Heterogeneous Transcriptional Landscapes in Human Sporadic Parathyroid Gland Tumors
Abstract
:1. Introduction
2. Results
2.1. Clinical Features of PHPT Patients Harbouring PAds
2.2. Gene Expression by Real-Time PCR
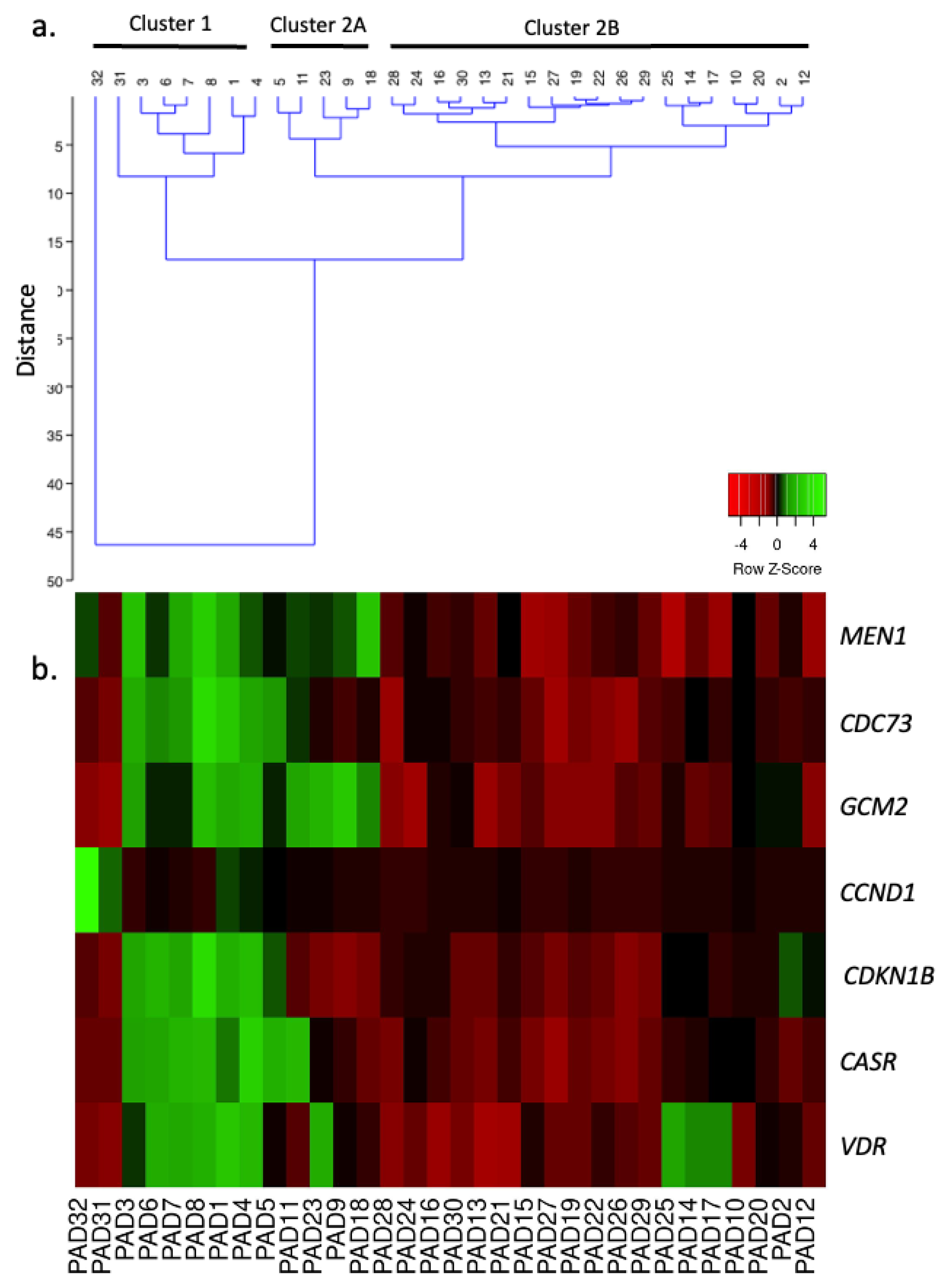
2.3. Unsupervised Clustering Analysis
2.4. Selected mRNA, miRNA, and lncRNA Expression Profiles in the Different PAd Clusters
2.5. Clinical Features Associated with the Different Gene Profiles
2.6. Prediction of Intracellular Pathway Deregulated in PAds
3. Discussion
4. Materials and Methods
4.1. Parathyroid Tumor Samples
4.2. RNA Extraction
4.3. Real-Time PCR
4.4. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Agarwal, S.K. Exploring the tumors of multiple endocrine neoplasia type 1 in mouse models for basic and preclinical studies. Int. J. Endocr. Oncol. 2014, 1, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Harding, B.; Lemos, M.C.; Reed, A.A.; Walls, G.V.; Jeyabalan, J.; Bowl, M.R.; Tateossian, H.; Sullivan, N.; Hough, T.; Fraser, W.D.; et al. Multiple endocrine neoplasia type 1 knockout mice develop parathyroid, pancreatic, pituitary and adrenal tumours with hypercalcaemia, hypophosphataemia and hypercorticosteronaemia. Endocr. Relat. Cancer 2009, 16, 1313–1327. [Google Scholar] [CrossRef] [PubMed]
- Walls, G.V.; Stevenson, M.; Lines, K.E.; Newey, P.J.; Reed, A.A.C.; Bowl, M.R.; Jeyabalan, J.; Harding, B.; Bradley, K.J.; Manek, S.; et al. Mice deleted for cell division cycle 73 gene develop parathyroid and uterine tumours: Model for the hyperparathyroidism-jaw tumour syndrome. Oncogene 2017, 36, 4025–4036. [Google Scholar] [CrossRef] [PubMed]
- Canaff, L.; Guarnieri, V.; Kim, Y.; Wong, B.Y.L.; Nolin-Lapalme, A.; Cole, D.E.C.; Minisola, S.; Eller-Vainicher, C.; Cetani, F.; Repaci, A.; et al. Novel Glial Cells Missing-2 (GCM2) variants in parathyroid disorders. Eur. J. Endocrinol. 2022, 186, 351–366. [Google Scholar] [CrossRef] [PubMed]
- Guan, B.; Welch, J.M.; Sapp, J.C.; Ling, H.; Li, Y.; Johnston, J.J.; Kebebew, E.; Biesecker, L.G.; Simonds, W.F.; Marx, S.J.; et al. GCM2-activating mutations in familial isolated hyperparathyroidism. Am. J. Hum. Genet. 2016, 99, 1034–1044. [Google Scholar] [CrossRef]
- Riccardi, A.; Aspir, T.; Shen, L.; Kuo, C.L.; Brown, T.C.; Korah, R.; Murtha, T.D.; Bellizzi, J.; Parham, K.; Carling, T.; et al. Analysis of activating GCM2 sequence variants in sporadic parathyroid adenomas. J. Clin. Endocrinol. Metab. 2019, 104, 1948–1952. [Google Scholar] [CrossRef]
- Song, A.; Yang, Y.; Wang, Y.; Liu, S.; Nie, M.; Jiang, Y.; Li, M.; Xia, W.; Wang, O.; Xing, X. Germline GCM2 mutation screening in Chinese primary hyperparathyroidism patients. Endocr. Pract. 2020, 26, 1093–1104. [Google Scholar] [CrossRef]
- Vincze, S.; Peters, N.V.; Kuo, C.L.; Brown, T.C.; Korah, R.; Murtha, T.D.; Bellizzi, J.; Riccardi, A.; Parham, K.; Carling, T.; et al. GCM2 variants in familial and multiglandular primary hyperparathyroidism. J. Clin. Endocrinol. Metab. 2022, 107, e2021–e2026. [Google Scholar] [CrossRef]
- Pellegata, N.S.; Quintanilla-Martinez, L.; Siggelkow, H.; Samson, E.; Bink, K.; Höfler, H.; Fend, F.; Graw, J.; Atkinson, M.J. Germ-line mutations in p27Kip1 cause a multiple endocrine neoplasia syndrome in rats and humans. Proc. Natl. Acad. Sci. USA 2006, 103, 15558–15563. [Google Scholar] [CrossRef]
- Lee, M.; Pellegata, N.S. Multiple endocrine neoplasia syndromes associated with mutation of p27. J. Endocrinol. Investig. 2013, 36, 781–787. [Google Scholar] [CrossRef]
- Mallya, S.M.; Gallagher, J.J.; Wild, Y.K.; Kifor, O.; Costa-Guda, J.; Saucier, K.; Brown, E.M.; Arnold, A. Abnormal parathyroid cell proliferation precedes biochemical abnormalities in a mouse model of primary hyperparathyroidism. Mol. Endocrinol. 2005, 19, 2603–2609. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Liu, W.; Bi, R.; Densmore, M.J.; Sato, T.; Mannstadt, M.; Yuan, Q.; Zhou, X.; Olauson, H.; Larsson, T.E.; et al. Interrelated role of Klotho and calcium-sensing receptor in parathyroid hormone synthesis and parathyroid hyperplasia. Proc. Natl. Acad. Sci. USA 2018, 115, E3749–E3758. [Google Scholar] [CrossRef] [PubMed]
- Ho, C.; Conner, D.A.; Pollak, M.R.; Ladd, D.J.; Kifor, O.; Warren, H.B.; Brown, E.M.; Seidman, J.G.; Seidman, C.E. A mouse model of human familial hypocalciuric hypercalcemia and neonatal severe hyperparathyroidism. Nat. Genet. 1995, 11, 389–394. [Google Scholar] [CrossRef] [PubMed]
- Meir, T.; Levi, R.; Lieben, L.; Libutti, S.; Carmeliet, G.; Bouillon, R.; Silver, J.; Naveh-Many, T. Deletion of the vitamin D receptor specifically in the parathyroid demonstrates a limited role for the receptor in parathyroid physiology. Am. J. Physiol. Ren. Physiol. 2009, 297, F1192–F1198. [Google Scholar] [CrossRef]
- Chorti, A.; Cheva, A.; Chatzikyriakidou, A.; Achilla, C.; Boulogeorgou, K.; Despoina, K.; Milias, S.; Zarampoukas, T.; Papavramidis, T. Sporadic parathyroid adenoma: An updated review of molecular genetics. Front. Endocrinol. 2023, 14, 1180211. [Google Scholar] [CrossRef]
- Marini, F.; Giusti, F.; Iantomasi, T.; Brandi, M.L. Parathyroid tumors: Molecular signatures. Int. J. Mol. Sci. 2021, 22, 11206. [Google Scholar] [CrossRef]
- Aurilia, C.; Donati, S.; Palmini, G.; Miglietta, F.; Falsetti, I.; Iantomasi, T.; Brandi, M.L. Are non-coding RNAs useful biomarkers in parathyroid tumorigenesis? Int. J. Mol. Sci. 2021, 22, 10465. [Google Scholar] [CrossRef] [PubMed]
- Levey, A.S.; Eckardt, K.U.; Tsukamoto, Y.; Levin, A.; Coresh, J.; Rossert, J.; De Zeeuw, D.; Hostetter, T.H.; Lameire, N.; Eknoyan, G. Definition and classification of chronic kidney disease: A position statement from Kidney Disease: Improving Global Outcomes (KDIGO). Kidney Int. 2005, 67, 2089–2100. [Google Scholar] [CrossRef]
- Erickson, L.A.; Mete, O.; Juhlin, C.C.; Perren, A.; Gill, A.J. Overview of the 2022 WHO Classification of parathyroid tumors. Endocr. Pathol. 2022, 33, 64–89. [Google Scholar] [CrossRef]
- Perfetto, L.; Briganti, L.; Calderone, A.; Cerquone Perpetuini, A.; Iannuccelli, M.; Langone, F.; Licata, L.; Marinkovic, M.; Mattioni, A.; Pavlidou, T.; et al. SIGNOR: A database of causal relationships between biological entities. Nucleic Acids Res. 2016, 44, D548–D554. [Google Scholar] [CrossRef]
- Medina, J.E.; Randolph, G.W.; Angelos, P.; Zafereo, M.E.; Tufano, R.P.; Kowalski, L.P.; Montenegro, F.L.M.; Owen, R.P.; Khafif, A.; Suárez, C.; et al. Primary hyperparathyroidism: Disease of diverse genetic, symptomatic, and biochemical phenotypes. Head Neck 2021, 43, 3996–4009. [Google Scholar] [CrossRef] [PubMed]
- Rizk, Y.; Saad, N.; Arnaout, W.; Chalah, M.A.; Farah, S. Primary hyperparathyroidism in older adults: A narrative review of the most recent literature on epidemiology, diagnosis and management. J. Clin. Med. 2023, 12, 6321. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Zhang, X.; Wang, O.; Cui, M.; Li, X.; Wang, M.; Hua, S.; Liao, Q. Integrated whole-exome and transcriptome sequencing of sporadic parathyroid adenoma. Front. Endocrinol. 2021, 12, 631680. [Google Scholar] [CrossRef]
- Sweeney, K.; Cameron, E.R.; Blyth, K. Complex interplay between the RUNX transcription factors and Wnt/beta-catenin pathway in cancer: A tango in the night. Mol. Cells 2020, 43, 188–197. [Google Scholar] [CrossRef] [PubMed]
- Rozenberg, J.M.; Zvereva, S.; Dalina, A.; Blatov, I.; Zubarev, I.; Luppov, D.; Bessmertnyi, A.; Romanishin, A.; Alsoulaiman, L.; Kumeiko, V.; et al. Dual Role of p73 in Cancer Microenvironment and DNA Damage Response. Cells 2021, 10, 3516. [Google Scholar] [CrossRef]
- Yao, Q.; Wang, C.; Wang, Y.; Zhang, X.; Jiang, H.; Chen, D. The integrated comprehension of lncRNA HOXA-AS3 implication on human diseases. Clin. Transl. Oncol. 2022, 24, 2342–2350. [Google Scholar] [CrossRef]
- Lu, B.; Liu, Z.; Ji, X.; Zhang, X. Assessment of the prognostic and clinicopathological significance of HOXA-AS2 in human cancers: A systematic review and meta-analysis. Transl. Cancer Res. 2023, 12, 605–615. [Google Scholar] [CrossRef]
- Waters, E.; Pucci, P.; Hirst, M.; Chapman, S.; Wang, Y.; Crea, F.; Heath, C.J. HAR1: An insight into lncRNA genetic evolution. Epigenomics 2021, 13, 1831–1843. [Google Scholar] [CrossRef]
- Verdelli, C.; Morotti, A.; Tavanti, G.S.; Silipigni, R.; Guerneri, S.; Ferrero, S.; Vicentini, L.; Vaira, V.; Corbetta, S. The core stem genes SOX2, POU5F1/OCT4, and NANOG are expressed in human parathyroid tumors and modulated by MEN1, YAP1, and β-catenin pathways activation. Biomedicines 2021, 9, 637. [Google Scholar] [CrossRef]
- Mizobuchi, M.; Ritter, C.S.; Krits, I.; Slatopolsky, E.; Sicard, G.; Brown, A.J. Calcium-sensing receptor expression is regulated by glial cells missing-2 in human parathyroid cells. J. Bone Min. Res. 2009, 24, 1173–1179. [Google Scholar] [CrossRef]
- Hussain, M.S.; Gupta, G.; Afzal, M.; Alqahtani, S.M.; Samuel, V.P.; Hassan Almalki, W.; Kazmi, I.; Alzarea, S.I.; Saleem, S.; Dureja, H.; et al. Exploring the role of lncrna neat1 knockdown in regulating apoptosis across multiple cancer types: A review. Pathol. Res. Pract. 2023, 252, 154908. [Google Scholar] [CrossRef] [PubMed]
- Ji, G.; Ren, R.; Fang, X. Identification and characterization of non-coding RNAs in thymoma. Med. Sci. Monit. 2021, 27, e929727. [Google Scholar] [CrossRef] [PubMed]
- Guarnieri, V.; Muscarella, L.A.; Verdelli, C.; Corbetta, S. Alterations of DNA methylation in parathyroid tumors. Mol. Cell. Endocrinol. 2018, 469, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Vaira, V.; Elli, F.; Forno, I.; Guarnieri, V.; Verdelli, C.; Ferrero, S.; Scillitani, A.; Vicentini, L.; Cetani, F.; Mantovani, G.; et al. The microRNA cluster C19MC is deregulated in parathyroid tumours. J. Mol. Endocrinol. 2012, 49, 115–124. [Google Scholar] [CrossRef]
- Bilezikian, J.P.; Khan, A.A.; Clarke, B.L.; Mannstadt, M.; Potts, J.T.; Brandi, M.L. The Fifth International Workshop on the evaluation and management of primary hyperparathyroidism. J. Bone Miner. Res. 2022, 37, 2290–2292. [Google Scholar] [CrossRef]
| Gene | Cluster 1 | Cluster 2A | Cluster 2B | p |
|---|---|---|---|---|
| Oncosuppressors | ||||
| MEN1 | 1.921 ± 0.718 | 1.706 ± 0.555 | 0.770 ± 0.290 *,** | <0.0001 |
| CDC73 | 2.034 ± 1.046 | 1.252 ± 0.441 * | 0.806 ± 0.269 * | 0.0001 |
| RASSF1A | 1.417 (0.788, 2.022) | 1.596 (1.155, 2.578) | 0.836 (0.542, 1.048) ** | 0.0439 |
| YAP1 | 2.033 ± 1.210 | 1.403 ± 0.450 | 0.788 ± 0.324 * | 0.0004 |
| CTNNB1 | 2.198 ± 1.290 | 1.511 ± 1.028 | 0.877 ± 0.421 * | 0.0021 |
| Transcription factors | ||||
| GCM2 | 1.693 ± 0.955 | 2.315 ± 0.713 | 0.713 ± 0.357 * | <0.0001 |
| GATA3 | 1.629 (1.260, 2.498) | 1.202 (0.819, 1.462) | 0.857 (0.580, 1.040) * | 0.0041 |
| TBX1 | 1.157 (0.704, 2.010) | 1.711 (1.027, 2.211) | 0.806 (0.540, 1.091) | 0.1292 |
| PAX1 | 2.058 (0.854, 2.884) | 0.803 (0.533, 2.012) | 1.089 (0.515, 1.308) | 0.0742 |
| Parathyroid-specific genes | ||||
| PTH | 2.577 (1.393, 7.689) | 1.218 (0.488, 3.108) | 0.811 (0.509, 1.123) * | 0.0052 |
| CASR | 2.380 ± 1.252 | 1.833 ± 1.252 | 0.756 ± 0.313 *,** | 0.0001 |
| VDR | 2.972 (0.868, 3.417) | 1.298 (0.915, 2.201) | 0.713 (0.498, 1.206) * | 0.0252 |
| GPRC6A | 0.999 (0.542, 2.569) | 2.092 (0.931, 4.779) | 0.983 (0.364, 2.262) | 0.4753 |
| Cell cycle genes | ||||
| CCND1 | 2.340 (0.685, 5.710) | 1.484 (0.873, 2.008) | 0.676 (0.502, 0.954) *,** | 0.0058 |
| CDKN1B/p27 | 3.723 ± 2.229 | 0.809 ± 0.982 * | 1.039 ± 0.634* | <0.0001 |
| CDKN1A/p21 | 2.342 (1.186, 4.633) | 1.227 (0.193, 2.820) | 1.025 (0.292, 1.681) | 0.0679 |
| TP73 | 1.232 (0.579, 14.48) | 0.213 (0.213, 6.900) | 0.511 (0.213, 5.485) | 0.0930 |
| MicroRNAs | ||||
| miR-372-5p | 0.268 (0.194, 2.463) | 0.532 (0.304, 24.51) | 0.353 (0.231, 0.927) | 0.5641 |
| miR-517c | 0.217 (0.052, 0.752) | 0.098 (0.061, 7674) | 0.424 (0.135, 1.628) | 0.2115 |
| miR-126-3p | 0.744 ± 0.504 | 0.951 ± 0.498 | 1.270 ± 0.502 | 0.0507 |
| miR-93-5p | 0.681 ± 0.423 | 1.173 ± 0.412 | 1.068 ± 0.396 | 0.0560 |
| Long non-coding RNAs | ||||
| NEAT1 | 2.386 (1.636, 6.208) | 2.285 (1.629, 3.635) | 0.641 (0.402, 0.966) *,** | <0.0001 |
| HAR1B | 2.059 (0.519, 15.45) | 0.130 (0.075, 0.698) * | 2.507 (0.467, 3.592) ** | 0.0371 |
| HOXA-AS2 | 0.348 (0.092, 1.871) | 9.099 (3.443, 16.30) * | 0.866 (0.079, 1.496) ** | 0.0125 |
| HOXA-AS3 | 0.458 (0.458, 1.049) | 4.784 (2.910, 11.92) * | 0.458 (0.458, 1.302) ** | 0.0028 |
| VLDLR-AS1 | 2.279 (1.278, 3.423) | 1.976 (0.812, 2.556) | 0.760 (0.303, 1.521) * | 0.0139 |
| SNHG6 | 1.681 (0.875, 2.607) | 0.795 (0.509, 1.917) | 0.971 (0.513, 1.383) | 0.1202 |
| Features | n.v. | Cluster 1 | Cluster 2A | Cluster 2B | p |
|---|---|---|---|---|---|
| General and tumor features | |||||
| Female/male (%) | - | 8/0 (100.0) * | 2/3 (40.0) | 15/4 (78.9) | 0.0388 |
| Age (years) | - | 55.3 ± 7.8 | 59.8 ± 12.3 | 58.3 ± 11.2 | 0.2583 |
| BMI (kg/m2) | - | 23.7 ± 2.6 | 24.7 ± 1.8 | 25.6 ± 6.0 | 0.2863 |
| Tumor size (cm) | - | 1.88 ± 1.03 | 1.98 ± 0.38 | 1.27 ± 0.51 | 0.0376 |
| Biochemical and hormonal parameters | |||||
| Ca2+ (mmol/L) | 1.15–1.29 | 1.46 (1.35, 1.63) | 1.62 (1.53, 2.33) | 1.41 (1.38, 1.49) * | 0.0398 |
| S Total Ca (mg/dL) | 8.4–10.4 | 11.9 ± 1.2 | 14.7 ± 4.5 | 11.3 ± 0.7 * | 0.0097 |
| S Phosphate (mg/dL) | 3.5–5.0 | 2.7 ± 0.7 | 2.5 ± 0.6 | 2.1 ± 0.7 | 0.2479 |
| PTH (pg/mL) | 10.0–65.0 | 170.5 (110.5, 372.5) | 301.0 (195.0, 1031.0) | 135.0 (88.1, 173.8) * | 0.0427 |
| eGFR (mL/min) | >90.0 | 85.8 ± 18.6 | 85.7 ± 16.5 | 83.2 ± 13.0 | 0.9278 |
| UCa (mg/24 h) | <300.0 | 430.4 ± 183.6 | 451.4 ± 65.4 | 360.6 ± 128.9 | 0.3214 |
| PHPT-related complications | |||||
| Kidney stones (y/n, %) | - | 4/4 (50.0) | 3/2 (60.0) | 8/11 (42.1) | 0.7593 |
| Osteoporosis (y/n, %) | - | 6/2 (75.0) | 4/1 (80.0) | 10/9 (52.6) | 0.3723 |
| Hypertension (y/n, %) | - | 4/4 (50.0) | 1/4 (20.0) | 5/14 (26.3) | 0.4027 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Verdelli, C.; Carrara, S.; Maggiore, R.; Dalino Ciaramella, P.; Corbetta, S. Heterogeneous Transcriptional Landscapes in Human Sporadic Parathyroid Gland Tumors. Int. J. Mol. Sci. 2024, 25, 10782. https://doi.org/10.3390/ijms251910782
Verdelli C, Carrara S, Maggiore R, Dalino Ciaramella P, Corbetta S. Heterogeneous Transcriptional Landscapes in Human Sporadic Parathyroid Gland Tumors. International Journal of Molecular Sciences. 2024; 25(19):10782. https://doi.org/10.3390/ijms251910782
Chicago/Turabian StyleVerdelli, Chiara, Silvia Carrara, Riccardo Maggiore, Paolo Dalino Ciaramella, and Sabrina Corbetta. 2024. "Heterogeneous Transcriptional Landscapes in Human Sporadic Parathyroid Gland Tumors" International Journal of Molecular Sciences 25, no. 19: 10782. https://doi.org/10.3390/ijms251910782






