Six-Gene Signature for Differential Diagnosis and Therapeutic Decisions in Non-Small-Cell Lung Cancer—A Validation Study
Abstract
1. Introduction
2. Results
2.1. Patients’ Characteristics
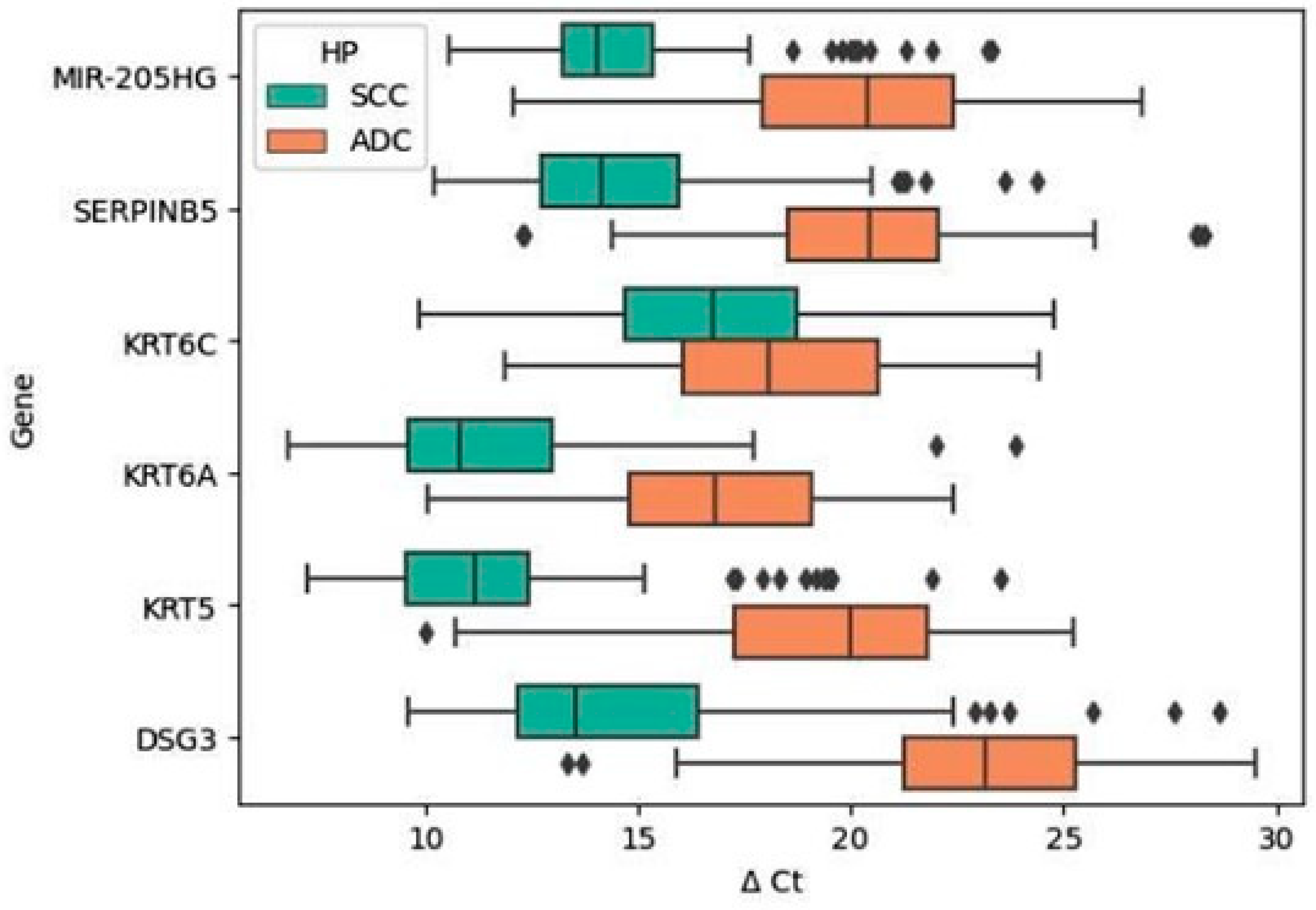
2.2. Differences in the Expression Profiles of Validated Genes between the Histological Subtypes of NSCLC (SCC vs. ADC)
2.3. Relationships between Expression Levels of Validated Genes and Clinicopathological Characteristics of NSCLC Patients
2.4. Visualizing Decision Boundary for Logistic Regression Models of Individual Validated Genes
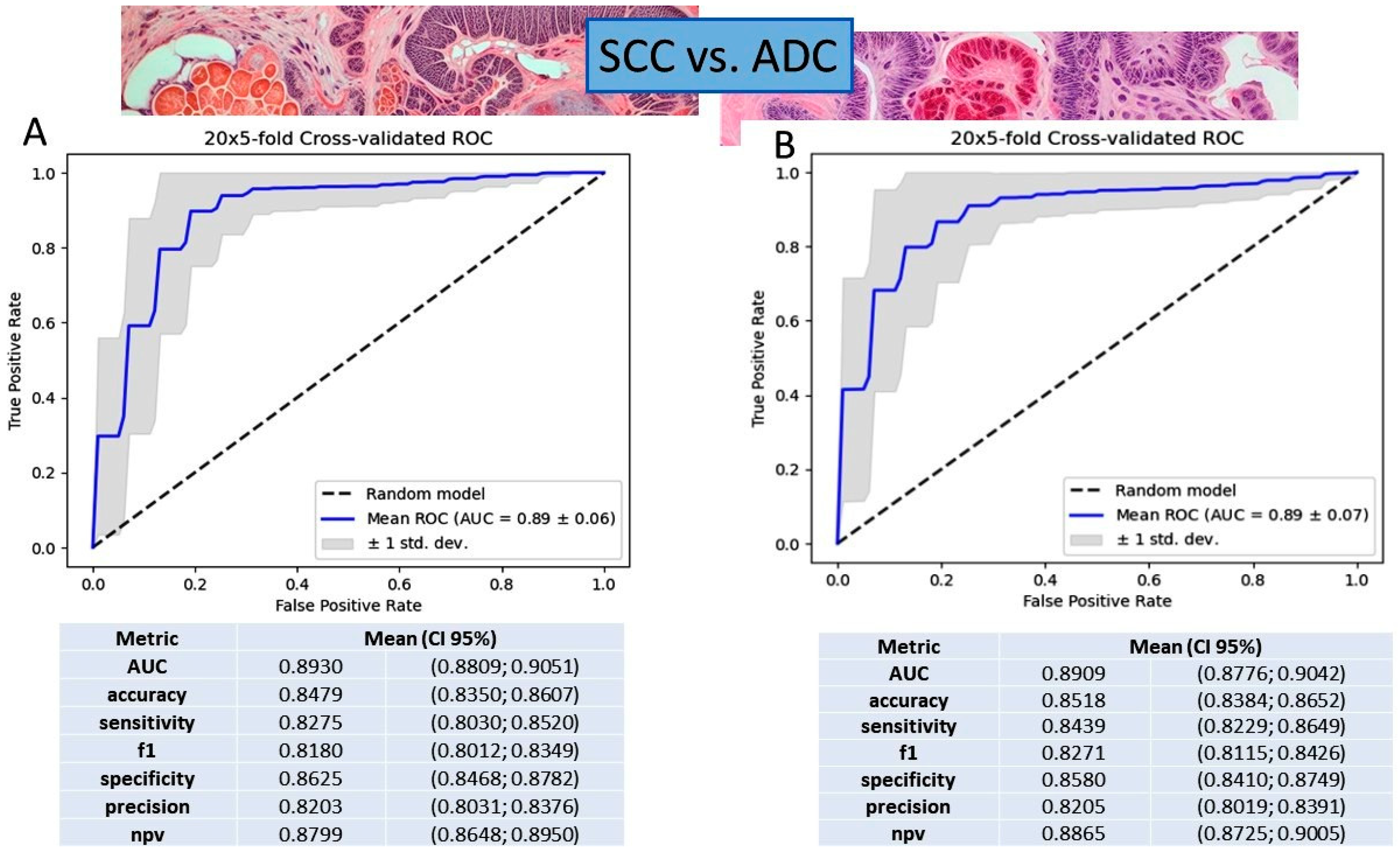
2.5. Gene Expression Value in NSCLC Subtyping Based on Logistic Regression and Gradient Boosting Decision Tree Models
2.6. Prognostic Significance of Validated Gene Expression in NSCLC Patients
2.7. Gene Ontology Analyses
3. Discussion
4. Materials and Methods
4.1. Patients and Samples
4.2. Histopathological Diagnosis
4.3. RNA Extraction and Quality Control
4.4. Quantitative Real-Time PCR Analysis
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sulewska, A.; Pilz, L.; Manegold, C.; Ramlau, R.; Charkiewicz, R.; Niklinski, J. A Systematic Review of Progress toward Unlocking the Power of Epigenetics in NSCLC: Latest Updates and Perspectives. Cells 2023, 12, 905. [Google Scholar] [CrossRef]
- Charkiewicz, R.; Pilz, L.; Sulewska, A.; Kozlowski, M.; Niklinska, W.; Moniuszko, M.; Reszec, J.; Manegold, C.; Niklinski, J. Validation for histology-driven diagnosis in non-small cell lung cancer using hsa-miR-205 and hsa-miR-21 expression by two different normalization strategies. Int. J. Cancer 2016, 138, 689–697. [Google Scholar] [CrossRef]
- Tane, S.; Nishio, W.; Ogawa, H.; Hokka, D.; Tane, K.; Tanaka, Y.; Tauchi, S.; Uchino, K.; Sakai, Y.; Ohbayashi, C.; et al. Clinical significance of the “not otherwise specified” subtype in candidates for resectable non-small cell lung cancer. Oncol. Lett. 2014, 8, 1017–1024. [Google Scholar] [CrossRef]
- Al-Farsi, A.; Ellis, P.M. Treatment paradigms for patients with metastatic non-small cell lung cancer, squamous lung cancer: First, second, and third-line. Front. Oncol. 2014, 4, 157. [Google Scholar] [CrossRef]
- Uramoto, H.; Tanaka, F. Recurrence after surgery in patients with NSCLC. Transl. Lung Cancer Res. 2014, 3, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Molina, J.R.; Yang, P.; Cassivi, S.D.; Schild, S.E.; Adjei, A.A. Non-small cell lung cancer: Epidemiology, risk factors, treatment, and survivorship. Mayo Clin. Proc. 2008, 83, 584–594. [Google Scholar] [CrossRef]
- Herbst, R.S.; Heymach, J.V.; Lippman, S.M. Lung cancer. N. Engl. J. Med. 2008, 359, 1367–1380. [Google Scholar] [CrossRef]
- Shroff, G.S.; de Groot, P.M.; Papadimitrakopoulou, V.A.; Truong, M.T.; Carter, B.W. Targeted Therapy and Immunotherapy in the Treatment of Non-Small Cell Lung Cancer. Radiol. Clin. N. Am. 2018, 56, 485–495. [Google Scholar] [CrossRef]
- Pal, S.K.; Figlin, R.A.; Reckamp, K. Targeted therapies for non-small cell lung cancer: An evolving landscape. Mol. Cancer Ther. 2010, 9, 1931–1944. [Google Scholar] [CrossRef] [PubMed]
- Lauro, S.; Onesti, C.E.; Righini, R.; Marchetti, P. The use of bevacizumab in non-small cell lung cancer: An update. Anticancer Res. 2014, 34, 1537–1545. [Google Scholar] [PubMed]
- Chen, Z.; Akbay, E.; Mikse, O.; Tupper, T.; Cheng, K.; Wang, Y.; Tan, X.; Altabef, A.; Woo, S.-A.; Chen, L.; et al. Co-clinical trials demonstrate superiority of crizotinib to chemotherapy in ALK-rearranged non-small cell lung cancer and predict strategies to overcome resistance. Clin. Cancer Res. 2014, 20, 1204–1211. [Google Scholar] [CrossRef]
- Thakur, M.K.; Wozniak, A.J. Spotlight on necitumumab in the treatment of non-small-cell lung carcinoma. Lung Cancer 2017, 8, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.-L.; Herbst, R.S.; Mann, H.; Rukazenkov, Y.; Marotti, M.; Tsuboi, M. ADAURA: Phase III, Double-blind, Randomized Study of Osimertinib Versus Placebo in EGFR Mutation-positive Early-stage NSCLC After Complete Surgical Resection. Clin. Lung Cancer 2018, 19, E533–E536. [Google Scholar] [CrossRef] [PubMed]
- Charkiewicz, R.; Sulewska, A.; Charkiewicz, A.; Gyenesei, A.; Galik, B.; Ramlau, R.; Piwkowski, C.; Stec, R.; Biecek, P.; Karabowicz, P.; et al. miRNA-Seq Tissue Diagnostic Signature: A Novel Model for NSCLC Subtyping. Int. J. Mol. Sci. 2023, 24, 13318. [Google Scholar] [CrossRef]
- Sulewska, A.; Niklinski, J.; Charkiewicz, R.; Karabowicz, P.; Biecek, P.; Baniecki, H.; Kowalczuk, O.; Kozlowski, M.; Modzelewska, P.; Majewski, P.; et al. A Signature of 14 Long Non-Coding RNAs (lncRNAs) as a Step towards Precision Diagnosis for NSCLC. Cancers 2022, 14, 439. [Google Scholar] [CrossRef] [PubMed]
- Charkiewicz, R.; Sulewska, A.; Mroz, R.; Charkiewicz, A.; Naumnik, W.; Kraska, M.; Gyenesei, A.; Galik, B.; Junttila, S.; Miskiewicz, B.; et al. Serum Insights: Leveraging the Power of miRNA Profiling as an Early Diagnostic Tool for Non-Small Cell Lung Cancer. Cancers 2023, 15, 4910. [Google Scholar] [CrossRef] [PubMed]
- Charkiewicz, R.; Niklinski, J.; Claesen, J.; Sulewska, A.; Kozlowski, M.; Michalska-Falkowska, A.; Reszec, J.; Moniuszko, M.; Naumnik, W.; Niklinska, W. Gene Expression Signature Differentiates Histology but Not Progression Status of Early-Stage NSCLC. Transl. Oncol. 2017, 10, 450–458. [Google Scholar] [CrossRef] [PubMed]
- Tibshirani, R.; Hastie, T.; Narasimhan, B.; Chu, G. Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc. Natl. Acad. Sci. USA 2002, 99, 6567–6572. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Yang, W.; Zhang, Y.; Yang, J.Y.; Guan, R.; Xu, D.; Yang, M.Q. Genomic analyses based on pulmonary adenocarcinoma in situ reveal early lung cancer signature. BMC Med. Genom. 2018, 11, 106. [Google Scholar] [CrossRef]
- Niemira, M.; Collin, F.; Szalkowska, A.; Bielska, A.; Chwialkowska, K.; Reszec, J.; Niklinski, J.; Kwasniewski, M.; Kretowski, A. Molecular Signature of Subtypes of Non-Small-Cell Lung Cancer by Large-Scale Transcriptional Profiling: Identification of Key Modules and Genes by Weighted Gene Co-Expression Network Analysis (WGCNA). Cancers 2020, 12, 37. [Google Scholar] [CrossRef]
- Song, J.-Y.; Li, X.-P.; Qin, X.-J.; Zhang, J.-D.; Zhao, J.-Y.; Wange, R. A fourteen-lncRNA risk score system for prognostic prediction of patients with non-small cell lung cancer. Cancer Biomark. 2020, 29, 493–508. [Google Scholar] [CrossRef]
- Girard, L.; Rodriguez-Canales, J.; Behrens, C.; Thompson, D.M.; Botros, I.W.; Tang, H.; Xie, Y.; Rekhtman, N.; Travis, W.D.; Wistuba, I.I.; et al. An Expression Signature as an Aid to the Histologic Classification of Non-Small Cell Lung Cancer. Clin. Cancer Res. 2016, 22, 4880–4889. [Google Scholar] [CrossRef]
- Hou, J.; Aerts, J.; den Hamer, B.; van Ijcken, W.; den Bakker, M.; Riegman, P.; Van Der Leest, C.; Van Der Spek, P.; Foekens, J.A.; Hoogsteden, H.C.; et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLoS ONE 2010, 5, e10312. [Google Scholar] [CrossRef] [PubMed]
- Wilkerson, M.D.; Schallheim, J.M.; Hayes, D.N.; Roberts, P.J.; Bastien, R.R.L.; Mullins, M.; Yin, X.; Miller, C.R.; Thorne, L.B.; Geiersbach, K.B.; et al. Prediction of lung cancer histological types by RT-qPCR gene expression in FFPE specimens. J. Mol. Diagn. 2013, 15, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Chen, H.; Shan, Z.; Yang, L. Identification of differentially expressed genes between lung adenocarcinoma and lung squamous cell carcinoma by gene expression profiling. Mol. Med. Rep. 2016, 14, 1483–1490. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Chen, X.; Lu, D.; Diao, D.; Liu, X.; Mai, S.; Feng, S.; Xiong, G. LncRNA miR205HG hinders HNRNPA0 translation: Anti-oncogenic effects in esophageal carcinoma. Mol. Oncol. 2021, 16, 795–812. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Liang, F.; Li, Q.; Sun, H.; Li, F.; Jiao, Z.; Lei, J. LncRNA MIR205HG accelerates cell proliferation, migration and invasion in hepatoblastoma through the activation of MAPK signaling pathway and PI3K/AKT signaling pathway. Biol. Direct 2022, 17, 2. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Zhang, Y.; Zheng, L. Analysis of differentially expressed long non-coding RNAs revealed a pro-tumor role of MIR205HG in cervical cancer. Mol. Med. Rep. 2022, 25, 42. [Google Scholar] [CrossRef] [PubMed]
- Di Agostino, S.; Valenti, F.; Sacconi, A.; Fontemaggi, G.; Pallocca, M.; Pulito, C.; Ganci, F.; Muti, P.; Strano, S.; Blandino, G. Long Non-coding MIR205HG Depletes Hsa-miR-590-3p Leading to Unrestrained Proliferation in Head and Neck Squamous Cell Carcinoma. Theranostics 2018, 8, 1850–1868. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Yuan, C.; He, X.; Wang, M.; Zhang, H.; Cheng, J.; Wang, H. Identification and in vitro validation of diagnostic and prognostic biomarkers for lung squamous cell carcinoma. J. Thorac. Dis. 2022, 14, 1243–1255. [Google Scholar] [CrossRef]
- Liu, L.; Li, Y.; Zhang, R.; Li, C.; Xiong, J.; Wei, Y. MIR205HG acts as a ceRNA to expedite cell proliferation and progression in lung squamous cell carcinoma via targeting miR-299-3p/MAP3K2 axis. BMC Pulm. Med. 2020, 20, 163. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Xue, X.; Li, C.; Zhao, W.; Ma, Y.; Xu, F.; Wu, Z.; Dai, Y.; Li, Y.; Liu, Y.; et al. MIR205HG facilitates carcinogenesis of lung squamous cell carcinoma in vitro revealed by long noncoding RNA profiling. Acta Biochim. Biophys. Sin. 2020, 52, 371–381. [Google Scholar] [CrossRef] [PubMed]
- Larzabal, L.; de Aberasturi, A.L.; Redrado, M.; Rueda, P.; Rodriguez, M.J.; Bodegas, M.E.; Montuenga, L.M.; Calvo, A. TMPRSS4 regulates levels of integrin α5 in NSCLC through miR-205 activity to promote metastasis. Br. J. Cancer 2014, 110, 764–774. [Google Scholar] [CrossRef] [PubMed]
- Che, D.; Wang, M.; Sun, J.; Li, B.; Xu, T.; Lu, Y.; Pan, H.; Lu, Z.; Gu, X. KRT6A Promotes Lung Cancer Cell Growth and Invasion through MYC-Regulated Pentose Phosphate Pathway. Front. Cell Dev. Biol. 2021, 9, 2021. [Google Scholar] [CrossRef]
- Xiao, J.; Kuang, X.; Dai, L.; Zhang, L.; He, B. Anti-tumour effects of Keratin 6A in lung adenocarcinoma. Clin. Respir. J. 2020, 14, 667–674. [Google Scholar] [CrossRef]
- Yang, B.; Zhang, W.; Zhang, M.; Wang, X.; Peng, S.; Zhang, R. KRT6A Promotes EMT and Cancer Stem Cell Transformation in Lung Adenocarcinoma. Technol. Cancer Res. Treat. 2020, 19, 1533033820921248. [Google Scholar] [CrossRef]
- Chen, S.; Hong, K.; Zhou, L.; Ran, R.; Huang, J.; Zheng, Y.; Xing, M.; Cai, Y. Hsa_circRNA_0017620 regulated cell progression of non-small-cell lung cancer via miR-520a-5p/KRT5 axis. J. Clin. Lab. Anal. 2022, 36, e24347. [Google Scholar] [CrossRef]
- Xiao, K.; Wang, Y.; Zhou, L.; Wang, J.; Wang, Y.; Tong, D.; Zhu, Z.; Jiang, J. Construction of ceRNA network to identify the lncRNA and mRNA related to non-small cell lung cancer. PLoS ONE 2021, 16, e0259091. [Google Scholar] [CrossRef]
- Pan, B.; Wei, Z.-X.; Zhang, J.-X.; Li, X.; Meng, Q.-W.; Cao, Y.-Y.; Qi, L.-S.; Yu, Y. The value of AGR2 and KRT5 as an immunomarker combination in distinguishing lung squamous cell carcinoma from adenocarcinoma. Am. J. Transl. Res. 2021, 13, 4464–4476. [Google Scholar]
- Wu, X.; Wang, L.; Feng, F.; Tian, S. Weighted gene expression profiles identify diagnostic and prognostic genes for lung adenocarcinoma and squamous cell carcinoma. J. Int. Med. Res. 2020, 48, 300060519893837. [Google Scholar] [CrossRef]
- Hamaneh, M.; Yu, Y.-K. An 8-Gene Signature for Classifying Major Subtypes of Non-Small-Cell Lung Cancer. Cancer Inform. 2022, 21, 11769351221100718. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Pang, Z.; Li, G.; Gu, T. Bioinformatics analysis of differentially expressed miRNAs in non-small cell lung cancer. J. Clin. Lab. Anal. 2021, 35, e23588. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Hou, Z.; Liu, W.; Yu, Z.; Liang, Z.; Chen, S. circ-Keratin 6c Promotes Malignant Progression and Immune Evasion of Colorectal Cancer through microRNA-485-3p/Programmed Cell Death Receptor Ligand 1 Axis. J. Pharmacol. Exp. Ther. 2021, 377, 358–367. [Google Scholar] [CrossRef]
- Lei, K.-F.; Liu, B.-Y.; Wang, Y.-F.; Chen, X.-H.; Yu, B.-Q.; Guo, Y.; Zhu, Z.-G. SerpinB5 interacts with KHDRBS3 and FBXO32 in gastric cancer cells. Oncol. Rep. 2011, 26, 1115–1120. [Google Scholar] [CrossRef] [PubMed]
- Mardin, W.A.; Ntalos, D.; Mees, S.T.; Spieker, T.; Senninger, N.; Haier, J.; Dhayat, S.A. SERPINB5 Promoter Hypomethylation Differentiates Pancreatic Ductal Adenocarcinoma from Pancreatitis. Pancreas 2016, 45, 743–747. [Google Scholar] [CrossRef]
- Su, C.; Liu, W.-X.; Wu, L.-S.; Dong, T.-J.; Liu, J.-F. Screening of Hub Gene Targets for Lung Cancer via Microarray Data. Comb. Chem. High Throughput Screen. 2021, 24, 269–285. [Google Scholar] [CrossRef]
- Xiao, L.; Li, Q.; Huang, Y.; Fan, Z.; Qin, W.; Liu, B.; Yuan, X. Integrative Analysis Constructs an Extracellular Matrix-Associated Gene Signature for the Prediction of Survival and Tumor Immunity in Lung Adenocarcinoma. Front. Cell Dev. Biol. 2022, 10, 835043. [Google Scholar] [CrossRef]
- Abula, Y.; Su, Y.; Tuniyazi, D.; Yi, C. Desmoglein 3 contributes to tumorigenicity of pancreatic ductal adenocarcinoma through activating Src-FAK signaling. Anim. Cells Syst. 2021, 25, 195–202. [Google Scholar] [CrossRef]
- Dong, Y.; Li, S.; Sun, X.; Wang, Y.; Lu, T.; Wo, Y.; Leng, X.; Kong, D.; Jiao, W. Desmoglein 3 and Keratin 14 for Distinguishing Between Lung Adenocarcinoma and Lung Squamous Cell Carcinoma. Onco Targets Ther. 2020, 13, 11111–11124. [Google Scholar] [CrossRef]
- Sepesi, B.; Mehran, R.; Spicer, J.; Cascone, T. NEOSTAR trial and the current status of neoadjuvant therapy in non-small cell lung cancer. J. Thorac. Cardiovasc. Surg. 2023, in press. [Google Scholar] [CrossRef]
- Chen, L.N.; Wei, A.Z.; Shu, C.A. Neoadjuvant immunotherapy in resectable non-small-cell lung cancer. Ther. Adv. Med. Oncol. 2023, 15, 17588359231163798. [Google Scholar] [CrossRef]
- Provencio, M.; Nadal, E.; Insa, A.; García-Campelo, M.R.; Casal-Rubio, J.; Dómine, M.; Majem, M.; Rodríguez-Abreu, D.; Martínez-Martí, A.; Carpeño, J.D.C.; et al. Neoadjuvant chemotherapy and nivolumab in resectable non-small-cell lung cancer (NADIM): An open-label, multicentre, single-arm, phase 2 trial. Lancet Oncol. 2020, 21, 1413–1422. [Google Scholar] [CrossRef] [PubMed]
- Niklinski, J.; Kretowski, A.; Moniuszko, M.; Reszec, J.; Michalska-Falkowska, A.; Niemira, M.; Ciborowski, M.; Charkiewicz, R.; Jurgilewicz, D.; Kozlowski, M.; et al. Systematic biobanking, novel imaging techniques, and advanced molecular analysis for precise tumor diagnosis and therapy: The Polish MOBIT project. Adv. Med. Sci. 2017, 62, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Michalska-Falkowska, A.; Niklinski, J.; Juhl, H.; Sulewska, A.; Kisluk, J.; Charkiewicz, R.; Ciborowski, M.; Ramlau, R.; Gryczka, R.; Piwkowski, C.; et al. Applied Molecular-Based Quality Control of Biobanked Samples for Multi-Omics Approach. Cancers 2023, 15, 3742. [Google Scholar] [CrossRef] [PubMed]
- Travis, W.; Brambilla, E.; Muller-Hermelink, H.; Harris, C. Pathology and Genetics of Tumours of the Lung, Pleura, Thymus and Heart. In WHO Classification of Tumours; WHO Press: Geneva, Switzerland, 2004. [Google Scholar]
- Travis, W.D.; Brambilla, E.; Noguchi, M.; Nicholson, A.G.; Geisinger, K.R.; Yatabe, Y.; Beer, D.G.; Powell, C.A.; Riely, G.J.; Van Schil, P.E. International association for the study of lung cancer/american thoracic society/european respiratory society international multidisciplinary classification of lung adenocarcinoma. J. Thorac. Oncol. 2011, 6, 244–285. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, New York, NY, USA, 13–17 August 2016; pp. 785–794. [Google Scholar]


| Gender | N | % |
|---|---|---|
| Male | 107 | 76.43 |
| Female | 33 | 23.57 |
| Age at Diagnosis (years) | ||
| Range | 40–81 | |
| Mean | 63.33 | |
| Histological Type | ||
| Squamous cell carcinoma | 79 | 56.42 |
| Adenocarcinoma | 61 | 43.57 |
| TNM Stage | ||
| IA | 33 | 23.57 |
| IB | 32 | 22.86 |
| IIA | 28 | 20.0 |
| IIB | 38 | 27.14 |
| IIIA | 8 | 5.71 |
| Information not available | 1 | 0.7 |
| Recurrence | ||
| Present | 54 | 38.57 |
| Absent | 82 | 58.57 |
| Information not available | 4 | 2.86 |
| Genes ∆Ct SCC | Genes ∆Ct ADC | Expression Profile | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| N | Mean | SD | 25% | 75% | N | Mean | SD | 25% | 75% | p-Value | SCC vs. ADC | |
| MIR-205HG | 81 | 14.808 | 2.769 | 13.226 | 15.362 | 59 | 20.099 | 3.426 | 17.940 | 22.425 | <0.001 | up |
| SERPINB5 | 81 | 14.850 | 2.990 | 12.725 | 15.977 | 59 | 20.305 | 3.640 | 18.546 | 22.048 | <0.001 | up |
| KRT6C | 81 | 16.697 | 3.104 | 14.670 | 18.749 | 59 | 18.422 | 3.041 | 16.061 | 20.628 | 0.003381 | up |
| KRT6A | 81 | 11.574 | 3.145 | 9.580 | 13.001 | 59 | 16.578 | 2.858 | 14.772 | 19.071 | <0.001 | up |
| KRT5 | 81 | 11.968 | 3.492 | 9.543 | 12.405 | 59 | 19.383 | 3.369 | 17.277 | 21.823 | <0.001 | up |
| DSG | 81 | 14.864 | 4.012 | 12.172 | 16.386 | 59 | 23.050 | 3.559 | 21.261 | 25.255 | <0.001 | up |
| GO Category | GO Term Id | GO Term Label | p-Value | Genes Involved |
|---|---|---|---|---|
| Cellular component | GO:0001533 | cornified envelope | 8.86 × 10−5 | DSG3, SERPINB5 |
| GO:0045095 | keratin filament | 1.10 × 10−6 | KRT6A, KRT5, KRT6C | |
| GO:0005882 | intermediate filament | 1.35 × 10−5 | KRT6A | |
| GO:0045111 | intermediate filament cytoskeleton | 2.09 × 10−5 | KRT6A, KRT5, KRT6C | |
| GO:0005615 | extracellular space | 0.00011 | DSG3, KRT6A, KRT5, KRT6C, SERPINB5 | |
| Biological process | GO:0045109 | intermediate filament organization | 4.99 × 10−7 | KRT6A, KRT5, KRT6C |
| GO:0045104 | intermediate filament cytoskeleton organization | 1.04 × 10−6 | KRT6A, KRT5, KRT6C | |
| GO:0045103 | intermediate filament-based process | 1.07 × 10−6 | KRT6A, KRT5, KRT6C | |
| GO:0031424 | keratinization | 6.99 × 10−7 | KRT6A, KRT5, KRT6C | |
| GO:0030216 | keratinocyte differentiation | 3.67 × 10−6 | KRT6A, KRT5, KRT6C | |
| GO:0009913 | epidermal cell differentiation | 1.10 × 10−5 | KRT6A, KRT5, KRT6C | |
| Molecular function | GO:0030280 | structural constituent of skin epidermis | 6.77 × 10−8 | KRT6A, KRT5, KRT6C |
| RefSeq | Gene Name, Abbreviation | Assay IDs from Applied Biosystems (Primers + Probe) |
|---|---|---|
| NM_000424.3 | Keratin 5 (KRT5) | Hs00361185_m1 |
| NM_005554.3 | Keratin 6A (KRT6A) | Hs01699178_g1 |
| NM_002639.4 XM_006722483.2 | Serpin Family B Member 5 (SERPINB5) | Hs00985285_m1 |
| NM_001104548.1 XM_017002058.1 | MIR205 Host Gene (MIR205HG) | Hs03405498_m1 |
| NM_173086.4 | Keratin 6C (KRT6C) | Hs00752476_s1 |
| NM_001944.2 XM_011525850.1 | Desmoglein 3 (DSG3) | Hs00951897_m1 |
| X03205.1 | 18S rRNA (Reference Gene) | Hs99999901_s1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Charkiewicz, R.; Sulewska, A.; Karabowicz, P.; Lapuc, G.; Charkiewicz, A.; Kraska, M.; Pancewicz, J.; Lukasik, M.; Kozlowski, M.; Stec, R.; et al. Six-Gene Signature for Differential Diagnosis and Therapeutic Decisions in Non-Small-Cell Lung Cancer—A Validation Study. Int. J. Mol. Sci. 2024, 25, 3607. https://doi.org/10.3390/ijms25073607
Charkiewicz R, Sulewska A, Karabowicz P, Lapuc G, Charkiewicz A, Kraska M, Pancewicz J, Lukasik M, Kozlowski M, Stec R, et al. Six-Gene Signature for Differential Diagnosis and Therapeutic Decisions in Non-Small-Cell Lung Cancer—A Validation Study. International Journal of Molecular Sciences. 2024; 25(7):3607. https://doi.org/10.3390/ijms25073607
Chicago/Turabian StyleCharkiewicz, Radoslaw, Anetta Sulewska, Piotr Karabowicz, Grzegorz Lapuc, Alicja Charkiewicz, Marcin Kraska, Joanna Pancewicz, Malgorzata Lukasik, Miroslaw Kozlowski, Rafal Stec, and et al. 2024. "Six-Gene Signature for Differential Diagnosis and Therapeutic Decisions in Non-Small-Cell Lung Cancer—A Validation Study" International Journal of Molecular Sciences 25, no. 7: 3607. https://doi.org/10.3390/ijms25073607
APA StyleCharkiewicz, R., Sulewska, A., Karabowicz, P., Lapuc, G., Charkiewicz, A., Kraska, M., Pancewicz, J., Lukasik, M., Kozlowski, M., Stec, R., Ziembicka, D., Piszcz, W., Miltyk, W., & Niklinska, W. (2024). Six-Gene Signature for Differential Diagnosis and Therapeutic Decisions in Non-Small-Cell Lung Cancer—A Validation Study. International Journal of Molecular Sciences, 25(7), 3607. https://doi.org/10.3390/ijms25073607






