]
- Cohen, T.J.; Guo, J.L.; Hurtado, D.E.; Kwong, L.K.; Mills, I.P.; Trojanowski, J.Q.; Lee, V.M. The acetylation of tau inhibits its function and promotes pathological tau aggregation. Nat. Commun. 2011, 2, 252. [Google Scholar] [CrossRef] [PubMed]
- Min, S.W.; Chen, X.; Tracy, T.E.; Li, Y.; Zhou, Y.; Wang, C.; Shirakawa, K.; Minami, S.S.; Defensor, E.; Mok, S.A.; et al. Critical role of acetylation in tau-mediated neurodegeneration and cognitive deficits. Nat. Med. 2015, 21, 1154–1162. [Google Scholar] [CrossRef]
- Plass, P. Home-care services: How many can they help? Health Soc. Work 1978, 3, 182–189. [Google Scholar] [CrossRef]
- Zhao, K.; Lim, Y.J.; Liu, Z.; Long, H.; Sun, Y.; Hu, J.J.; Zhao, C.; Tao, Y.; Zhang, X.; Li, D.; et al. Parkinson’s disease-related phosphorylation at Tyr39 rearranges alpha-synuclein amyloid fibril structure revealed by cryo-EM. Proc. Natl. Acad. Sci. USA 2020, 117, 20305–20315. [Google Scholar] [CrossRef]
- Olteanu, A.; Pielak, G.J. Peroxidative aggregation of alpha-synuclein requires tyrosines. Protein Sci. 2004, 13, 2852–3856. [Google Scholar] [CrossRef] [PubMed]
- Okochi, M.; Walter, J.; Koyama, A.; Nakajo, S.; Baba, M.; Iwatsubo, T.; Meijer, L.; Kahle, P.J.; Haass, C. Constitutive phosphorylation of the Parkinson’s disease associated alpha-synuclein. J. Biol. Chem. 2000, 275, 390–397. [Google Scholar] [CrossRef]
- Inglis, K.J.; Chereau, D.; Brigham, E.F.; Chiou, S.S.; Schobel, S.; Frigon, N.L.; Yu, M.; Caccavello, R.J.; Nelson, S.; Motter, R.; et al. Polo-like kinase 2 (PLK2) phosphorylates alpha-synuclein at serine 129 in central nervous system. J. Biol. Chem. 2009, 284, 2598–2602. [Google Scholar] [CrossRef]
- Paleologou, K.E.; Oueslati, A.; Shakked, G.; Rospigliosi, C.C.; Kim, H.Y.; Lamberto, G.R.; Fernandez, C.O.; Schmid, A.; Chegini, F.; Gai, W.P.; et al. Phosphorylation at S87 is enhanced in synucleinopathies, inhibits alpha-synuclein oligomerization, and influences synuclein-membrane interactions. J. Neurosci. 2010, 30, 3184–3198. [Google Scholar] [CrossRef]
- Shin, Y.; Klucken, J.; Patterson, C.; Hyman, B.T.; McLean, P.J. The co-chaperone carboxyl terminus of Hsp70-interacting protein (CHIP) mediates alpha-synuclein degradation decisions between proteasomal and lysosomal pathways. J. Biol. Chem. 2005, 280, 23727–23734. [Google Scholar] [CrossRef]
- Lee, J.T.; Wheeler, T.C.; Li, L.; Chin, L.S. Ubiquitination of alpha-synuclein by Siah-1 promotes alpha-synuclein aggregation and apoptotic cell death. Hum. Mol. Genet. 2008, 17, 906–917. [Google Scholar] [CrossRef] [PubMed]
- Rott, R.; Szargel, R.; Haskin, J.; Shani, V.; Shainskaya, A.; Manov, I.; Liani, E.; Avraham, E.; Engelender, S. Monoubiquitylation of alpha-synuclein by seven in absentia homolog (SIAH) promotes its aggregation in dopaminergic cells. J. Biol. Chem. 2008, 283, 3316–3328. [Google Scholar] [CrossRef]
- Tetzlaff, J.E.; Putcha, P.; Outeiro, T.F.; Ivanov, A.; Berezovska, O.; Hyman, B.T.; McLean, P.J. CHIP targets toxic alpha-Synuclein oligomers for degradation. J. Biol. Chem. 2008, 283, 17962–17968. [Google Scholar] [CrossRef] [PubMed]
- Lewis, Y.E.; Galesic, A.; Levine, P.M.; De Leon, C.A.; Lamiri, N.; Brennan, C.K.; Pratt, M.R. O-GlcNAcylation of alpha-Synuclein at Serine 87 Reduces Aggregation without Affecting Membrane Binding. ACS Chem. Biol. 2017, 12, 1020–1027. [Google Scholar] [CrossRef]
- Savyon, M.; Engelender, S. SUMOylation in alpha-Synuclein Homeostasis and Pathology. Front. Aging Neurosci. 2020, 12, 167. [Google Scholar] [CrossRef]
- Zhu, L.N.; Qiao, H.H.; Chen, L.; Sun, L.P.; Hui, J.L.; Lian, Y.L.; Xie, W.B.; Ding, J.Y.; Meng, Y.L.; Zhu, B.F.; et al. SUMOylation of Alpha-Synuclein Influences on Alpha-Synuclein Aggregation Induced by Methamphetamine. Front. Cell Neurosci. 2018, 12, 262. [Google Scholar] [CrossRef] [PubMed]
- Hassanzadeh, K.; Morrone, C.; Akhtari, K.; Gerhardt, E.; Zaccagnini, L.; Outeiro, T.F.; Feligioni, M. Non-SUMOylated alternative spliced isoforms of alpha-synuclein are more aggregation-prone and toxic. Mech. Ageing Dev. 2023, 209, 111759. [Google Scholar] [CrossRef] [PubMed]
- Hegde, R.N.; Chiki, A.; Petricca, L.; Martufi, P.; Arbez, N.; Mouchiroud, L.; Auwerx, J.; Landles, C.; Bates, G.P.; Singh-Bains, M.K.; et al. TBK1 phosphorylates mutant Huntingtin and suppresses its aggregation and toxicity in Huntington’s disease models. EMBO J. 2020, 39, e104671. [Google Scholar] [CrossRef] [PubMed]
- Ratovitski, T.; Jiang, M.; O’Meally, R.N.; Rauniyar, P.; Chighladze, E.; Farago, A.; Kamath, S.V.; Jin, J.; Shevelkin, A.V.; Cole, R.N.; et al. Interaction of huntingtin with PRMTs and its subsequent arginine methylation affects HTT solubility, phase transition behavior and neuronal toxicity. Hum. Mol. Genet. 2022, 31, 1651–1672. [Google Scholar] [CrossRef]
- Bhat, K.P.; Yan, S.; Wang, C.E.; Li, S.; Li, X.J. Differential ubiquitination and degradation of huntingtin fragments modulated by ubiquitin-protein ligase E3A. Proc. Natl. Acad. Sci. USA 2014, 111, 5706–5711. [Google Scholar] [CrossRef]
- Banks, C.J.; Andersen, J.L. Mechanisms of SOD1 regulation by post-translational modifications. Redox Biol. 2019, 26, 101270. [Google Scholar] [CrossRef]
- Stoica, T.; Sorescu, I.; Stoica, D.T. [Role of “minimal” pharnygostomy in the re-equilibration of patients after extensive exeresis of the digestive tract (original method)]. Rev. Chir. Oncol. Radiol. ORL Oftalmol. Stomatol. Chir. 1978, 27, 153–156. [Google Scholar]
- Eck, R.J.; Kraemer, B.C.; Liachko, N.F. Regulation of TDP-43 phosphorylation in aging and disease. Geroscience 2021, 43, 1605–1614. [Google Scholar] [CrossRef] [PubMed]
- Hebron, M.L.; Lonskaya, I.; Sharpe, K.; Weerasinghe, P.P.; Algarzae, N.K.; Shekoyan, A.R.; Moussa, C.E. Parkin ubiquitinates Tar-DNA binding protein-43 (TDP-43) and promotes its cytosolic accumulation via interaction with histone deacetylase 6 (HDAC6). J. Biol. Chem. 2013, 288, 4103–4115. [Google Scholar] [CrossRef]
- Motaln, H.; Cercek, U.; Yamoah, A.; Tripathi, P.; Aronica, E.; Goswami, A.; Rogelj, B. Abl kinase-mediated FUS Tyr526 phosphorylation alters nucleocytoplasmic FUS localization in FTLD-FUS. Brain 2023, 146, 4088–4104. [Google Scholar] [CrossRef]
- Arenas, A.; Chen, J.; Kuang, L.; Barnett, K.R.; Kasarskis, E.J.; Gal, J.; Zhu, H. Lysine acetylation regulates the RNA binding, subcellular localization and inclusion formation of FUS. Hum. Mol. Genet. 2020, 29, 2684–2697. [Google Scholar] [CrossRef] [PubMed]
- Dada, S.T.; Hardenberg, M.C.; Toprakcioglu, Z.; Mrugalla, L.K.; Cali, M.P.; McKeon, M.O.; Klimont, E.; Michaels, T.C.T.; Knowles, T.P.J.; Vendruscolo, M. Spontaneous nucleation and fast aggregate-dependent proliferation of alpha-synuclein aggregates within liquid condensates at neutral pH. Proc. Natl. Acad. Sci. USA 2023, 120, e2208792120. [Google Scholar] [CrossRef]
- Agarwal, A.; Chandran, A.; Raza, F.; Ungureanu, I.M.; Hilcenko, C.; Stott, K.; Bright, N.A.; Morone, N.; Warren, A.J.; Lautenschlager, J. VAMP2 regulates phase separation of alpha-synuclein. Nat. Cell Biol. 2024, 26, 1296–1308. [Google Scholar] [CrossRef]
- Srivastava, T.; Raj, R.; Dubey, A.; Kumar, D.; Chaturvedi, R.K.; Sharma, S.K.; Priya, S. Fast kinetics of environmentally induced alpha-synuclein aggregation mediated by structural alteration in NAC region and result in structure dependent cytotoxicity. Sci. Rep. 2020, 10, 18412. [Google Scholar] [CrossRef]
- Shi, L.; Huang, C.; Luo, Q.; Rogers, E.; Xia, Y.; Liu, W.; Ma, W.; Zeng, W.; Gong, L.; Fang, J.; et al. The Association of Iron and the Pathologies of Parkinson’s Diseases in MPTP/MPP+-Induced Neuronal Degeneration in Non-human Primates and in Cell Culture. Front. Aging Neurosci. 2019, 11, 215. [Google Scholar] [CrossRef]
- Brown, D.R. alpha-Synuclein as a ferrireductase. Biochem. Soc. Trans. 2013, 41, 1513–1517. [Google Scholar] [CrossRef] [PubMed]
- McDowall, J.S.; Ntai, I.; Honeychurch, K.C.; Hart, J.P.; Colin, P.; Schneider, B.L.; Brown, D.R. Alpha-synuclein ferrireductase activity is detectible in vivo, is altered in Parkinson’s disease and increases the neurotoxicity of DOPAL. Mol. Cell Neurosci. 2017, 85, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Moller, H.E.; Bossoni, L.; Connor, J.R.; Crichton, R.R.; Does, M.D.; Ward, R.J.; Zecca, L.; Zucca, F.A.; Ronen, I. Iron, Myelin, and the Brain: Neuroimaging Meets Neurobiology. Trends Neurosci. 2019, 42, 384–401. [Google Scholar] [CrossRef]
- Crichton, R.R.; Dexter, D.T.; Ward, R.J. Brain iron metabolism and its perturbation in neurological diseases. J. Neural. Transm. 2011, 118, 301–314. [Google Scholar] [CrossRef]
- Oshiro, S.; Morioka, M.S.; Kikuchi, M. Dysregulation of iron metabolism in Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis. Adv. Pharmacol. Sci. 2011, 2011, 378278. [Google Scholar] [CrossRef]
- Ohshima, T.; Yamamoto, H.; Sakamaki, Y.; Saito, C.; Mizushima, N. NCOA4 drives ferritin phase separation to facilitate macroferritinophagy and microferritinophagy. J. Cell Biol. 2022, 221, e202203102. [Google Scholar] [CrossRef] [PubMed]
- Ohgita, T.; Namba, N.; Kono, H.; Shimanouchi, T.; Saito, H. Mechanisms of enhanced aggregation and fibril formation of Parkinson’s disease-related variants of alpha-synuclein. Sci. Rep. 2022, 12, 6770. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Garringer, H.J.; Shi, Y.; Lovestam, S.; Peak-Chew, S.; Zhang, X.; Kotecha, A.; Bacioglu, M.; Koto, A.; Takao, M.; et al. New SNCA mutation and structures of alpha-synuclein filaments from juvenile-onset synucleinopathy. Acta Neuropathol. 2023, 145, 561–572. [Google Scholar] [CrossRef]
- Proukakis, C.; Dudzik, C.G.; Brier, T.; MacKay, D.S.; Cooper, J.M.; Millhauser, G.L.; Houlden, H.; Schapira, A.H. A novel alpha-synuclein missense mutation in Parkinson disease. Neurology 2013, 80, 1062–1064. [Google Scholar] [CrossRef]
- Boyer, D.R.; Li, B.; Sun, C.; Fan, W.; Sawaya, M.R.; Jiang, L.; Eisenberg, D.S. Structures of fibrils formed by alpha-synuclein hereditary disease mutant H50Q reveal new polymorphs. Nat. Struct. Mol. Biol. 2019, 26, 1044–1052. [Google Scholar] [CrossRef] [PubMed]
- Chartier-Harlin, M.C.; Kachergus, J.; Roumier, C.; Mouroux, V.; Douay, X.; Lincoln, S.; Levecque, C.; Larvor, L.; Andrieux, J.; Hulihan, M.; et al. Alpha-synuclein locus duplication as a cause of familial Parkinson’s disease. Lancet 2004, 364, 1167–1169. [Google Scholar] [CrossRef]
- Singleton, A.B.; Farrer, M.; Johnson, J.; Singleton, A.; Hague, S.; Kachergus, J.; Hulihan, M.; Peuralinna, T.; Dutra, A.; Nussbaum, R.; et al. alpha-Synuclein locus triplication causes Parkinson’s disease. Science 2003, 302, 841. [Google Scholar] [CrossRef]
- Puspita, L.; Chung, S.Y.; Shim, J.W. Oxidative stress and cellular pathologies in Parkinson’s disease. Mol. Brain 2017, 10, 53. [Google Scholar] [CrossRef]
- Hijaz, B.A.; Volpicelli-Daley, L.A. Initiation and propagation of alpha-synuclein aggregation in the nervous system. Mol. Neurodegener. 2020, 15, 19. [Google Scholar] [CrossRef]
- Roberts, H.L.; Schneider, B.L.; Brown, D.R. alpha-Synuclein increases beta-amyloid secretion by promoting beta-/gamma-secretase processing of APP. PLoS ONE 2017, 12, e0171925. [Google Scholar] [CrossRef]
- Manzanza, N.O.; Sedlackova, L.; Kalaria, R.N. Alpha-Synuclein Post-translational Modifications: Implications for Pathogenesis of Lewy Body Disorders. Front. Aging Neurosci. 2021, 13, 690293. [Google Scholar] [CrossRef] [PubMed]
- Luo, S.; Wang, D.; Zhang, Z. Post-translational modification and mitochondrial function in Parkinson’s disease. Front. Mol. Neurosci. 2023, 16, 1329554. [Google Scholar] [CrossRef]
- Lu, Y.; Prudent, M.; Fauvet, B.; Lashuel, H.A.; Girault, H.H. Phosphorylation of alpha-Synuclein at Y125 and S129 alters its metal binding properties: Implications for understanding the role of alpha-Synuclein in the pathogenesis of Parkinson’s Disease and related disorders. ACS Chem. Neurosci. 2011, 2, 667–675. [Google Scholar] [CrossRef] [PubMed]
- Henderson, M.X.; Cornblath, E.J.; Darwich, A.; Zhang, B.; Brown, H.; Gathagan, R.J.; Sandler, R.M.; Bassett, D.S.; Trojanowski, J.Q.; Lee, V.M.Y. Spread of alpha-synuclein pathology through the brain connectome is modulated by selective vulnerability and predicted by network analysis. Nat. Neurosci. 2019, 22, 1248–1257. [Google Scholar] [CrossRef]
- Awa, S.; Suzuki, G.; Masuda-Suzukake, M.; Nonaka, T.; Saito, M.; Hasegawa, M. Phosphorylation of endogenous alpha-synuclein induced by extracellular seeds initiates at the pre-synaptic region and spreads to the cell body. Sci. Rep. 2022, 12, 1163. [Google Scholar] [CrossRef] [PubMed]
- Shimura, H.; Schlossmacher, M.G.; Hattori, N.; Frosch, M.P.; Trockenbacher, A.; Schneider, R.; Mizuno, Y.; Kosik, K.S.; Selkoe, D.J. Ubiquitination of a new form of alpha-synuclein by parkin from human brain: Implications for Parkinson’s disease. Science 2001, 293, 263–269. [Google Scholar] [CrossRef]
- McColgan, P.; Tabrizi, S.J. Huntington’s disease: A clinical review. Eur. J. Neurol. 2018, 25, 24–34. [Google Scholar] [CrossRef]
- Saudou, F.; Humbert, S. The Biology of Huntingtin. Neuron 2016, 89, 910–926. [Google Scholar] [CrossRef] [PubMed]
- Peskett, T.R.; Rau, F.; O’Driscoll, J.; Patani, R.; Lowe, A.R.; Saibil, H.R. A Liquid to Solid Phase Transition Underlying Pathological Huntingtin Exon1 Aggregation. Mol. Cell 2018, 70, 588–601.e6. [Google Scholar] [CrossRef]
- Aktar, F.; Burudpakdee, C.; Polanco, M.; Pei, S.; Swayne, T.C.; Lipke, P.N.; Emtage, L. The huntingtin inclusion is a dynamic phase-separated compartment. Life Sci. Alliance 2019, 2, e201900489. [Google Scholar] [CrossRef]
- Posey, A.E.; Ruff, K.M.; Harmon, T.S.; Crick, S.L.; Li, A.; Diamond, M.I.; Pappu, R.V. Profilin reduces aggregation and phase separation of huntingtin N-terminal fragments by preferentially binding to soluble monomers and oligomers. J. Biol. Chem. 2018, 293, 3734–3746. [Google Scholar] [CrossRef] [PubMed]
- Caron, N.S.; Hung, C.L.; Atwal, R.S.; Truant, R. Live cell imaging and biophotonic methods reveal two types of mutant huntingtin inclusions. Hum. Mol. Genet. 2014, 23, 2324–2338. [Google Scholar] [CrossRef] [PubMed]
- Ramdzan, Y.M.; Trubetskov, M.M.; Ormsby, A.R.; Newcombe, E.A.; Sui, X.; Tobin, M.J.; Bongiovanni, M.N.; Gras, S.L.; Dewson, G.; Miller, J.M.L.; et al. Huntingtin Inclusions Trigger Cellular Quiescence, Deactivate Apoptosis, and Lead to Delayed Necrosis. Cell Rep. 2017, 19, 919–927. [Google Scholar] [CrossRef] [PubMed]
- Jimenez-Sanchez, M.; Licitra, F.; Underwood, B.R.; Rubinsztein, D.C. Huntington’s Disease: Mechanisms of Pathogenesis and Therapeutic Strategies. Cold Spring Harb. Perspect. Med. 2017, 7, a024240. [Google Scholar] [CrossRef]
- Tsoi, H.; Chan, H.Y. Expression of expanded CAG transcripts triggers nucleolar stress in Huntington’s disease. Cerebellum 2013, 12, 310–312. [Google Scholar] [CrossRef] [PubMed]
- Lunkes, A.; Lindenberg, K.S.; Ben-Haiem, L.; Weber, C.; Devys, D.; Landwehrmeyer, G.B.; Mandel, J.L.; Trottier, Y. Proteases acting on mutant huntingtin generate cleaved products that differentially build up cytoplasmic and nuclear inclusions. Mol. Cell 2002, 10, 259–269. [Google Scholar] [CrossRef]
- DeGuire, S.M.; Ruggeri, F.S.; Fares, M.B.; Chiki, A.; Cendrowska, U.; Dietler, G.; Lashuel, H.A. N-terminal Huntingtin (Htt) phosphorylation is a molecular switch regulating Htt aggregation, helical conformation, internalization, and nuclear targeting. J. Biol. Chem. 2018, 293, 18540–18558. [Google Scholar] [CrossRef]
- Gottlieb, L.; Guo, L.; Shorter, J.; Marmorstein, R. N-alpha-acetylation of Huntingtin protein increases its propensity to aggregate. J. Biol. Chem. 2021, 297, 101363. [Google Scholar] [CrossRef]
- Chaibva, M.; Jawahery, S.; Pilkington, A.W.t.; Arndt, J.R.; Sarver, O.; Valentine, S.; Matysiak, S.; Legleiter, J. Acetylation within the First 17 Residues of Huntingtin Exon 1 Alters Aggregation and Lipid Binding. Biophys. J. 2016, 111, 349–362. [Google Scholar] [CrossRef]
- Sedighi, F.; Adegbuyiro, A.; Legleiter, J. SUMOylation Prevents Huntingtin Fibrillization and Localization onto Lipid Membranes. ACS Chem. Neurosci. 2020, 11, 328–343. [Google Scholar] [CrossRef]
- Dowling, J.J.; Gonorazky, H.D.; Cohn, R.D.; Campbell, C. Treating pediatric neuromuscular disorders: The future is now. Am. J. Med. Genet. A 2018, 176, 804–841. [Google Scholar] [CrossRef]
- Nijssen, J.; Comley, L.H.; Hedlund, E. Motor neuron vulnerability and resistance in amyotrophic lateral sclerosis. Acta Neuropathol. 2017, 133, 863–885. [Google Scholar] [CrossRef] [PubMed]
- Mejzini, R.; Flynn, L.L.; Pitout, I.L.; Fletcher, S.; Wilton, S.D.; Akkari, P.A. ALS Genetics, Mechanisms, and Therapeutics: Where Are We Now? Front. Neurosci. 2019, 13, 1310. [Google Scholar] [CrossRef] [PubMed]
- Neumann, M.; Sampathu, D.M.; Kwong, L.K.; Truax, A.C.; Micsenyi, M.C.; Chou, T.T.; Bruce, J.; Schuck, T.; Grossman, M.; Clark, C.M.; et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 2006, 314, 130–133. [Google Scholar] [CrossRef]
- Rosen, D.R.; Siddique, T.; Patterson, D.; Figlewicz, D.A.; Sapp, P.; Hentati, A.; Donaldson, D.; Goto, J.; O’Regan, J.P.; Deng, H.X.; et al. Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 1993, 362, 59–62. [Google Scholar] [CrossRef]
- Thomas, K.A.; Rubin, B.H.; Bier, C.J.; Richardson, J.S.; Richardson, D.C. The crystal structure of bovine Cu2+,Zn2+ superoxide dismutase at 5.5-A resolution. J. Biol. Chem. 1974, 249, 5677–5683. [Google Scholar] [CrossRef]
- Santamaria, N.; Alhothali, M.; Alfonso, M.H.; Breydo, L.; Uversky, V.N. Intrinsic disorder in proteins involved in amyotrophic lateral sclerosis. Cell Mol. Life Sci. 2017, 74, 1297–1318. [Google Scholar] [CrossRef] [PubMed]
- Das, B.; Roychowdhury, S.; Mohanty, P.; Rizuan, A.; Chakraborty, J.; Mittal, J.; Chattopadhyay, K. A Zn-dependent structural transition of SOD1 modulates its ability to undergo phase separation. EMBO J. 2023, 42, e111185. [Google Scholar] [CrossRef]
- Gu, S.; Xu, M.; Chen, L.; Shi, X.; Luo, S.Z. A liquid-to-solid phase transition of Cu/Zn superoxide dismutase 1 initiated by oxidation and disease mutation. J. Biol. Chem. 2023, 299, 102857. [Google Scholar] [CrossRef]
- Hallegger, M.; Chakrabarti, A.M.; Lee, F.C.Y.; Lee, B.L.; Amalietti, A.G.; Odeh, H.M.; Copley, K.E.; Rubien, J.D.; Portz, B.; Kuret, K.; et al. TDP-43 condensation properties specify its RNA-binding and regulatory repertoire. Cell 2021, 184, 4680–4696.e4622. [Google Scholar] [CrossRef]
- Buratti, E.; Baralle, F.E. Multiple roles of TDP-43 in gene expression, splicing regulation, and human disease. Front. Biosci. 2008, 13, 867–878. [Google Scholar] [CrossRef] [PubMed]
- Fujioka, Y.; Ishigaki, S.; Masuda, A.; Iguchi, Y.; Udagawa, T.; Watanabe, H.; Katsuno, M.; Ohno, K.; Sobue, G. FUS-regulated region- and cell-type-specific transcriptome is associated with cell selectivity in ALS/FTLD. Sci. Rep. 2013, 3, 2388. [Google Scholar] [CrossRef]
- Yang, L.; Zhang, J.; Kamelgarn, M.; Niu, C.; Gal, J.; Gong, W.; Zhu, H. Subcellular localization and RNAs determine FUS architecture in different cellular compartments. Hum. Mol. Genet. 2015, 24, 5174–5183. [Google Scholar] [CrossRef]
- Abel, O.; Powell, J.F.; Andersen, P.M.; Al-Chalabi, A. ALSoD: A user-friendly online bioinformatics tool for amyotrophic lateral sclerosis genetics. Hum. Mutat. 2012, 33, 1345–1351. [Google Scholar] [CrossRef]
- Berdynski, M.; Miszta, P.; Safranow, K.; Andersen, P.M.; Morita, M.; Filipek, S.; Zekanowski, C.; Kuzma-Kozakiewicz, M. SOD1 mutations associated with amyotrophic lateral sclerosis analysis of variant severity. Sci. Rep. 2022, 12, 103. [Google Scholar] [CrossRef] [PubMed]
- Culik, R.M.; Sekhar, A.; Nagesh, J.; Deol, H.; Rumfeldt, J.A.O.; Meiering, E.M.; Kay, L.E. Effects of maturation on the conformational free-energy landscape of SOD1. Proc. Natl. Acad. Sci. USA 2018, 115, E2546–E2555. [Google Scholar] [CrossRef]
- Hayward, L.J.; Rodriguez, J.A.; Kim, J.W.; Tiwari, A.; Goto, J.J.; Cabelli, D.E.; Valentine, J.S.; Brown, R.H., Jr. Decreased metallation and activity in subsets of mutant superoxide dismutases associated with familial amyotrophic lateral sclerosis. J. Biol. Chem. 2002, 277, 15923–15931. [Google Scholar] [CrossRef]
- Milicevic, K.; Rankovic, B.; Andjus, P.R.; Bataveljic, D.; Milovanovic, D. Emerging Roles for Phase Separation of RNA-Binding Proteins in Cellular Pathology of ALS. Front. Cell Dev. Biol. 2022, 10, 840256. [Google Scholar] [CrossRef]
- Fujita, Y.; Ikeda, M.; Yanagisawa, T.; Senoo, Y.; Okamoto, K. Different clinical and neuropathologic phenotypes of familial ALS with A315E TARDBP mutation. Neurology 2011, 77, 1427–1431. [Google Scholar] [CrossRef]
- Guo, W.; Chen, Y.; Zhou, X.; Kar, A.; Ray, P.; Chen, X.; Rao, E.J.; Yang, M.; Ye, H.; Zhu, L.; et al. An ALS-associated mutation affecting TDP-43 enhances protein aggregation, fibril formation and neurotoxicity. Nat. Struct. Mol. Biol. 2011, 18, 822–830. [Google Scholar] [CrossRef]
- Liu, X.; Lao, Z.; Li, X.; Dong, X.; Wei, G. ALS-associated A315E and A315pT variants exhibit distinct mechanisms in inducing irreversible aggregation of TDP-43312-317 peptides. Phys. Chem. Chem. Phys. 2022, 24, 16263–16273. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Li, X.; Qiao, Q.; Li, F.; Wei, G. ALS-Linked A315T and A315E Mutations Enhance beta-Barrel Formation of the TDP-43307-319 Hexamer: A REST2 Simulation Study. ACS Chem. Neurosci. 2023, 14, 1310–1320. [Google Scholar] [CrossRef]
- Sreedharan, J.; Blair, I.P.; Tripathi, V.B.; Hu, X.; Vance, C.; Rogelj, B.; Ackerley, S.; Durnall, J.C.; Williams, K.L.; Buratti, E.; et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 2008, 319, 1668–1672. [Google Scholar] [CrossRef] [PubMed]
- Johnson, B.S.; Snead, D.; Lee, J.J.; McCaffery, J.M.; Shorter, J.; Gitler, A.D. TDP-43 is intrinsically aggregation-prone, and amyotrophic lateral sclerosis-linked mutations accelerate aggregation and increase toxicity. J. Biol. Chem. 2009, 284, 20329–20339. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Zhao, X.; Wu, W.; Wang, Q.; Liu, J.; Zhang, W.; Yuan, Y.; Hong, D.; Wang, Z.; Deng, J. Widespread Mislocalization of FUS Is Associated With Mitochondrial Abnormalities in Skeletal Muscle in Amyotrophic Lateral Sclerosis with FUS Mutations. J. Neuropathol. Exp. Neurol. 2022, 81, 172–181. [Google Scholar] [CrossRef]
- Waibel, S.; Neumann, M.; Rabe, M.; Meyer, T.; Ludolph, A.C. Novel missense and truncating mutations in FUS/TLS in familial ALS. Neurology 2010, 75, 815–817. [Google Scholar] [CrossRef]
- Nakaya, T.; Maragkakis, M. Amyotrophic Lateral Sclerosis associated FUS mutation shortens mitochondria and induces neurotoxicity. Sci. Rep. 2018, 8, 15575. [Google Scholar] [CrossRef]
- Baumer, D.; Hilton, D.; Paine, S.M.; Turner, M.R.; Lowe, J.; Talbot, K.; Ansorge, O. Juvenile ALS with basophilic inclusions is a FUS proteinopathy with FUS mutations. Neurology 2010, 75, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Khare, S.D.; Caplow, M.; Dokholyan, N.V. The rate and equilibrium constants for a multistep reaction sequence for the aggregation of superoxide dismutase in amyotrophic lateral sclerosis. Proc. Natl. Acad. Sci. USA 2004, 101, 15094–15099. [Google Scholar] [CrossRef]
- Rakhit, R.; Crow, J.P.; Lepock, J.R.; Kondejewski, L.H.; Cashman, N.R.; Chakrabartty, A. Monomeric Cu, Zn-superoxide dismutase is a common misfolding intermediate in the oxidation models of sporadic and familial amyotrophic lateral sclerosis. J. Biol. Chem. 2004, 279, 15499–154504. [Google Scholar] [CrossRef]
- Wilcox, K.C.; Zhou, L.; Jordon, J.K.; Huang, Y.; Yu, Y.; Redler, R.L.; Chen, X.; Caplow, M.; Dokholyan, N.V. Modifications of superoxide dismutase (SOD1) in human erythrocytes: A possible role in amyotrophic lateral sclerosis. J. Biol. Chem. 2009, 284, 13940–13947. [Google Scholar] [CrossRef]
- Redler, R.L.; Wilcox, K.C.; Proctor, E.A.; Fee, L.; Caplow, M.; Dokholyan, N.V. Glutathionylation at Cys-111 induces dissociation of wild type and FALS mutant SOD1 dimers. Biochemistry 2011, 50, 7057–7066. [Google Scholar] [CrossRef] [PubMed]
- Pansarasa, O.; Bordoni, M.; Drufuca, L.; Diamanti, L.; Sproviero, D.; Trotti, R.; Bernuzzi, S.; La Salvia, S.; Gagliardi, S.; Ceroni, M.; et al. Lymphoblastoid cell lines as a model to understand amyotrophic lateral sclerosis disease mechanisms. Dis. Model. Mech. 2018, 11, dmm031625. [Google Scholar] [CrossRef] [PubMed]
- Cereda, C.; Leoni, E.; Milani, P.; Pansarasa, O.; Mazzini, G.; Guareschi, S.; Alvisi, E.; Ghiroldi, A.; Diamanti, L.; Bernuzzi, S.; et al. Altered intracellular localization of SOD1 in leukocytes from patients with sporadic amyotrophic lateral sclerosis. PLoS ONE 2013, 8, e75916. [Google Scholar] [CrossRef][Green Version]
- Gagliardi, S.; Cova, E.; Davin, A.; Guareschi, S.; Abel, K.; Alvisi, E.; Laforenza, U.; Ghidoni, R.; Cashman, J.R.; Ceroni, M.; et al. SOD1 mRNA expression in sporadic amyotrophic lateral sclerosis. Neurobiol. Dis. 2010, 39, 198–203. [Google Scholar] [CrossRef]
- Bordoni, M.; Pansarasa, O.; Dell’Orco, M.; Crippa, V.; Gagliardi, S.; Sproviero, D.; Bernuzzi, S.; Diamanti, L.; Ceroni, M.; Tedeschi, G.; et al. Nuclear Phospho-SOD1 Protects DNA from Oxidative Stress Damage in Amyotrophic Lateral Sclerosis. J. Clin. Med. 2019, 8, 729. [Google Scholar] [CrossRef] [PubMed]
- Fay, J.M.; Zhu, C.; Proctor, E.A.; Tao, Y.; Cui, W.; Ke, H.; Dokholyan, N.V. A Phosphomimetic Mutation Stabilizes SOD1 and Rescues Cell Viability in the Context of an ALS-Associated Mutation. Structure 2016, 24, 1898–1906. [Google Scholar] [CrossRef]
- Hasegawa, M.; Arai, T.; Nonaka, T.; Kametani, F.; Yoshida, M.; Hashizume, Y.; Beach, T.G.; Buratti, E.; Baralle, F.; Morita, M.; et al. Phosphorylated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Ann. Neurol. 2008, 64, 60–70. [Google Scholar] [CrossRef]
- Neumann, M.; Frick, P.; Paron, F.; Kosten, J.; Buratti, E.; Mackenzie, I.R. Antibody against TDP-43 phosphorylated at serine 375 suggests conformational differences of TDP-43 aggregates among FTLD-TDP subtypes. Acta Neuropathol. 2020, 140, 645–658. [Google Scholar] [CrossRef]
- Farrawell, N.E.; McAlary, L.; Lum, J.S.; Chisholm, C.G.; Warraich, S.T.; Blair, I.P.; Vine, K.L.; Saunders, D.N.; Yerbury, J.J. Ubiquitin Homeostasis Is Disrupted in TDP-43 and FUS Cell Models of ALS. iScience 2020, 23, 101700. [Google Scholar] [CrossRef]
- Riemenschneider, H.; Guo, Q.; Bader, J.; Frottin, F.; Farny, D.; Kleinberger, G.; Haass, C.; Mann, M.; Hartl, F.U.; Baumeister, W.; et al. Gel-like inclusions of C-terminal fragments of TDP-43 sequester stalled proteasomes in neurons. EMBO Rep. 2022, 23, e53890. [Google Scholar] [CrossRef] [PubMed]
- Rayner, S.L.; Yang, S.; Farrawell, N.E.; Jagaraj, C.J.; Cheng, F.; Davidson, J.M.; Luu, L.; Redondo, A.G.; Rabano, A.; Borrego-Hernandez, D.; et al. TDP-43 is a ubiquitylation substrate of the SCFcyclin F complex. Neurobiol. Dis. 2022, 167, 105673. [Google Scholar] [CrossRef]
- Rhoads, S.N.; Monahan, Z.T.; Yee, D.S.; Shewmaker, F.P. The Role of Post-Translational Modifications on Prion-Like Aggregation and Liquid-Phase Separation of FUS. Int. J. Mol. Sci. 2018, 19, 886. [Google Scholar] [CrossRef] [PubMed]
- Deng, Q.; Holler, C.J.; Taylor, G.; Hudson, K.F.; Watkins, W.; Gearing, M.; Ito, D.; Murray, M.E.; Dickson, D.W.; Seyfried, N.T.; et al. FUS is phosphorylated by DNA-PK and accumulates in the cytoplasm after DNA damage. J. Neurosci. 2014, 34, 7802–7813. [Google Scholar] [CrossRef]
- Mastrocola, A.S.; Kim, S.H.; Trinh, A.T.; Rodenkirch, L.A.; Tibbetts, R.S. The RNA-binding protein fused in sarcoma (FUS) functions downstream of poly(ADP-ribose) polymerase (PARP) in response to DNA damage. J. Biol. Chem. 2013, 288, 24731–24741. [Google Scholar] [CrossRef]
- Rulten, S.L.; Rotheray, A.; Green, R.L.; Grundy, G.J.; Moore, D.A.; Gomez-Herreros, F.; Hafezparast, M.; Caldecott, K.W. PARP-1 dependent recruitment of the amyotrophic lateral sclerosis-associated protein FUS/TLS to sites of oxidative DNA damage. Nucleic Acids Res. 2014, 42, 307–314. [Google Scholar] [CrossRef] [PubMed]
- Monahan, Z.; Ryan, V.H.; Janke, A.M.; Burke, K.A.; Rhoads, S.N.; Zerze, G.H.; O’Meally, R.; Dignon, G.L.; Conicella, A.E.; Zheng, W.; et al. Phosphorylation of the FUS low-complexity domain disrupts phase separation, aggregation, and toxicity. EMBO J. 2017, 36, 2951–2967. [Google Scholar] [CrossRef]
- Murray, D.T.; Kato, M.; Lin, Y.; Thurber, K.R.; Hung, I.; McKnight, S.L.; Tycko, R. Structure of FUS Protein Fibrils and Its Relevance to Self-Assembly and Phase Separation of Low-Complexity Domains. Cell 2017, 171, 615–627.e16. [Google Scholar] [CrossRef]
- Scaramuzzino, C.; Monaghan, J.; Milioto, C.; Lanson, N.A., Jr.; Maltare, A.; Aggarwal, T.; Casci, I.; Fackelmayer, F.O.; Pennuto, M.; Pandey, U.B. Protein arginine methyltransferase 1 and 8 interact with FUS to modify its sub-cellular distribution and toxicity in vitro and in vivo. PLoS ONE 2013, 8, e61576. [Google Scholar] [CrossRef]
- Du, K.; Arai, S.; Kawamura, T.; Matsushita, A.; Kurokawa, R. TLS and PRMT1 synergistically coactivate transcription at the survivin promoter through TLS arginine methylation. Biochem. Biophys. Res. Commun. 2011, 404, 991–996. [Google Scholar] [CrossRef]
- Sama, R.R.; Ward, C.L.; Kaushansky, L.J.; Lemay, N.; Ishigaki, S.; Urano, F.; Bosco, D.A. FUS/TLS assembles into stress granules and is a prosurvival factor during hyperosmolar stress. J. Cell Physiol. 2013, 228, 2222–2231. [Google Scholar] [CrossRef]
- Dormann, D.; Madl, T.; Valori, C.F.; Bentmann, E.; Tahirovic, S.; Abou-Ajram, C.; Kremmer, E.; Ansorge, O.; Mackenzie, I.R.; Neumann, M.; et al. Arginine methylation next to the PY-NLS modulates Transportin binding and nuclear import of FUS. EMBO J. 2012, 31, 4258–4275. [Google Scholar] [CrossRef] [PubMed]
- Suarez-Calvet, M.; Neumann, M.; Arzberger, T.; Abou-Ajram, C.; Funk, E.; Hartmann, H.; Edbauer, D.; Kremmer, E.; Gobl, C.; Resch, M.; et al. Monomethylated and unmethylated FUS exhibit increased binding to Transportin and distinguish FTLD-FUS from ALS-FUS. Acta Neuropathol. 2016, 131, 587–604. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.T.; Nazarov, S.; Porta, S.; Maharjan, N.; Cendrowska, U.; Kabani, M.; Finamore, F.; Xu, Y.; Lee, V.M.; Lashuel, H.A. Seeding the aggregation of TDP-43 requires post-fibrillization proteolytic cleavage. Nat. Neurosci. 2023, 26, 983–996. [Google Scholar] [CrossRef]
- Zhou, C.; Li, S.; Ye, L.; Chen, C.; Liu, S.; Yang, H.; Zhuang, P.; Liu, Z.; Jiang, H.; Han, J.; et al. Visual impairment and blindness caused by retinal diseases: A nationwide register-based study. J. Glob. Health 2023, 13, 04126. [Google Scholar] [CrossRef]
- Bharti, K.; den Hollander, A.I.; Lakkaraju, A.; Sinha, D.; Williams, D.S.; Finnemann, S.C.; Bowes-Rickman, C.; Malek, G.; D’Amore, P.A. Cell culture models to study retinal pigment epithelium-related pathogenesis in age-related macular degeneration. Exp. Eye Res. 2022, 222, 109170. [Google Scholar] [CrossRef]
- Vugler, A.A. Progress toward the maintenance and repair of degenerating retinal circuitry. Retina 2010, 30, 983–1001. [Google Scholar] [CrossRef]
- Wang, M.; Su, S.; Jiang, S.; Sun, X.; Wang, J. Role of amyloid beta-peptide in the pathogenesis of age-related macular degeneration. BMJ Open Ophthalmol. 2021, 6, e000774. [Google Scholar] [CrossRef] [PubMed]
- Wolf, G. Function of the protein RPE65 in the visual cycle. Nutr. Rev. 2005, 63, 97–100. [Google Scholar] [CrossRef]
- Chacon-Camacho, O.F.; Zenteno, J.C. Review and update on the molecular basis of Leber congenital amaurosis. World J. Clin. Cases 2015, 3, 112–124. [Google Scholar] [CrossRef]
- Hussain, R.M.; Gregori, N.Z.; Ciulla, T.A.; Lam, B.L. Pharmacotherapy of retinal disease with visual cycle modulators. Expert Opin. Pharmacother. 2018, 19, 471–481. [Google Scholar] [CrossRef] [PubMed]
- Bowne, S.J.; Humphries, M.M.; Sullivan, L.S.; Kenna, P.F.; Tam, L.C.; Kiang, A.S.; Campbell, M.; Weinstock, G.M.; Koboldt, D.C.; Ding, L.; et al. A dominant mutation in RPE65 identified by whole-exome sequencing causes retinitis pigmentosa with choroidal involvement. Eur. J. Hum. Genet. 2011, 19, 1074–1081. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Pozo, A.; Das, S.; Hyman, B.T. APOE and Alzheimer’s disease: Advances in genetics, pathophysiology, and therapeutic approaches. Lancet Neurol. 2021, 20, 68–80. [Google Scholar] [CrossRef] [PubMed]
- La Cunza, N.; Tan, L.X.; Thamban, T.; Germer, C.J.; Rathnasamy, G.; Toops, K.A.; Lakkaraju, A. Mitochondria-dependent phase separation of disease-relevant proteins drives pathological features of age-related macular degeneration. JCI Insight 2021, 6, e142254. [Google Scholar] [CrossRef] [PubMed]
- Napoli, J.L. Retinoic acid biosynthesis and metabolism. FASEB J. 1996, 10, 993–1001. [Google Scholar] [CrossRef]
- Rohm, T.V.; Meier, D.T.; Olefsky, J.M.; Donath, M.Y. Inflammation in obesity, diabetes, and related disorders. Immunity 2022, 55, 31–55. [Google Scholar] [CrossRef]
- Altmann, C.; Schmidt, M.H.H. The Role of Microglia in Diabetic Retinopathy: Inflammation, Microvasculature Defects and Neurodegeneration. Int. J. Mol. Sci. 2018, 19, 110. [Google Scholar] [CrossRef]
- Kang, Q.; Yang, C. Oxidative stress and diabetic retinopathy: Molecular mechanisms, pathogenetic role and therapeutic implications. Redox Biol. 2020, 37, 101799. [Google Scholar] [CrossRef]
- Tang, L.; Xu, G.T.; Zhang, J.F. Inflammation in diabetic retinopathy: Possible roles in pathogenesis and potential implications for therapy. Neural Regen. Res. 2023, 18, 976–982. [Google Scholar] [CrossRef]
- Zong, H.; Ward, M.; Stitt, A.W. AGEs, RAGE, and diabetic retinopathy. Curr. Diab. Rep. 2011, 11, 244–252. [Google Scholar] [CrossRef]
- Gerstberger, S.; Hafner, M.; Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 2014, 15, 829–845. [Google Scholar] [CrossRef] [PubMed]
- Ocharan-Mercado, A.; Loaeza-Loaeza, J.; Castro-Coronel, Y.; Acosta-Saavedra, L.C.; Hernandez-Kelly, L.C.; Hernandez-Sotelo, D.; Ortega, A. RNA-Binding Proteins: A Role in Neurotoxicity? Neurotox Res. 2023, 41, 681–697. [Google Scholar] [CrossRef] [PubMed]
- Langeberg, C.J.; Kieft, J.S. A generalizable scaffold-based approach for structure determination of RNAs by cryo-EM. Nucleic Acids Res. 2023, 51, e100. [Google Scholar] [CrossRef] [PubMed]
- Ottoz, D.S.M.; Berchowitz, L.E. The role of disorder in RNA binding affinity and specificity. Open Biol. 2020, 10, 200328. [Google Scholar] [CrossRef]
- Zhao, B.; Katuwawala, A.; Oldfield, C.J.; Hu, G.; Wu, Z.; Uversky, V.N.; Kurgan, L. Intrinsic Disorder in Human RNA-Binding Proteins. J. Mol. Biol. 2021, 433, 167229. [Google Scholar] [CrossRef] [PubMed]
- Patel, A.; Lee, H.O.; Jawerth, L.; Maharana, S.; Jahnel, M.; Hein, M.Y.; Stoynov, S.; Mahamid, J.; Saha, S.; Franzmann, T.M.; et al. A Liquid-to-Solid Phase Transition of the ALS Protein FUS Accelerated by Disease Mutation. Cell 2015, 162, 1066–1077. [Google Scholar] [CrossRef]
- Lovtrup, S. Philosophy and evolution. Comments on Michael Ruse: Karl Popper’s philosophy of biology. Riv. Biol. 1988, 81, 287–308. [Google Scholar]
- Cartegni, L.; Hastings, M.L.; Calarco, J.A.; de Stanchina, E.; Krainer, A.R. Determinants of exon 7 splicing in the spinal muscular atrophy genes, SMN1 and SMN2. Am. J. Hum. Genet. 2006, 78, 63–77. [Google Scholar] [CrossRef]
- Liu, Q.; Dreyfuss, G. A novel nuclear structure containing the survival of motor neurons protein. EMBO J. 1996, 15, 3555–3565. [Google Scholar] [CrossRef]
- Ottesen, E.W.; Singh, N.N.; Luo, D.; Singh, R.N. High-affinity RNA targets of the Survival Motor Neuron protein reveal diverse preferences for sequence and structural motifs. Nucleic Acids Res. 2018, 46, 10983–11001. [Google Scholar] [CrossRef]
- Monani, U.R.; Sendtner, M.; Coovert, D.D.; Parsons, D.W.; Andreassi, C.; Le, T.T.; Jablonka, S.; Schrank, B.; Rossoll, W.; Prior, T.W.; et al. The human centromeric survival motor neuron gene (SMN2) rescues embryonic lethality in Smn−/− mice and results in a mouse with spinal muscular atrophy. Hum. Mol. Genet. 2000, 9, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Smeyers, J.; Banchi, E.G.; Latouche, M. C9ORF72: What It Is, What It Does, and Why It Matters. Front. Cell Neurosci. 2021, 15, 661447. [Google Scholar] [CrossRef]
- Mori, K.; Weng, S.M.; Arzberger, T.; May, S.; Rentzsch, K.; Kremmer, E.; Schmid, B.; Kretzschmar, H.A.; Cruts, M.; Van Broeckhoven, C.; et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 2013, 339, 1335–1338. [Google Scholar] [CrossRef]
- Haeusler, A.R.; Donnelly, C.J.; Periz, G.; Simko, E.A.; Shaw, P.G.; Kim, M.S.; Maragakis, N.J.; Troncoso, J.C.; Pandey, A.; Sattler, R.; et al. C9orf72 nucleotide repeat structures initiate molecular cascades of disease. Nature 2014, 507, 195–200. [Google Scholar] [CrossRef] [PubMed]
- May, S.; Hornburg, D.; Schludi, M.H.; Arzberger, T.; Rentzsch, K.; Schwenk, B.M.; Grasser, F.A.; Mori, K.; Kremmer, E.; Banzhaf-Strathmann, J.; et al. C9orf72 FTLD/ALS-associated Gly-Ala dipeptide repeat proteins cause neuronal toxicity and Unc119 sequestration. Acta Neuropathol. 2014, 128, 485–503. [Google Scholar] [CrossRef]
- Teng, Y.; Zhu, M.; Qiu, Z. G-Quadruplexes in Repeat Expansion Disorders. Int. J. Mol. Sci. 2023, 24, 2375. [Google Scholar] [CrossRef] [PubMed]
- Sung, H.M.; Schott, J.; Boss, P.; Lehmann, J.A.; Hardt, M.R.; Lindner, D.; Messens, J.; Bogeski, I.; Ohler, U.; Stoecklin, G. Stress-induced nuclear speckle reorganization is linked to activation of immediate early gene splicing. J. Cell Biol. 2023, 222, e202111151. [Google Scholar] [CrossRef]
- Lee, Y.B.; Chen, H.J.; Peres, J.N.; Gomez-Deza, J.; Attig, J.; Stalekar, M.; Troakes, C.; Nishimura, A.L.; Scotter, E.L.; Vance, C.; et al. Hexanucleotide repeats in ALS/FTD form length-dependent RNA foci, sequester RNA binding proteins, and are neurotoxic. Cell Rep. 2013, 5, 1178–1186. [Google Scholar] [CrossRef]
- Zhang, K.; Donnelly, C.J.; Haeusler, A.R.; Grima, J.C.; Machamer, J.B.; Steinwald, P.; Daley, E.L.; Miller, S.J.; Cunningham, K.M.; Vidensky, S.; et al. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature 2015, 525, 56–61. [Google Scholar] [CrossRef]
- Jain, A.; Vale, R.D. RNA phase transitions in repeat expansion disorders. Nature 2017, 546, 243–247. [Google Scholar] [CrossRef]
- Ishiguro, A.; Kimura, N.; Watanabe, Y.; Watanabe, S.; Ishihama, A. TDP-43 binds and transports G-quadruplex-containing mRNAs into neurites for local translation. Genes Cells 2016, 21, 466–481. [Google Scholar] [CrossRef] [PubMed]
- Fakim, H.; Vande Velde, C. The implications of physiological biomolecular condensates in amyotrophic lateral sclerosis. Semin. Cell Dev. Biol. 2024, 156, 176–189. [Google Scholar] [CrossRef] [PubMed]
- Kamelgarn, M.; Chen, J.; Kuang, L.; Jin, H.; Kasarskis, E.J.; Zhu, H. ALS mutations of FUS suppress protein translation and disrupt the regulation of nonsense-mediated decay. Proc. Natl. Acad. Sci. USA 2018, 115, E11904–E11913. [Google Scholar] [CrossRef] [PubMed]
- Ishiguro, A.; Lu, J.; Ozawa, D.; Nagai, Y.; Ishihama, A. ALS-linked FUS mutations dysregulate G-quadruplex-dependent liquid-liquid phase separation and liquid-to-solid transition. J. Biol. Chem. 2021, 297, 101284. [Google Scholar] [CrossRef]
- Wang, S.; Xu, Y. RNA structure promotes liquid-to-solid phase transition of short RNAs in neuronal dysfunction. Commun. Biol. 2024, 7, 137. [Google Scholar] [CrossRef]
- Richter, J.D.; Zhao, X. The molecular biology of FMRP: New insights into fragile X syndrome. Nat. Rev. Neurosci. 2021, 22, 209–222. [Google Scholar] [CrossRef]
- Ashley, C.T., Jr.; Wilkinson, K.D.; Reines, D.; Warren, S.T. FMR1 protein: Conserved RNP family domains and selective RNA binding. Science 1993, 262, 563–566. [Google Scholar] [CrossRef] [PubMed]
- Schaeffer, C.; Bardoni, B.; Mandel, J.L.; Ehresmann, B.; Ehresmann, C.; Moine, H. The fragile X mental retardation protein binds specifically to its mRNA via a purine quartet motif. EMBO J. 2001, 20, 4803–4813. [Google Scholar] [CrossRef]
- Comery, T.A.; Harris, J.B.; Willems, P.J.; Oostra, B.A.; Irwin, S.A.; Weiler, I.J.; Greenough, W.T. Abnormal dendritic spines in fragile X knockout mice: Maturation and pruning deficits. Proc. Natl. Acad. Sci. USA 1997, 94, 5401–5404. [Google Scholar] [CrossRef]
- Deng, B.; Wan, G. Technologies for studying phase-separated biomolecular condensates. Adv. Biotechnol. 2024, 2, 10. [Google Scholar] [CrossRef]
- Ong, J.Y.; Torres, J.Z. Phase Separation in Cell Division. Mol. Cell 2020, 80, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Wiesenfarth, M.; Huppertz, H.J.; Dorst, J.; Lule, D.; Ludolph, A.C.; Muller, H.P.; Kassubek, J. Structural and microstructural neuroimaging signature of C9orf72-associated ALS: A multiparametric MRI study. Neuroimage Clin. 2023, 39, 103505. [Google Scholar] [CrossRef] [PubMed]
- Roytman, M.; Mashriqi, F.; Al-Tawil, K.; Schulz, P.E.; Zaharchuk, G.; Benzinger, T.L.S.; Franceschi, A.M. Amyloid-Related Imaging Abnormalities: An Update. AJR Am. J. Roentgenol. 2023, 220, 562–574. [Google Scholar] [CrossRef]
- Bidesi, N.S.R.; Vang Andersen, I.; Windhorst, A.D.; Shalgunov, V.; Herth, M.M. The role of neuroimaging in Parkinson’s disease. J. Neurochem. 2021, 159, 660–689. [Google Scholar] [CrossRef]
- Berkeley, R.F.; Kashefi, M.; Debelouchina, G.T. Real-time observation of structure and dynamics during the liquid-to-solid transition of FUS LC. Biophys. J. 2021, 120, 1276–1287. [Google Scholar] [CrossRef]
- Sakurai, K.; Morimoto, S.; Oguri, T.; Yuasa, H.; Uchida, Y.; Yamada, K.; Muto, M.; Saito, Y.; Aiba, I.; Takao, M.; et al. Multifaceted structural magnetic resonance imaging findings in demented patients with pathologically confirmed TDP-43 proteinopathy. Neuroradiology 2019, 61, 1333–1339. [Google Scholar] [CrossRef]
- Scheltens, P.; Blennow, K.; Breteler, M.M.; de Strooper, B.; Frisoni, G.B.; Salloway, S.; Van der Flier, W.M. Alzheimer’s disease. Lancet 2016, 388, 505–517. [Google Scholar] [CrossRef] [PubMed]
- Creekmore, B.C.; Chang, Y.W.; Lee, E.B. The Cryo-EM Effect: Structural Biology of Neurodegenerative Disease Aggregates. J. Neuropathol. Exp. Neurol. 2021, 80, 514–529. [Google Scholar] [CrossRef]
- Fitzpatrick, A.W.P.; Falcon, B.; He, S.; Murzin, A.G.; Murshudov, G.; Garringer, H.J.; Crowther, R.A.; Ghetti, B.; Goedert, M.; Scheres, S.H.W. Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 2017, 547, 185–190. [Google Scholar] [CrossRef]
- Schweighauser, M.; Shi, Y.; Tarutani, A.; Kametani, F.; Murzin, A.G.; Ghetti, B.; Matsubara, T.; Tomita, T.; Ando, T.; Hasegawa, K.; et al. Structures of alpha-synuclein filaments from multiple system atrophy. Nature 2020, 585, 464–469. [Google Scholar] [CrossRef]
- Kollmer, M.; Close, W.; Funk, L.; Rasmussen, J.; Bsoul, A.; Schierhorn, A.; Schmidt, M.; Sigurdson, C.J.; Jucker, M.; Fandrich, M. Cryo-EM structure and polymorphism of Abeta amyloid fibrils purified from Alzheimer’s brain tissue. Nat. Commun. 2019, 10, 4760. [Google Scholar] [CrossRef] [PubMed]
- Cao, Q.; Boyer, D.R.; Sawaya, M.R.; Ge, P.; Eisenberg, D.S. Cryo-EM structures of four polymorphic TDP-43 amyloid cores. Nat. Struct. Mol. Biol. 2019, 26, 619–627. [Google Scholar] [CrossRef] [PubMed]
- Espargaro, A.; Sabate, R. Phosphorylation-driven aggregative proteins in neurodegenerative diseases: Implications and therapeutics. Neural Regen. Res. 2024, 19, 966–968. [Google Scholar] [CrossRef] [PubMed]
- Ko, V.I.; Ong, K.; Cleveland, D.W.; Yu, H.; Ravits, J.M. CK1delta/epsilon kinases regulate TDP-43 phosphorylation and are therapeutic targets for ALS-related TDP-43 hyperphosphorylation. Neurobiol. Dis. 2024, 196, 106516. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Gonzalez, L.; Rodriguez-Cueto, C.; Cabezudo, D.; Bartolome, F.; Andres-Benito, P.; Ferrer, I.; Gil, C.; Martin-Requero, A.; Fernandez-Ruiz, J.; Martinez, A.; et al. Motor neuron preservation and decrease of in vivo TDP-43 phosphorylation by protein CK-1delta kinase inhibitor treatment. Sci. Rep. 2020, 10, 4449. [Google Scholar] [CrossRef]
- Leblond, J.; Petitjean, A. Molecular tweezers: Concepts and applications. Chemphyschem 2011, 12, 1043–1051. [Google Scholar] [CrossRef]
- Sinha, S.; Lopes, D.H.; Du, Z.; Pang, E.S.; Shanmugam, A.; Lomakin, A.; Talbiersky, P.; Tennstaedt, A.; McDaniel, K.; Bakshi, R.; et al. Lysine-specific molecular tweezers are broad-spectrum inhibitors of assembly and toxicity of amyloid proteins. J. Am. Chem. Soc. 2011, 133, 16958–16969. [Google Scholar] [CrossRef]
- Zheng, X.; Liu, D.; Klarner, F.G.; Schrader, T.; Bitan, G.; Bowers, M.T. Amyloid beta-protein assembly: The effect of molecular tweezers CLR01 and CLR03. J. Phys. Chem. B 2015, 119, 4831–4841. [Google Scholar] [CrossRef]
- Di, J.; Siddique, I.; Li, Z.; Malki, G.; Hornung, S.; Dutta, S.; Hurst, I.; Ishaaya, E.; Wang, A.; Tu, S.; et al. The molecular tweezer CLR01 improves behavioral deficits and reduces tau pathology in P301S-tau transgenic mice. Alzheimers Res. Ther. 2021, 13, 6. [Google Scholar] [CrossRef]
- Bengoa-Vergniory, N.; Faggiani, E.; Ramos-Gonzalez, P.; Kirkiz, E.; Connor-Robson, N.; Brown, L.V.; Siddique, I.; Li, Z.; Vingill, S.; Cioroch, M.; et al. CLR01 protects dopaminergic neurons in vitro and in mouse models of Parkinson’s disease. Nat. Commun. 2020, 11, 4885. [Google Scholar] [CrossRef]
- Vopel, T.; Bravo-Rodriguez, K.; Mittal, S.; Vachharajani, S.; Gnutt, D.; Sharma, A.; Steinhof, A.; Fatoba, O.; Ellrichmann, G.; Nshanian, M.; et al. Inhibition of Huntingtin Exon-1 Aggregation by the Molecular Tweezer CLR01. J. Am. Chem. Soc. 2017, 139, 5640–5643. [Google Scholar] [CrossRef] [PubMed]
- Malik, R.; Meng, H.; Wongkongkathep, P.; Corrales, C.I.; Sepanj, N.; Atlasi, R.S.; Klarner, F.G.; Schrader, T.; Spencer, M.J.; Loo, J.A.; et al. The molecular tweezer CLR01 inhibits aberrant superoxide dismutase 1 (SOD1) self-assembly in vitro and in the G93A-SOD1 mouse model of ALS. J. Biol. Chem. 2019, 294, 3501–3513. [Google Scholar] [CrossRef]
- Samanta, N.; Ruiz-Blanco, Y.B.; Fetahaj, Z.; Gnutt, D.; Lantz, C.; Loo, J.A.; Sanchez-Garcia, E.; Ebbinghaus, S. Superoxide Dismutase 1 Folding Stability as a Target for Molecular Tweezers in SOD1-Related Amyotrophic Lateral Sclerosis. Chembiochem 2022, 23, e202200396. [Google Scholar] [CrossRef]
- Ekman, F.K.; Ojala, D.S.; Adil, M.M.; Lopez, P.A.; Schaffer, D.V.; Gaj, T. CRISPR-Cas9-Mediated Genome Editing Increases Lifespan and Improves Motor Deficits in a Huntington’s Disease Mouse Model. Mol. Ther. Nucleic Acids 2019, 17, 829–839. [Google Scholar] [CrossRef] [PubMed]
- Elden, A.C.; Kim, H.J.; Hart, M.P.; Chen-Plotkin, A.S.; Johnson, B.S.; Fang, X.; Armakola, M.; Geser, F.; Greene, R.; Lu, M.M.; et al. Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature 2010, 466, 1069–1075. [Google Scholar] [CrossRef] [PubMed]
- Becker, L.A.; Huang, B.; Bieri, G.; Ma, R.; Knowles, D.A.; Jafar-Nejad, P.; Messing, J.; Kim, H.J.; Soriano, A.; Auburger, G.; et al. Therapeutic reduction of ataxin-2 extends lifespan and reduces pathology in TDP-43 mice. Nature 2017, 544, 367–371. [Google Scholar] [CrossRef]
- Nussbacher, J.K.; Tabet, R.; Yeo, G.W.; Lagier-Tourenne, C. Disruption of RNA Metabolism in Neurological Diseases and Emerging Therapeutic Interventions. Neuron 2019, 102, 294–320. [Google Scholar] [CrossRef]
- Hua, Y.; Vickers, T.A.; Okunola, H.L.; Bennett, C.F.; Krainer, A.R. Antisense masking of an hnRNP A1/A2 intronic splicing silencer corrects SMN2 splicing in transgenic mice. Am. J. Hum. Genet. 2008, 82, 834–848. [Google Scholar] [CrossRef]
- Hua, Y.; Sahashi, K.; Hung, G.; Rigo, F.; Passini, M.A.; Bennett, C.F.; Krainer, A.R. Antisense correction of SMN2 splicing in the CNS rescues necrosis in a type III SMA mouse model. Genes Dev. 2010, 24, 1634–1644. [Google Scholar] [CrossRef]
- Chiriboga, C.A.; Swoboda, K.J.; Darras, B.T.; Iannaccone, S.T.; Montes, J.; De Vivo, D.C.; Norris, D.A.; Bennett, C.F.; Bishop, K.M. Results from a phase 1 study of nusinersen (ISIS-SMNRx) in children with spinal muscular atrophy. Neurology 2016, 86, 890–897. [Google Scholar] [CrossRef]
- Finkel, R.S.; Chiriboga, C.A.; Vajsar, J.; Day, J.W.; Montes, J.; De Vivo, D.C.; Yamashita, M.; Rigo, F.; Hung, G.; Schneider, E.; et al. Treatment of infantile-onset spinal muscular atrophy with nusinersen: A phase 2, open-label, dose-escalation study. Lancet 2016, 388, 3017–3026. [Google Scholar] [CrossRef] [PubMed]
- Finkel, R.S.; Mercuri, E.; Darras, B.T.; Connolly, A.M.; Kuntz, N.L.; Kirschner, J.; Chiriboga, C.A.; Saito, K.; Servais, L.; Tizzano, E.; et al. Nusinersen versus Sham Control in Infantile-Onset Spinal Muscular Atrophy. N. Engl. J. Med. 2017, 377, 1723–1732. [Google Scholar] [CrossRef] [PubMed]
- Mercuri, E.; Darras, B.T.; Chiriboga, C.A.; Day, J.W.; Campbell, C.; Connolly, A.M.; Iannaccone, S.T.; Kirschner, J.; Kuntz, N.L.; Saito, K.; et al. Nusinersen versus Sham Control in Later-Onset Spinal Muscular Atrophy. N. Engl. J. Med. 2018, 378, 625–635. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.S.; Wu, H.; Krzisch, M.; Wu, X.; Graef, J.; Muffat, J.; Hnisz, D.; Li, C.H.; Yuan, B.; Xu, C.; et al. Rescue of Fragile X Syndrome Neurons by DNA Methylation Editing of the FMR1 Gene. Cell 2018, 172, 979–992.e6. [Google Scholar] [CrossRef]
- Xie, N.; Gong, H.; Suhl, J.A.; Chopra, P.; Wang, T.; Warren, S.T. Reactivation of FMR1 by CRISPR/Cas9-Mediated Deletion of the Expanded CGG-Repeat of the Fragile X Chromosome. PLoS ONE 2016, 11, e0165499. [Google Scholar] [CrossRef]
- Cummings, J.L.; Morstorf, T.; Zhong, K. Alzheimer’s disease drug-development pipeline: Few candidates, frequent failures. Alzheimers Res. Ther. 2014, 6, 37. [Google Scholar] [CrossRef]
- Yoo, Y.; Neumayer, G.; Shibuya, Y.; Mader, M.M.; Wernig, M. A cell therapy approach to restore microglial Trem2 function in a mouse model of Alzheimer’s disease. Cell Stem Cell 2023, 30, 1043–1053.e6. [Google Scholar] [CrossRef]
- Mishra, P.; Silva, A.; Sharma, J.; Nguyen, J.; Pizzo, D.P.; Hinz, D.; Sahoo, D.; Cherqui, S. Rescue of Alzheimer’s disease phenotype in a mouse model by transplantation of wild-type hematopoietic stem and progenitor cells. Cell Rep. 2023, 42, 112956. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Sun, Z.; Peng, D.; Gianchandani, S.; Le, W.; Boltze, J.; Li, S. Cell-therapy for Parkinson’s disease: A systematic review and meta-analysis. J. Transl. Med. 2023, 21, 601. [Google Scholar] [CrossRef]
- Inuzuka, H.; Liu, J.; Wei, W.; Rezaeian, A.H. PROTACs technology for treatment of Alzheimer’s disease: Advances and perspectives. Acta Mater. Med. 2022, 1, 24–41. [Google Scholar] [CrossRef]
- Tseng, Y.L.; Lu, P.C.; Lee, C.C.; He, R.Y.; Huang, Y.A.; Tseng, Y.C.; Cheng, T.R.; Huang, J.J.; Fang, J.M. Degradation of neurodegenerative disease-associated TDP-43 aggregates and oligomers via a proteolysis-targeting chimera. J. Biomed. Sci. 2023, 30, 27. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Zhang, W.; Chen, J.; Tong, Y.; Xu, F.; Pang, J. Discovery of Effective Dual PROTAC Degraders for Neurodegenerative Disease-Associated Aggregates. J. Med. Chem. 2024, 67, 3448–3466. [Google Scholar] [CrossRef] [PubMed]
- Miller, L.V.C.; Papa, G.; Vaysburd, M.; Cheng, S.; Sweeney, P.W.; Smith, A.; Franco, C.; Katsinelos, T.; Huang, M.; Sanford, S.A.I.; et al. Co-opting templated aggregation to degrade pathogenic tau assemblies and improve motor function. Cell, 2024; online ahead of print. [Google Scholar] [CrossRef]
- Schenk, D.; Barbour, R.; Dunn, W.; Gordon, G.; Grajeda, H.; Guido, T.; Hu, K.; Huang, J.; Johnson-Wood, K.; Khan, K.; et al. Immunization with amyloid-beta attenuates Alzheimer-disease-like pathology in the PDAPP mouse. Nature 1999, 400, 173–177. [Google Scholar] [CrossRef] [PubMed]
- Janus, C.; Pearson, J.; McLaurin, J.; Mathews, P.M.; Jiang, Y.; Schmidt, S.D.; Chishti, M.A.; Horne, P.; Heslin, D.; French, J.; et al. A beta peptide immunization reduces behavioural impairment and plaques in a model of Alzheimer’s disease. Nature 2000, 408, 979–982. [Google Scholar] [CrossRef]
- Hou, J.; Chen, Y.; Cai, Z.; Heo, G.S.; Yuede, C.M.; Wang, Z.; Lin, K.; Saadi, F.; Trsan, T.; Nguyen, A.T.; et al. Antibody-mediated targeting of human microglial leukocyte Ig-like receptor B4 attenuates amyloid pathology in a mouse model. Sci. Transl. Med. 2024, 16, eadj9052. [Google Scholar] [CrossRef]
- Wal, A.; Wal, P.; Vig, H.; Jain, N.K.; Rathore, S.; Krishnan, K.; Srivastava, A. Treatment of Parkinson’s Disease: Current Treatments and Recent Therapeutic Developments. Curr. Drug Discov. Technol. 2023, 20, e120523216834. [Google Scholar] [CrossRef]
- Shan, L.; Wang, W.; Du, L.; Li, D.; Wang, Y.; Xie, Y.; Li, H.; Wang, J.; Shi, Z.; Zhou, Y.; et al. SP1 undergoes phase separation and activates RGS20 expression through super-enhancers to promote lung adenocarcinoma progression. Proc. Natl. Acad. Sci. USA 2024, 121, e2401834121. [Google Scholar] [CrossRef]
- Wang, W.; Li, D.; Xu, Q.; Cheng, J.; Yu, Z.; Li, G.; Qiao, S.; Pan, J.; Wang, H.; Shi, J.; et al. G-quadruplexes promote the motility in MAZ phase-separated condensates to activate CCND1 expression and contribute to hepatocarcinogenesis. Nat. Commun. 2024, 15, 1045. [Google Scholar] [CrossRef]
- Huang, L.K.; Kuan, Y.C.; Lin, H.W.; Hu, C.J. Clinical trials of new drugs for Alzheimer disease: A 2020–2023 update. J. Biomed. Sci. 2023, 30, 83. [Google Scholar] [CrossRef] [PubMed]
- Urrestizala-Arenaza, N.; Cerchio, S.; Cavaliere, F.; Magliaro, C. Limitations of human brain organoids to study neurodegenerative diseases: A manual to survive. Front. Cell Neurosci. 2024, 18, 1419526. [Google Scholar] [CrossRef]
| Diseases | Key Proteins | Genes | Functions | Consequence of Mutations | References |
|---|---|---|---|---|---|
| AD | Aβ (or APP) | APP | Cell surface receptor | Increase of Aβ, especially the Aβ-42 peptide | [23] |
| γ-secretase | PSEN1/ PSEN2 | APP cleavage to form Aβ | Increased ratio of Aβ-42/Aβ-40 to form amyloid plaques | [24] | |
| tau | MAPT | A microtubule- associated protein | Formation of tau aggregates toxic to neuronal cells | [25] | |
| PD | α-syn | SNCA | A neuronal protein for synaptic vesicle trafficking and neurotransmitter release | Formation of insoluble amyloid fibrils in Lewy bodies | [26] |
| GBA1 | GBA1 | Glucocerebrosidase β1 | Accumulation of glycolipid substrates | [27] | |
| ferritin | FTH, FTL | Conversion of Fe2+ to Fe3+ iron | Generation of hydroxyl radicals for PD pathogenesis | [28,29] | |
| HD | HTT | HTT | Cellular signaling and axonal transport | Formation of aggregates to generate Huntington’s bodies | [30,31] |
| ALS | SOD1 | SOD1 | Destruction of free superoxide radicals | Aggregation leading to ALS development | [32] |
| TDP-43 | TDP-43 | A transcriptional factor | Altered subcellular localization to form aggregates | [33,34] | |
| FUS | FUS | Transcription, RNA processing, DNA repair | Altered subcellular localization to form cytoplasmic inclusions | [35] | |
| AMD | ApoE | APOE | A lipoprotein as a cholesterol transporter | ApoE polymorphism isoforms with different risks for AMD development | [36,37] |
| RPE65 | RPE65 | A retinoid isomerohydrolase | Increase of RPE65 aggregation propensity | [38,39] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, Y.; Ma, B.; Liu, C.; Li, D.; Sui, G. Pathological Involvement of Protein Phase Separation and Aggregation in Neurodegenerative Diseases. Int. J. Mol. Sci. 2024, 25, 10187. https://doi.org/10.3390/ijms251810187
Wu Y, Ma B, Liu C, Li D, Sui G. Pathological Involvement of Protein Phase Separation and Aggregation in Neurodegenerative Diseases. International Journal of Molecular Sciences. 2024; 25(18):10187. https://doi.org/10.3390/ijms251810187
Chicago/Turabian StyleWu, Yinuo, Biao Ma, Chang Liu, Dangdang Li, and Guangchao Sui. 2024. "Pathological Involvement of Protein Phase Separation and Aggregation in Neurodegenerative Diseases" International Journal of Molecular Sciences 25, no. 18: 10187. https://doi.org/10.3390/ijms251810187
APA StyleWu, Y., Ma, B., Liu, C., Li, D., & Sui, G. (2024). Pathological Involvement of Protein Phase Separation and Aggregation in Neurodegenerative Diseases. International Journal of Molecular Sciences, 25(18), 10187. https://doi.org/10.3390/ijms251810187






