Effects of Regulating Hippo and Wnt on the Development and Fate Differentiation of Bovine Embryo
Abstract
1. Introduction
2. Results
2.1. Effect of LPA and DKK1 on the Development of Bovine Embryos
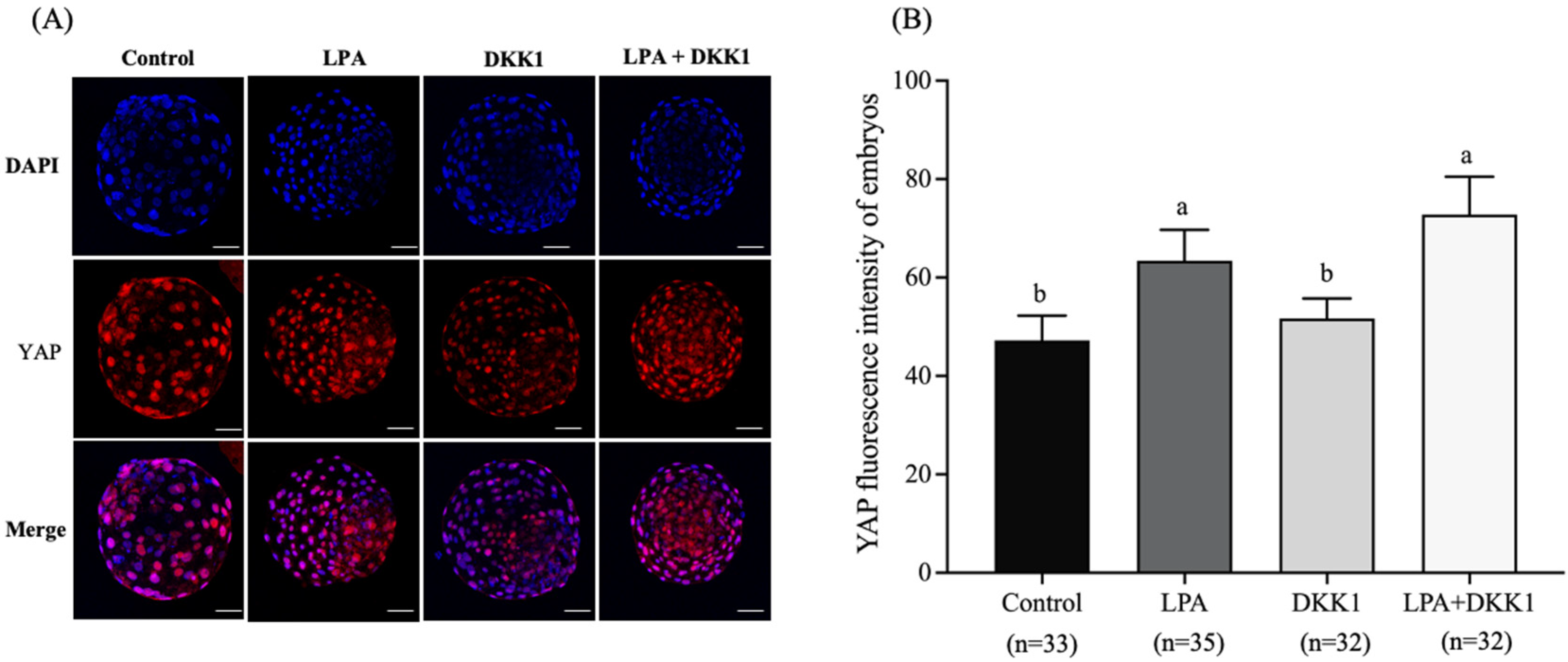
2.2. Effect of LPA and DKK1 on the YAP Fluorescence Intensity of Bovine Embryos
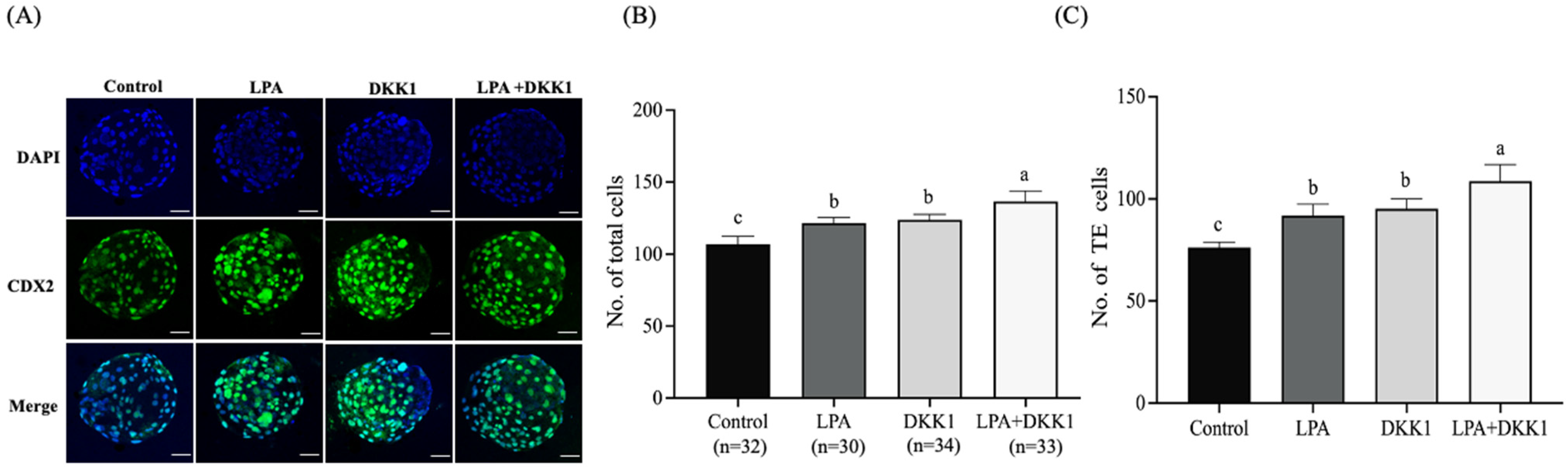
2.3. LPA and DKK1 Enhanced the Number of TE Cells of Bovine Embryos
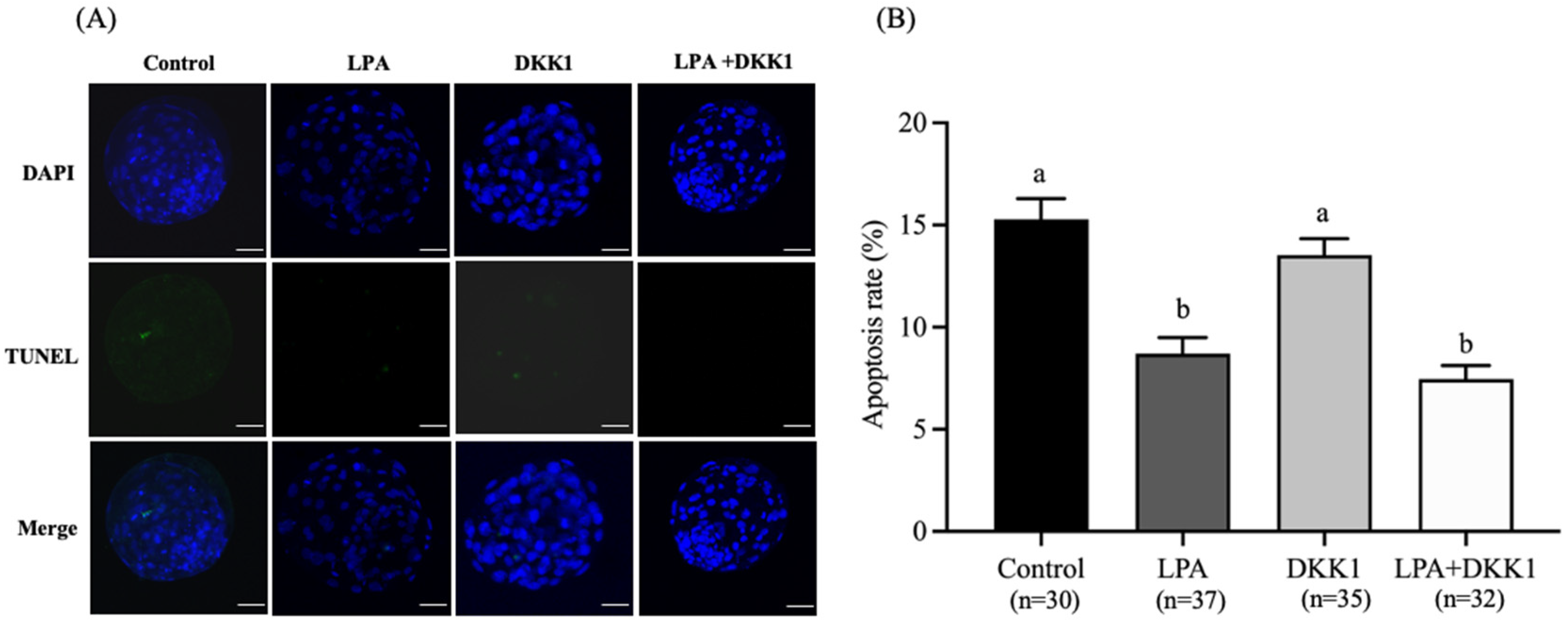
2.4. LPA and DKK1 Suppressed the Apoptosis of Bovine Embryos
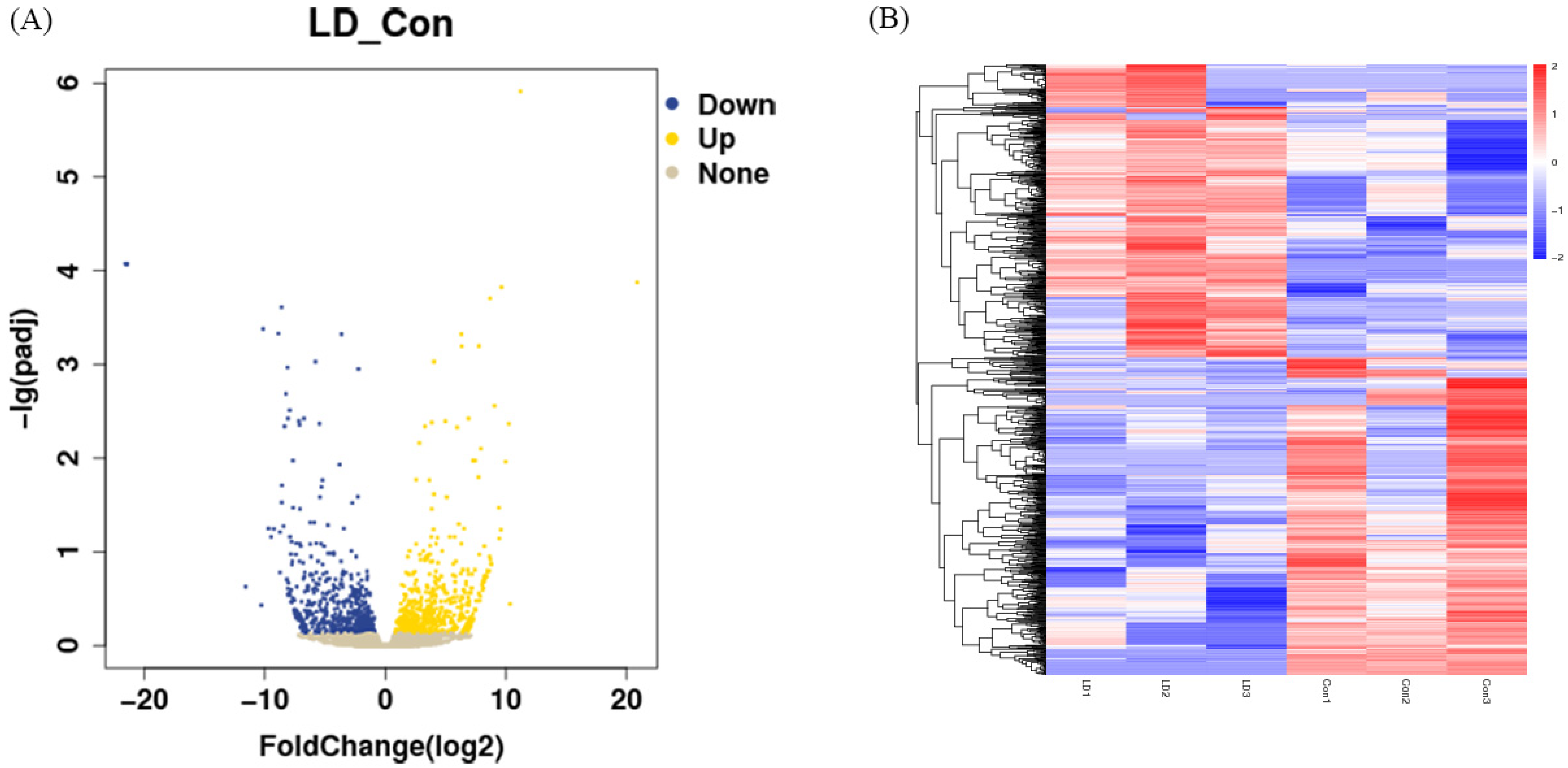
2.5. Transcriptomic Analysis of LPA- and DKK1-Treated Bovine Embryos
2.6. Screening of Differentially Expressed Genes (DEGs) in LPA- and DKK1-Treated Bovine Embryos
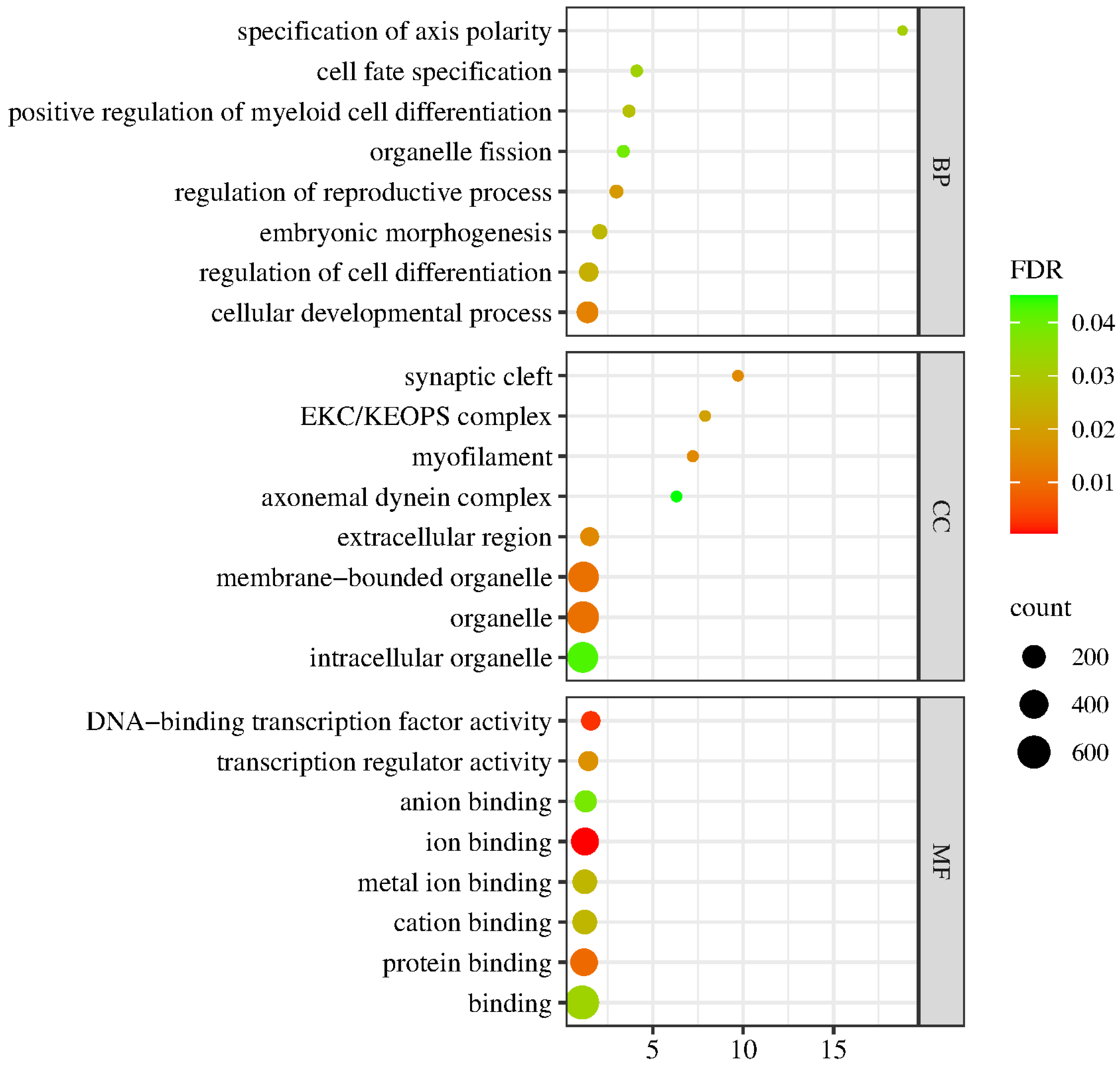
2.7. GO Enrichment Analysis of the DEGs
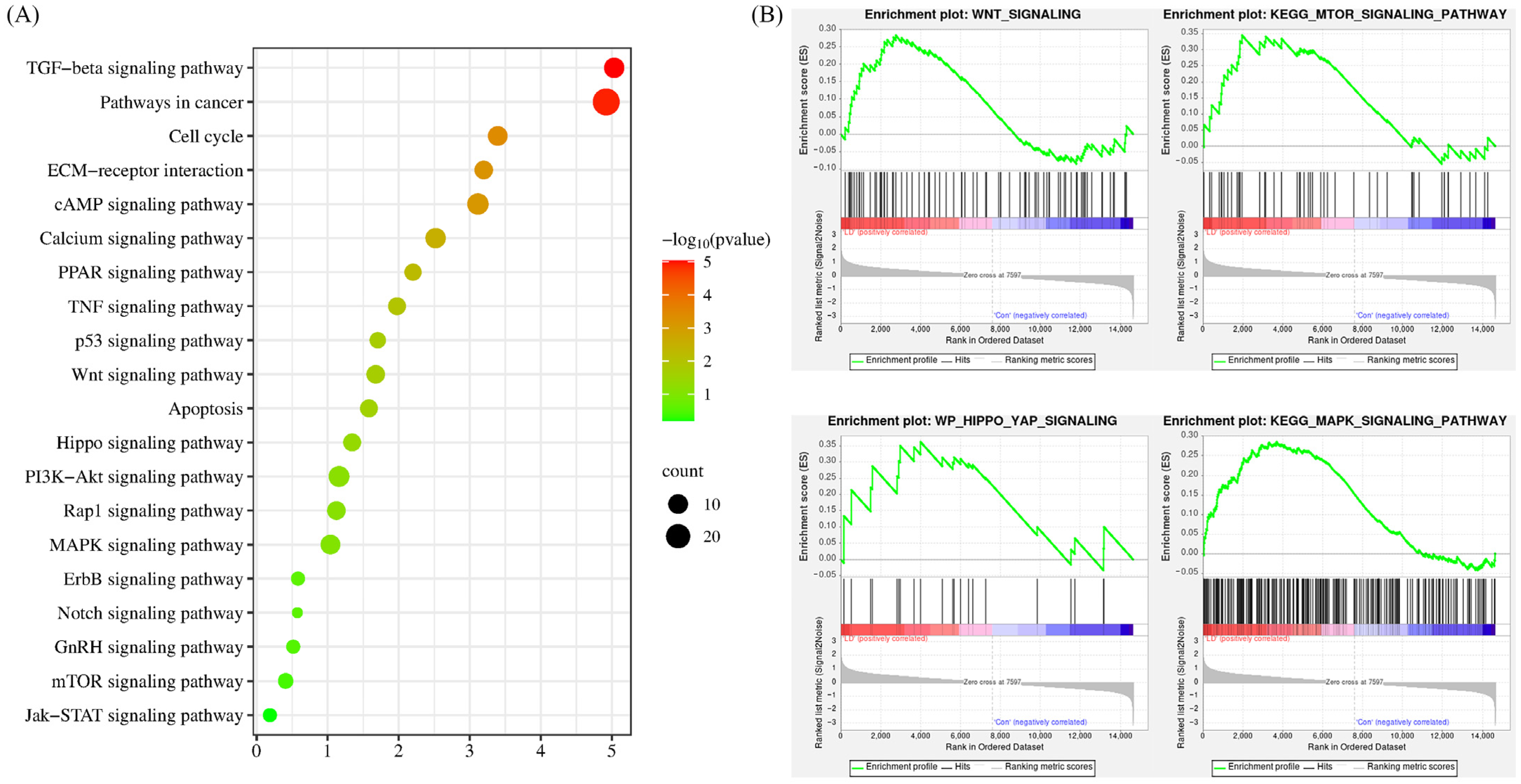
2.8. Significantly Regulated Biological Signaling Pathways in Response to LPA and DKK1 Treatment in Bovine Embryos
3. Discussion
4. Materials and Methods
4.1. In Vitro Maturation of Oocytes
4.2. In Vitro Fertilization (IVF) of Oocytes
4.3. IVC
4.4. Immunofluorescence Staining of Embryo
4.5. Detection of DNA Fragmentation by TUNEL
4.6. Sample Preparation, Library Construction and RNA Sequencing
4.7. Functional Enrichment Analysis
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wei, Q.Q.; Zhong, L.; Zhang, S.P.; Mu, H.Y.; Xiang, J.Z.; Yue, L.; Dai, Y.P.; Han, J.Y. Bovine lineage specification revealed by single-cell gene expression analysis from zygote to blastocyst. Biol. Reprod. 2017, 97, 5–17. [Google Scholar] [CrossRef]
- Gassler, J.; Kobayashi, W.; Gáspár, I.; Ruangroengkulrith, S.; Mohanan, A.; Gómez Hernández, L.; Kravchenko, P.; Kümmecke, M.; Lalic, A.; Rifel, N.; et al. Zygotic genome activation by the totipotency pioneer factor Nr5a2. Science 2022, 378, 1305–1315. [Google Scholar] [CrossRef]
- Underhill, L.A.; Robins, J.C. Trophoblast development in the murine preimplantation embryo. Semin. Reprod. Med. 2016, 34, 57–62. [Google Scholar]
- Yao, C.; Zhang, W.; Shuai, L. The first cell fate decision in pre-implantation mouse embryos. Cell Regen. 2019, 8, 51–57. [Google Scholar] [CrossRef]
- Garg, V.; Yang, Y.; Nowotschin, S.; Setty, M.; Kuo, Y.Y.; Sharma, R.; Polyzos, A.; Salataj, E.; Murphy, D.; Jang, A.; et al. Single-cell analysis of bidirectional reprogramming between early embryonic states reveals mechanisms of differential lineage plasticities. bioRxiv 2023, 534648. [Google Scholar] [CrossRef]
- Perera, M.; Brickman, J.M. In vitro models of human hypoblast and mouse primitive endoderm. Curr. Opin. Genet. Dev. 2023, 83, 102115. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, B.; Haider, S.; Meinhardt, G.; Pollheimer, J.; Knöfler, M. WNT and NOTCH signaling in human trophoblast development and differentiation. Cell Mol. Life Sci. 2022, 79, 292. [Google Scholar] [CrossRef]
- Fan, R.; Kim, Y.S.; Wu, J.; Chen, R.; Zeuschner, D.; Mildner, K.; Adachi, K.; Wu, G.; Galatidou, S.; Li, J.; et al. Wnt/Beta-catenin/Esrrb signalling controls the tissue-scale reorganization and maintenance of the pluripotent lineage during murine embryonic diapause. Nat. Commun. 2020, 11, 5499. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhang, E.; Yang, M.; Lu, L. Overexpression of Wnt11 promotes chondrogenic differentiation of bone marrow-derived mesenchymal stem cells in synergism with TGF-β. Mol. Cell Biochem. 2014, 390, 123–131. [Google Scholar] [CrossRef]
- Logan, C.Y.; Nusse, R. The Wnt signaling pathway in development and disease. Annu. Rev. Cell Dev. Biol. 2004, 20, 781–810. [Google Scholar] [CrossRef]
- Hayat, R.; Manzoor, M.; Hussain, A. Wnt signaling pathway: A comprehensive review. Cell Biol. Int. 2021, 46, 863–877. [Google Scholar] [CrossRef] [PubMed]
- Zinovyeva, A.Y.; Yamamoto, Y.; Sawa, H.; Forrester, W.C. Complex network of Wnt signaling regulates neuronal migrations during Caenorhabditis elegans development. Genetics 2008, 179, 1357–1371. [Google Scholar] [CrossRef] [PubMed]
- Steinhart, Z.; Angers, S. Wnt signaling in development and tissue homeostasis. Development 2018, 146, dev146589. [Google Scholar] [CrossRef] [PubMed]
- Denicol, A.C.; Block, J.; Kelley, D.E.; Pohler, K.G.; Dobbs, K.B.; Mortensen, C.J.; Ortega, M.S.; Hansen, P.J. The WNT signaling antagonist Dickkopf-1 directs lineage commitment and promotes survival of the preimplantation embryo. FASEB J. 2014, 28, 3975–3986. [Google Scholar] [CrossRef] [PubMed]
- Minten, M.A.; Bilby, T.R.; Bruno, R.G.; Allen, C.C.; Madsen, C.A.; Wang, Z.; Sawyer, J.E.; Tibary, A.; Neibergs, H.L.; Geary, T.W.; et al. Effects of fertility on gene expression and function of the bovine endometrium. PLoS ONE 2013, 8, 69444. [Google Scholar] [CrossRef]
- Kawashima, I.; Kawamura, K. Regulation of follicle growth through hormonal factors and mechanical cues mediated by Hippo signaling pathway. Syst. Biol. Reprod. Med. 2018, 64, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Grosbois, J.; Demeestere, I. Dynamics of PI3K and Hippo signaling pathways during in vitro human follicle activation. Hum. Reprod. 2018, 33, 1705–1714. [Google Scholar] [CrossRef] [PubMed]
- Masciangelo, R.; Hossay, C.; Chiti, M.C.; Manavella, D.D.; Amorim, C.A.; Donnez, J.; Dolmans, M.M. Role of the PI3K and Hippo pathways in follicle activation after grafting of human ovarian tissue. J. Assist. Reprod. Genet. 2020, 37, 101–108. [Google Scholar] [CrossRef]
- Hashimoto, M.; Sasaki, H. Epiblast formation by TEAD-YAP-dependent expression of pluripotency factors and competitive elimination of unspecified cells. Dev. Cell 2019, 50, 139–154. [Google Scholar] [CrossRef]
- Karasek, C.; Ashry, M.; Driscoll, C.S.; Knott, J.G. A tale of two cell-fates: Role of the Hippo signaling pathway and transcription factors in early lineage formation in mouse preimplantation embryos. Mol. Hum. Reprod. 2020, 26, 653–664. [Google Scholar] [CrossRef]
- Dong, J.; Feldmann, G.; Huang, J.; Wu, S.; Zhang, N.; Comerford, S.A.; Gayyed, M.F.; Anders, R.A.; Maitra, A.; Pan, D. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell 2007, 130, 1120–1133. [Google Scholar] [CrossRef] [PubMed]
- Sheng, X.; Yung, Y.C.; Chen, A.; Chun, J. Lysophosphatidic acid signalling in development. Development 2015, 142, 1390–1395. [Google Scholar] [CrossRef]
- Hama, K.; Aoki, J.; Inoue, A.; Endo, T.; Amano, T.; Motoki, R.; Kanai, M.; Ye, X.; Chun, J.; Matsuki, N.; et al. Embryo Spacing and Implantation Timing Are Differentially Regulated by LPA3-Mediated Lysophosphatidic Acid Signaling in Mice1. Biol. Reprod. 2007, 77, 954–959. [Google Scholar] [CrossRef]
- Lin, M.E.; Herr, D.R.; Chun, J. Lysophosphatidic acid (LPA) receptors: Signaling properties and disease relevance. Prostaglandins Other Lipid Mediat. 2010, 91, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Teo, S.T.; Yung, Y.C.; Herr, D.R.; Chun, J. Lysophosphatidic acid in vascular development and disease. IUBMB Life 2009, 61, 791–799. [Google Scholar] [CrossRef]
- Kim, D.; Li, H.Y.; Lee, J.H.; Oh, Y.S.; Jun, H.S. Lysophosphatidic acid increases mesangial cell proliferation in models of diabetic nephropathy via Rac1/MAPK/KLF5 signaling. Exp. Mol. Med. 2019, 51, 1–10. [Google Scholar] [CrossRef]
- Macosko, E.Z.; Basu, A.; Satija, R.; Nemesh, J.; Shekhar, K.; Goldman, M.; Tirosh, I.; Bialas, A.R.; Kamitaki, N.; Martersteck, E.M.; et al. Highly Parallel Genome-wide Expression Profiling of Individual Cells Using Nanoliter Droplets. Cell 2015, 161, 1202–1214. [Google Scholar] [CrossRef] [PubMed]
- Zilionis, R.; Nainys, J.; Veres, A.; Savova, V.; Zemmour, D.; Klein, A.M.; Mazutis, L. Single-cell barcoding and sequencing using droplet microfluidics. Nat. Protoc. 2017, 12, 44–73. [Google Scholar] [CrossRef] [PubMed]
- Pijuan-Sala, B.; Griffiths, J.A.; Guibentif, C.; Hiscock, T.W.; Jawaid, W.; Calero-Nieto, F.J.; Mulas, C.; Ibarra-Soria, X.; Tyser, R.C.V.; Ho, D.L.L.; et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 2019, 566, 490–495. [Google Scholar] [CrossRef]
- Argelaguet, R.; Clark, S.J.; Mohammed, H.; Stapel, L.C.; Krueger, C.; Kapourani, C.A.; Imaz-Rosshandler, I.; Lohoff, T.; Xiang, Y.; Hanna, C.W.; et al. Multiomics profiling of mouse gastrulation at single-cell resolution. Nature 2019, 576, 487–491. [Google Scholar] [CrossRef]
- Ji, P.; Liu, Y.; Yan, L.; Jia, Y.; Zhao, M.; Lv, D.; Yao, Y.; Ma, W.; Yin, D.; Liu, F.; et al. Melatonin improves the vitrification of sheep morulae by modulating transcriptome. Front. Vet. Sci. 2023, 10, 1212047. [Google Scholar] [CrossRef] [PubMed]
- Im, G.S.; Lai, L.; Liu, Z.; Hao, Y.; Wax, D.; Bonk, A.; Prather, R.S. In vitro development of preimplantation porcine nuclear transfer embryos cultured in different media and gas atmospheres. Theriogenology 2004, 61, 1125–1135. [Google Scholar] [CrossRef]
- Shin, M.Y.; Lee, S.E.; Son, Y.J.; Park, Y.G.; Jeong, S.G.; Kim, E.Y.; Park, S.P. Lysophosphatidic acid accelerates development of porcine embryos by activating formation of the blastocoel. Mol. Reprod. Dev. 2018, 85, 62–71. [Google Scholar] [CrossRef]
- Jo, J.W.; Jee, B.C.; Suh, C.S.; Kim, S.H. Addition of lysophosphatidic acid to mouse oocyte maturation media can enhance fertilization and developmental competence. Hum. Reprod. 2014, 29, 234–241. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.Y.; Jiang, Y.; Lin, T.; Kang, J.W.; Lee, J.E.; Jin, D.I. Lysophosphatidic acid improves porcine oocyte maturation and embryo development in vitro. Mol. Reprod. Dev. 2015, 82, 66–77. [Google Scholar] [CrossRef]
- Zhu, X.; Li, L.; Gao, B.; Zhang, D.; Ren, Y.; Zheng, B.; Li, M.; Shi, D.; Huang, B. Early development of porcine parthenogenetic embryos and reduced expression of primed pluripotent marker genes under the effect of lysophosphatidic acid. Reprod. Domest. Anim. 2018, 53, 1191–1199. [Google Scholar] [CrossRef]
- Torres, A.C.; Boruszewska, D.; Batista, M.; Kowalczyk-Zieba, I.; Diniz, P.; Sinderewicz, E.; Saulnier-Blache, J.S.; Woclawek-Potocka, I.; Lopes-da-Costa, L. Lysophosphatidic acid signaling in late cleavage and blastocyst stage bovine embryos. Mediat. Inflamm. 2014, 2014, 678968. [Google Scholar] [CrossRef]
- Boruszewska, D.; Sinderewicz, E.; Kowalczyk-Zieba, I.; Grycmacher, K.; Woclawek-Potocka, I. Studies on lysophosphatidic acid action during in vitro preimplantation embryo development. Domest. Anim. Endocrinol. 2016, 54, 15–29. [Google Scholar] [CrossRef] [PubMed]
- Shyam, S.; Goel, P.; Kumar, D.; Malpotra, S.; Singh, M.K.; Lathwal, S.S.; Chand, S.; Palta, P. Effect of Dickkopf-1 and colony stimulating factor-2 on the developmental competence, quality, gene expression and live birth rate of buffalo (Bubalus bubalis) embryos produced by hand-made cloning. Theriogenology 2020, 157, 254–262. [Google Scholar] [CrossRef]
- Qin, H.; Hejna, M.; Liu, Y.; Percharde, M.; Wossidlo, M.; Blouin, L.; Durruthy-Durruthy, J.; Wong, P.; Qi, Z.; Yu, J.; et al. YAP Induces Human Naive Pluripotency. Cell Rep. 2016, 14, 2301–2312. [Google Scholar] [CrossRef]
- Emura, N.; Saito, Y.; Miura, R.; Sawai, K. Effect of downregulating the Hippo pathway members YAP1 and LATS2 transcripts on early development and gene expression involved in differentiation in porcine embryos. Cell Reprogr. 2020, 22, 62–70. [Google Scholar] [CrossRef] [PubMed]
- Nishioka, N.; Inoue, K.I.; Adachi, K.; Kiyonari, H.; Ota, M.; Ralston, A.; Yabuta, N.; Hirahara, S.; Stephenson, R.O.; Ogonuki, N.; et al. The Hippo Signaling Pathway Components Lats and Yap Pattern Tead4 Activity to Distinguish Mouse Trophectoderm from Inner Cell Mass. Dev. Cell 2009, 16, 398–410. [Google Scholar] [CrossRef] [PubMed]
- Van Sciver, N.; Ohashi, M.; Pauly, N.P.; Bristol, J.A.; Nelson, S.E.; Johannsen, E.C.; Kenney, S.C. Hippo signaling effectors YAP and TAZ induce Epstein-Barr Virus (EBV) lytic reactivation through TEADs in epithelial cells. PLoS Pathog. 2021, 17, e1009783. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Lin, L.; Zhu, H.; Wu, Z.; Ding, X.; Hu, R.; Jiang, Y.; Tang, C.; Ding, S.; Guo, R. YAP Promotes Cell Proliferation and Stemness Maintenance of Porcine Muscle Stem Cells under High-Density Condition. Cells 2021, 10, 3069. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.X.; Zhao, B.; Panupinthu, N.; Jewell, J.L.; Lian, I.; Wang, L.H.; Zhao, J.; Yuan, H.; Tumaneng, K.; Li, H.; et al. Regulation of the Hippo-YAP Pathway by G-Protein-Coupled Receptor Signaling. Cell 2012, 150, 780–791. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Gentile, L.; Fuchikami, T.; Sutter, J.; Psathaki, K.; Esteves, T.C.; Araúzo-Bravo, M.J.; Ortmeier, C.; Verberk, G.; Abe, K.; et al. Initiation of trophectoderm lineage specification in mouse embryos is independent of Cdx2. Development 2010, 137, 4159–4169. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Armant, D.R. Lysophosphatidic acid regulates murine blastocyst development by transactivation of receptors for heparin- binding EGF-like growth factor. Exp. Cell Res. 2004, 296, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, D.; Okura, K.; Nagakura, S.; Ogawa, H. CDX2 downregulation in mouse mural trophectoderm during peri-implantation is heteronomous, dependent on the YAP-TEAD pathway and controlled by estrogen-induced factors. Reprod. Med. Biol. 2022, 21, e12446. [Google Scholar] [CrossRef] [PubMed]
- Liu-chittenden, Y.; Huang, B.; Shim, J.S.; Dev, G.; Chen, Q.; Lee, S.; Anders, R.A.; Liu, J.O.; Pan, D. Genetic and pharmacological disruption of the TEAD-YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 2012, 26, 1300–1305. [Google Scholar] [CrossRef]
- Negrón-Pérez, V.M.; Hansen, P.J. Role of yes-associated protein 1, angiomotin, and mitogen-activated kinase kinase 1/2 in development of the bovine blastocyst. Biol. Reprod. 2018, 98, 170–183. [Google Scholar] [CrossRef]
- Xie, H.; Tranguch, S.; Jia, X.; Zhang, H.; Das, S.K.; Dey, S.K.; Kuo, C.J.; Wang, H. Inactivation of nuclear Wnt-β-catenin signaling limits blastocyst competency for implantation. Development 2008, 135, 717–727. [Google Scholar] [CrossRef] [PubMed]
- Amaral, T.F.; Gonella-Diaza, A.; Heredia, D.; Melo, G.D.; Estrada-Cortés, E.; Jensen, L.M.; Pohler, K.; Hansen, P.J. Actions of DKK1 on the preimplantation bovine embryo to affect pregnancy establishment, placental function, and postnatal phenotype†. Biol. Reprod. 2022, 107, 945–955. [Google Scholar] [CrossRef]
- Lim, K.T.; Gupta, M.K.; Lee, S.H.; Jung, Y.H.; Han, D.W.; Lee, H.T. Possible involvement of Wnt/β-catenin signaling pathway in hatching and trophectoderm differentiation of pig blastocysts. Theriogenology 2013, 79, 284–290. [Google Scholar] [CrossRef] [PubMed]
- Denicol, A.C.; Dobbs, K.B.; McLean, K.M.; Carambula, S.F.; Loureiro, B.; Hansen, P.J. Canonical WNT signaling regulates development of bovine embryos to the blastocyst stage. Sci. Rep. 2013, 3, 1266. [Google Scholar] [CrossRef] [PubMed]
- Tribulo, P.; Bernal Ballesteros, B.H.; Ruiz, A.; Tríbulo, A.; Tríbulo, R.J.; Tríbulo, H.E.; Bo, G.A.; Hansen, P.J. Consequences of exposure of embryos produced in vitro in a serum-containing medium to dickkopf-related protein 1 and colony stimulating factor 2 on blastocyst yield, pregnancy rate, and birth weight. J. Anim. Sci. 2017, 95, 4407–4412. [Google Scholar] [CrossRef] [PubMed]
- Nabenishi, H.; Takagi, S.; Kamata, H.; Nishimoto, T.; Morita, T.; Ashizawa, K.; Tsuzuki, Y. The role of mitochondrial transition pores on bovine oocyte competence after heat stress, as determined by effects of cyclosporin A. Mol. Reprod. Dev. 2012, 79, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Han, X.; Zhou, Z.; Uyunbilig, B.; Huang, X.; Li, R.; Li, X. Wnt3a Activates the WNT-YAP/TAZ Pathway to Sustain CDX2 Expression in Bovine Trophoblast Stem Cells. DNA Cell Biol. 2019, 38, 410–422. [Google Scholar] [CrossRef] [PubMed]
- Hirate, Y.; Hirahara, S.; Inoue, K.I.; Suzuki, A.; Alarcon, V.B.; Akimoto, K.; Hirai, T.; Hara, T.; Adachi, M.; Chida, K.; et al. Polarity-dependent distribution of angiomotin localizes hippo signaling in preimplantation embryos. Curr. Biol. 2013, 23, 1181–1194. [Google Scholar] [CrossRef] [PubMed]
- Panda, B.S.K.; Mohapatra, S.K.; Chaudhary, D.; Alhussien, M.N.; Kapila, R.; Dang, A.K. Proteomics and transcriptomics study reveals the utility of ISGs as novel molecules for early pregnancy diagnosis in dairy cows. J. Reprod. Immunol. 2020, 140, 103148. [Google Scholar] [CrossRef]
- Casano, A.B.; Menchetti, L.; Trabalza-Marinucci, M.; Riva, F.; De Matteis, G.; Brecchia, G.; Inglesi, A.; Rossi, E.; Signorelli, F.; Barile, V.L.; et al. Gene expression of pregnancy-associated glycoproteins-1 (PAG-1), interferon-tau (IFNt) and interferon stimulated genes (ISGs) as diagnostic and prognostic markers of maternal-fetal cellular interaction in buffalo cows. Theriogenology 2023, 209, 89–97. [Google Scholar] [CrossRef]
- Adhikari, B.; Lee, C.N.; Khadka, V.S.; Deng, Y.; Fukumoto, G.; Thorne, M.; Caires, K.; Odani, J.; Mishra, B. RNA-Sequencing based analysis of bovine endometrium during the maternal recognition of pregnancy. BMC Genom. 2022, 23, 494. [Google Scholar] [CrossRef]
- Constantinou, P.E.; Morgado, M.; Carson, D.D. Transmembrane Mucin Expression and Function in Embryo Implantation and Placentation. Adv. Anat. Embryol. Cell Biol. 2015, 216, 51–68. [Google Scholar] [PubMed]
- Wang, X.; Zhu, B.; Xiong, S.; Sheng, X.; Qi, X.; Huang, Q.; Chen, C.; Guo, Y.; Ni, H. Expression and function of MUC1 in uterine tissues during early pregnancy in sheep after natural oestrous or artificially-induced oestrous. Theriogenology 2018, 108, 339–347. [Google Scholar] [CrossRef] [PubMed]
- Sponchiado, M.; Gomes, N.S.; Fontes, P.K.; Martins, T.; Del Collado, M.; Pastore, A.A.; Pugliesi, G.; Nogueira, M.F.G.; Binelli, M. Pre-hatching embryo-dependent and -independent programming of endometrial function in cattle. PLoS ONE 2017, 12, e0175954. [Google Scholar] [CrossRef] [PubMed]
- Cormier, S.; Vandormael-Pournin, S.; Babinet, C.; Cohen-Tannoudji, M. Developmental expression of the Notch signaling pathway genes during mouse preimplantation development. Gene Expr. Patterns 2004, 4, 713–717. [Google Scholar] [CrossRef] [PubMed]
- Rayon, T.; Menchero, S.; Nieto, A.; Xenopoulos, P.; Crespo, M.; Cockburn, K.; Canon, S.; Sasaki, H.; Hadjantonakis, A.K.; de la Pompa, J.L.; et al. Notch and hippo converge on Cdx2 to specify the trophectoderm lineage in the mouse blastocyst. Dev. Cell 2014, 30, 410–422. [Google Scholar] [CrossRef] [PubMed]
- Menchero, S.; Rollan, I.; Lopez-Izquierdo, A.; Andreu, M.J.; de Aja, J.S.; Kang, M.J.; Adan, J.; Benedito, R.; Rayon, T.; Hadjantonakis, A.K.; et al. Transitions in cell potency during early mouse development are driven by Notch. Elife 2019, 8, e42930. [Google Scholar] [CrossRef] [PubMed]
- Ni, N.; Li, Q. TGFβ superfamily signaling and uterine decidualization. Reprod. Biol. Endocrinol. 2017, 15, 84. [Google Scholar] [CrossRef] [PubMed]
- Hwang, M.; Perez, C.A.; Moretti, L.; Lu, B. The mTOR signaling network: Insights from its role during embryonic development. Curr. Med. Chem. 2008, 15, 1192–1208. [Google Scholar] [CrossRef]
- Hentges, K.E.; Sirry, B.; Gingras, A.C.; Sarbassov, D.; Sonenberg, N.; Sabatini, D.; Peterson, A.S. FRAP/mTOR is required for proliferation and patterning during embryonic development in the mouse. Proc. Natl. Acad. Sci. USA 2001, 98, 13796–13801. [Google Scholar] [CrossRef]
- Murakami, M.; Ichisaka, T.; Maeda, M.; Oshiro, N.; Hara, K.; Edenhofer, F.; Kiyama, H.; Yonezawa, K.; Yamanaka, S. mTOR is essential for growth and proliferation in early mouse embryos and embryonic stem cells. Mol. Cell Biol. 2004, 24, 6710–6718. [Google Scholar] [CrossRef]
- Ma, C.; Li, Q.; Yang, Y.; Ge, L.; Cai, J.; Wang, J.; Zhu, M.; Xiong, Y.; Zhang, W.; Xie, J.; et al. mTOR hypoactivity leads to trophectoderm cell failure by enhancing lysosomal activation and disrupting the cytoskeleton in preimplantation embryo. Cell Biosci. 2023, 13, 219. [Google Scholar] [CrossRef] [PubMed]
- Radi, Z.A.; Marusak, R.A.; Morris, D.L. Species comparison of the role of p38 MAP kinase in the female reproductive system. J. Toxicol. Pathol. 2009, 22, 109–124. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bell, C.E.; Watson, A.J. p38 MAPK regulates cavitation and tight junction function in the mouse blastocyst. PLoS ONE 2013, 8, e59528. [Google Scholar] [CrossRef]
- Mudgett, J.S.; Ding, J.; Guh-Siesel, L.; Chartrain, N.A.; Yang, L.; Gopal, S.; Shen, M.M. Essential role for p38alpha mitogen-activated protein kinase in placental angiogenesis. Proc. Natl. Acad. Sci. USA 2000, 97, 10454–10459. [Google Scholar] [CrossRef] [PubMed]
- Jamioł, M.; Sozoniuk, M.; Wawrzykowski, J.; Kankofer, M. Effect of Sex Steroids and PGF2α on the Expression of Their Receptors and Decorin in Bovine Caruncular Epithelial Cells in Early-Mid Pregnancy. Molecules 2022, 27, 7420. [Google Scholar] [CrossRef]
- McBeath, R.; Pirone, D.M.; Nelson, C.M.; Bhadriraju, K.; Chen, C.S. Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev. Cell 2004, 6, 483–495. [Google Scholar] [CrossRef]
- Brackett, B.G.; Oliphant, G. Capacitation of Rabbit Spermatozoa in vitro. Biol. Reprod. 1975, 12, 260–274. [Google Scholar] [CrossRef]
- Yu, B.; van Tol, H.T.A.; Oei, C.H.Y.; Stout, T.A.E.; Roelen, B.A.J. Lysophosphatidic Acid Accelerates Bovine In Vitro-Produced Blastocyst Formation through the Hippo/YAP Pathway. Int. J. Mol. Sci. 2021, 22, 5915. [Google Scholar] [CrossRef]
- Gao, S.; Yan, L.; Wang, R.; Li, J.; Yong, J.; Zhou, X.; Wei, Y.; Wu, X.; Wang, X.; Fan, X.; et al. Tracing the temporal-spatial transcriptome landscapes of the human fetal digestive tract using single-cell RNA-sequencing. Nat. Cell Biol. 2018, 20, 721–734. [Google Scholar] [CrossRef]
| Groups | NO. of COCs | NO. of Embryos Cleaved | Blastocyst Rate |
|---|---|---|---|
| Control | 133 | 99 (74.18 ± 3.44) c | 30 (30.02 ± 3.30) c |
| LPA | 138 | 115 (83.57 ± 3.60) b | 46 (40.04 ± 2.54) b |
| DKK1 | 145 | 119 (81.97 ± 3.38) b | 46 (38.91 ± 3.10) b |
| LPA + DKK1 | 127 | 114 (89.67 ± 4.53) a | 54 (47.11 ± 4.23) a |
| Index | Sample | |||||
|---|---|---|---|---|---|---|
| Con1 | Con2 | Con3 | LD1 | LD2 | LD3 | |
| Raw Reads/bp | 63,756,616 | 59,220,724 | 45,639,012 | 55,652,282 | 53,065,386 | 41,746,418 |
| Clean Reads/bp | 62,536,622 | 58,162,374 | 44,891,384 | 54,470,954 | 52,075,814 | 40,869,626 |
| Clean Reads Rate (%) | 98.09 | 98.21 | 98.36 | 97.88 | 98.14 | 97.90 |
| Clean Q30 (%) | 96.00 | 95.97 | 95.89 | 96.41 | 96.11 | 96.13 |
| Mapped Reads/bp | 60,980,959 | 56,889,681 | 43,953,245 | 53,453,414 | 50,735,875 | 39,926,192 |
| Mapping Rate (%) | 97.51 | 97.81 | 97.91 | 98.13 | 97.43 | 97.69 |
| Gene | Sample | Up/Down | |||||
|---|---|---|---|---|---|---|---|
| LD1 | LD2 | LD3 | Con1 | Con2 | Con3 | ||
| GPX3 | 5 | 47 | 38 | 6 | 3 | 0 | up |
| SORBS3 | 74 | 179 | 267 | 10 | 29 | 40 | up |
| MX2 | 26 | 286 | 74 | 0 | 25 | 0 | up |
| FAM46A | 748 | 710 | 172 | 57 | 198 | 107 | up |
| FOS | 3482 | 9111 | 5408 | 1770 | 777 | 2426 | up |
| ASTL | 17 | 36 | 127 | 5 | 0 | 12 | up |
| ID1 | 51 | 220 | 98 | 13 | 6 | 9 | up |
| MAP1LC3A | 363 | 248 | 214 | 138 | 130 | 133 | up |
| C15H11orf34 | 293 | 5806 | 473 | 11 | 69 | 8 | up |
| SOX6 | 79 | 20 | 17 | 11 | 14 | 79 | up |
| OAS1Y | 25 | 152 | 0 | 0 | 0 | 0 | up |
| RICTOR | 1089 | 1323 | 1061 | 748 | 924 | 432 | up |
| TUNAR | 14 | 53 | 0 | 0 | 0 | 0 | up |
| TGFBR2 | 99 | 308 | 420 | 192 | 16 | 56 | up |
| RGS4 | 6960 | 1438 | 0 | 34 | 11 | 9 | up |
| FAS | 480 | 1454 | 1385 | 357 | 190 | 0 | up |
| MUC1 | 992 | 1502 | 556 | 50 | 294 | 34 | up |
| OVOL1 | 475 | 553 | 292 | 86 | 288 | 48 | up |
| BEX2 | 17 | 187 | 7 | 7 | 3 | 3 | up |
| AMOT | 2114 | 2437 | 727 | 276 | 1038 | 216 | up |
| IFNGR2 | 8 | 197 | 41 | 0 | 1 | 0 | up |
| ABCB4 | 150 | 450 | 27 | 2 | 9 | 0 | up |
| IL1RAP | 18 | 140 | 106 | 33 | 33 | 17 | up |
| KRT7 | 2200 | 5046 | 7411 | 3081 | 2034 | 1022 | up |
| MGST1 | 434 | 1348 | 58 | 31 | 248 | 31 | up |
| IGFBP7 | 22 | 108 | 118 | 8 | 2 | 0 | up |
| WNT3A | 4 | 33 | 12 | 0 | 0 | 0 | up |
| COL5A3 | 17 | 111 | 44 | 7 | 12 | 1 | up |
| HMGB2 | 3389 | 2589 | 2258 | 6630 | 4773 | 5665 | down |
| OTX1 | 37 | 22 | 0 | 282 | 2 | 1442 | down |
| RSPO3 | 268 | 230 | 77 | 523 | 380 | 680 | down |
| RASSF2 | 50 | 65 | 52 | 1377 | 15 | 799 | down |
| LRP4 | 63 | 38 | 12 | 61 | 94 | 291 | down |
| SLC25A27 | 105 | 54 | 47 | 464 | 190 | 153 | down |
| GCM1 | 32 | 2 | 0 | 117 | 320 | 90 | down |
| BAD | 331 | 513 | 326 | 1242 | 398 | 1843 | down |
| FAM53B | 23 | 4 | 1 | 53 | 41 | 64 | down |
| EED | 1068 | 704 | 811 | 2493 | 1260 | 3958 | down |
| WNT2 | 0 | 0 | 0 | 0 | 4 | 64 | down |
| NANOG | 185 | 381 | 68 | 50 | 2645 | 14,549 | down |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, P.; Zhang, H.; Li, C.; Yang, B.; Feng, X.; Cao, J.; Du, W.; Shahzad, M.; Khan, A.; Sun, S.-C.; et al. Effects of Regulating Hippo and Wnt on the Development and Fate Differentiation of Bovine Embryo. Int. J. Mol. Sci. 2024, 25, 3912. https://doi.org/10.3390/ijms25073912
Zhang P, Zhang H, Li C, Yang B, Feng X, Cao J, Du W, Shahzad M, Khan A, Sun S-C, et al. Effects of Regulating Hippo and Wnt on the Development and Fate Differentiation of Bovine Embryo. International Journal of Molecular Sciences. 2024; 25(7):3912. https://doi.org/10.3390/ijms25073912
Chicago/Turabian StyleZhang, Peipei, Hang Zhang, Chongyang Li, Baigao Yang, Xiaoyi Feng, Jianhua Cao, Weihua Du, Muhammad Shahzad, Adnan Khan, Shao-Chen Sun, and et al. 2024. "Effects of Regulating Hippo and Wnt on the Development and Fate Differentiation of Bovine Embryo" International Journal of Molecular Sciences 25, no. 7: 3912. https://doi.org/10.3390/ijms25073912
APA StyleZhang, P., Zhang, H., Li, C., Yang, B., Feng, X., Cao, J., Du, W., Shahzad, M., Khan, A., Sun, S.-C., & Zhao, X. (2024). Effects of Regulating Hippo and Wnt on the Development and Fate Differentiation of Bovine Embryo. International Journal of Molecular Sciences, 25(7), 3912. https://doi.org/10.3390/ijms25073912







