Structural and Dynamic-Based Characterization of the Recognition Patterns of E7 and TRP-2 Epitopes by MHC Class I Receptors through Computational Approaches
Abstract
:1. Introduction
2. Results
2.1. Quality Assessment of AlphaFold Predictions
2.1.1. GP33Db System
2.1.2. TRP-2A68 System
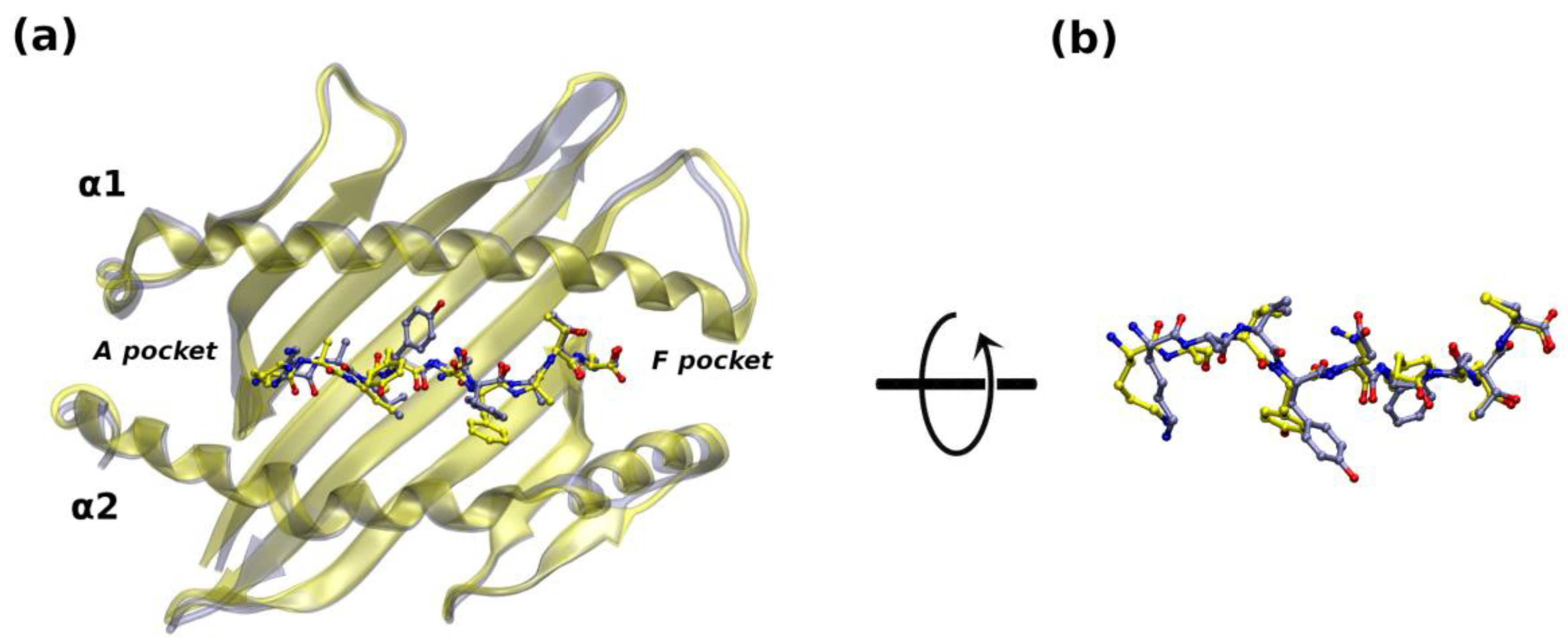
2.2. Ab Initio Prediction of E7 Interaction with H2Db
2.2.1. AF Modeling: Evaluation of the Structural Models
2.2.2. A Dynamic View of the Interaction: MD Study
2.3. Ab Initio Prediction of TRP-2 Interaction with MHC Receptors
2.3.1. Selection of MHC-Peptide Systems
2.3.2. AF Modeling: Evaluation of the Structural Models
2.3.3. A Dynamic View of the Interaction: MD Study
3. Discussion

4. Materials and Methods
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Bouvier, M.; Wiley, D.C. Importance of Peptide Amino and Carboxyl Termini to the Stability of MHC Class I Molecules. Science 1994, 265, 398–402. [Google Scholar] [CrossRef] [PubMed]
- Sieker, F.; Springer, S.; Zacharias, M. Comparative Molecular Dynamics Analysis of Tapasin-Dependent and -Independent MHC Class I Alleles. Protein Sci. 2007, 16, 299–308. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Peng, X.; Batliwala, M.; Bouvier, M. Crystal Structures of MHC Class I Complexes Reveal the Elusive Intermediate Conformations Explored during Peptide Editing. Nat. Commun. 2023, 14, 5020. [Google Scholar] [CrossRef] [PubMed]
- Wieczorek, M.; Abualrous, E.T.; Sticht, J.; Álvaro-Benito, M.; Stolzenberg, S.; Noé, F.; Freund, C. Major Histocompatibility Complex (MHC) Class I and MHC Class II Proteins: Conformational Plasticity in Antigen Presentation. Front. Immunol. 2017, 8, 292. [Google Scholar] [CrossRef] [PubMed]
- Zacharias, M.; Springer, S. Conformational Flexibility of the MHC Class I Alpha1-Alpha2 Domain in Peptide Bound and Free States: A Molecular Dynamics Simulation Study. Biophys. J. 2004, 87, 2203–2214. [Google Scholar] [CrossRef]
- Abualrous, E.T.; Saini, S.K.; Ramnarayan, V.R.; Ilca, F.T.; Zacharias, M.; Springer, S. The Carboxy Terminus of the Ligand Peptide Determines the Stability of the MHC Class I Molecule H-2Kb: A Combined Molecular Dynamics and Experimental Study. PLoS ONE 2015, 10, e0135421. [Google Scholar] [CrossRef]
- Panahi, H.A.; Bolhassani, A.; Javadi, G.; Noormohammadi, Z. A Comprehensive in Silico Analysis for Identification of Therapeutic Epitopes in HPV16, 18, 31 and 45 Oncoproteins. PLoS ONE 2018, 13, e0205933. [Google Scholar] [CrossRef]
- Jabbar, B.; Rafique, S.; Salo-Ahen, O.M.H.; Ali, A.; Munir, M.; Idrees, M.; Mirza, M.U.; Vanmeert, M.; Shah, S.Z.; Jabbar, I.; et al. Antigenic Peptide Prediction From E6 and E7 Oncoproteins of HPV Types 16 and 18 for Therapeutic Vaccine Design Using Immunoinformatics and MD Simulation Analysis. Front. Immunol. 2018, 9, 3000. [Google Scholar] [CrossRef]
- Ochoa, R.; Laio, A.; Cossio, P. Predicting the Affinity of Peptides to Major Histocompatibility Complex Class II by Scoring Molecular Dynamics Simulations. J. Chem. Inf. Model. 2019, 59, 3464–3473. [Google Scholar] [CrossRef]
- Parizi, F.M.; Marzella, D.F.; Ramakrishnan, G.; AC’t Hoen, P.; Karimi-Jafari, M.H.; Xue, L.C. PANDORA v2.0: Benchmarking Peptide-MHC II Models and Software Improvements. Front. Immunol. 2023, 14, 1285899. [Google Scholar] [CrossRef]
- Amari, S.; Kataoka, R.; Ikegami, T.; Hirayama, N. HLA-Modeler: Automated Homology Modeling of Human Leukocyte Antigens. Int. J. Med. Chem. 2013, 2013, 690513. [Google Scholar] [CrossRef] [PubMed]
- Tran, N.H.; Xu, J.; Li, M. A Tale of Solving Two Computational Challenges in Protein Science: Neoantigen Prediction and Protein Structure Prediction. Brief. Bioinform. 2022, 23, bbab493. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Ip, P.; Nijman, H.; Daemen, T. Epitope Prediction Assays Combined with Validation Assays Strongly Narrows down Putative Cytotoxic T Lymphocyte Epitopes. Vaccines 2015, 3, 203–220. [Google Scholar] [CrossRef] [PubMed]
- Peng, S.; Xing, D.; Ferrall, L.; Tsai, Y.-C.; Roden, R.B.S.; Hung, C.-F.; Wu, T.-C. Development of a Spontaneous HPV16 E6/E7-Expressing Head and Neck Squamous Cell Carcinoma in HLA-A2 Transgenic Mice. mBio 2022, 13, e03252-21. [Google Scholar] [CrossRef]
- Smahel, M.; Tejklova, P.; Smahelova, J.; Polakova, I.; Mackova, J. Mutation in the Immunodominant Epitope of the HPV16 E7 Oncoprotein as a Mechanism of Tumor Escape. Cancer Immunol. Immunother. CII 2008, 57, 823–831. [Google Scholar] [CrossRef]
- Tagliamonte, M.; Mauriello, A.; Cavalluzzo, B.; Ragone, C.; Manolio, C.; Luciano, A.; Barbieri, A.; Palma, G.; Scognamiglio, G.; Di Mauro, A.; et al. MHC-Optimized Peptide Scaffold for Improved Antigen Presentation and Anti-Tumor Response. Front. Immunol. 2021, 12, 769799. [Google Scholar] [CrossRef]
- Cavalluzzo, B.; Ragone, C.; Mauriello, A.; Petrizzo, A.; Manolio, C.; Caporale, A.; Vitagliano, L.; Ruvo, M.; Buonaguro, L.; Tagliamonte, M. Identification and Characterization of Heteroclitic Peptides in TCR-Binding Positions with Improved HLA-Binding Efficacy. J. Transl. Med. 2021, 19, 89. [Google Scholar] [CrossRef]
- Bloom, M.B.; Perry-Lalley, D.; Robbins, P.F.; Li, Y.; el-Gamil, M.; Rosenberg, S.A.; Yang, J.C. Identification of Tyrosinase-Related Protein 2 as a Tumor Rejection Antigen for the B16 Melanoma. J. Exp. Med. 1997, 185, 453–459. [Google Scholar] [CrossRef]
- Capasso, C.; Magarkar, A.; Cervera-Carrascon, V.; Fusciello, M.; Feola, S.; Muller, M.; Garofalo, M.; Kuryk, L.; Tähtinen, S.; Pastore, L.; et al. A Novel in Silico Framework to Improve MHC-I Epitopes and Break the Tolerance to Melanoma. Oncoimmunology 2017, 6, e1319028. [Google Scholar] [CrossRef]
- Yang, X.; Nishimiya, D.; Löchte, S.; Jude, K.M.; Borowska, M.; Savvides, C.S.; Dougan, M.; Su, L.; Zhao, X.; Piehler, J.; et al. Facile Repurposing of Peptide-MHC-Restricted Antibodies for Cancer Immunotherapy. Nat. Biotechnol. 2023, 41, 932–943. [Google Scholar] [CrossRef] [PubMed]
- Sewell, A.K. Why Must T Cells Be Cross-Reactive? Nat. Rev. Immunol. 2012, 12, 669–677. [Google Scholar] [CrossRef] [PubMed]
- Tissot, A.C.; Ciatto, C.; Mittl, P.R.E.; Grütter, M.G.; Plückthun, A. Viral Escape at the Molecular Level Explained by Quantitative T-Cell Receptor/Peptide/MHC Interactions and the Crystal Structure of a Peptide/MHC Complex. J. Mol. Biol. 2000, 302, 873–885. [Google Scholar] [CrossRef] [PubMed]
- Niu, L.; Cheng, H.; Zhang, S.; Tan, S.; Zhang, Y.; Qi, J.; Liu, J.; Gao, G.F. Structural Basis for the Differential Classification of HLA-A*6802 and HLA-A*6801 into the A2 and A3 Supertypes. Mol. Immunol. 2013, 55, 381–392. [Google Scholar] [CrossRef] [PubMed]
- Hoppes, R.; Oostvogels, R.; Luimstra, J.J.; Wals, K.; Toebes, M.; Bies, L.; Ekkebus, R.; Rijal, P.; Celie, P.H.N.; Huang, J.H.; et al. Altered Peptide Ligands Revisited: Vaccine Design through Chemically Modified HLA-A2-Restricted T Cell Epitopes. J. Immunol. 2014, 193, 4803–4813. [Google Scholar] [CrossRef] [PubMed]
- Wang, E.H.C.; Yu, M.; Breitkopf, T.; Akhoundsadegh, N.; Wang, X.; Shi, F.-T.; Leung, G.; Dutz, J.P.; Shapiro, J.; McElwee, K.J. Identification of Autoantigen Epitopes in Alopecia Areata. J. Investig. Dermatol. 2016, 136, 1617–1626. [Google Scholar] [CrossRef]
- You, S.; Choi, Y.S.; Hong, S.; Shin, E.-C. Priming of Autoreactive CD8(+) T Cells Is Inhibited by Immunogenic Peptides Which Are Competitive for Major Histocompatibility Complex Class I Binding. Immune Netw. 2013, 13, 86–93. [Google Scholar] [CrossRef]
- Hudrisier, D.; Mazarguil, H.; Laval, F.; Oldstone, M.B.A.; Gairin, J.E. Binding of Viral Antigens to Major Histocompatibility Complex Class I H-2Db Molecules Is Controlled by Dominant Negative Elements at Peptide Non-Anchor Residues. J. Biol. Chem. 1996, 271, 17829–17836. [Google Scholar] [CrossRef]
- Schuster, H.; Shao, W.; Weiss, T.; Pedrioli, P.G.A.; Roth, P.; Weller, M.; Campbell, D.S.; Deutsch, E.W.; Moritz, R.L.; Planz, O.; et al. A Tissue-Based Draft Map of the Murine MHC Class I Immunopeptidome. Sci. Data 2018, 5, 180157. [Google Scholar] [CrossRef]
- Zhao, R.; Loftus, D.J.; Appella, E.; Collins, E.J. Structural Evidence of T Cell Xeno-Reactivity in the Absence of Molecular Mimicry. J. Exp. Med. 1999, 189, 359–370. [Google Scholar] [CrossRef]
- La Gruta, N.L.; Thomas, P.G.; Webb, A.I.; Dunstone, M.A.; Cukalac, T.; Doherty, P.C.; Purcell, A.W.; Rossjohn, J.; Turner, S.J. Epitope-Specific TCRbeta Repertoire Diversity Imparts No Functional Advantage on the CD8+ T Cell Response to Cognate Viral Peptides. Proc. Natl. Acad. Sci. USA 2008, 105, 2034–2039. [Google Scholar] [CrossRef]
- Valkenburg, S.A.; Gras, S.; Guillonneau, C.; Hatton, L.A.; Bird, N.A.; Twist, K.-A.; Halim, H.; Jackson, D.C.; Purcell, A.W.; Turner, S.J.; et al. Preemptive Priming Readily Overcomes Structure-Based Mechanisms of Virus Escape. Proc. Natl. Acad. Sci. USA 2013, 110, 5570–5575. [Google Scholar] [CrossRef] [PubMed]
- Van Braeckel-Budimir, N.; Gras, S.; Ladell, K.; Josephs, T.M.; Pewe, L.; Urban, S.L.; Miners, K.L.; Farenc, C.; Price, D.A.; Rossjohn, J.; et al. A T Cell Receptor Locus Harbors a Malaria-Specific Immune Response Gene. Immunity 2017, 47, 835–847.e4. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold—Making Protein Folding Accessible to All. Nat. Methods 2021, 19, 679–682. [Google Scholar] [CrossRef] [PubMed]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J.C. GROMACS: Fast, Flexible, and Free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef] [PubMed]
- Lindorff-Larsen, K.; Piana, S.; Palmo, K.; Maragakis, P.; Klepeis, J.L.; Dror, R.O.; Shaw, D.E. Improved Side-Chain Torsion Potentials for the Amber ff99SB Protein Force Field: Improved Protein Side-Chain Potentials. Proteins Struct. Funct. Bioinform. 2010, 78, 1950–1958. [Google Scholar] [CrossRef] [PubMed]
- Paladino, A.; Zangi, R. Ribose 2′-Hydroxyl Groups Stabilize RNA Hairpin Structures Containing GCUAA Pentaloop. J. Chem. Theory Comput. 2013, 9, 1214–1221. [Google Scholar] [CrossRef]
- Paladino, A.; D’Angelo, F.; Noviello, T.M.R.; Iavarone, A.; Ceccarelli, M. Structural Model for Recruitment of RIT1 to the LZTR1 E3 Ligase: Evidences from an Integrated Computational Approach. J. Chem. Inf. Model. 2021, 61, 1875–1888. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of Simple Potential Functions for Simulating Liquid Water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Parrinello, M.; Rahman, A. Polymorphic Transitions in Single Crystals: A New Molecular Dynamics Method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical Sampling through Velocity Rescaling. J. Chem. Phys. 2007, 126, 014101. [Google Scholar] [CrossRef] [PubMed]
- Darden, T.; York, D.; Pedersen, L. Particle Mesh Ewald: An N⋅log(N) Method for Ewald Sums in Large Systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.C.; Fraaije, J.G.E.M. LINCS: A Linear Constraint Solver for Molecular Simulations. J. Comput. Chem. 1997, 18, 1463–1472. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Swindells, M.B. LigPlot+: Multiple Ligand-Protein Interaction Diagrams for Drug Discovery. J. Chem. Inf. Model. 2011, 51, 2778–2786. [Google Scholar] [CrossRef] [PubMed]
- Schrödinger, L.; DeLano, W. PyMOL. 2020. Available online: http://www.pymol.org/pymol (accessed on 1 November 2023).
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual Molecular Dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Nielsen, M.; Lundegaard, C.; Worning, P.; Lauemøller, S.L.; Lamberth, K.; Buus, S.; Brunak, S.; Lund, O. Reliable Prediction of T-Cell Epitopes Using Neural Networks with Novel Sequence Representations. Protein Sci. 2003, 12, 1007–1017. [Google Scholar] [CrossRef]
- Andreatta, M.; Nielsen, M. Gapped Sequence Alignment Using Artificial Neural Networks: Application to the MHC Class I System. Bioinform. Oxf. Engl. 2016, 32, 511–517. [Google Scholar] [CrossRef]
- Thomsen, M.C.F.; Nielsen, M. Seq2Logo: A Method for Construction and Visualization of Amino Acid Binding Motifs and Sequence Profiles Including Sequence Weighting, Pseudo Counts and Two-Sided Representation of Amino Acid Enrichment and Depletion. Nucleic Acids Res. 2012, 40, W281–W287. [Google Scholar] [CrossRef]





| EPITOPE SEQUENCE | H2 CLASS I MHC | ||
|---|---|---|---|
| HLA-A68 UniProtKB ID P04439 | H2-Db UniProtKB ID P01899 | H2-Kb UniProtKB ID P01901 | |
| GP33 KAVYNFATC | GP33Db * (1FG2) | ||
| E7 RAHYNIVTF | E7Db | ||
| TRP-2 SVYDFFVWL | TRP-2A68 *(4HX1) | TRP-2Db | TRP-2Kb |
| GP33 | H2-Db | E7 | H2-Db | |||
|---|---|---|---|---|---|---|
| Residue | Atom | PDB ID 1FG2 * | AF Model | Residue | Atom | AF Model |
| K1 | N | Y172OH | Y8OH | R1 | N | Y8OH Y172OH |
| O | Y160OH | |||||
| NZ | E164OE2 | NH | R63O | |||
| A2 | N | E64OE1 | A2 | |||
| V3 | O | Q71NE2 | H3 | O | Q71NE2 | |
| NE2 | H156NE2 | |||||
| Y4 | O | H156NE2 | H156NE2 | Y4 | ||
| N5 | N | Q71OE1 | N5 | ND2 | Q98OE1 | |
| OD1 | Q98OE1 Q98NE2 | Y157OH Q98OE1 Q98NE2 | ||||
| ND2 | Q98OE1 Q98NE2 | Q71OE1 Q98OE1 Q98NE2 | ||||
| F6 | N | W74NE1 | I6 | |||
| O | Y157OH | |||||
| A7 | O | W148NE1 | V7 | O | W148NE1 | |
| T8 | O | W148NE1 K147NZ | W148NE1 | T8 | O | K147NZ |
| OG1 | K147NZ | |||||
| C9 | N | S78OG | S78OG | F9 | O | N81ND2 Y85OH K147NZ |
| O | N81ND2 | |||||
| TRP-2 | HLA-A68 | H2-Db | H2-Kb | ||
|---|---|---|---|---|---|
| Residue | Atom | PDB ID 4HX1 | AF Model | AF Model | AF Model |
| S1 | N | Y171OH Y7OH | Y171OH | E164OE | |
| O | Y159OH | Y159OH | |||
| OG | R62NH N63ND2 | K66NZ | |||
| V2 | N | N63OD1 | N63OD1 | ||
| O | N66ND2 | ||||
| Y3 | N | Y99OH | |||
| O | N66ND2 | ||||
| OH | Q70OE1 | R97NH | |||
| D4 | O | R155NH | |||
| F5 | O | Q114NE2 | |||
| F6 | O | R155NH | |||
| V7 | O | Y116OH | |||
| W8 | O | W147NE1 | W147NE1 K146NZ | W147NE1 | |
| L9 | N | D77OD | D77OD | D77OD | |
| O | T143OG K146NZ | T143OG K146NZ | N81ND2 Y85OH | K146NZ | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Balasco, N.; Tagliamonte, M.; Buonaguro, L.; Vitagliano, L.; Paladino, A. Structural and Dynamic-Based Characterization of the Recognition Patterns of E7 and TRP-2 Epitopes by MHC Class I Receptors through Computational Approaches. Int. J. Mol. Sci. 2024, 25, 1384. https://doi.org/10.3390/ijms25031384
Balasco N, Tagliamonte M, Buonaguro L, Vitagliano L, Paladino A. Structural and Dynamic-Based Characterization of the Recognition Patterns of E7 and TRP-2 Epitopes by MHC Class I Receptors through Computational Approaches. International Journal of Molecular Sciences. 2024; 25(3):1384. https://doi.org/10.3390/ijms25031384
Chicago/Turabian StyleBalasco, Nicole, Maria Tagliamonte, Luigi Buonaguro, Luigi Vitagliano, and Antonella Paladino. 2024. "Structural and Dynamic-Based Characterization of the Recognition Patterns of E7 and TRP-2 Epitopes by MHC Class I Receptors through Computational Approaches" International Journal of Molecular Sciences 25, no. 3: 1384. https://doi.org/10.3390/ijms25031384










