An Efficient Expression and Purification Protocol for SpCas9 Nuclease and Evaluation of Different Delivery Methods of Ribonucleoprotein
Abstract
1. Introduction
2. Results
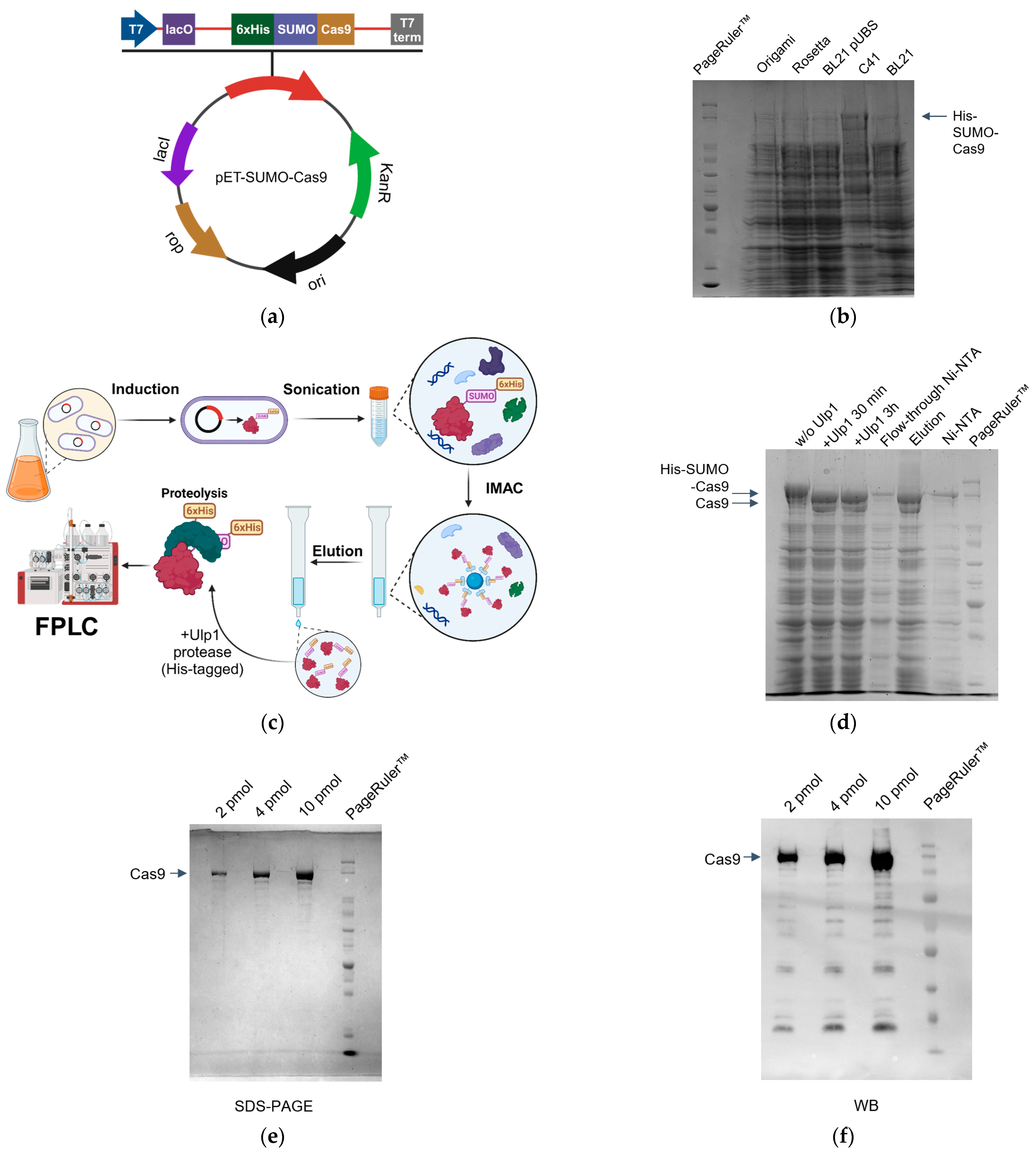
2.1. Optimization of SpCas9 Expression
2.2. SpCas9 Purification Protocol
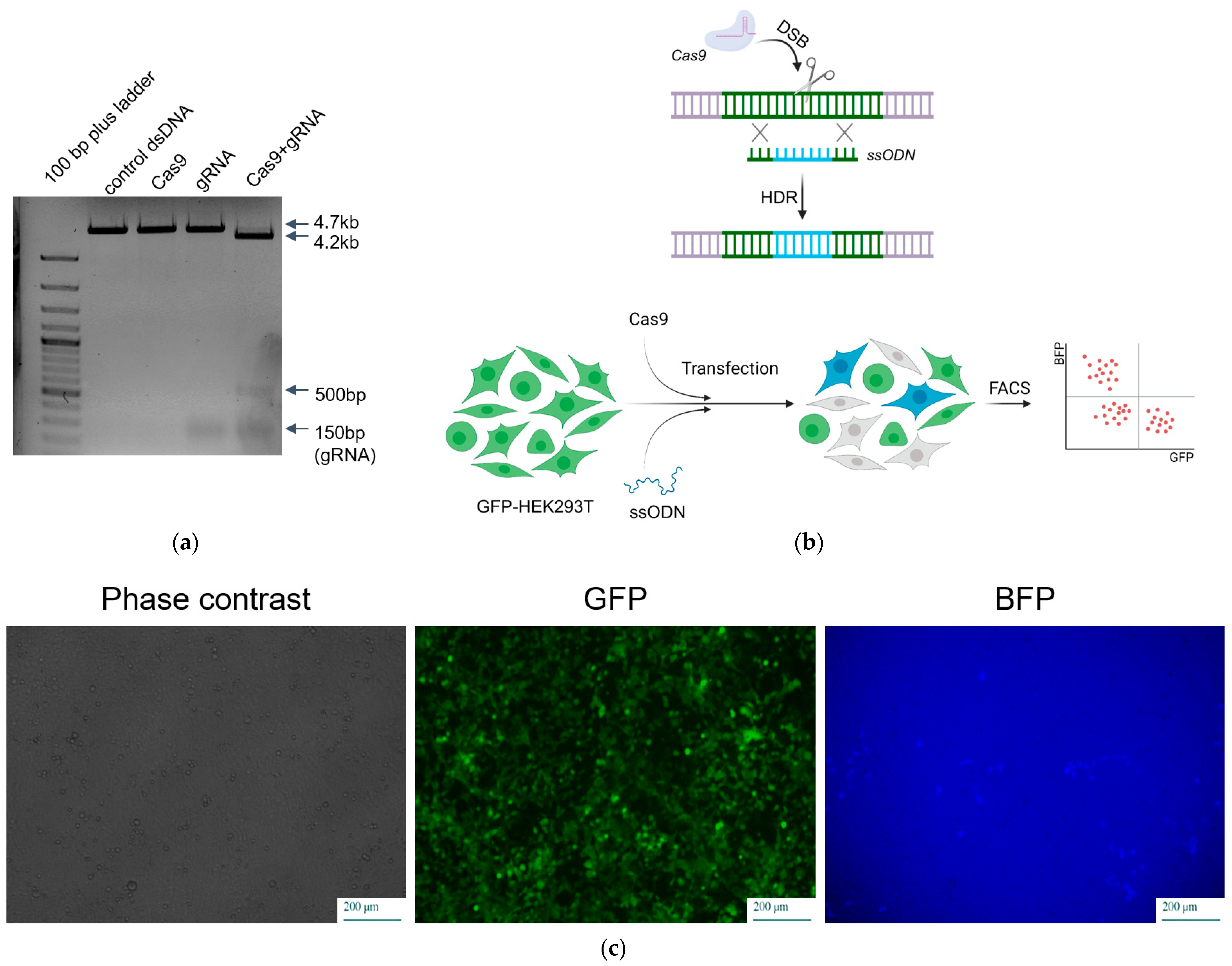
2.3. RNP Assembly and Evaluation of Its Activity In Vitro
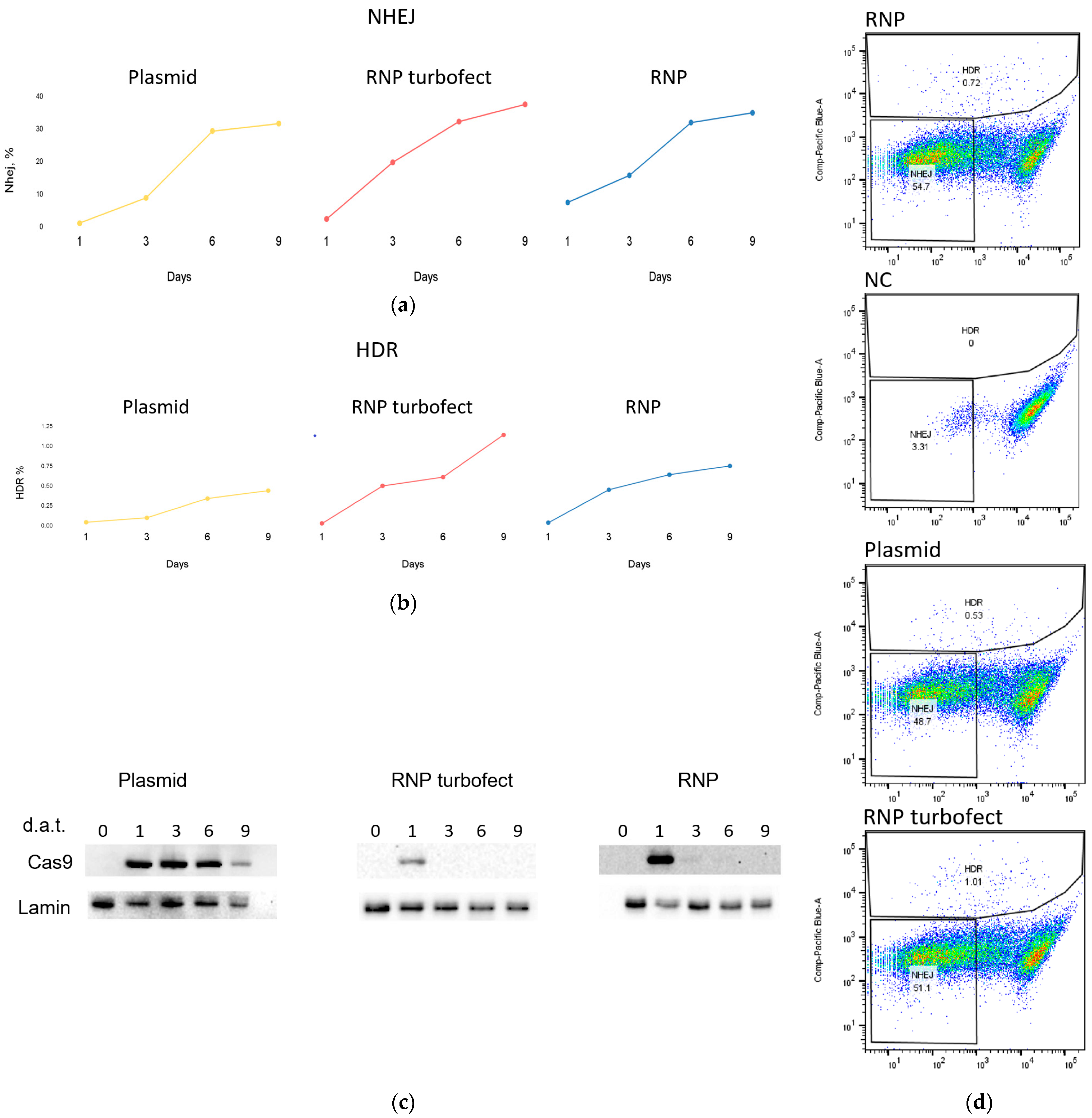
2.4. Comparison of Delivery Methods of SpCas9 into Mammalian Cells
3. Discussion
4. Materials and Methods
4.1. Plasmids Construction
4.2. Expression and Purification of SpCas9 Protein
4.3. gRNA Preparation
4.4. RNP Complexes Preparation
4.5. Mammalian Cell Line and Transfection
4.6. Flow Cytometry
4.7. Western Blot Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gupta, D.; Bhattacharjee, O.; Mandal, D.; Sen, M.K.; Dey, D.; Dasgupta, A.; Kazi, T.A.; Gupta, R.; Sinharoy, S.; Acharya, K.; et al. CRISPR-Cas9 System: A New-Fangled Dawn in Gene Editing. Life Sci. 2019, 232, 116636. [Google Scholar] [CrossRef] [PubMed]
- Le Rhun, A.; Escalera-Maurer, A.; Bratovič, M.; Charpentier, E. CRISPR-Cas in Streptococcus Pyogenes. RNA Biol. 2019, 16, 380–389. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Li, J. Review, Analysis, and Optimization of the CRISPR Streptococcus Pyogenes Cas9 System. Med. Drug Discov. 2021, 9, 100080. [Google Scholar] [CrossRef]
- Gillmore, J.D.; Gane, E.; Taubel, J.; Kao, J.; Fontana, M.; Maitland, M.L.; Seitzer, J.; O’Connell, D.; Walsh, K.R.; Wood, K.; et al. CRISPR-Cas9 In Vivo Gene Editing for Transthyretin Amyloidosis. N. Engl. J. Med. 2021, 385, 493–502. [Google Scholar] [CrossRef]
- Zhen, S.; Hua, L.; Takahashi, Y.; Narita, S.; Liu, Y.-H.; Li, Y. In Vitro and in Vivo Growth Suppression of Human Papillomavirus 16-Positive Cervical Cancer Cells by CRISPR/Cas9. Biochem. Biophys. Res. Commun. 2014, 450, 1422–1426. [Google Scholar] [CrossRef]
- Goyal, S.; Tisdale, J.; Schmidt, M.; Kanter, J.; Jaroscak, J.; Whitney, D.; Bitter, H.; Gregory, P.D.; Parsons, G.; Foos, M.; et al. Acute Myeloid Leukemia Case after Gene Therapy for Sickle Cell Disease. N. Engl. J. Med. 2022, 386, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Alayoubi, A.M.; Khawaji, Z.Y.; Mohammed, M.A.; Mercier, F.E. CRISPR-Cas9 System: A Novel and Promising Era of Genotherapy for Beta-Hemoglobinopathies, Hematological Malignancy, and Hemophilia. Ann. Hematol. 2023; Online ahead of print. [Google Scholar] [CrossRef] [PubMed]
- Chu, V.T.; Weber, T.; Wefers, B.; Wurst, W.; Sander, S.; Rajewsky, K.; Kühn, R. Increasing the Efficiency of Homology-Directed Repair for CRISPR-Cas9-Induced Precise Gene Editing in Mammalian Cells. Nat. Biotechnol. 2015, 33, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Doench, J.G.; Fusi, N.; Sullender, M.; Hegde, M.; Vaimberg, E.W.; Donovan, K.F.; Smith, I.; Tothova, Z.; Wilen, C.; Orchard, R.; et al. Optimized sgRNA Design to Maximize Activity and Minimize Off-Target Effects of CRISPR-Cas9. Nat. Biotechnol. 2016, 34, 184–191. [Google Scholar] [CrossRef] [PubMed]
- Kulcsár, P.I.; Tálas, A.; Tóth, E.; Nyeste, A.; Ligeti, Z.; Welker, Z.; Welker, E. Blackjack Mutations Improve the On-Target Activities of Increased Fidelity Variants of SpCas9 with 5′G-Extended sgRNAs. Nat. Commun. 2020, 11, 1223. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Yuan, H.; Du, W.; Li, G.; Xue, D.; Huang, Q. Active-Site Models of Streptococcus Pyogenes Cas9 in DNA Cleavage State. Front. Mol. Biosci. 2021, 8, 653262. [Google Scholar] [CrossRef] [PubMed]
- Rostain, W.; Grebert, T.; Vyhovskyi, D.; Pizarro, P.T.; Tshinsele-Van Bellingen, G.; Cui, L.; Bikard, D. Cas9 Off-Target Binding to the Promoter of Bacterial Genes Leads to Silencing and Toxicity. Nucleic Acids Res. 2023, 51, 3485–3496. [Google Scholar] [CrossRef] [PubMed]
- Rajagopalan, N.; Kagale, S.; Bhowmik, P.; Song, H. A Two-Step Method for Obtaining Highly Pure Cas9 Nuclease for Genome Editing, Biophysical, and Structural Studies. Methods Protoc. 2018, 1, 17. [Google Scholar] [CrossRef] [PubMed]
- Teng, A.C.T.; Tavassoli, M.; Shrestha, S.; Marks, R.M.; McFadden, M.J.; Evagelou, S.L.; Lindsay, K.; Vandenbelt, A.; Li, W.; Ivakine, E.; et al. An Efficient and Cost-Effective Purification Protocol for Staphylococcus Aureus Cas9 Nuclease. STAR Protoc. 2023, 4, 101933. [Google Scholar] [CrossRef]
- Flottmann, F.; Pohl, G.M.; Gummert, J.; Milting, H.; Brodehl, A. A Detailed Protocol for Expression, Purification, and Activity Determination of Recombinant SaCas9. STAR Protoc. 2022, 3, 101276. [Google Scholar] [CrossRef] [PubMed]
- Medeiros, L.C.S.; South, L.; Peng, D.; Bustamante, J.M.; Wang, W.; Bunkofske, M.; Perumal, N.; Sanchez-Valdez, F.; Tarleton, R.L. Rapid, Selection-Free, High-Efficiency Genome Editing in Protozoan Parasites Using CRISPR-Cas9 Ribonucleoproteins. mBio 2017, 8, e01788-17. [Google Scholar] [CrossRef] [PubMed]
- Fleitas, A.L.; Señorale, M.; Vidal, S. A Robust Expression and Purification Method for Production of SpCas9-GFP-MBP Fusion Protein for In Vitro Applications. Methods Protoc. 2022, 5, 44. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Zhang, F.; Ding, Y. CRISPR/Cas9 Delivery System Engineering for Genome Editing in Therapeutic Applications. Pharmaceutics 2021, 13, 1649. [Google Scholar] [CrossRef]
- Yip, B.H. Recent Advances in CRISPR/Cas9 Delivery Strategies. Biomolecules 2020, 10, 839. [Google Scholar] [CrossRef]
- Fajrial, A.K.; He, Q.Q.; Wirusanti, N.I.; Slansky, J.E.; Ding, X. A Review of Emerging Physical Transfection Methods for CRISPR/Cas9-Mediated Gene Editing. Theranostics 2020, 10, 5532–5549. [Google Scholar] [CrossRef]
- Cradick, T.J.; Fine, E.J.; Antico, C.J.; Bao, G. CRISPR/Cas9 Systems Targeting β-Globin and CCR5 Genes Have Substantial off-Target Activity. Nucleic Acids Res. 2013, 41, 9584–9592. [Google Scholar] [CrossRef]
- Fu, Y.; Foden, J.A.; Khayter, C.; Maeder, M.L.; Reyon, D.; Joung, J.K.; Sander, J.D. High-Frequency off-Target Mutagenesis Induced by CRISPR-Cas Nucleases in Human Cells. Nat. Biotechnol. 2013, 31, 822–826. [Google Scholar] [CrossRef]
- Pattanayak, V.; Lin, S.; Guilinger, J.P.; Ma, E.; Doudna, J.A.; Liu, D.R. High-Throughput Profiling of off-Target DNA Cleavage Reveals RNA-Programmed Cas9 Nuclease Specificity. Nat. Biotechnol. 2013, 31, 839–843. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Kim, D.; Cho, S.W.; Kim, J.; Kim, J.-S. Highly Efficient RNA-Guided Genome Editing in Human Cells via Delivery of Purified Cas9 Ribonucleoproteins. Genome Res. 2014, 24, 1012–1019. [Google Scholar] [CrossRef] [PubMed]
- Asmamaw Mengstie, M. Viral Vectors for the in Vivo Delivery of CRISPR Components: Advances and Challenges. Front. Bioeng. Biotechnol. 2022, 10, 895713. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.B.; Zhang, S.; Kos, P.; Xiong, H.; Zhou, K.; Perelman, S.S.; Zhu, H.; Siegwart, D.J. Non-Viral CRISPR/Cas Gene Editing In Vitro and In Vivo Enabled by Synthetic Nanoparticle Co-Delivery of Cas9 mRNA and sgRNA. Angew. Chem. 2017, 129, 1079–1083. [Google Scholar] [CrossRef]
- Leonhardt, C.; Schwake, G.; Stögbauer, T.R.; Rappl, S.; Kuhr, J.-T.; Ligon, T.S.; Rädler, J.O. Single-Cell mRNA Transfection Studies: Delivery, Kinetics and Statistics by Numbers. Nanomed. Nanotechnol. Biol. Med. 2014, 10, 679–688. [Google Scholar] [CrossRef]
- Kim, T.K.; Eberwine, J.H. Mammalian Cell Transfection: The Present and the Future. Anal. Bioanal. Chem. 2010, 397, 3173–3178. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.H.; Lee, J.; Choi, S.A.; Kim, Y.-S.; Koo, O.; Choi, S.H.; Ahn, W.S.; Jie, E.Y.; Kim, S.W. Efficient Genome Editing Using CRISPR–Cas9 RNP Delivery into Cabbage Protoplasts via Electro-Transfection. Plant Biotechnol. Rep. 2020, 14, 695–702. [Google Scholar] [CrossRef]
- Seki, A.; Rutz, S. Optimized RNP Transfection for Highly Efficient CRISPR/Cas9-Mediated Gene Knockout in Primary T Cells. J. Exp. Med. 2018, 215, 985–997. [Google Scholar] [CrossRef]
- Zhang, S.; Shen, J.; Li, D.; Cheng, Y. Strategies in the Delivery of Cas9 Ribonucleoprotein for CRISPR/Cas9 Genome Editing. Theranostics 2021, 11, 614–648. [Google Scholar] [CrossRef] [PubMed]
- Tyumentseva, M.A.; Tyumentsev, A.I.; Akimkin, V.G. Protocol for Assessment of the Efficiency of CRISPR/Cas RNP Delivery to Different Types of Target Cells. PLoS ONE 2021, 16, e0259812. [Google Scholar] [CrossRef] [PubMed]
- Marblestone, J.G.; Edavettal, S.C.; Lim, Y.; Lim, P.; Zuo, X.; Butt, T.R. Comparison of SUMO Fusion Technology with Traditional Gene Fusion Systems: Enhanced Expression and Solubility with SUMO. Protein Sci. 2006, 15, 182–189. [Google Scholar] [CrossRef] [PubMed]
- Chen, R. Bacterial Expression Systems for Recombinant Protein Production: E. Coli and Beyond. Biotechnol. Adv. 2012, 30, 1102–1107. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.-J.; Lin, H.; Yang, X. Industrial Production of Recombinant Therapeutics in Escherichia coli and Its Recent Advancements. J. Ind. Microbiol. Biotechnol. 2012, 39, 383–399. [Google Scholar] [CrossRef] [PubMed]
- Schlegel, S.; Genevaux, P.; de Gier, J.-W. Isolating Escherichia coli Strains for Recombinant Protein Production. Cell. Mol. Life Sci. 2017, 74, 891–908. [Google Scholar] [CrossRef] [PubMed]
- Butt, T.R.; Edavettal, S.C.; Hall, J.P.; Mattern, M.R. SUMO Fusion Technology for Difficult-to-Express Proteins. Protein Expr. Purif. 2005, 43, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Carmignotto, G.P.; Azzoni, A.R. On the Expression of Recombinant Cas9 Protein in E. Coli BL21(DE3) and BL21(DE3) Rosetta Strains. J. Biotechnol. 2019, 306, 62–70. [Google Scholar] [CrossRef]
- Glaser, A.; McColl, B.; Vadolas, J. GFP to BFP Conversion: A Versatile Assay for the Quantification of CRISPR/Cas9-Mediated Genome Editing. Mol. Ther.-Nucleic Acids 2016, 5, e334. [Google Scholar] [CrossRef]
- Bravo, J.P.K.; Liu, M.-S.; Hibshman, G.N.; Dangerfield, T.L.; Jung, K.; McCool, R.S.; Johnson, K.A.; Taylor, D.W. Structural Basis for Mismatch Surveillance by CRISPR–Cas9. Nature 2022, 603, 343–347. [Google Scholar] [CrossRef]
- Pacesa, M.; Lin, C.-H.; Cléry, A.; Saha, A.; Arantes, P.R.; Bargsten, K.; Irby, M.J.; Allain, F.H.-T.; Palermo, G.; Cameron, P.; et al. Structural Basis for Cas9 Off-Target Activity. Cell 2022, 185, 4067–4081.e21. [Google Scholar] [CrossRef]
- Zhao, J.; Fang, H.; Zhang, D. Expanding Application of CRISPR-Cas9 System in Microorganisms. Synth. Syst. Biotechnol. 2020, 5, 269–276. [Google Scholar] [CrossRef]
- Kane, J.F. Effects of Rare Codon Clusters on High-Level Expression of Heterologous Proteins in Escherichia Coli. Curr. Opin. Biotechnol. 1995, 6, 494–500. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.-W.; Nguyen, V.Q.; Lin, S. Preparation of Cas9 Ribonucleoproteins for Genome Editing. Bio Protoc. 2022, 12, e4420. [Google Scholar] [CrossRef] [PubMed]
- Chaverra-Rodriguez, D.; Macias, V.M.; Hughes, G.L.; Pujhari, S.; Suzuki, Y.; Peterson, D.R.; Kim, D.; McKeand, S.; Rasgon, J.L. Targeted Delivery of CRISPR-Cas9 Ribonucleoprotein into Arthropod Ovaries for Heritable Germline Gene Editing. Nat. Commun. 2018, 9, 3008. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Lee, B.; Lee, A.Y.-F.; Modzelewski, A.J.; He, L. Highly Efficient Mouse Genome Editing by CRISPR Ribonucleoprotein Electroporation of Zygotes. J. Biol. Chem. 2016, 291, 14457–14467. [Google Scholar] [CrossRef] [PubMed]
- Ramakrishna, S.; Kwaku Dad, A.-B.; Beloor, J.; Gopalappa, R.; Lee, S.-K.; Kim, H. Gene Disruption by Cell-Penetrating Peptide-Mediated Delivery of Cas9 Protein and Guide RNA. Genome Res. 2014, 24, 1020–1027. [Google Scholar] [CrossRef] [PubMed]
- Suresh, B.; Ramakrishna, S.; Kim, H. Cell-Penetrating Peptide-Mediated Delivery of Cas9 Protein and Guide RNA for Genome Editing. Methods Mol. Biol. 2017, 1507, 81–94. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Ji, W.; Hall, J.M.; Hu, Q.; Wang, C.; Beisel, C.L.; Gu, Z. Self-Assembled DNA Nanoclews for the Efficient Delivery of CRISPR-Cas9 for Genome Editing. Angew. Chem. Int. Ed. Engl. 2015, 54, 12029–12033. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Conboy, M.; Park, H.M.; Jiang, F.; Kim, H.J.; Dewitt, M.A.; Mackley, V.A.; Chang, K.; Rao, A.; Skinner, C.; et al. Nanoparticle Delivery of Cas9 Ribonucleoprotein and Donor DNA in Vivo Induces Homology-Directed DNA Repair. Nat. Biomed. Eng. 2017, 1, 889–901. [Google Scholar] [CrossRef] [PubMed]
- D’Astolfo, D.S.; Pagliero, R.J.; Pras, A.; Karthaus, W.R.; Clevers, H.; Prasad, V.; Lebbink, R.J.; Rehmann, H.; Geijsen, N. Efficient Intracellular Delivery of Native Proteins. Cell 2015, 161, 674–690. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Liang, X.; Xie, H.; Kumar, S.; Ravinder, N.; Potter, J.; de Mollerat du Jeu, X.; Chesnut, J.D. Improved Delivery of Cas9 Protein/gRNA Complexes Using Lipofectamine CRISPRMAX. Biotechnol. Lett. 2016, 38, 919–929. [Google Scholar] [CrossRef] [PubMed]
- Bruce, V.J.; McNaughton, B.R. Inside Job: Methods for Delivering Proteins to the Interior of Mammalian Cells. Cell Chem. Biol. 2017, 24, 924–934. [Google Scholar] [CrossRef] [PubMed]
| Stain | Source | Features |
|---|---|---|
| Rosetta | #70954 Novagen, Darmstadt, Germany | Expression of eukaryotic proteins containing codons rarely used in E. coli (AGG, AGA, AUA, CUA, CCC, GGA) is enhanced. |
| Origami B | #70836 Novagen, Darmstadt, Germany | Efficiently expresses and folds disulfide bond-containing proteins. |
| BL21 | #200133 Stratagene, La Jolla, CA, USA | The regular BL21 (DE3) strain contains the lambda DE3 prophage that carries the gene for T7 RNA polymerase under control of a lacUV5 promoter, allowing expression of the T7 RNA polymerase to be induced with IPTG. |
| BL21-pUBS | Lab collection | Plasmid pUBS-520 was introduced, supplying recombinant bacteria with high levels of rare tRNA-Arg. |
| C41 | #CMC0017 Sigma-Aldrich, Burlington, MA, USA | Expression of toxic proteins. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Evmenov, K.; Pustogarov, N.; Panteleev, D.; Safin, A.; Alkalaeva, E. An Efficient Expression and Purification Protocol for SpCas9 Nuclease and Evaluation of Different Delivery Methods of Ribonucleoprotein. Int. J. Mol. Sci. 2024, 25, 1622. https://doi.org/10.3390/ijms25031622
Evmenov K, Pustogarov N, Panteleev D, Safin A, Alkalaeva E. An Efficient Expression and Purification Protocol for SpCas9 Nuclease and Evaluation of Different Delivery Methods of Ribonucleoprotein. International Journal of Molecular Sciences. 2024; 25(3):1622. https://doi.org/10.3390/ijms25031622
Chicago/Turabian StyleEvmenov, Konstantin, Nikolay Pustogarov, Dmitri Panteleev, Artur Safin, and Elena Alkalaeva. 2024. "An Efficient Expression and Purification Protocol for SpCas9 Nuclease and Evaluation of Different Delivery Methods of Ribonucleoprotein" International Journal of Molecular Sciences 25, no. 3: 1622. https://doi.org/10.3390/ijms25031622
APA StyleEvmenov, K., Pustogarov, N., Panteleev, D., Safin, A., & Alkalaeva, E. (2024). An Efficient Expression and Purification Protocol for SpCas9 Nuclease and Evaluation of Different Delivery Methods of Ribonucleoprotein. International Journal of Molecular Sciences, 25(3), 1622. https://doi.org/10.3390/ijms25031622






