Mapping and Screening of Candidate Gene Regulating the Biomass Yield of Sorghum (Sorghum bicolor L.)
Abstract
:1. Introduction
2. Results
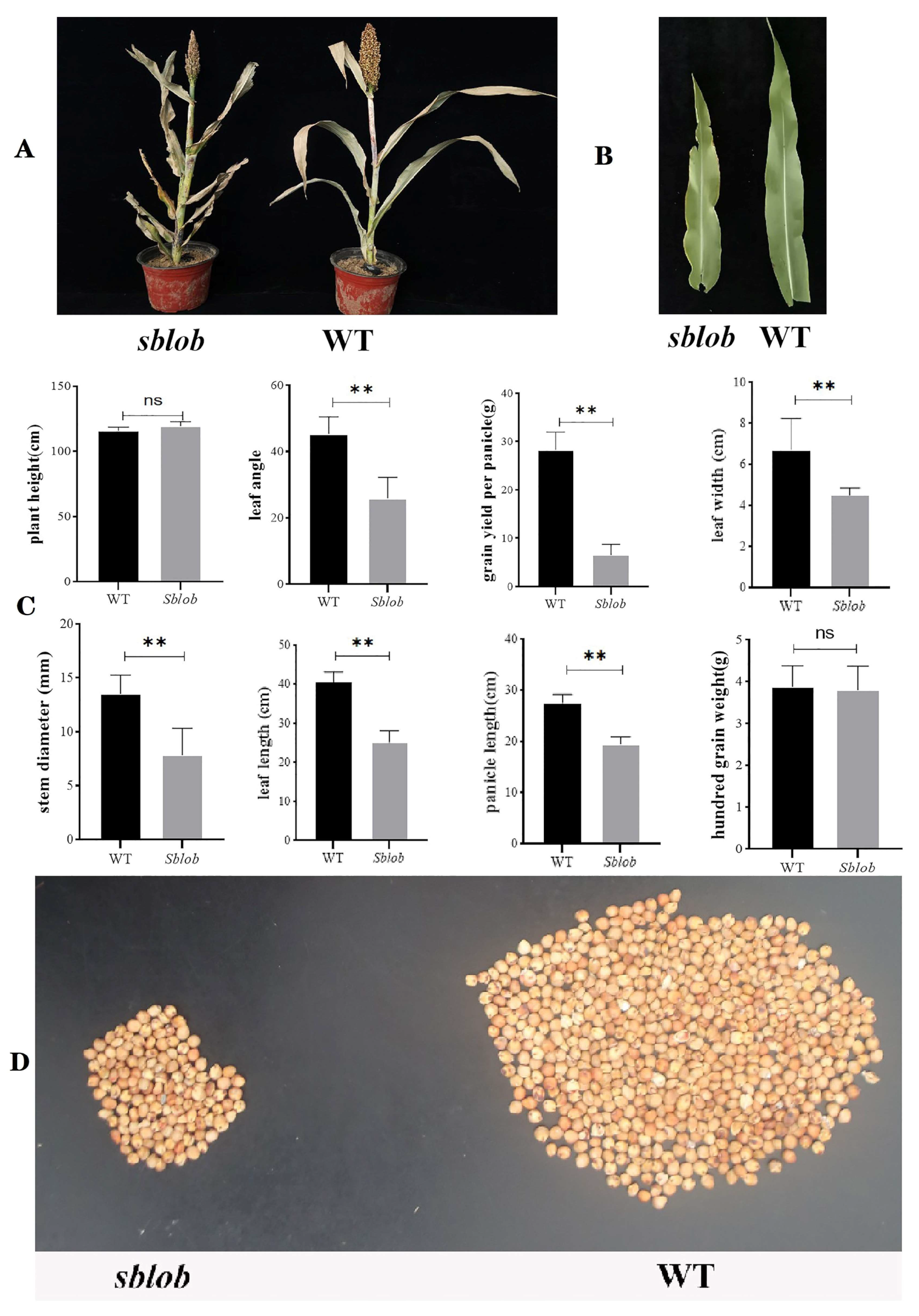
2.1. Phenotypic and Physiology Analysis of sblob and WT Plants
2.2. Genetic Analysis of the sblob Mutant
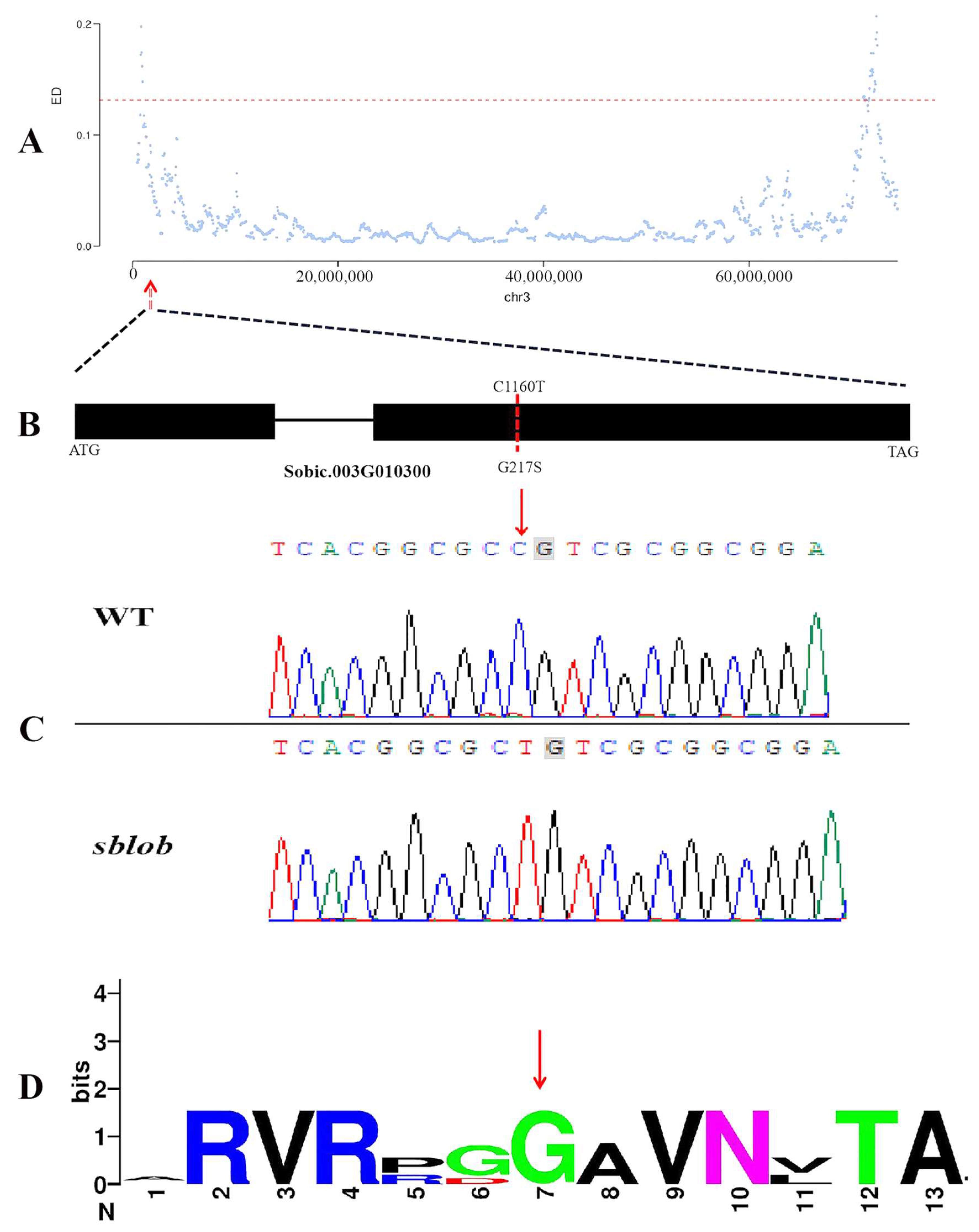
2.3. Mapping of the sblob1 Gene by Whole-Genome Resequencing
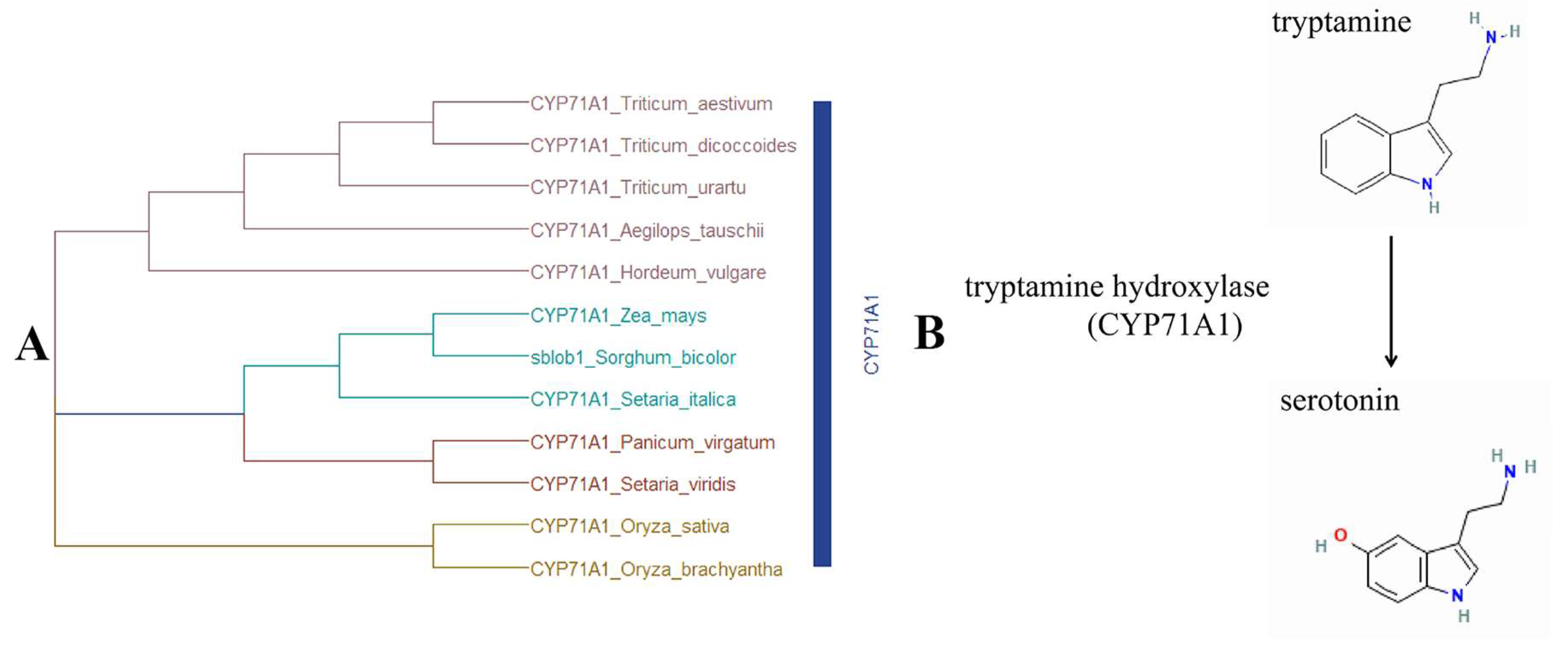
2.4. Identification and Validation of Candidate Gene Accounting for sblob Mutant
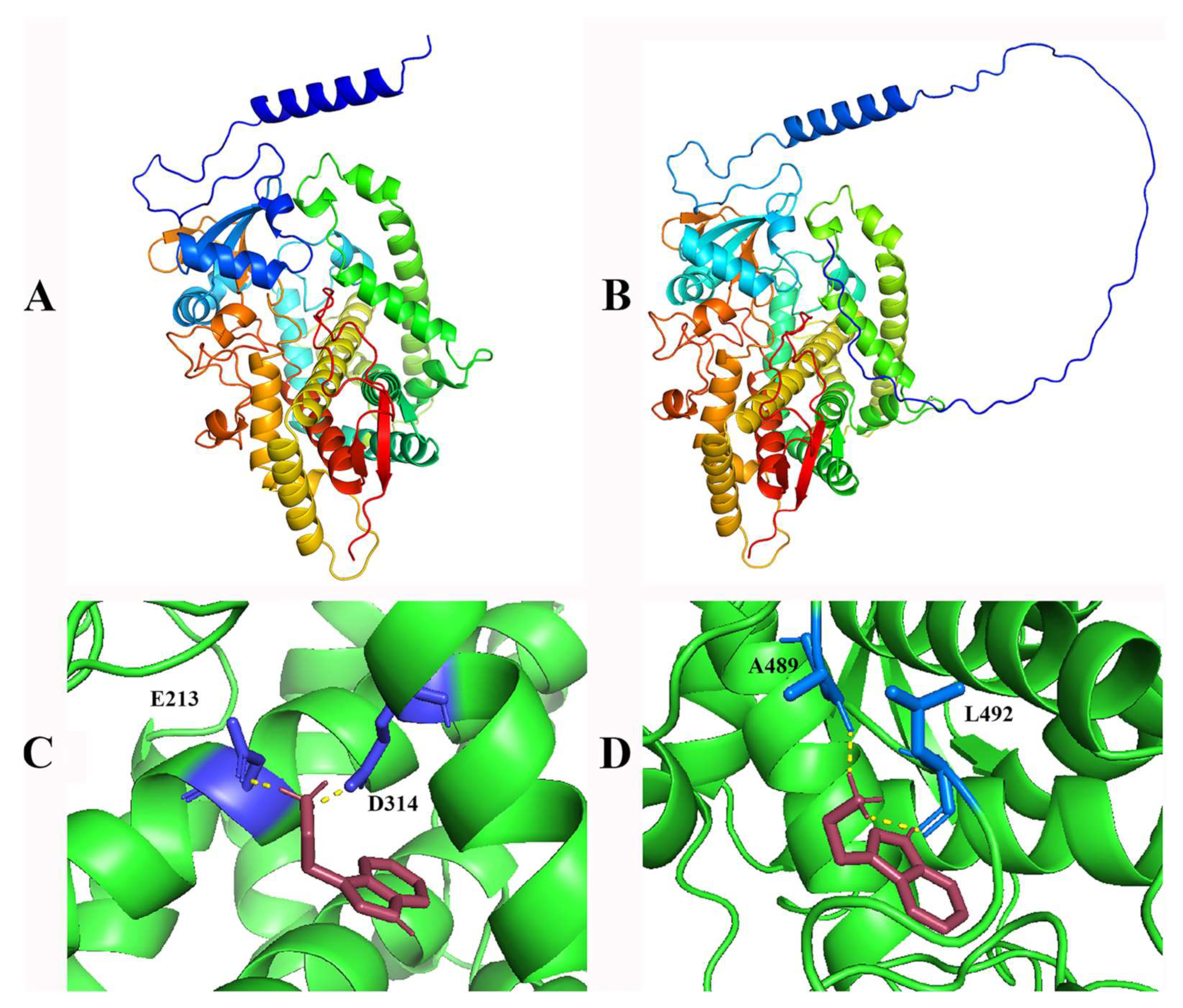
2.5. Structure Prediction and Molecular Docking
2.6. Determination of the Melatonin and Auxin Content in Both the WT Plant and sblob Mutant
3. Discussion
3.1. Abnormal Leaf Phenotype and Cell Development Are Responsible for sblob Mutant
3.2. Detection of Candidate Gene Accounting for sblob by MutMap
3.3. Single Nucleotide Mutations Result in Gene Functional Inactivation of sblob1
3.4. Sblob1 Regulates Sorghum Biomass by Influencing Melatonin Synthesis
4. Materials and Methods
4.1. Construction of Mapping Population
4.2. Field Experiment and Phenotype Analysis
4.3. Histological Analysis
4.4. DNA Extraction and Whole-Genome Sequence
4.5. SNP Calling and Annotation
4.6. Association Analysis Based on Euclidean Distance (ED)
4.7. Candidate Mutation Site Sequence
4.8. Bioinformatics Analysis and Function Prediction of Candidate Genes
4.9. 3D Structural Prediction and Molecular Docking
4.10. Detection of Tryptamine, Melatonin and Various Hormone Contents
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Silva, T.N.; Thomas, J.B.; Dahlberg, J.; Rhee, S.Y.; Mortimer, J.C. Progress and challenges in sorghum biotechnology, a multipurpose feedstock for the bioeconomy. J. Exp. Bot. 2021, 73, 646–664. [Google Scholar] [CrossRef] [PubMed]
- Ostmeyer, T.J.; Bahuguna, R.N.; Kirkham, M.B.; Bean, S.; Jagadish, S.V.K. Enhancing Sorghum Yield Through Efficient Use of Nitrogen—Challenges and Opportunities. Front. Plant Sci. 2022, 13, 845443. [Google Scholar] [CrossRef]
- Bollam, S.; Romana, K.K.; Rayaprolu, L.; Vemula, A.; Das, R.R.; Rathore, A.; Gandham, P.; Chander, G.; Deshpande, S.P.; Gupta, R. Nitrogen Use Efficiency in Sorghum: Exploring Native Variability for Traits Under Variable N-Regimes. Front. Plant Sci. 2021, 12, 643192. [Google Scholar] [CrossRef]
- Yang, K.-W.; Chapman, S.; Carpenter, N.; Hammer, G.; McLean, G.; Zheng, B.; Chen, Y.; Delp, E.; Masjedi, A.; Crawford, M.; et al. Integrating crop growth models with remote sensing for predicting biomass yield of sorghum. In Silico Plants 2021, 3, diab001. [Google Scholar] [CrossRef]
- Chiluwal, A.; Bheemanahalli, R.; Perumal, R.; Asebedo, A.R.; Bashir, E.; Lamsal, A.; Sebela, D.; Shetty, N.J.; Krishna Jagadish, S.V. Integrated aerial and destructive phenotyping differentiates chilling stress tolerance during early seedling growth in sorghum. Field Crops Res. 2018, 227, 1–10. [Google Scholar] [CrossRef]
- Baye, W.; Xie, Q.; Xie, P. Genetic Architecture of Grain Yield-Related Traits in Sorghum and Maize. Int. J. Mol. Sci. 2022, 23, 2405. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Luo, H.; Xu, J.; Cruickshank, A.; Zhao, X.; Teng, F.; Hathorn, A.; Wu, X.; Liu, Y.; Shatte, T.; et al. Extensive variation within the pan-genome of cultivated and wild sorghum. Nat. Plants 2021, 7, 766–773. [Google Scholar] [CrossRef]
- Zhi, X.; Tao, Y.; Jordan, D.; Borrell, A.; Hunt, C.; Cruickshank, A.; Potgieter, A.; Wu, A.; Hammer, G.; George-Jaeggli, B.; et al. Genetic control of leaf angle in sorghum and its effect on light interception. J. Exp. Bot. 2021, 73, 801–816. [Google Scholar] [CrossRef]
- Zhang, B.; Qi, F.; Hu, G.; Yang, Y.; Zhang, L.; Meng, J.; Han, Z.; Zhou, X.; Liu, H.; Ayaad, M.; et al. BSA-seq-based identification of a major additive plant height QTL with an effect equivalent to that of Semi-dwarf 1 in a large rice F2 population. Crop J. 2021, 9, 1428–1437. [Google Scholar] [CrossRef]
- Ochar, K.; Su, B.-H.; Zhou, M.-M.; Liu, Z.-X.; Gao, H.-W.; Lamlom, S.F.; Qiu, L.-J. Identification of the genetic locus associated with the crinkled leaf phenotype in a soybean (Glycine max L.) mutant by BSA-Seq technology. J. Integr. Agric. 2022, 21, 3524–3539. [Google Scholar] [CrossRef]
- Chen, J.; Zhu, M.; Liu, R.; Zhang, M.; Lv, Y.; Liu, Y.; Xiao, X.; Yuan, J.; Cai, H. BIOMASS YIELD 1 regulates sorghum biomass and grain yield via the shikimate pathway. J. Exp. Bot. 2020, 71, 5506–5520. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Liu, H.; Li, K.; Li, S.; Tao, Y. Genetic analysis and gene mapping of a narrow leaf mutant in rice (Oryza sativa L.). Chin. Sci. Bull. 2009, 54, 752–758. [Google Scholar] [CrossRef]
- Wang, T.; Crawford, M.M.; Tuinstra, M.R. A novel transfer learning framework for sorghum biomass prediction using UAV-based remote sensing data and genetic markers. Front. Plant Sci. 2023, 14, 1138479. [Google Scholar] [CrossRef] [PubMed]
- Chai, L.; Feng, B.; Liu, X.; Jiang, L.; Yuan, S.; Zhang, Z.; Li, H.; Zhang, J.; Fernando, D.; Xu, C.; et al. Fine Mapping of a Locus Underlying the Ectopic Blade-Like Outgrowths on Leaf and Screening Its Candidate Genes in Rapeseed (Brassica napus L.). Front. Plant Sci. 2021, 11, 616844. [Google Scholar] [CrossRef]
- Xia, W.; Xiao, Z.a.; Cao, P.; Zhang, Y.; Du, K.; Wang, N. Construction of a high-density genetic map and its application for leaf shape QTL mapping in poplar. Planta 2018, 248, 1173–1185. [Google Scholar] [CrossRef] [PubMed]
- Prigge, M.J.; Wagner, D.R. The Arabidopsis SERRATE Gene Encodes a Zinc-Finger Protein Required for Normal Shoot Development. Plant Cell 2001, 13, 1263–1280. [Google Scholar] [CrossRef] [PubMed]
- Saddic, L.A.; Huvermann, B.r.; Bezhani, S.; Su, Y.; Winter, C.M.; Kwon, C.S.; Collum, R.P.; Wagner, D. The LEAFY target LMI1 is a meristem identity regulator and acts together with LEAFY to regulate expression of CAULIFLOWER. Development 2006, 133, 1673–1682. [Google Scholar] [CrossRef]
- Lin, L.; Zhao, Y.; Liu, F.; Chen, Q.; Qi, J. Narrow leaf 1 (NAL1) regulates leaf shape by affecting cell expansion in rice (Oryza sativa L.). Biochem. Biophys. Res. Commun. 2019, 516, 957–962. [Google Scholar] [CrossRef]
- Abe, A.; Kosugi, S.; Yoshida, K.; Natsume, S.; Takagi, H.; Kanzaki, H.; Matsumura, H.; Yoshida, K.; Mitsuoka, C.; Tamiru, M.; et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 2012, 30, 174–178. [Google Scholar] [CrossRef]
- Fekih, R.; Takagi, H.; Tamiru, M.; Abe, A.; Natsume, S.; Yaegashi, H.; Sharma, S.; Sharma, S.; Kanzaki, H.; Matsumura, H.; et al. MutMap+: Genetic mapping and mutant identification without crossing in rice. PLoS ONE 2013, 8, e68529. [Google Scholar] [CrossRef]
- Takagi, H.; Tamiru, M.; Abe, A.; Yoshida, K.; Uemura, A.; Yaegashi, H.; Obara, T.; Oikawa, K.; Utsushi, H.; Kanzaki, E.; et al. MutMap accelerates breeding of a salt-tolerant rice cultivar. Nat. Biotechnol. 2015, 33, 445–449. [Google Scholar] [CrossRef]
- Zhang, L.; You, R.; Chen, H.; Zhu, J.; Lin, L.; Wei, C. A New SNP in AGPL2, Associated with Floury Endosperm in Rice, Is Identified Using a Modified MutMap Method. Agronomy 2023, 13, 1381. [Google Scholar] [CrossRef]
- Lyu, J.; Wang, D.; Duan, P.; Liu, Y.; Huang, K.; Zeng, D.; Zhang, L.; Dong, G.; Li, Y.; Xu, R.; et al. Control of Grain Size and Weight by the GSK2-LARGE1/OML4 Pathway in Rice. Plant Cell 2020, 32, 1905–1918. [Google Scholar] [CrossRef] [PubMed]
- Zegeye, W.A.; Zhang, Y.; Cao, L.; Cheng, S. Whole Genome Resequencing from Bulked Populations as a Rapid QTL and Gene Identification Method in Rice. Int. J. Mol. Sci. 2018, 19, 4000. [Google Scholar] [CrossRef]
- Liu, D.; Zhao, H.; Xiao, Y.; Zhang, G.; Cao, S.; Yin, W.; Qian, Y.; Yin, Y.; Zhang, J.; Chen, S.; et al. A cryptic inhibitor of cytokinin phosphorelay controls rice grain size. Mol. Plant 2022, 15, 293–307. [Google Scholar] [CrossRef]
- Zhu, X.; Pan, Y.; Liu, Z.; Liu, Y.; Zhong, D.; Duan, Z.; Tian, Z.; Zhu, B.; Zhou, G. Mutation of YL Results in a Yellow Leaf with Chloroplast RNA Editing Defect in Soybean. Int. J. Mol. Sci. 2020, 21, 4275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Han, M.; Liu, Y.; Lin, X.; Liu, X.; Zhu, H.; He, Y.; Zhang, Q.; Liu, J. Whole-genome resequencing from bulked-segregant analysis reveals gene set based association analyses for the Vibrio anguillarum resistance of turbot (Scophthalmus maximus). Fish Shellfish Immunol. 2019, 88, 76–83. [Google Scholar] [CrossRef] [PubMed]
- Shen, S.; Xu, S.; Wang, M.; Ma, T.; Chen, N.; Wang, J.; Zheng, H.; Yang, L.; Zou, D.; Xin, W.; et al. BSA-Seq for the Identification of Major Genes for EPN in Rice. Int. J. Mol. Sci. 2023, 24, 14838. [Google Scholar] [CrossRef]
- Schneeberger, K. Using next-generation sequencing to isolate mutant genes from forward genetic screens. Nat. Rev. Genet. 2014, 15, 662–676. [Google Scholar] [CrossRef]
- Bhowal, B.; Bhattacharjee, A.; Goswami, K.; Sanan-Mishra, N.; Singla-Pareek, S.L.; Kaur, C.; Sopory, S. Serotonin and Melatonin Biosynthesis in Plants: Genome-Wide Identification of the Genes and Their Expression Reveal a Conserved Role in Stress and Development. Int. J. Mol. Sci. 2021, 22, 11034. [Google Scholar] [CrossRef]
- Shekhar, S.; Prasad, A.S.; Banjare, K.; Kaushik, A.; Mannade, A.K.; Dubey, M.; Patil, A.; Premi, V.; Vishwakarma, A.K.; Sao, A.; et al. LMT(db): A comprehensive transcriptome database for climate-resilient, nutritionally rich little millet (Panicum sumatrense). Front. Plant Sci. 2023, 14, 1106104. [Google Scholar] [CrossRef]
- Song, Z.; Zhang, L.; Wang, Y.; Li, H.; Li, S.; Zhao, H.; Zhang, H. Constitutive Expression of miR408 Improves Biomass and Seed Yield in Arabidopsis. Front. Plant Sci. 2017, 8, 2114. [Google Scholar] [CrossRef] [PubMed]
- Borrell, A.K.; Mullet, J.E.; George-Jaeggli, B.; van Oosterom, E.J.; Hammer, G.L.; Klein, P.E.; Jordan, D.R. Drought adaptation of stay-green sorghum is associated with canopy development, leaf anatomy, root growth, and water uptake. J. Exp. Bot. 2014, 65, 6251–6263. [Google Scholar] [CrossRef] [PubMed]
- Borrell, A.K.; van Oosterom, E.J.; Mullet, J.E.; George-Jaeggli, B.; Jordan, D.R.; Klein, P.E.; Hammer, G.L. Stay-green alleles individually enhance grain yield in sorghum under drought by modifying canopy development and water uptake patterns. New Phytol. 2014, 203, 817–830. [Google Scholar] [CrossRef] [PubMed]
- Mantilla Perez, M.B.; Zhao, J.; Yin, Y.; Hu, J.; Salas Fernandez, M.G. Association mapping of brassinosteroid candidate genes and plant architecture in a diverse panel of Sorghum bicolor. Theor. Appl. Genet. 2014, 127, 2645–2662. [Google Scholar] [CrossRef]
- Kersey, P.J. Plant genome sequences: Past, present, future. Curr. Opin. Plant Biol. 2019, 48, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Shang, L.; Zhu, Q.-H.; Fan, L.; Guo, L. Twenty years of plant genome sequencing: Achievements and challenges. Trends Plant Sci. 2022, 27, 391–401. [Google Scholar] [CrossRef]
- Huo, H.; Henry, I.M.; Coppoolse, E.R.; Verhoef-Post, M.; Schut, J.W.; de Rooij, H.; Vogelaar, A.; Joosen, R.V.; Woudenberg, L.; Comai, L.; et al. Rapid identification of lettuce seed germination mutants by bulked segregant analysis and whole genome sequencing. Plant J. 2016, 88, 345–360. [Google Scholar] [CrossRef]
- Wang, H.; Tang, S.; Zhi, H.; Xing, L.; Zhang, H.; Tang, C.; Wang, E.; Zhao, M.; Jia, G.; Feng, B.; et al. The boron transporter SiBOR1 functions in cell wall integrity, cellular homeostasis, and panicle development in foxtail millet. Crop J. 2022, 10, 342–353. [Google Scholar] [CrossRef]
- Cao, Z.Z.; Lin, X.Y.; Yang, Y.J.; Guan, M.Y.; Xu, P.; Chen, M.X. Gene identification and transcriptome analysis of low cadmium accumulation rice mutant (lcd1) in response to cadmium stress using MutMap and RNA-seq. BMC Plant Biol. 2019, 19, 250. [Google Scholar] [CrossRef]
- Jiao, Y.; Burow, G.; Gladman, N.; Acosta-Martinez, V.; Chen, J.; Burke, J.; Ware, D.; Xin, Z. Efficient Identification of Causal Mutations through Sequencing of Bulked F2 from Two Allelic Bloomless Mutants of Sorghum bicolor. Front. Plant Sci. 2018, 8, 2267. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Abdelghany, A.M.; Azam, M.; Qi, J.; Li, J.; Feng, Y.; Liu, Y.; Feng, H.; Ma, C.; Gebregziabher, B.S.; et al. Mining candidate genes underlying seed oil content using BSA-seq in soybean. Ind. Crops Prod. 2023, 194, 116308. [Google Scholar] [CrossRef]
- Sun, C.; Liu, L.; Wang, L.; Li, B.; Jin, C.; Lin, X. Melatonin: A master regulator of plant development and stress responses. J. Integr. Plant Biol. 2021, 63, 126–145. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhang, Y. Melatonin in plants: What we know and what we don’t. Food Qual. Saf. 2021, 5, fyab009. [Google Scholar] [CrossRef]
- Byeon, Y.; Back, K. An increase in melatonin in transgenic rice causes pleiotropic phenotypes, including enhanced seedling growth, delayed flowering, and low grain yield. J. Pineal Res. 2014, 56, 408–414. [Google Scholar] [CrossRef] [PubMed]
- Heydarnajad Giglou, R.; Torabi Giglou, M.; Esmaeilpour, B.; Padash, A.; Ghahremanzadeh, S.; Sobhanizade, A.; Hatami, M. Exogenous melatonin differentially affects biomass, total carbohydrates, and essential oil production in peppermint upon simultaneous exposure to chitosan-coated Fe3O4 NPs. S. Afr. J. Bot. 2023, 163, 135–144. [Google Scholar] [CrossRef]
- Agathokleous, E.; Zhou, B.; Xu, J.; Ioannou, A.; Feng, Z.; Saitanis, C.J.; Frei, M.; Calabrese, E.J.; Fotopoulos, V. Exogenous application of melatonin to plants, algae, and harvested products to sustain agricultural productivity and enhance nutritional and nutraceutical value: A meta-analysis. Environ. Res. 2021, 200, 111746. [Google Scholar] [CrossRef]
- Huang, Y.-H.; Liu, S.-J.; Yuan, S.; Guan, C.; Tian, D.-Y.; Cui, X.; Zhang, Y.-W.; Yang, F.-Y. Overexpression of ovine AANAT and HIOMT genes in switchgrass leads to improved growth performance and salt-tolerance. Sci. Rep. 2017, 7, 12212. [Google Scholar] [CrossRef]
- Ke, S.; Liu, S.; Luan, X.; Xie, X.M.; Hsieh, T.F.; Zhang, X.Q. Mutation in a putative glycosyltransferase-like gene causes programmed cell death and early leaf senescence in rice. Rice 2019, 12, 7. [Google Scholar] [CrossRef]
- Lee, K.; Choi, G.H.; Back, K. Inhibition of Rice Serotonin N-Acetyltransferases by MG149 Decreased Melatonin Synthesis in Rice Seedlings. Biomolecules 2021, 11, 658. [Google Scholar] [CrossRef]
- Mannino, G.; Pernici, C.; Serio, G.; Gentile, C.; Bertea, C.M. Melatonin and Phytomelatonin: Chemistry, Biosynthesis, Metabolism, Distribution and Bioactivity in Plants and Animals—An Overview. Int. J. Mol. Sci. 2021, 22, 9996. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Nigam, D.; Barry, K.; Daum, C.; Yoshinaga, Y.; Lipzen, A.; Khan, A.; Parasa, S.-P.; Wei, S.; Lu, Z.; et al. A large sequenced mutant library–valuable reverse genetic resource that covers 98% of sorghum genes. Plant J. 2023. Online ahead of print. [Google Scholar] [CrossRef]
- Jiao, Y.; Burke, J.; Chopra, R.; Burow, G.; Chen, J.; Wang, B.; Hayes, C.; Emendack, Y.; Ware, D.; Xin, Z. A Sorghum Mutant Resource as an Efficient Platform for Gene Discovery in Grasses. Plant Cell 2016, 28, 1551–1562. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, J.; Chen, L.; Meng, X.; Zhen, X.; Liang, Y.; Han, Y.; Li, H.; Zhang, B. Identification and function analysis of yellow-leaf mutant (YX-yl) of broomcorn millet. BMC Plant Biol. 2022, 22, 463. [Google Scholar] [CrossRef] [PubMed]
- Chai, L.; Li, H.; Zhang, J.; Wu, L.; Zheng, B.; Cui, C.; Jiang, J.; Zuo, S.; Jiang, L. Rapid Identification of a Genomic Region Conferring Dwarfism in Rapeseed (Brassica napus L.) YA2016-12. Agronomy 2019, 9, 129. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Hill, J.T.; Demarest, B.L.; Bisgrove, B.W.; Gorsi, B.; Su, Y.C.; Yost, H.J. MMAPPR: Mutation mapping analysis pipeline for pooled RNA-seq. Genome Res. 2013, 23, 687–697. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Xu, X.; Murphy, L.A. Fast and sensitive LC-MS/MS method for quantification of cannabinoids and their metabolites in plasma of cattle fed hemp. J. Sep. Sci. 2023, 2300630. [Google Scholar] [CrossRef]
- Li, Y.; Zhou, C.; Yan, X.; Zhang, J.; Xu, J. Simultaneous analysis of ten phytohormones in Sargassum horneri by high-performance liquid chromatography with electrospray ionization tandem mass spectrometry. J. Sep. Sci. 2016, 39, 1804–1813. [Google Scholar] [CrossRef]


| WT | sblob | Total | χ2 (df = 1) | |
|---|---|---|---|---|
| actual | 118 | 294 | 412 | |
| expected | 103 | 309 | 412 | 2.63 |
| Clean Reads | Clean Bases (bp) | Clean Q30 (%) | Mapped Ratio (%) | Proper Ratio (%) | Average Depths | |
|---|---|---|---|---|---|---|
| WT | 190,778,008 | 28,728,254,222 | 94.49 | 99.12 | 95.68 | 44.16 |
| sblob | 177,743,206 | 26,767,472,954 | 94.75 | 98.49 | 95.13 | 40.98 |
| Region | Start Position (Mb) | End Position (Mb) | Region Size | Transcripts | Gene Number |
|---|---|---|---|---|---|
| chr10 | 57.91461 | 58.149969 | 0.235359 | 30 | 24 |
| chr1 | 1.154111 | 1.34952 | 0.195409 | 46 | 39 |
| chr1 | 1.409762 | 1.532592 | 0.12283 | 49 | 30 |
| chr1 | 1.623681 | 1.691337 | 0.067656 | 15 | 14 |
| chr1 | 1.812801 | 1.938655 | 0.125854 | 38 | 31 |
| chr1 | 4.068754 | 4.498218 | 0.429464 | 63 | 52 |
| chr1 | 58.306682 | 59.217158 | 0.910476 | 87 | 65 |
| chr1 | 65.020891 | 65.164125 | 0.143234 | 16 | 15 |
| chr1 | 6.504078 | 6.639419 | 0.135341 | 24 | 16 |
| chr1 | 67.295568 | 67.722791 | 0.427223 | 57 | 49 |
| chr1 | 72.061557 | 72.655387 | 0.59383 | 85 | 67 |
| chr1 | 8.567085 | 8.898033 | 0.330948 | 63 | 29 |
| chr2 | 4.668026 | 5.149803 | 0.481777 | 76 | 59 |
| chr2 | 5.400013 | 5.487665 | 0.087652 | 7 | 5 |
| chr2 | 5.664728 | 5.793508 | 0.12878 | 21 | 11 |
| chr2 | 71.663212 | 72.648182 | 0.98497 | 190 | 116 |
| chr3 | 0.703708 | 1.323768 | 0.62006 | 104 | 85 |
| chr3 | 70.882922 | 72.678683 | 1.795761 | 285 | 199 |
| chr4 | 60.551245 | 60.866205 | 0.31496 | 45 | 29 |
| chr6 | 23.890052 | 24.617971 | 0.727919 | 0 | 0 |
| chr6 | 59.233376 | 59.338064 | 0.104688 | 22 | 20 |
| chr8 | 58.306666 | 58.339277 | 0.032611 | 5 | 5 |
| total | 8.996802 | 1328 | 960 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, M.; Cai, Q.; Liang, Y.; Zhao, Y.; Hao, Y.; Qin, Y.; Qiao, X.; Han, Y.; Li, H. Mapping and Screening of Candidate Gene Regulating the Biomass Yield of Sorghum (Sorghum bicolor L.). Int. J. Mol. Sci. 2024, 25, 796. https://doi.org/10.3390/ijms25020796
Li M, Cai Q, Liang Y, Zhao Y, Hao Y, Qin Y, Qiao X, Han Y, Li H. Mapping and Screening of Candidate Gene Regulating the Biomass Yield of Sorghum (Sorghum bicolor L.). International Journal of Molecular Sciences. 2024; 25(2):796. https://doi.org/10.3390/ijms25020796
Chicago/Turabian StyleLi, Mao, Qizhe Cai, Yinpei Liang, Yaofei Zhao, Yaoshan Hao, Yingying Qin, Xinrui Qiao, Yuanhuai Han, and Hongying Li. 2024. "Mapping and Screening of Candidate Gene Regulating the Biomass Yield of Sorghum (Sorghum bicolor L.)" International Journal of Molecular Sciences 25, no. 2: 796. https://doi.org/10.3390/ijms25020796





