Circulating microRNA miR-425-5p Associated with Brain White Matter Lesions and Inflammatory Processes
Abstract
1. Introduction
2. Results
2.1. Clinical Impact of WML
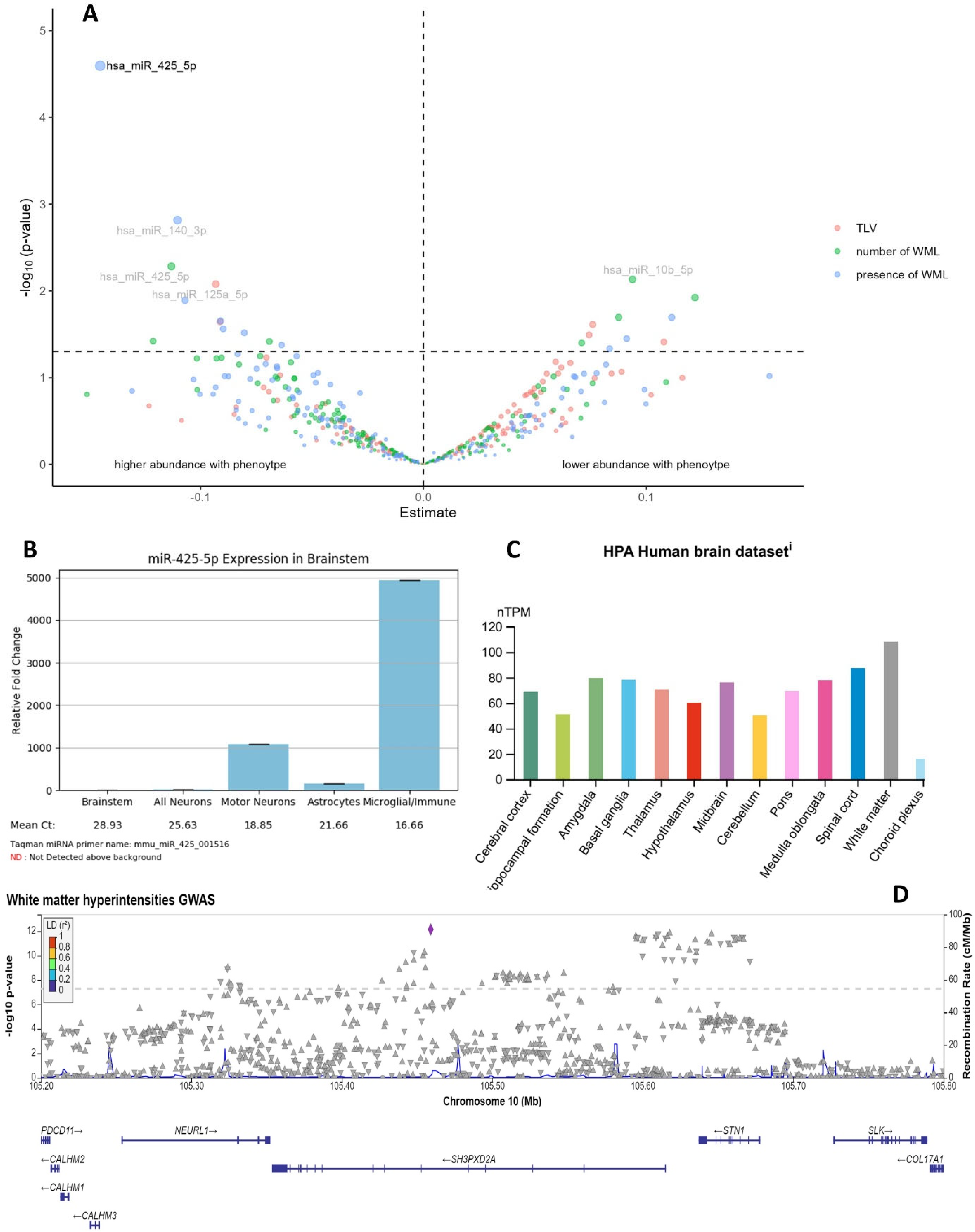
2.2. hsa-miR-425-5p Is Associated with WMLs
2.2.1. Association of Target miRNAs with Structural AD-Related MRI Phenotypes
2.2.2. The Role of hsa-miR-425-5p in Inflammation
2.2.3. Target Genes of hsa-miR-425-5p in WMLs GWAS
2.3. Influence of Moderating Factors
- Moderation by sex: In sex interaction models, 19 and 18 nominally significant miRNAs were identified for total WML volume and number of WMLs, respectively (Table S5, Figure S14). Only for the number of WMLs, two miRNAs, hsa-miR-126-3p and hsa-miR-374a-5p, survived multiple tests (Table 2). In both cases, lower values of miRNA abundance (reflected in higher ΔCt values) were associated with a higher WML burden in females but with a reduced burden in males (Figure S16). Both miRNAs are expressed in the brain (Figure S15A,B) and the EGFL7 (EGF-like domain multiple 7) gene harboring MIR126 on chromosome 9 has been associated with AD as well as white matter growth (Table S4).
- Moderation by APOE ε4: In interaction analyses with the APOE ε4 carrier status, 13 and 19 nominally significant miRNAs were identified for total WML volume and number of WMLs, respectively (Table S6, Figure S14). None of them survived multiple testing corrections. The lowest p-value was observed for hsa-miR-140-5p on the number of WMLs (p = 0.0011). Carriers of the APOE ε4 allele had a beneficial outcome for WML burden in the case of high levels of hsa-miR-140-5p, whereas in the case of lower levels, the WML burden increased (Figure S17). Hsa-miR-140-5p is only slightly expressed in the brain (Figure S18) and its harboring gene WWP2 (WW domain containing E3 ubiquitin protein ligase 2) has been associated with addictive behavior and cognitive traits (Table S4).
- Moderation by smoking status: Interaction with smoking status revealed 13 and 15 nominal significant miRNAs for total WML volume and number of WMLs, respectively (Table S7, Figure S14). Three miRNAs reached BH-corrected significance in the latter model (hsa-miR-885-5p, hsa-miR-199a-5p, hsa-miR-194-5p; Table 3, Figure S19) with hsa-miR-199a-5p reaching significance in both WML models. Interestingly, only hsa-miR-885-5p shows a substantial expression in brain tissues (Figure S20A–C) and the strongest link towards neurodegenerative endpoints concerning its harboring gene ATP2B2 (ATPase plasma membrane Ca2+ transporting 2) (Table S4).
2.4. Significant Plasma-Circulating miRNAs Are Enriched in Neurodegeneration
3. Discussion
| miRNA | Previous Results Regarding AD and/or Cognition from Pubmed |
|---|---|
| hsa-miR-425-5p | Regulation of AD pathogenic genes [23] Interacting with BACE1 [35] Upregulated in AD [24,25] Association with memory and learning disorders [36] Promotes formation of Aβ plaques [37] |
| hsa-miR-126-3p | Overexpression could reduce Aβ plaque area and neuroinflammation in the hippocampus [38] Associated with inflammation in the pathogenesis of AD [39] Involved in neurogenesis [40] Upregulated in plasma of AD patients [41] Altered regulation in brain of AD male rats [42] Involved in neuronal accumulation of AD [43] Decreased in plasma of AD subjects [44] Part of a nine-miRNA signature as potential biomarker for AD [45] Dysregulated in plasma of AMD rats [46] Associated with stroke recovery [47] Negative correlation with cognitive function [48] Dysregulated in AD NMV [49] Cardiovascular events (including stroke) [50] Regulation of BDNF synthesis [51] |
| hsa-miR-374a-5p | Part of plasma signature of obstructive sleep apnea in AD [52] Cardiovascular events (including stroke) [50] Overexpression reduces cell apoptosis [53] |
| hsa-miR-885-5p | Regulating neuronal cell injury [54] Serum biomarker for AD [55] Upregulated in AD [56] Associated with higher metabolic risk profile in older subjects [57] |
| hsa-miR-199a-5p | Involved in AD development [58] Related to cognitive impairment [59] Link between AD and diabetes [60] Protects cognitive function in ischemic stroke [61] |
| hsa-miR-194-5p | Association with WML and cognitive impairment [16] Associated with higher metabolic risk profile in older subjects [57] Downregulated in blood of AD patients [62] Inhibit apoptosis of hippocampal neurons [63] |
| hsa-miR-140-5p | Risk factor for memory impairment induced by Aβ [64] Associated with neurodegenerative diseases in general [65] Associated with vascular cognitive impairment [66] Associated with cognitive performance in healthy older adults [67] Associated with AD risk gene ADAM10 [68] Neuroprotective effects [69] |
4. Materials and Methods
4.1. SHIP Sample
4.2. Verbal Memory Scores
4.3. Brain Imaging Data
4.4. Plasma-Circulating miRNAs
4.5. Immunological Markers
4.6. Additional Variables
4.7. Statistical Analyses
- Clinical impact of WMLs: The following generalized linear models (GLM) were calculated with WMLs as predictors and verbal memory scores as outcomes.
+ hypertension + ICV
- 2.
- Impact of miRNAs on WMLs: GLMs were calculated with miRNA levels as predictors and WMLs (structural MRI markers) as outcome.
ICV + batch
- 3.
- Impact of target miRNAs and inflammatory markers: In GLMs, the association between circulating inflammatory markers (CRP, fibrinogen) and significant miRNAs was investigated and corrected for age, sex, miRNA batch, smoking, BMI, education, HCT, and PLT. CRP was log-transformed prior to analyses.
- 4.
- Moderation effect of sex, APOE ε4, smoking: Similar models as in 2. were performed, additionally including an interaction term between miRNA levels and sex, APOE ε4, or ever smoking.
4.8. Post Hoc In Silico Analysis
- Using GTExPortal (https://www.gtexportal.org/home/, accessed on 6 June 2023), miRNA TissueAtlas [79], Human Protein Atlas [80], and CNS microRNA Profiles database for mice [81], we investigated the expression of miRNAs and genes in different brain tissues of human and mouse samples.
- We used the over-representation analysis implemented in the miRNA Enrichment Analysis and Annotation Tool (miEAA 2.0) [85] to search for significant associations between sets of target miRNAs and disease outcomes, incorporating data from large miRNA, tissue, and pathway databases.
- Comparison with GWAS results: results for SNPs within target genes of significant miRNAs were looked up in publicly available GWAS summary statistics on white matter hyperintensity burden (dbGaP: phs002227.v1.p1 [11]).
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Castro-Aldrete, L.; Moser, M.V.; Putignano, G.; Ferretti, M.T.; Schumacher Dimech, A.; Santuccione Chadha, A. Sex and gender considerations in Alzheimer’s disease: The Women’s Brain Project contribution. Front. Aging Neurosci. 2023, 15, 1105620. [Google Scholar] [CrossRef] [PubMed]
- Bidzan, L. Cardiovascular factors in dementia. Psychiatr. Pol. 2022, 56, 991–1001. [Google Scholar] [CrossRef] [PubMed]
- Esparza, T.J.; Gangolli, M.; Cairns, N.J.; Brody, D.L. Soluble amyloid-beta buffering by plaques in Alzheimer disease dementia versus high-pathology controls. PLoS ONE 2018, 13, e0200251. [Google Scholar] [CrossRef] [PubMed]
- Hase, Y.; Horsburgh, K.; Ihara, M.; Kalaria, R.N. White matter degeneration in vascular and other ageing-related dementias. J. Neurochem. 2018, 144, 617–633. [Google Scholar] [CrossRef] [PubMed]
- Inoue, Y.; Shue, F.; Bu, G.; Kanekiyo, T. Pathophysiology and probable etiology of cerebral small vessel disease in vascular dementia and Alzheimer’s disease. Mol. Neurodegener. 2023, 18, 46. [Google Scholar] [CrossRef] [PubMed]
- Debette, S.; Markus, H.S. The clinical importance of white matter hyperintensities on brain magnetic resonance imaging: Systematic review and meta-analysis. BMJ 2010, 341, c3666. [Google Scholar] [CrossRef]
- Habes, M.; Erus, G.; Toledo, J.B.; Zhang, T.; Bryan, N.; Launer, L.J.; Rosseel, Y.; Janowitz, D.; Dosho, J.; Van der Auwera, S.; et al. White matter hyperintensities and imaging patterns of brain ageing in the general population. Brain 2016, 139, 1164–1179. [Google Scholar] [CrossRef] [PubMed]
- Alqarni, A.; Jiang, J.; Crawford, J.D.; Koch, F.; Brodaty, H.; Sachdev, P.; Wen, W. Sex differences in risk factors for white matter hyperintensities in non-demented older individuals. Neurobiol. Aging 2021, 98, 197–204. [Google Scholar] [CrossRef]
- Botz, J.; Lohner, V.; Schirmer, M.D. Spatial patterns of white matter hyperintensities: A systematic review. Front. Aging Neurosci. 2023, 15, 1165324. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Chen, Q.; Chen, J.; Yang, N.; Zheng, K. Risk factors of cerebral small vessel disease: A systematic review and meta-analysis. Medicine 2021, 100, e28229. [Google Scholar] [CrossRef]
- Sargurupremraj, M.; Suzuki, H.; Jian, X.; Sarnowski, C.; Evans, T.E.; Bis, J.C.; Eiriksdottir, G.; Sakave, S.; Terzikhan, N.; Habes, M.; et al. Cerebral small vessel disease genomics and its implications across the lifespan. Nat. Commun. 2020, 11, 6285. [Google Scholar] [CrossRef]
- Li, Y.; Zheng, J.; Li, T.; Zhang, J. White Matter and Alzheimer’s Disease: A Bidirectional Mendelian Randomization Study. Neurol. Ther. 2022, 11, 881–892. [Google Scholar] [CrossRef] [PubMed]
- Samadian, M.; Gholipour, M.; Hajiesmaeili, M.; Taheri, M.; Ghafouri-Fard, S. The Eminent Role of microRNAs in the Pathogenesis of Alzheimer’s Disease. Front. Aging Neurosci. 2021, 13, 641080. [Google Scholar] [CrossRef]
- Singh, R.; Hussain, J.; Kaur, A.; Jamdare, B.G.; Pathak, D.; Garg, K.; Kaur, R.; Shankar, S.; Sunkaria, A. The Hidden Players: Shedding Light on the Significance of Post-Translational Modifications and miRNAs in Alzheimer’s Disease Development. Ageing Res. Rev. 2023, 90, 102002. [Google Scholar] [CrossRef] [PubMed]
- Bhatnagar, D.; Ladhe, S.; Kumar, D. Discerning the Prospects of miRNAs as a Multi-Target Therapeutic and Diagnostic for Alzheimer’s Disease. Mol. Neurobiol. 2023, 60, 5954–5974. [Google Scholar] [CrossRef]
- Dong, X.; Sun, H.; Mao, J.; Zhang, S.; Meng, C. Differential expression of circular RNA in patients with white matter hyperintensity and cognitive impairment. J. Zhejiang Univ. Med. Sci. 2021, 46, 1080–1089. [Google Scholar] [CrossRef]
- Huang, W.-Q.; Lin, Q.; Chen, S.; Sun, L.; Chen, Q.; Yi, K.; Li, Z.; Ma, Q.; Tzeng, C. Integrated analysis of microRNA and mRNA expression profiling identifies BAIAP3 as a novel target of dysregulated hsa-miR-1972 in age-related white matter lesions. Aging 2021, 13, 4674–4695. [Google Scholar] [CrossRef]
- Badimon, A.; Torrente, D.; Norris, E.H. Vascular Dysfunction in Alzheimer’s Disease: Alterations in the Plasma Contact and Fibrinolytic Systems. Int. J. Mol. Sci. 2023, 24, 7046. [Google Scholar] [CrossRef]
- Wen, T.; Zhang, Z. Cellular mechanisms of fibrin (ogen): Insight from neurodegenerative diseases. Front. Neurosci. 2023, 17, 1197094. [Google Scholar] [CrossRef]
- Tian, J.; Liu, Y.; Wang, Z.; Zhang, S.; Yang, Y.; Zhu, Y.; Yang, C. LncRNA Snhg8 attenuates microglial inflammation response and blood-brain barrier damage in ischemic stroke through regulating miR-425-5p mediated SIRT1/NF-κB signaling. J. Biochem. Mol. Toxicol. 2021, 35, e22724. [Google Scholar] [CrossRef]
- Meng, F.; Yang, Y.; Jin, G. Research Progress on MRI for White Matter Hyperintensity of Presumed Vascular Origin and Cognitive Impairment. Front. Neurol. 2022, 13, 865920. [Google Scholar] [CrossRef]
- Cascella, M.; Al Khalili, Y. Short-Term Memory Impairment; StatPearls Publishing: Treasure Island, FL, USA, 2023. [Google Scholar]
- Zhang, Q.; Yang, P.; Pang, X.; Guo, W.; Sun, Y.; Wie, Y.; Pang, C. Preliminary exploration of the co-regulation of Alzheimer’s disease pathogenic genes by microRNAs and transcription factors. Front. Aging Neurosci. 2022, 14, 1069606. [Google Scholar] [CrossRef] [PubMed]
- Satoh, J.-I.; Kino, Y.; Niida, S. MicroRNA-Seq Data Analysis Pipeline to Identify Blood Biomarkers for Alzheimer’s Disease from Public Data. Biomark. Insights 2015, 10, 21–31. [Google Scholar] [CrossRef]
- Yuan, J.; Wu, Y.; Li, L.; Liu, C. MicroRNA-425-5p promotes tau phosphorylation and cell apoptosis in Alzheimer’s disease by targeting heat shock protein B8. J. Neural. Transm. 2020, 127, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Knol, M.J.; Wang, R.; Mishra, A.; Liu, D.; Luciano, M.; Teumer, A.; Armstrong, N.; Bis, J.C.; Jhun, M.A.; et al. Epigenetic and integrative cross-omics analyses of cerebral white matter hyperintensities on MRI. Brain 2023, 146, 492–506. [Google Scholar] [CrossRef]
- Laumet, G.; Petitprez, V.; Sillaire, A.; Ayral, A.-M.; Hansmannel, F.; Chapuis, J.; Hannequin, D.; Pasquier, F.; Scarpini, E.; Galimberti, D.; et al. A study of the association between the ADAM12 and SH3PXD2A (SH3MD1) genes and Alzheimer’s disease. Neurosci. Lett. 2010, 468, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Hsia, H.-E.; Tüshaus, J.; Brummer, T.; Zheng, Y.; Scilabra, S.D.; Lichtenthaler, S.F. Functions of ‘A disintegrin and metalloproteases (ADAMs)’ in the mammalian nervous system. Cell Mol. Life Sci. 2019, 76, 3055–3081. [Google Scholar] [CrossRef]
- Sun, T.; Tan, L.; Liu, M.; Zheng, L.; Zhao, K.; Cai, Z.; Sun, S.; Li, Z.; Liu, R. Tilianin improves cognition in a vascular dementia rodent model by targeting miR-193b-3p/CaM- and miR-152-3p/CaMKIIα-mediated inflammatory and apoptotic pathways. Front. Immunol. 2023, 14, 1118808. [Google Scholar] [CrossRef]
- Liu, Q.-S.; Jiang, H.-L.; Wang, Y.; Wang, L.-L.; Zhang, J.-X.; He, C.-H.; Shao, S.; Zhang, T.-T.; Xing, J.-G.; Liu, R. Total flavonoid extract from Dracoephalum moldavica L. attenuates β-amyloid-induced toxicity through anti-amyloidogenesic and neurotrophic pathways. Life Sci. 2018, 193, 214–225. [Google Scholar] [CrossRef]
- Jiang, H.; Ashraf, G.M.; Liu, M.; Zhao, K.; Wang, Y.; Wang, L.; Xing, J.; Alghamdi, B.S.; Li, Z.; Liu, R. Tilianin Ameliorates Cognitive Dysfunction and Neuronal Damage in Rats with Vascular Dementia via p-CaMKII/ERK/CREB and ox-CaMKII-Dependent MAPK/NF-κB Pathways. Oxid. Med. Cell Longev. 2021, 2021, 6673967. [Google Scholar] [CrossRef]
- Bourquard, T.; Lee, K.; Al-Ramahi, I.; Pham, M.; Shapiro, D.; Lagiselly, Y.; Soleimani, S.; Mota, S.; Wilhelm, K.; Samieinasab, M.; et al. Functional variants identify sex-specific genes and pathways in Alzheimer’s Disease. Nat. Commun. 2023, 14, 2765. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Cao, J.; Hou, J.; Li, Y.; Huang, M.; Zhu, L.; Zhang, L.; Lee, Y.; Duarte, M.L.; Zhou, X.; et al. Sex specific molecular networks and key drivers of Alzheimer’s disease. Mol. Neurodegener. 2023, 18, 39. [Google Scholar] [CrossRef]
- Hu, W.T.; Nayyar, A.; Kaluzova, M. Charting the Next Road Map for CSF Biomarkers in Alzheimer’s Disease and Related Dementias. Neurotherapeutics 2023, 20, 955–974. [Google Scholar] [CrossRef]
- Ren, R.-J.; Zhang, Y.-F.; Dammer, E.B.; Zhou, Y.; Wang, L.-L.; Liu, X.-H.; Feng, B.-L.; Jiang, G.-X.; Chen, S.-D.; Wang, G.; et al. Peripheral Blood MicroRNA Expression Profiles in Alzheimer’s Disease: Screening, Validation, Association with Clinical Phenotype and Implications for Molecular Mechanism. Mol. Neurobiol. 2016, 53, 5772–5781. [Google Scholar] [CrossRef]
- Sun, H.; Hu, H.; Xu, X.; Tao, T.; Liang, Z. Key miRNAs associated with memory and learning disorder upon exposure to sevoflurane determined by RNA sequencing. Mol. Med. Rep. 2020, 22, 1567–1575. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.-B.; Zhang, Y.-F.; Ren, R.-J.; Dammer, E.B.; Xie, X.-Y.; Chen, S.-W.; Huang, Q.; Huang, W.-Y.; Zhang, R.; Chen, H.-Z.; et al. microRNA-425 loss mediates amyloid plaque microenvironment heterogeneity and promotes neurodegenerative pathologies. Aging Cell 2021, 20, e13454. [Google Scholar] [CrossRef]
- Xue, B.; Qu, Y.; Zhang, X.; Xu, X.-F. miRNA-126a-3p participates in hippocampal memory via alzheimer’s disease-related proteins. Cereb. Cortex. 2022, 32, 4763–4781. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.; Noh, H.; Lee, Y.; Jeon, J.; Shanmugavadivu, A.; McPhie, D.L.; Kim, K.-S.; Cohen, B.M.; Seo, H.; Sonntag, K.C. MiR-126 Regulates Growth Factor Activities and Vulnerability to Toxic Insult in Neurons. Mol. Neurobiol. 2016, 53, 95–108. [Google Scholar] [CrossRef]
- Bicker, F.; Vasic, V.; Horta, G.; Ortega, F.; Nolte, H.; Kavyanifar, A.; Keller, S.; Stankovic, N.D.; Harter, P.N.; Benedito, R.; et al. Neurovascular EGFL7 regulates adult neurogenesis in the subventricular zone and thereby affects olfactory perception. Nat. Commun. 2017, 8, 15922. [Google Scholar] [CrossRef] [PubMed]
- Giuliani, A.; Gaetani, S.; Sorgentoni, G.; Agarbati, S.; Laggetta, M.; Matacchione, G.; Gobbi, M.; Rossi, T.; Galeazzi, R.; Piccinini, G.; et al. Circulating Inflamma-miRs as Potential Biomarkers of Cognitive Impairment in Patients Affected by Alzheimer’s Disease. Front. Aging Neurosci. 2021, 13, 647015. [Google Scholar] [CrossRef]
- Chum, P.P.; Hakim, M.A.; Behringer, E.J. Cerebrovascular microRNA Expression Profile During Early Development of Alzheimer’s Disease in a Mouse Model. J. Alzheimers Dis. 2022, 85, 91–113. [Google Scholar] [CrossRef]
- Gwon, Y.; Kam, T.-I.; Kim, S.-H.; Song, S.; Park, H.; Lim, B.; Lee, H.; Lee, W.; Jo, D.-G.; Jung, Y.-K. TOM1 Regulates Neuronal Accumulation of Amyloid-β Oligomers by FcγRIIb2 Variant in Alzheimer’s Disease. J. Neurosci. 2018, 38, 9001–9018. [Google Scholar] [CrossRef]
- Gámez-Valero, A.; Campdelacreu, J.; Vilas, D.; Ispierto, L.; Rene, R.; Alvarez, R.; Armengol, M.P.; Borras, F.E.; Beyer, K. Exploratory study on microRNA profiles from plasma-derived extracellular vesicles in Alzheimer’s disease and dementia with Lewy bodies. Transl. Neurodegener. 2019, 8, 31. [Google Scholar] [CrossRef] [PubMed]
- Guo, R.; Fan, G.; Zhang, J.; Wu, C.; Du, Y.; Ye, H.; Li, Z.; Wang, L.; Zhang, Z.; Zhang, L.; et al. A 9-microRNA Signature in Serum Serves as a Noninvasive Biomarker in Early Diagnosis of Alzheimer’s Disease. J. Alzheimers Dis. 2017, 60, 1365–1377. [Google Scholar] [CrossRef] [PubMed]
- Romano, G.L.; Platania, C.B.M.; Drago, F.; Salomone, S.; Ragusa, M.; Barbagallo, C.; Di Pietro, C.; Purrello, M.; Reibaldi, M.; Avitabile, T.; et al. Retinal and Circulating miRNAs in Age-Related Macular Degeneration: An In vivo Animal and Human Study. Front. Pharmacol. 2017, 8, 168. [Google Scholar] [CrossRef] [PubMed]
- Burlacu, C.-C.; Ciobanu, D.; Badulescu, A.-V.; Chelaru, V.-F.; Mitre, A.-O.; Capitanescu, B.; Hermann, D.M.; Popa-Wagner, A. Circulating MicroRNAs and Extracellular Vesicle-Derived MicroRNAs as Predictors of Functional Recovery in Ischemic Stroke Patients: A Systematic Review and Meta-Analysis. Int. J. Mol. Sci. 2022, 24, 251. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Mozo, M.I.; Casanova, I.; de Torres, L.; Aladro-Benito, Y.; Perez-Perez, S.; Garcia-Martinez, A.; Gomez, P.; Abellan, S.; De Antonio, E.; Lopez-De-Silanes, C.; et al. microRNA Expression and Its Association with Disability and Brain Atrophy in Multiple Sclerosis Patients Treated with Glatiramer Acetate. Front. Immunol. 2022, 13, 904683. [Google Scholar] [CrossRef]
- Vázquez-Villaseñor, I.; Smith, C.I.; Thang, Y.J.R.; Hearth, P.R.; Wharton, S.B.; Blackburn, D.J.; Ridger, V.C.; Simpson, J.E. RNA-Seq Profiling of Neutrophil-Derived Microvesicles in Alzheimer’s Disease Patients Identifies a miRNA Signature That May Impact Blood-Brain Barrier Integrity. Int. J. Mol. Sci. 2022, 23, 5913. [Google Scholar] [CrossRef]
- Martinez-Arroyo, O.; Ortega, A.; Flores-Chova, A.; Sanches-Garcia, B.; Garcia-Garcia, A.B.; Chaves, F.J.; Martin-Escudero, J.C.; Forner, M.J.; Redon, J.; Cortes, R. High miR-126-3p levels associated with cardiovascular events in a general population. Eur. J. Intern. Med. 2023, 113, 49–56. [Google Scholar] [CrossRef]
- Małczyńska, P.; Piotrowicz, Z.; Drabarek, D.; Langfort, J.; Chalimoniuk, M. Rola mózgowego czynnika neurotroficznego (BDNF) w procesach neurodegeneracji oraz w mechanizmach neuroregeneracji wywołanej wzmożoną aktywnością fizyczną [The role of the brain-derived neurotrophic factor (BDNF) in neurodegenerative processes and in the neuroregeneration mechanisms induced by increased physical activity]. Postepy Biochem. 2019, 65, 2–8. [Google Scholar] [CrossRef]
- Targa, A.; Dakterzada, F.; Benítez, I.D.; de Gonzalo-Calvo, D.; Moncusí-Moix, A.; López, R.; Pujol, M.; Arias, A.; de Batlle, J.; Sánchez-de-la-Torre, M.; et al. Circulating MicroRNA Profile Associated with Obstructive Sleep Apnea in Alzheimer’s Disease. Mol. Neurobiol. 2020, 57, 4363–4372. [Google Scholar] [CrossRef]
- Jiang, F.; Yang, M.; Wu, C.; Wang, J. Potential Roles of miR-374a-5p in Mediating Neuroprotective Effects and Related Molecular Mechanism. J. Mol. Neurosci. 2019, 69, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Pan, W.; Hu, Y.; Wang, L.; Li, J. Circ_0003611 acts as a miR-885-5p sponge to aggravate the amyloid-β-induced neuronal injury in Alzheimer’s disease. Metab. Brain Dis. 2022, 37, 961–971. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Yu, J.-T.; Tan, M.-S.; Liu, Q.-Y.; Wang, W.-F.; Zhang, W.; Jiang, T.; Tan, L. Genome-wide serum microRNA expression profiling identifies serum biomarkers for Alzheimer’s disease. J. Alzheimers Dis. 2014, 40, 1017–1027. [Google Scholar] [CrossRef] [PubMed]
- Guévremont, D.; Tsui, H.; Knight, R.; Fowler, C.J.; Masters, C.L.; Martins, R.N.; Abraham, W.C.; Tate, W.P.; Cutfield, M.J.; Williams, J.M. Plasma microRNA vary in association with the progression of Alzheimer’s disease. Alzheimers Dement. 2022, 14, e12251. [Google Scholar] [CrossRef] [PubMed]
- Streese, L.; Demougin, P.; Iborra, P.; Kanitz, A.; Deiseroth, A.; Kröpfl, J.M.; Schmidt-Trucksäss, A.; Zavolan, M.; Hanssen, H. Untargeted sequencing of circulating microRNAs in a healthy and diseased older population. Sci. Rep. 2022, 12, 2991. [Google Scholar] [CrossRef] [PubMed]
- Song, D.; Li, G.; Hong, Y.; Zhang, P.; Zhu, J.; Yang, L.; Huang, J. miR-199a decreases Neuritin expression involved in the development of Alzheimer’s disease in APP/PS1 mice. Int. J. Mol. Med. 2020, 46, 384–396. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.D.; Kim, M.-S. Exposure to a mixture of heavy metals induces cognitive impairment: Genes and microRNAs involved. Toxicology 2022, 471, 153164. [Google Scholar] [CrossRef]
- Ghiam, S.; Eslahchi, C.; Shahpasand, K.; Habibi-Rezaei, M.; Gharaghani, S. Exploring the role of non-coding RNAs as potential candidate biomarkers in the cross-talk between diabetes mellitus and Alzheimer’s disease. Front. Aging Neurosci. 2022, 14, 955461. [Google Scholar] [CrossRef]
- Zhang, X.; Zhou, G. MiR-199a-5p inhibition protects cognitive function of ischemic stroke rats by AKT signaling pathway. Am. J. Transl. Res. 2020, 12, 6549–6558. [Google Scholar]
- Sørensen, S.S.; Nygaard, A.-B.; Christensen, T. miRNA expression profiles in cerebrospinal fluid and blood of patients with Alzheimer’s disease and other types of dementia—An exploratory study. Transl. Neurodegener. 2016, 5, 6. [Google Scholar] [CrossRef]
- Wang, T.; Cheng, Y.; Han, H.; Liu, J.; Tian, B.; Liu, X. miR-194 Accelerates Apoptosis of Aβ1⁻42-Transduced Hippocampal Neurons by Inhibiting Nrn1 and Decreasing PI3K/Akt Signaling Pathway Activity. Genes 2019, 10, 313. [Google Scholar] [CrossRef] [PubMed]
- Khodabakhsh, P.; Bazrgar, M.; Mohagheghi, F.; Parvardeh, S.; Ahmadiani, A. MicroRNA-140-5p inhibitor attenuates memory impairment induced by amyloid-ß oligomer in vivo possibly through Pin1 regulation. CNS Neurosci. Ther. 2023, 29, 91–103. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.P.N.; Kumar, M.; Fedele, E.; Bonanno, G.; Bonifacino, T. MicroRNA Alteration, Application as Biomarkers, and Therapeutic Approaches in Neurodegenerative Diseases. Int. J. Mol. Sci. 2022, 23, 4718. [Google Scholar] [CrossRef] [PubMed]
- Liang, H.-B.; Lai, Z.-H.; Tu, X.-Q.; Ding, K.-Q.; He, J.-R.; Yang, G.-Y.; Sheng, H.; Zheng, L.-L. MicroRNA-140-5p exacerbates vascular cognitive impairment by inhibiting neurogenesis in the adult mouse hippocampus after global cerebral ischemia. Brain Res. Bull. 2022, 183, 73–83. [Google Scholar] [CrossRef]
- Gullett, J.M.; Chen, Z.; O’Shea, A.; Akbar, M.; Bian, J.; Rani, A.; Porges, E.C.; Foster, T.C.; Woods, A.J.; Modave, F.; et al. MicroRNA predicts cognitive performance in healthy older adults. Neurobiol. Aging 2020, 95, 186–194. [Google Scholar] [CrossRef]
- Akhter, R.; Shao, Y.; Shaw, M.; Formica, S.; Khrestian, M.; Leverenz, J.B.; Bekris, L.M. Regulation of ADAM10 by miR-140-5p and potential relevance for Alzheimer’s disease. Neurobiol. Aging 2018, 63, 110–119. [Google Scholar] [CrossRef]
- Song, W.; Wang, T.; Shi, B.; Wu, Z.; Wang, W.; Yang, Y. Neuroprotective effects of microRNA-140-5p on ischemic stroke in mice via regulation of the TLR4/NF-κB axis. Brain Res. Bull. 2021, 168, 8–16. [Google Scholar] [CrossRef]
- Völzke, H.; Schössow, J.; Schmidt, C.O.; Jürgens, C.; Richter, A.; Werner, A.; Werner, N.; Radke, D.; Teumer, A.; Ittermann, T.; et al. Cohort Profile Update: The Study of Health in Pomerania (SHIP). Int. J. Epidemiol. 2022, 51, e372–e383. [Google Scholar] [CrossRef]
- Oswald, W.D.; Fleischmann, U.M. (Eds.) NAI-Testmanual und Textband. In Nürnberger-Alters-Inventar: (NAI); Hogrefe: Boston, MA, USA, 1999. [Google Scholar]
- van der Auwera, S.; Garvert, L.; Ameling, S.; Völzke, H.; Nauck, M.; Völker, U.; Grabe, H.J. The interplay between micro RNAs and genetic liability to Alzheimer’s Disease on memory trajectories in the general population. Psychiatry Res. 2023, 323, 115141. [Google Scholar] [CrossRef]
- Kirchner, K.; Garvert, L.; Wittfeld, K.; Ameling, S.; Bülow, R.; Meyer Zu Schwabedissen, H.; Nauck, M.; Völzke, H.; Grabe, H.J.; Van der Auwera, S. Deciphering the Effect of Different Genetic Variants on Hippocampal Subfield Volumes in the General Population. Int. J. Mol. Sci. 2023, 24, 1120. [Google Scholar] [CrossRef] [PubMed]
- Frenzel, S.; Wittfeld, K.; Habes, M.; Klinger-König, J.; Bülow, R.; Völzke, H.; Grabe, H.J. A Biomarker for Alzheimer’s Disease Based on Patterns of Regional Brain Atrophy. Front. Psychiatry 2019, 10, 953. [Google Scholar] [CrossRef]
- Sollis, E.; Mosaku, A.; Abid, A.; Buniello, A.; Cerezo, M.; Gil, L.; Groza, T.; Güneş, O.; Hall, P.; Hayhurst, J.; et al. The NHGRI-EBI GWAS Catalog: Knowledgebase and deposition resource. Nucleic Acids Res. 2023, 51, D977–D985. [Google Scholar] [CrossRef]
- Kamat, M.A.; Blackshaw, J.A.; Young, R.; Surendran, P.; Burgess, S.; Danesh, J.; Butterworth, A.S.; Staley, J.R. PhenoScanner V2: An expanded tool for searching human genotype-phenotype associations. Bioinformatics 2019, 35, 4851–4853. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.M.; Douaud, G.; Chen, W.; Hanayik, T.; Alfaro-Almagro, F.; Sharp, K.; Elliott, L.T. An expanded set of genome-wide association studies of brain imaging phenotypes in, U.K.; Biobank. Nat. Neurosci. 2021, 24, 737–745. [Google Scholar] [CrossRef]
- Liu, C.-J.; Fu, X.; Xia, M.; Zhang, Q.; Gu, Z.; Guo, A.-Y. miRNASNP-v3: A comprehensive database for SNPs and disease-related variations in miRNAs and miRNA targets. Nucleic Acids Res. 2021, 49, D1276–D1281. [Google Scholar] [CrossRef] [PubMed]
- Keller, A.; Gröger, L.; Tschernig, T.; Solomon, J.; Laham, O.; Schaum, N.; Wagner, V.; Kern, F.; Schmartz, G.P.; Li, Y.; et al. miRNATissueAtlas2: An update to the human miRNA tissue atlas. Nucleic Acids Res. 2022, 50, D211–D221. [Google Scholar] [CrossRef]
- Thul, P.J.; Lindskog, C. The human protein atlas: A spatial map of the human proteome. Protein Sci. 2018, 27, 233–244. [Google Scholar] [CrossRef]
- Pomper, N.; Liu, Y.; Hoye, M.L.; Dougherty, J.D.; Miller, T.M. CNS microRNA profiles: A database for cell type enriched microRNA expression across the mouse central nervous system. Sci. Rep. 2020, 10, 4921. [Google Scholar] [CrossRef]
- Huang, H.-Y.; Lin, Y.-C.-D.; Cui, S.; Huang, Y.; Tang, Y.; Xu, J.; Bao, J.; Li, Y.; Wen, J.; Zuo, H.; et al. miRTarBase update 2022: An informative resource for experimentally validated miRNA-target interactions. Nucleic Acids Res. 2022, 50, D222–D230. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, X. miRDB: An online database for prediction of functional microRNA targets. Nucleic Acids Res. 2020, 48, D127–D131. [Google Scholar] [CrossRef] [PubMed]
- McGeary, S.E.; Lin, K.S.; Shi, C.Y.; Pham, T.M.; Bisaria, N.; Kelley, G.M.; Bartel, D.P. The biochemical basis of microRNA targeting efficacy. Science 2019, 366, eaav1741. [Google Scholar] [CrossRef] [PubMed]
- Aparicio-Puerta, E.; Hirsch, P.; Schmartz, G.P.; Kern, F.; Fehlmann, T.; Keller, A. miEAA 2023: Updates, new functional microRNA sets and improved enrichment visualizations. Nucleic Acids Res. 2023, 51, W319–W325. [Google Scholar] [CrossRef] [PubMed]
- Hosten, N.; Bülow, R.; Völzke, H.; Domin, M.; Schmidt, C.O.; Teumer, A.; Ittermann, T.; Nauck, M.; Felix, S.; Dörr, M.; et al. SHIP-MR and Radiology: 12 Years of Whole-Body Magnetic Resonance Imaging in a Single Center. Healthcare 2021, 10. [Google Scholar] [CrossRef]
- Pitchika, A.; Markus, M.R.P.; Schipf, S.; Teumer, A.; van der Auwera, S.; Nauck, M.; Dörr, M.; Felix, S.; Grabe, H.-J.; Völzke, H.; et al. Effects of Apolipoprotein E polymorphism on carotid intima-media thickness, incident myocardial infarction and incident stroke. Sci. Rep. 2022, 12, 5142. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, P.; Gaser, C.; Arsic, M.; Buck, D.; Förschler, A.; Berthele, A.; Hoshi, M.; Ilg, R.; Schmid, V.J.; Zimmer, C.; et al. An automated tool for detection of FLAIR-hyperintense white-matter lesions in Multiple Sclerosis. NeuroImage 2012, 59, 3774–3783. [Google Scholar] [CrossRef]
| TREND-0 miRNA/MRI Sample * (n = 648) | TREND-0 MRI Sample (n = 1854) | |
|---|---|---|
| Sex | ||
| Males | 328 (50.6%) | 890 (48%) |
| Females | 320 (49.4%) | 964 (52%) |
| Age in years | 50.3 (13.7), [21–79] | 51.0 (13.9), [21–81] |
| BMI | 27.2 (4.2), [17.7–48.0] | 27.5 (4.4), [17.7–48.0] |
| Systolic blood pressure (mmHg) | 124.9 (16.3), [88–196] | 126.2 (17.2), [84–196] |
| diastolic blood pressure (mmHg) | 76.5 (9.6), [51–115] | 77.1 (9.9), [47–118] |
| Hypertension | 251 (38.8%) | 790 (42.7%) |
| Current depressive symptoms (PHQ-9) | 12.7 (3.4), [9–35] | 12.8 (3.5), [9–35] |
| Education | ||
| <10 years | 68 (10.5%) | 273 (14.7%) |
| =10 years | 369 (57%) | 1011 (54.5%) |
| >10 years | 211 (32.5%) | 570 (30.8%) |
| Smoking | ||
| Never | 267 (41.2%) | 732 (39.5%) |
| Former | 247 (38.1%) | 684 (36.9%) |
| Current | 134 (20.7%) | 438 (23.6%) |
| Verbal memory immediate recall | 5.4 (1.2), [0–8] | 5.4 (1.3), [0–8] |
| Verbal memory delayed recall | 5.7 (1.6), [−3–8] | 5.8 (1.7), [−3–8] |
| APOE ε4 carrier | 155 (24.0%) | 448 (24.2%) |
| ICV in cm3 | 1563 (148), [1016–2040] | 1560 (145), [1016–2040] |
| WMLV in cm3 | 0.56 (2.1), [0–27] | 0.68 (2.4), [0–43.8] |
| Number of WMLs | 3.0 (4.2), [0–37] | 3.2 (4.4), [0–37] |
| Presence of lesions | 454 (70.1%) | 1320 (71.2%) |
| miRNA | WML Volume | Number of WMLs | Presence of WMLs | n |
|---|---|---|---|---|
| Direct effects | ||||
| hsa-miR-425-5p | Pos, 0.46, 0.94 | Pos, 0.005, 0.63 | Pos, 5.9 × 10−5, 0.01 | 638 |
| Interaction with sex | ||||
| hsa-miR-126-3p | Neg, 1.7 × 10−3, 0.13 | Neg, 2.6 × 10−4, 0.044 | - | 641 |
| hsa-miR-374a-5p | Neg, 0.03, 0.39 | Neg, 9.4 × 10−4, 0.08 | - | 410 |
| Interaction with APOE ε4 carrier status | ||||
| hsa-miR-140-5p * | Neg, 0.024, 0.58 | Neg, 0.001, 0.12 | - | 497 |
| Interaction with smoking status | ||||
| hsa-miR-199a-5p | Neg, 1.0 × 10−4, 0.018 | Neg, 2.6 × 10−4, 0.022 | - | 516 |
| hsa-miR-885-5p | Pos, 9.6 × 10−4, 0.083 | Pos, 1.2 × 10−4, 0.022 | - | 544 |
| hsa-miR-194-5p | Pos, 0.011, 0.32 | Pos, 7.8 × 10−4, 0.045 | - | 619 |
| miR-425-5p | miR-126-3p | miR-374a-5p | miR-199a-5p | miR-885-5p | miR-194-5p | FDR Corrected p-Value | |
|---|---|---|---|---|---|---|---|
| Neurodegenerative diseases | x | x | x | x | x | x | 0.005 |
| Niemann-pick disease | x | x | x | x | - | - | 3.5 × 10−4 |
| Alzheimer’s disease | x | x | x | x | x | x | 0.008 |
| Multiple sclerosis | x | x | x | x | - | x | 0.003 |
| Amyotrophic lateral sclerosis | x | x | - | x | x | x | 0.006 |
| Intellectual disability | x | x | - | x | x | x | 0.004 |
| Vascular disease | x | - | x | x | x | x | 0.018 |
| Brain disease | x | x | - | x | x | x | 0.012 |
| Huntington’s disease | - | x | - | x | x | x | 0.009 |
| Stroke | - | x | x | x | - | x | 0.011 |
| Schizophrenia | - | x | - | x | x | - | 0.011 |
| Hypertension | x | x | - | x | - | - | 0.023 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Van der Auwera, S.; Ameling, S.; Wittfeld, K.; Frenzel, S.; Bülow, R.; Nauck, M.; Völzke, H.; Völker, U.; Grabe, H.J. Circulating microRNA miR-425-5p Associated with Brain White Matter Lesions and Inflammatory Processes. Int. J. Mol. Sci. 2024, 25, 887. https://doi.org/10.3390/ijms25020887
Van der Auwera S, Ameling S, Wittfeld K, Frenzel S, Bülow R, Nauck M, Völzke H, Völker U, Grabe HJ. Circulating microRNA miR-425-5p Associated with Brain White Matter Lesions and Inflammatory Processes. International Journal of Molecular Sciences. 2024; 25(2):887. https://doi.org/10.3390/ijms25020887
Chicago/Turabian StyleVan der Auwera, Sandra, Sabine Ameling, Katharina Wittfeld, Stefan Frenzel, Robin Bülow, Matthias Nauck, Henry Völzke, Uwe Völker, and Hans J. Grabe. 2024. "Circulating microRNA miR-425-5p Associated with Brain White Matter Lesions and Inflammatory Processes" International Journal of Molecular Sciences 25, no. 2: 887. https://doi.org/10.3390/ijms25020887
APA StyleVan der Auwera, S., Ameling, S., Wittfeld, K., Frenzel, S., Bülow, R., Nauck, M., Völzke, H., Völker, U., & Grabe, H. J. (2024). Circulating microRNA miR-425-5p Associated with Brain White Matter Lesions and Inflammatory Processes. International Journal of Molecular Sciences, 25(2), 887. https://doi.org/10.3390/ijms25020887





