NMR Studies of the Interactions between Sialyllactoses and the Polysialytransferase Domain for Polysialylation Inhibition
Abstract
1. Introduction
2. Methods and Materials
2.1. Material Sources
2.2. Preparations of Circular Dichroism (CD) Samples and the Measurements and Recorded Methods of CD Spectra
2.3. Preparation of NMR Samples
2.4. NMR Spectroscopic Measurements
3. Results
3.1. CD Data
3.2. NMR Results
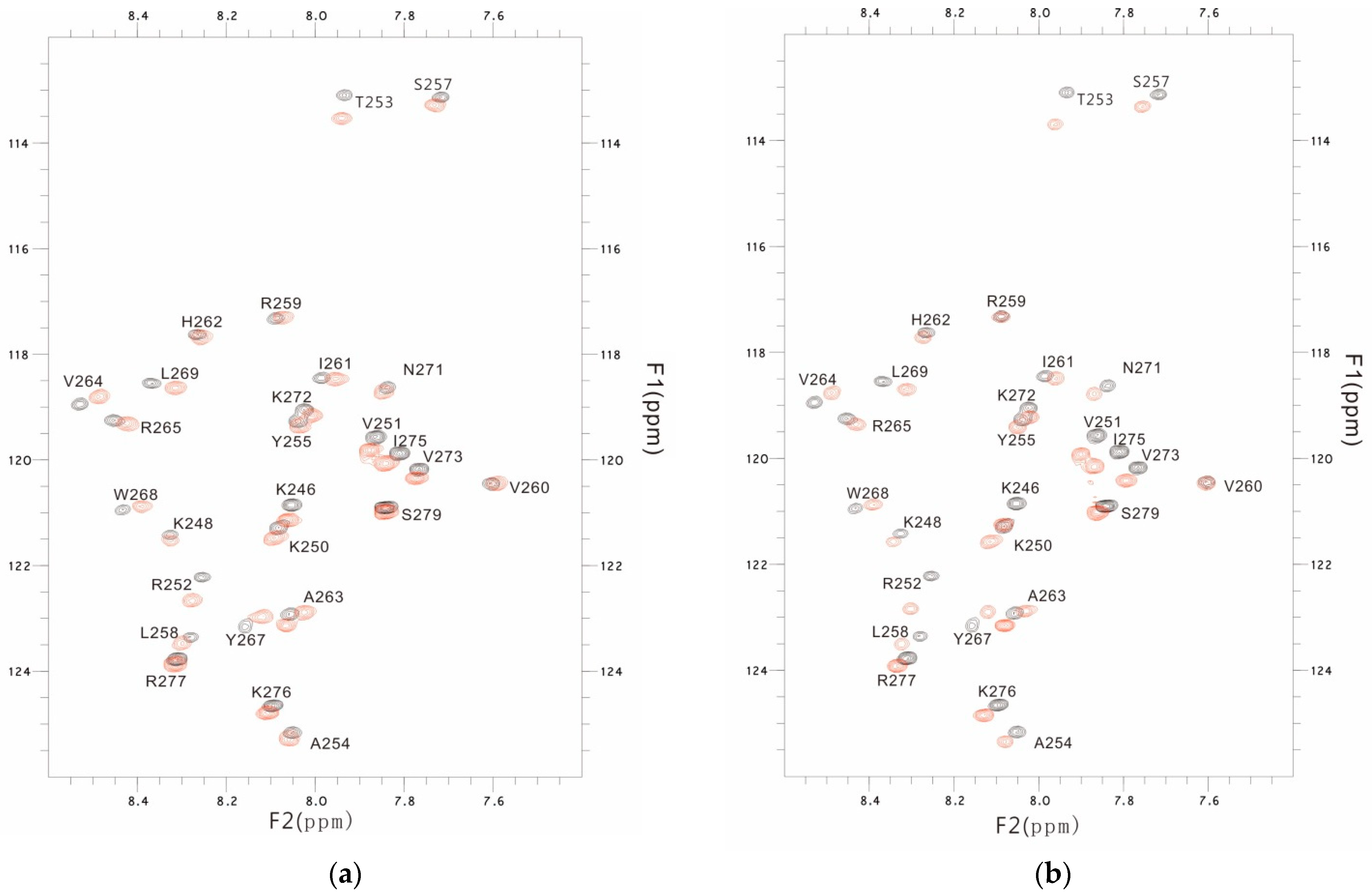
3.3. The PSTD-(6′-SL) Interaction
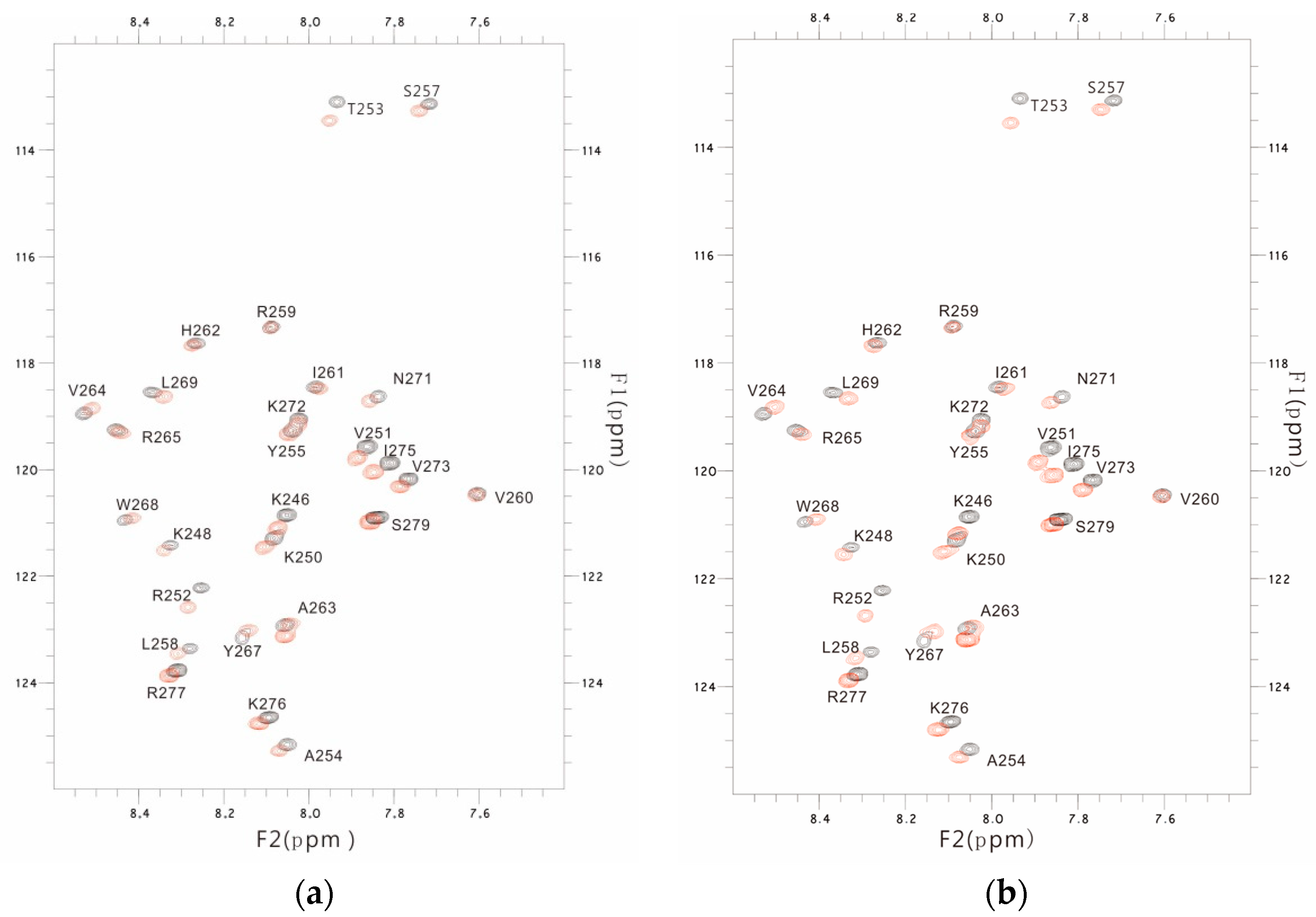
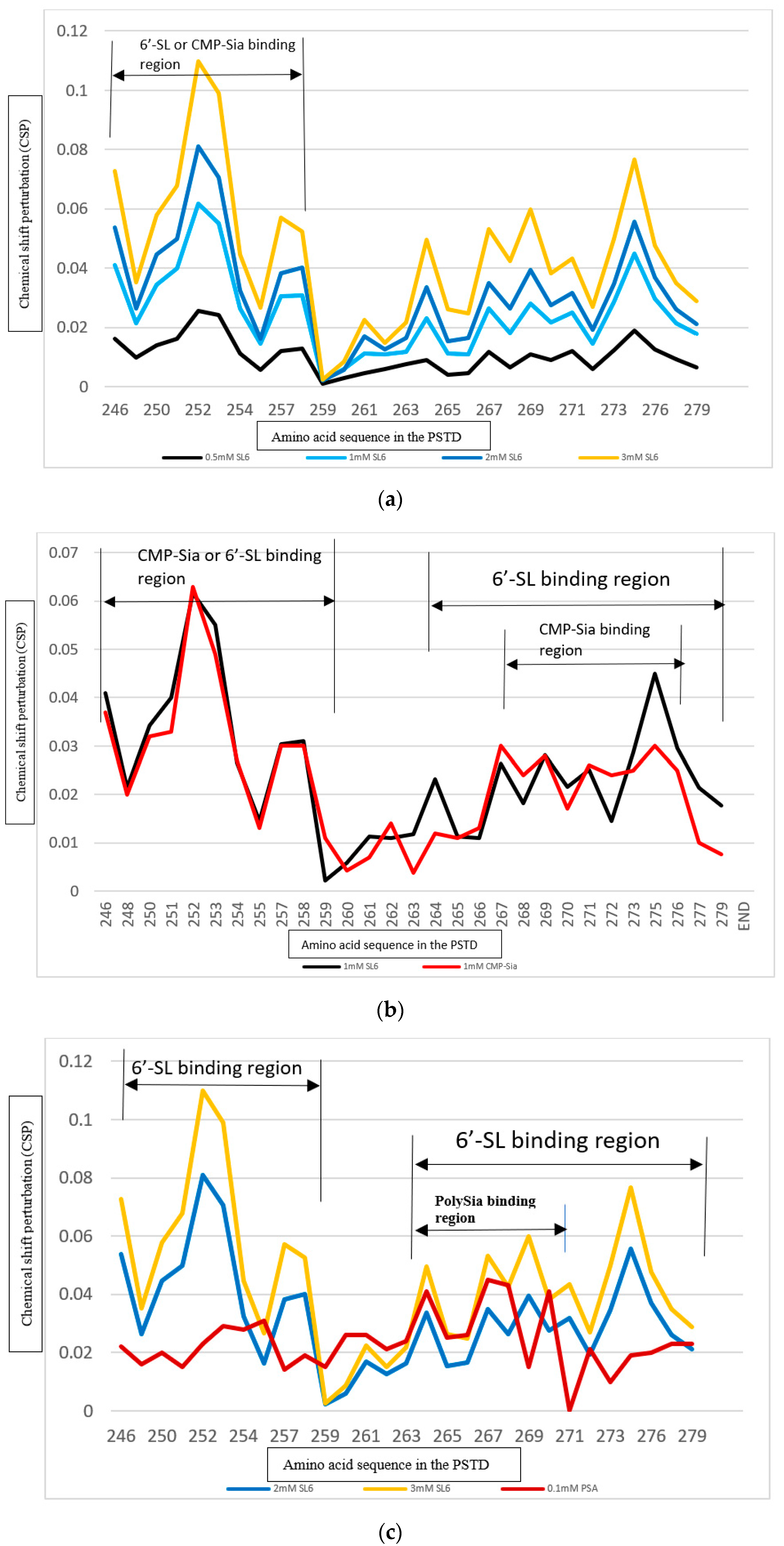
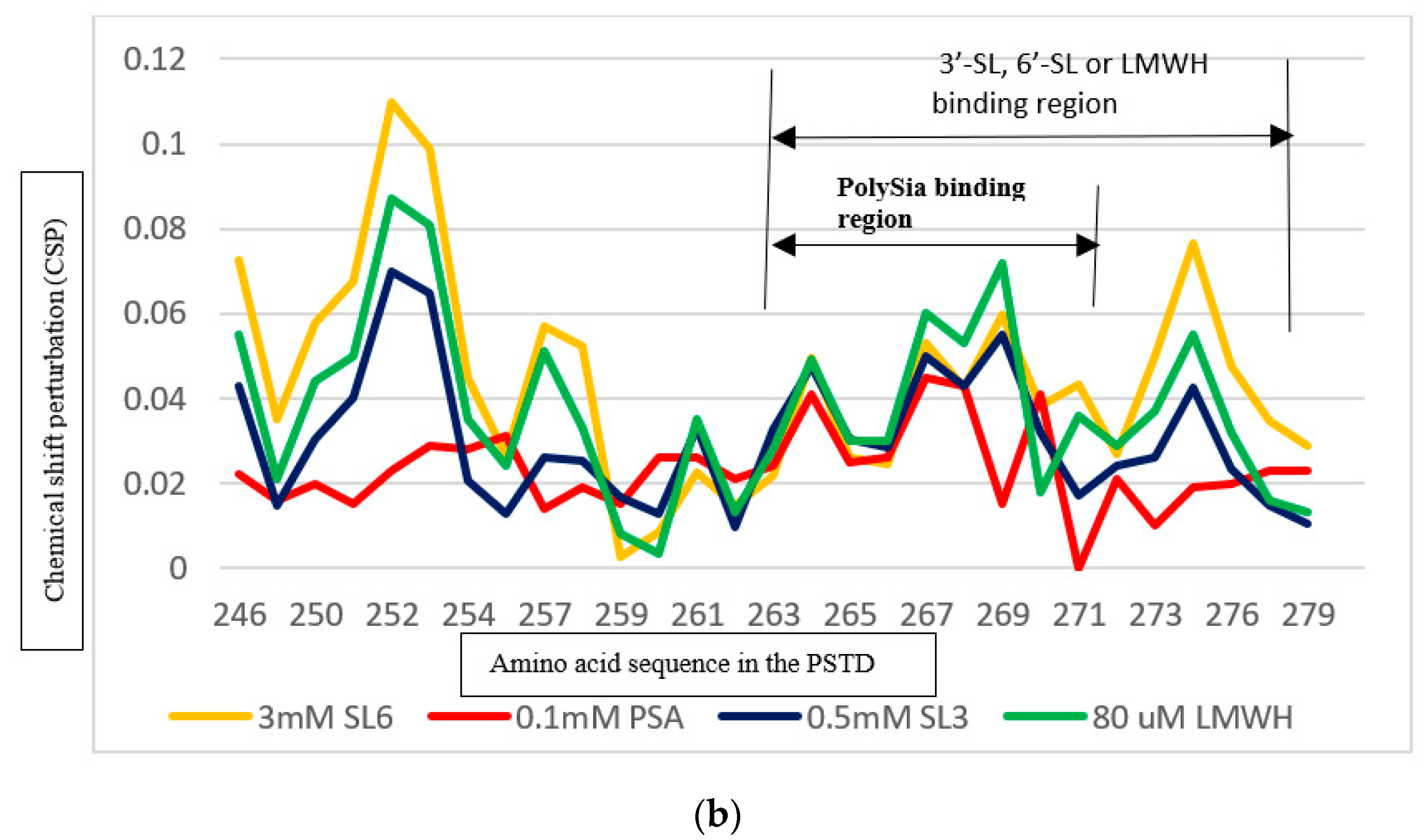
3.4. Comparison of the Inhibition of the SLs and Heparin (LMWH) on Polysialylation
3.4.1. Comparison of the Inhibition of the SLs and Heparin (LMWH) on the PSTD-CMP-Sia Interaction
3.4.2. Comparison of the Inhibition of the SLs and Heparin (LMWH) on the PSTD-polySia Interaction
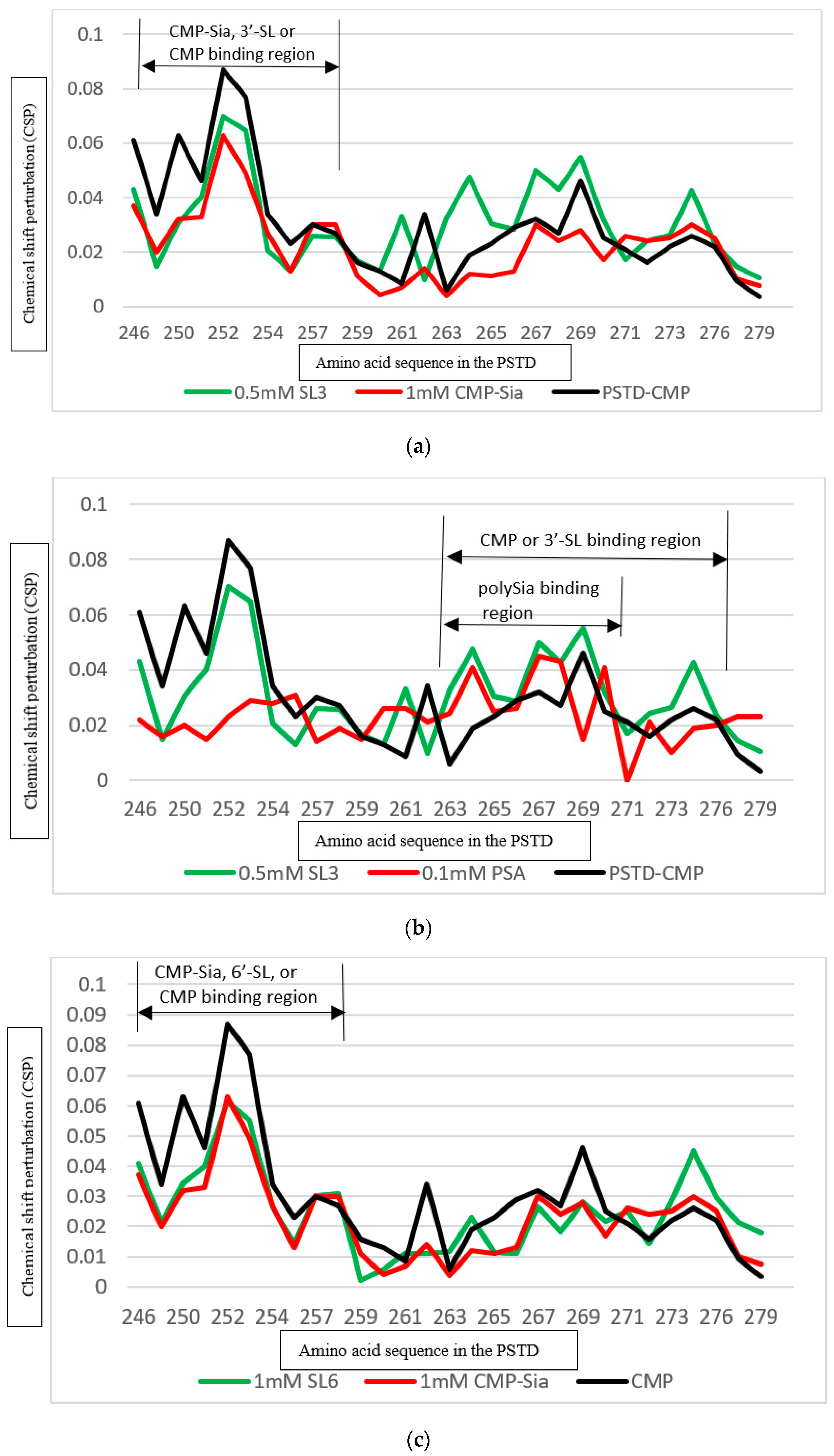
3.5. Comparison of the Inhibition of the SLs and CMP on Polysialylation
3.5.1. Comparison of the Inhibition of the 3′-SL and CMP on the PSTD-(CMP-Sia) Interaction
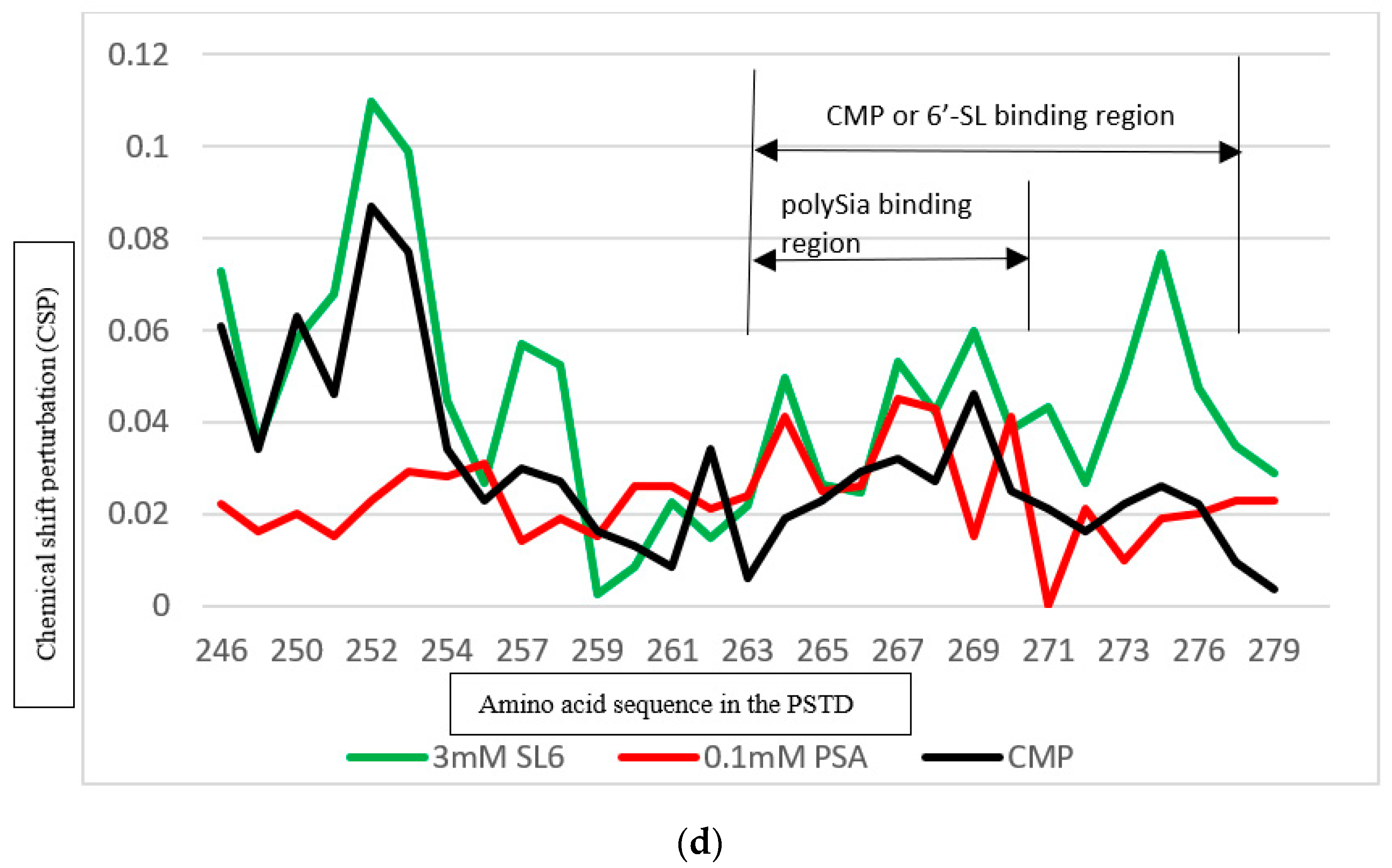
3.5.2. Comparison of the Inhibition of the 6′-SL and CMP on Polysialylation
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| polySia | polysialic acid |
| Sia | mono-sialic acid |
| CMP-Sia | cytidine monophosphate-sialic acid |
| NCAMs | neural cell adhesion molecules |
| polySTs | polysialyltransferases (ST8Sia II (STX) & ST8Sia IV (PST) |
| PSTD | polysialyltransferase domain |
| SL | sialyllactose |
| SMO | sialylated milk oligosaccharide |
| HMO | human milk oligosaccharide |
| CSP | chemical shift perturbation |
References
- Craft, K.M.; Townsend, S.D. The Human Milk Glycome as a Defense Against Infectious Diseases: Rationale, Challenges, and Opportunities. ACS Infect. Dis. 2017, 4, 77–83. [Google Scholar] [CrossRef] [PubMed]
- ten Bruggencate, S.J.; Bovee-Oudenhoven, I.M.; Feitsma, A.L.; van Hoffen, E.; Schoterman, M.H. Functional role and mechanisms of sialyllactose and other sialylated milk oligosaccharides. Nutr. Rev. 2014, 72, 377–389. [Google Scholar] [CrossRef] [PubMed]
- Martín-Sosa, S.; Martín, M.-J.; García-Pardo, L.-A.; Hueso, P. Sialyloligosaccharides in Human and Bovine Milk and in Infant Formulas: Variations with the Progression of Lactation. J. Dairy Sci. 2003, 86, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Ding, J.; Jin, G.; Yu, D.; Yu, L.; Long, Z.; Guo, Z.; Chai, W.; Liang, X. Profiling of Sialylated Oligosaccharides in Mammalian Milk Using Online Solid Phase Extraction-Hydrophilic Interaction Chromatography Coupled with Negative-Ion Electrospray Mass Spectrometry. Anal. Chem. 2018, 90, 3174–3182. [Google Scholar] [CrossRef] [PubMed]
- Facinelli, B.; Marini, E.; Magi, G.; Zampini, L.; Santoro, L.; Catassi, C.; Monachesi, C.; Gabrielli, O.; Coppa, G.V. Breast milk oligo-saccharides: Effects of 2’-fucosyllactose and 6′-sialyllactose on the adhesion of Escherichia coli and Salmonella fyris to Caco-2 cells. J. Matern. Fetal. Neonatal. Med. 2019, 32, 2950–2952. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.-T.; Chen, C.; Newburg, D.S. Utilization of major fucosylated and sialylated human milk oligosaccharides by isolated human gut microbes. Glycobiology 2013, 23, 1281–1292. [Google Scholar] [CrossRef] [PubMed]
- Kawashima, N.; Yoon, S.-J.; Itoh, K.; Nakayama, K.-I. Tyrosine Kinase Activity of Epidermal Growth Factor Receptor Is Regulated by GM3 Binding through Carbohydrate to Carbohydrate Interactions. J. Biol. Chem. 2009, 284, 6147–6155. [Google Scholar] [CrossRef] [PubMed]
- Kuntz, S.; Rudloff, S.; Kunz, C. Oligosaccharides from human milk influence growth-related characteristics of intestinally transformed and non-transformed intestinal cells. Br. J. Nutr. 2008, 99, 462–471. [Google Scholar] [CrossRef]
- Yang, C.; Zhang, P.; Fang, W.; Chen, Y.; Zhang, N.; Qiao, Z.; Troy, F.A.; Wang, B. Molecular Mechanisms Underlying How Sialyllactose Intervention Promotes Intestinal Maturity by Upregulating GDNF Through a CREB-Dependent Pathway in Neonatal Piglets. Mol. Neurobiol. 2019, 56, 7994–8007. [Google Scholar] [CrossRef]
- Chung, T.-W.; Kim, E.-Y.; Kim, S.-J.; Choi, H.-J.; Jang, S.B.; Kim, K.-J.; Ha, S.-H.; Abekura, F.; Kwak, C.-H.; Kim, C.-H.; et al. Sialyllactose suppresses angiogenesis by inhibiting VEGFR-2 activation, and tumor progression. Oncotarget 2017, 8, 58152–58162. [Google Scholar] [CrossRef]
- Alassane-Kpembi, I.; Canlet, C.; Tremblay-Franco, M.; Jourdan, F.; Chalzaviel, M.; Pinton, P.; Cossalter, A.M.; Achard, C.; Castex, M.; Combes, S.; et al. 1H-NMR metabolomics response to a realistic diet contamination with the mycotoxin deoxynivalenol: Effect of probiotics supplementation. Food Chem. Toxicol. 2020, 138, 111222. [Google Scholar] [CrossRef] [PubMed]
- Angata, T.; Varki, A. Chemical diversity in the sialic acids and related r-keto acids: An evolutionary perspective. Chem. Rev. 2002, 102, 439–469. [Google Scholar] [CrossRef] [PubMed]
- Angata, K.; Suzuki, M.; McAuliffe, J.; Ding, Y.; Hindsgaul, O.; Fukuda, M. Differential biosynthesis of polysialic acid on neural cell adhesion molecule (NCAM) and oligosaccharide acceptors by three distinct alpha 2,8-sialyltransferases, ST8Sia IV (PST), ST8Sia II (STX), and ST8Sia III. J. Biol. Chem. 2000, 275, 18594–18601. [Google Scholar] [CrossRef] [PubMed]
- Troy, F.A. Polysialylation: From bacteria to brains. Glycobiology 1992, 2, 5–23. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, A. Book: Biology of Sialic Acid; Springer: Berlin/Heidelberg, Germany, 1995. [Google Scholar]
- Munro, S. Essentials of Glycobiology. Trends Cell Biol. 2000, 10, 552–553. [Google Scholar] [CrossRef]
- Zhou, G.-P.; Huang, R.-B.; Ii, F.A.T. 3D Structural Conformation and Functional Domains of Polysialyltransferase ST8Sia IV Required for Polysialylation of Neural Cell Adhesion Molecules. Protein Pept. Lett. 2015, 22, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Nakata, D.; Zhang, L.; Troy, F.A. Molecular basis for polysialylation: A novel polybasic polysialyltransferase domain (PSTD) of 32 amino acids unique to the α2,8-polysialyltransferases is essential for polysialylation. Glycoconj. J. 2006, 23, 423–436. [Google Scholar] [CrossRef] [PubMed]
- Foley, D.A.; Swartzentruber, K.G.; Colley, K.J. Identification of Sequences in the Polysialyltransferases ST8Sia II and ST8Sia IV That Are Required for the Protein-specific Polysialylation of the Neural Cell Adhesion Molecule, NCAM. J. Biol. Chem. 2009, 284, 15505–15516. [Google Scholar] [CrossRef]
- Bhide, G.P.; Prehna, G.; Ramirez, B.E.; Colley, K.J. The Polybasic Region of the Polysialyltransferase ST8Sia-IV Binds Directly to the Neural Cell Adhesion Molecule, NCAM. Biochemistry 2017, 56, 1504–1517. [Google Scholar] [CrossRef]
- Zhou, G.-P.; Liao, S.-M.; Chen, D.; Huang, R.-B. The Cooperative Effect between Polybasic Region (PBR) and Polysialyltransferase Domain (PSTD) within Tumor-Target Polysialyltranseferase ST8Sia II. Curr. Top. Med. Chem. 2020, 19, 2831–2841. [Google Scholar] [CrossRef]
- Huang, R.-B.; Cheng, D.; Liao, S.-M.; Lu, B.; Wang, Q.-Y.; Xie, N.-Z.; Ii, F.A.T.; Zhou, G.-P. The Intrinsic Relationship Between Structure and Function of the Sialyltransferase ST8Sia Family Members. Curr. Top. Med. Chem. 2017, 17, 2359–2369. [Google Scholar] [CrossRef] [PubMed]
- Liao, S.-M.; Lu, B.; Liu, X.-H.; Lu, Z.-L.; Liang, S.-J.; Chen, D.; Troy, F.A.; Huang, R.-B.; Zhou, G.-P. Molecular Interactions of the Polysialytransferase Domain (PSTD) in ST8Sia IV with CMP-Sialic Acid and Polysialic Acid Required for Polysialylation of the Neural Cell Adhesion Molecule Proteins: An NMR Study. Int. J. Mol. Sci. 2020, 21, 1590. [Google Scholar] [CrossRef] [PubMed]
- Liao, S.-M.; Liu, X.-H.; Peng, L.-X.; Lu, B.; Huang, R.-B.; Zhou, G.-P. Molecular Mechanism of Inhibition of Polysialyltransferase Domain (PSTD) by Heparin. Curr. Top. Med. Chem. 2021, 21, 1113–1120. [Google Scholar] [CrossRef] [PubMed]
- Lu, B.; Liu, X.-H.; Liao, S.-M.; Lu, Z.-L.; Chen, D.; Ii, F.A.T.; Huang, R.-B.; Zhou, G.-P. A Possible Modulation Mechanism of Intramolecular and Intermolecular Interactions for NCAM Polysialylation and Cell Migration. Curr. Top. Med. Chem. 2019, 19, 2271–2282. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.-X.; Liu, X.-H.; Lu, B.; Liao, S.-M.; Zhou, F.; Huang, J.-M.; Chen, D.; Ii, F.A.T.; Zhou, G.-P.; Huang, R.-B. The Inhibition of Polysialyltranseferase ST8SiaIV Through Heparin Binding to Polysialyltransferase Domain (PSTD). Med. Chem. 2019, 15, 486–495. [Google Scholar] [CrossRef] [PubMed]
- Lu, B.; Liao, S.-M.; Liu, X.-H.; Liang, S.-J.; Huang, J.; Lin, M.; Meng, L.; Wang, Q.-Y.; Huang, R.-B.; Zhou, G.-P. The NMR studies of CMP inhibition of polysialylation. J. Enzym. Inhib. Med. Chem. 2023, 38, 2248411. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.-P. The Latest Researches of Enzyme Inhibition and Multi-Target Drug Predictors in Medicinal Chemistry. Med. Chem. 2019, 15, 572–573. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.-P. Current Advances of Drug Target Research in Medicinal Chemistry. Curr. Top. Med. Chem. 2019, 19, 2269–2270. [Google Scholar] [CrossRef] [PubMed]
- Duncan, P.I.; Raymond, F.; Fuerholz, A.; Sprenger, N. Sialic Acid Utilisation and Synthesis in the Neonatal Rat Revisited. PLoS ONE 2009, 4, e8241. [Google Scholar] [CrossRef]
- Sprenger, N.; Duncan, P.I. Sialic Acid Utilization. Adv. Nutr. Int. Rev. J. 2012, 3, 392S–397S. [Google Scholar] [CrossRef]
- Lu, B.; Liao, S.-M.; Liang, S.-J.; Peng, L.-X.; Li, J.-X.; Liu, X.-H.; Huang, R.-B.; Zhou, G.-P. The Bifunctional Effects of Lactoferrin (LFcinB11) in Inhibiting Neural Cell Adhesive Molecule (NCAM) Polysialylation and the Release of Neutrophil Extracellular Traps (NETs). Int. J. Mol. Sci. 2024, 25, 4641. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.K.; Zhou, G.P.; Kupferman, J.; Surks, H.K.; Christensen, E.N.; Chou, J.J.; Mendelsohn, M.E.; Rigby, A.C. Probing the interaction between the coiled coil leucine zipper of cGMP-dependent protein kinase Ialpha and the C terminus of the myosin binding subunit of the myosin light chain phosphatase. J. Biol. Chem. 2008, 283, 32860–32869. [Google Scholar] [CrossRef] [PubMed]
- Berardi, M.J.; Shih, W.M.; Harrison, S.C.; Chou, J.J. Mitochondrial uncoupling protein 2 structure determined by NMR molecular fragment searching. Nature 2011, 476, 109–113. [Google Scholar] [CrossRef]
- OuYang, B.; Xie, S.; Berardi, M.J.; Zhao, X.; Dev, J.; Yu, W.; Sun, B.; Chou, J.J. Unusual architecture of the p7 channel from hepatitis C virus. Nature 2013, 498, 521–525. [Google Scholar] [CrossRef]
- Oxenoid, K.; Dong, Y.; Cao, C.; Cui, T.; Sancak, Y.; Markhard, A.L.; Grabarek, Z.; Kong, L.; Liu, Z.; Ouyang, B.; et al. Architecture of the mitochondrial calcium uniporter. Nature 2016, 533, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; OuYang, B.; Chou, J.J. Critical Effect of the Detergent: Protein Ratio on the Formation of the Hepatitis C Virus p7 Channel. Biochemistry 2019, 58, 3834–3837. [Google Scholar] [CrossRef]
- Fu, Q.; Fu, T.M.; Cruz, A.C.; Sengupta, P.; Thomas, S.K.; Wang, S.; Siegel, R.M.; Wu, H.; Chou, J.J. Structural Basis and Functional Role of Intramembrane Trimerization of the Fas/CD95 Death Receptor. Mol Cell. 2016, 61, 602–613. [Google Scholar] [CrossRef]
- Du, G.; Zhao, L.; Zheng, Y.; Belfetmi, A.; Cai, T.; Xu, B.; Heyninck, K.; Heede, K.V.D.; Buyse, M.-A.; Fontana, P.; et al. Autoinhibitory structure of preligand association state implicates a new strategy to attain effective DR5 receptor activation. Cell Res. 2023, 33, 131–146. [Google Scholar] [CrossRef] [PubMed]
- Cai, T.; Capello, R.L.; Pi, X.; Wu, H.; Chou, J.J. Structural basis of γ chain family receptor sharing at the membrane level. Science 2023, 381, 569–576. [Google Scholar] [CrossRef]
- Selvaratnam, R.; Chowdhury, S.; VanSchouwen, B.; Melacini, G. Mapping allostery through the covariance analysis of NMR chemical shifts. Proc. Natl. Acad. Sci. USA 2011, 108, 6133–6138. [Google Scholar] [CrossRef]
- Selvaratnam, R.; Mazhab-Jafari, M.T.; Das, R.; Melacini, G. The Auto-Inhibitory Role of the EPAC Hinge Helix as Mapped by NMR. PLoS ONE 2012, 7, e48707. [Google Scholar] [CrossRef]
- Boulton, S.; Melacini, G. Advances in NMR Methods To Map Allosteric Sites: From Models to Translation. Chem. Rev. 2016, 116, 6267–6304. [Google Scholar] [CrossRef]
- Krishnamoorthy, J.; Yu, V.C.; Mok, Y.K. Auto-FACE: An NMR based binding site mapping program for fast chemical exchange pro-tein-ligand systems. PLoS ONE 2010, 5, e8943. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, A.; Konuma, T.; Yanaka, S.; Sugase, K. Quantitative analysis of protein–ligand interactions by NMR. Prog. Nucl. Magn. Reson. Spectrosc. 2016, 96, 47–57. [Google Scholar] [CrossRef]
- González-Ruiz, D.; Gohlke, H. Steering protein-ligand docking with quantitative NMR chemical shift perturbations. J. Chem. Inf. Model. 2009, 49, 2260–2271. [Google Scholar] [CrossRef] [PubMed]
- Markin, C.J.; Spyracopoulos, L. Accuracy and precision of protein–ligand interaction kinetics determined from chemical shift titrations. J. Biomol. NMR 2012, 54, 355–376. [Google Scholar] [CrossRef]
- Loria, J.P.; Berlow, R.B.; Watt, E.D. Characterization of Enzyme Motions by Solution NMR Relaxation Dispersion. Accounts Chem. Res. 2008, 41, 214–221. [Google Scholar] [CrossRef]
- Joseph, P.R.; Poluri, K.M.; Sepuru, K.M.; Rajarathnam, K. Characterizing protein-glycosaminoglycan interactions using solution NMR spectroscopy. Methods Mol. Biol. 2015, 1229, 325–333. [Google Scholar]
- Bjorndahl, T.C.; Zhou, G.P.; Liu, X.; Perez-Pineiro, R.; Semenchenko, V.; Saleem, F.; Acharya, S.; Bujold, A.; Sobsey, C.A.; Wishart, D.S. Detailed biophysical characterization of the acid-induced PrP(c) to PrP(beta) conversion process. Biochemistry 2011, 50, 1162–1173. [Google Scholar] [CrossRef]
- Xu, J.; Sarma, A.V.S.; Wei, Y.; Beamer, L.J.; Van Doren, S.R. Multiple Ligand-Bound States of a Phosphohexomutase Revealed by Principal Component Analysis of NMR Peak Shifts. Sci. Rep. 2017, 7, 5343. [Google Scholar] [CrossRef]
- Sarma, A.V.S.; Anbanandam, A.; Kelm, A.; Mehra-Chaudhary, R.; Wei, Y.; Qin, P.; Lee, Y.; Berjanskii, M.V.; Mick, J.A.; Beamer, L.J.; et al. Solution NMR of a 463-Residue Phosphohexomutase: Domain 4 Mobility, Substates, and Phosphoryl Transfer Defect. Biochemistry 2012, 51, 807–819. [Google Scholar] [CrossRef]
- Skeensa, E.; Lisia, G.P. Analysis of coordinated NMR chemical shifts to map allosteric regulatory networks in proteins. Methods 2023, 209, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Williamson, M.P. Using chemical shift perturbation to characterise ligand binding. Prog. Nucl. Magn. Reson. Spectrosc. 2013, 73, 1–16. [Google Scholar] [CrossRef]
- Al-Saraireh, Y.M.J.; Sutherland, M.; Springett, B.R.; Freiberger, F.; Morais, G.R.; Loadman, P.M.; Errington, R.J.; Smith, P.J.; Fukuda, M.; Gerardy-Schahn, R.; et al. Pharmacological Inhibition of polysialyltransferase ST8SiaII Modulates Tumour Cell Migration. PLoS ONE 2013, 8, e73366. [Google Scholar] [CrossRef]
- Ortiz, A.I.; Reglero, A.; Rodriguez-Aparicio, L.B.; Luengo, J.M. In vitro synthesis of colominic acid by membrane-bound sialyl-transferase of Escherichia coli K-235. Kinetic properties of this enzyme and inhibition by CMP and other cytidine nucleotides. Eur. J. Biochem. 1989, 178, 741–749. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.-P.; Huang, R.-B. The Graphical Studies of the Major Molecular Interactions for Neural Cell Adhesion Molecule (NCAM) Polysialylation by Incorporating Wenxiang Diagram into NMR Spectroscopy. Int. J. Mol. Sci. 2022, 23, 15128. [Google Scholar] [CrossRef]
- Zhou, G.-P.; Chen, D.; Liao, S.; Huang, R.-B. Recent Progresses in Studying Helix-Helix Interactions in Proteins by Incorporating the Wenxiang Diagram into the NMR Spectroscopy. Curr. Top. Med. Chem. 2015, 16, 581–590. [Google Scholar] [CrossRef]
- Zhou, G.-P. The disposition of the LZCC protein residues in wenxiang diagram provides new insights into the protein–protein interaction mechanism. J. Theor. Biol. 2011, 284, 142–148. [Google Scholar] [CrossRef] [PubMed]
- Jungck, J.R.; Cebeci, M. Wenxiang 3.0: Evolutionary Visualization of α, π, and 3/10 Helices. Evol. Bioinform. 2022, 18, 11769343221101014. [Google Scholar] [CrossRef]
- Chou, K.-C. The Significant and Profound Impacts of Chou’s “wenxiang” Diagram. Voice Publ. 2020, 06, 102–103. [Google Scholar] [CrossRef]
- Chou, K.-C.; Lin, W.-Z.; Xiao, X. Wenxiang: A web-server for drawing wenxiang diagrams. Nat. Sci. 2011, 03, 862–865. [Google Scholar] [CrossRef]
- Chou, K.-C.; Zhang, C.-T.; Maggiora, G.M. Disposition of amphiphilic helices in heteropolar environments. Proteins Struct. Funct. Genet. 1997, 28, 99–108. [Google Scholar] [CrossRef]
- Sprenger, N.; Lee, L.Y.; De Castro, C.A.; Steenhout, P.; Thakkar, S.K. Longitudinal change of selected human milk oligosaccharides and association to infants’ growth, an observatory, single center, longitudinal cohort study. PLoS ONE 2017, 12, e0171814. [Google Scholar] [CrossRef]
- Hauser, J.; Pisa, E.; Arias Vasquez, A.; Tomasi, F.; Traversa, A.; Chiodi, V.; Martin, F.P.; Sprenger, N.; Lukjancenko, O.; Zollinger, A.; et al. Sialylated human milk oligosaccharides program cognitive development through a non-genomic transmission mode. Mol. Psychiatry 2021, 26, 2854–2871. [Google Scholar] [CrossRef]
- Gray, E.; Hogwood, J.; Mulloy, B. The anticoagulant and antithrombotic mechanisms of heparin. Handb. Exp. Pharmacol. 2012, 207, 43–61. [Google Scholar]
- EFSA Panel on Nutrition, Novel Foods and Food Allergens (NDA); Turck, D.; Bohn, T.; Castenmiller, J.; De Henauw, S.; Hirsch-Ernst, K.I.; Maciuk, A.; Mangelsdorf, I.; McArdle, H.J.; Naska, A.; et al. Safety of 6′-sialyllactose (6′-SL) sodium salt produced by derivative strains of Escherichia coli BL21 (DE3) as a novel food pursuant to Regulation (EU) 2015/2283. EFSA J. 2022, 20, e07645. [Google Scholar] [CrossRef]
- EFSA Panel on Nutrition, Novel Foods and Food Allergens (NDA); Turck, D.; Bohn, T.; Castenmiller, J.; De Henauw, S.; Hirsch-Ernst, K.I.; Maciuk, A.; Mangelsdorf, I.; McArdle, H.J.; Naska, A.; et al. Safety of 3′-sialyllactose (3′-SL) sodium salt produced by derivative strains of Escherichia coli BL21 (DE3) as a Novel Food pursuant to Regulation (EU) 2015/2283. EFSA J. 2022, 20, e07331. [Google Scholar] [CrossRef]
- Schauer, R.; Kamerling, J.P. Exploration of the Sialic Acid World. Adv. Carbohydr. Chem. Biochem. 2018, 75, 1–213. [Google Scholar]
- Zhang, R.; Loers, G.; Schachner, M.; Boelens, R.; Wienk, H.; Siebert, S.; Eckert, T.; Kraan, S.; Rojas-Macias, M.A.; Lutteke, T.; et al. Molecular Basis of the Receptor Interactions of Polysialic Acid (polySia), polySia Mimetics, and Sulfated Polysaccharides. ChemMedChem 2016, 11, 990–1002. [Google Scholar] [CrossRef]
- Fu, Q.; Piai, A.; Chen, W.; Xia, K.; Chou, J.J. Structure determination protocol for transmembrane domain oligomers. Nat. Protoc. 2019, 14, 2483–2520. [Google Scholar] [CrossRef]
- Schnell, J.R.; Zhou, G.P.; Zweckstetter, M.; Rigby, A.C.; Chou, J.J. Rapid and accurate structure determination of coiled-coil domains using NMR dipolar couplings: Application to cGMP-dependent protein kinase Ialpha. Protein Sci. 2005, 14, 2421–2428. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Dai, G.; Cheng, Y.B.; Qi, X.; Geng, M.Y. Polysialylation promotes neural cell adhesion molecule-mediated cell migration in a fibroblast growth factor receptor-dependent manner, but independent of adhesion capability. Glycobiology 2011, 21, 1010–1018. [Google Scholar] [CrossRef] [PubMed]
- Chou, K.C.; Liu, W.; Maggiora, G.M.; Zhang, C.T. Prediction and Classification of Domain Structural Classes. Proteins Struct. Funct. Bioinform. 1998, 31, 97–103. [Google Scholar] [CrossRef]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Saisho, S.; Takashima, S.; Ohsumi, S.; Saeki, H.; Aogi, K.; Saeki, T.; Mandai, K.; Iwata, S.; Takeda, T. Two cases with long-term disease-free survival after resection and radiotherapy for solitary brain metastasis from breast cancer with extensive nodal metastases. Breast Cancer 2005, 12, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Chou, T.M.; Demer, J.L. Isolated inferior rectus palsy caused by a metastasis to the oculomotor nucleus. Am. J. Ophthalmol. 1998, 126, 737–740. [Google Scholar] [CrossRef] [PubMed]
- Koller, M.; Ram, Z.; Findler, G.; Lipshitz, M. Brain metastasis: A rare manifestation of adenoid cystic carcinoma of the breast. Surg. Neurol. 1986, 26, 470–472. [Google Scholar] [CrossRef] [PubMed]
- Paget, S. The distribution of secondary growths in cancer of the breast. Cancer Metastasis Rev. 1989, 8, 98–101. [Google Scholar] [CrossRef]
- Talmadge, J.E.; Fidler, I.J. AACR centennial series: The biology of cancer metastasis: Historical perspective. Cancer Res. 2010, 70, 5649–5669. [Google Scholar] [CrossRef]
- Virchow, R. Cellular pathology. As based upon physiological and pathological histology. Lecture XVI-Atheromatous affection of arteries. 1858. Nutr. Rev. 1989, 47, 23–25. [Google Scholar] [CrossRef]
- Langley, R.R.; Fidler, I.J. Tumor cell-organ microenvironment interactions in the pathogenesis of cancer metastasis. Endocr. Rev. 2007, 28, 297–321. [Google Scholar] [CrossRef] [PubMed]
- Ribatti, D.; Mangialardi, G.; Vacca, A. Stephen Paget and the ‘seed and soil’ theory of metastatic dissemination. Clin. Exp. Med. 2006, 6, 145–149. [Google Scholar] [CrossRef] [PubMed]
- Di Micco, R.; Santurro, L.; Gasparri, M.L.; Zuber, V.; Fiacco, E.; Gazzetta, G.; Smart, C.E.; Valentini, A.; Gentilini, O.D. Rare sites of breast cancer metastasis: A review. Transl. Cancer Res. 2019, 8, S518–S552. [Google Scholar] [CrossRef] [PubMed]
- Cichoń, S.; Anielski, R.; Konturek, A.; Barczyński, M.; Cichoń, W. Metastases to the thyroid gland: Seventeen cases operated on in a single clinical center. Langenbecks Arch. Surg. 2006, 391, 581–587. [Google Scholar] [CrossRef]
- Vaynberg, J.; Qin, J. Weak protein-protein interactions as probed by NMR spectroscopy. Trends Biotechnol. 2006, 24, 22–27. [Google Scholar] [CrossRef]
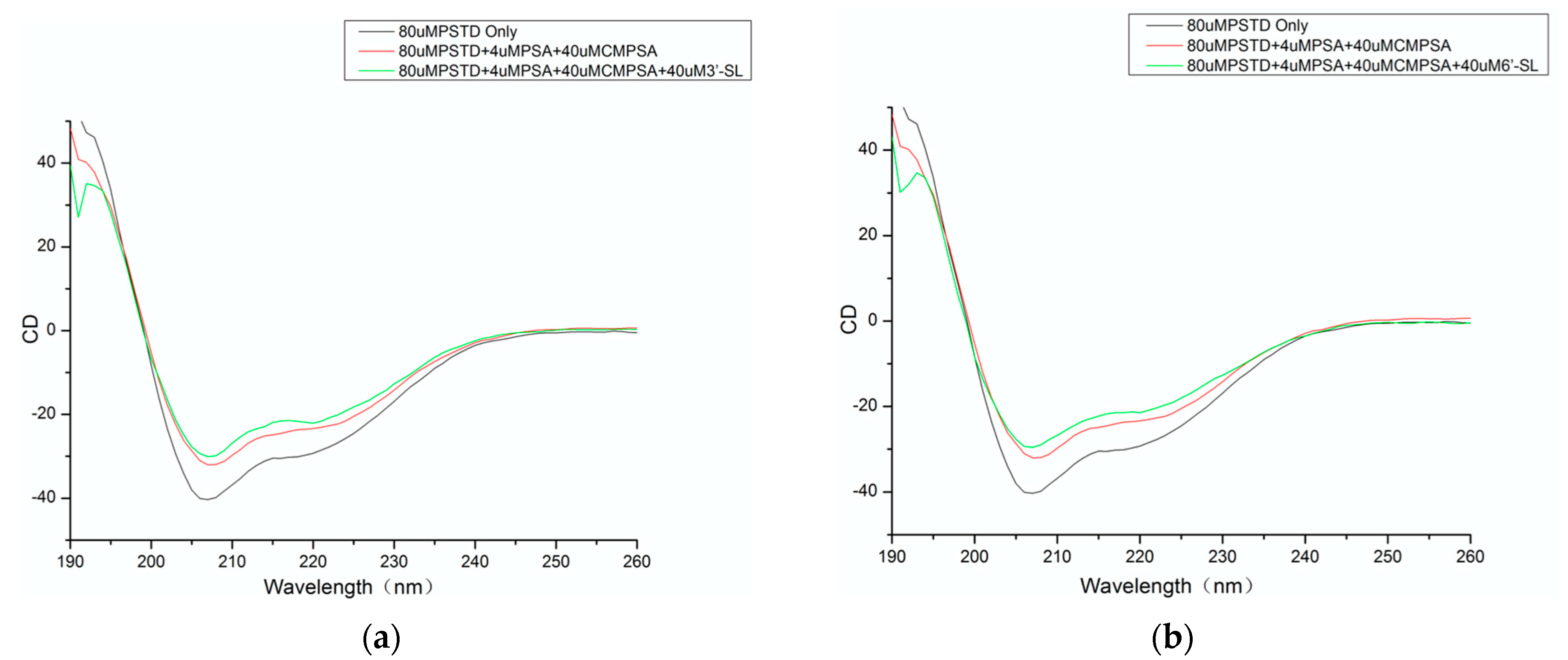



| The helix content of the PSTD alone (%): 26.5 | The helix content of the PSTD after CMP-Sia and polySia were added (%): 18.4 |
| 26.5 | The helix content of the PSTD after CMP-Sia, polySia and 3′-SL were added (%): 17.7 |
| 26.5 | The helix content of the PSTD after CMP-Sia, polySia and 3′-SL were added (%): 14.5 |
| Sialylactose (SL) interacted with the PSTD | SL’s concentration when CSP values of PSTD-SL binding are close to that of PSTD-(CMP-Sia) binding in same amino acid range of the PSTD (mM) | SL’s concentration when CSP values of PSTD-SL binding are close to that of PSTD-polySia binding in same amino acid range of the PSTD (mM) | The maximum concentration of the SL in human milk during 1st month of lactation (mM) [9] |
| 3′-SL | 0.5 | 0.5 | Ca. 0.5 |
| 6′-SL | 1.0 | Ca. 3.0 | Ca. 1.0 |
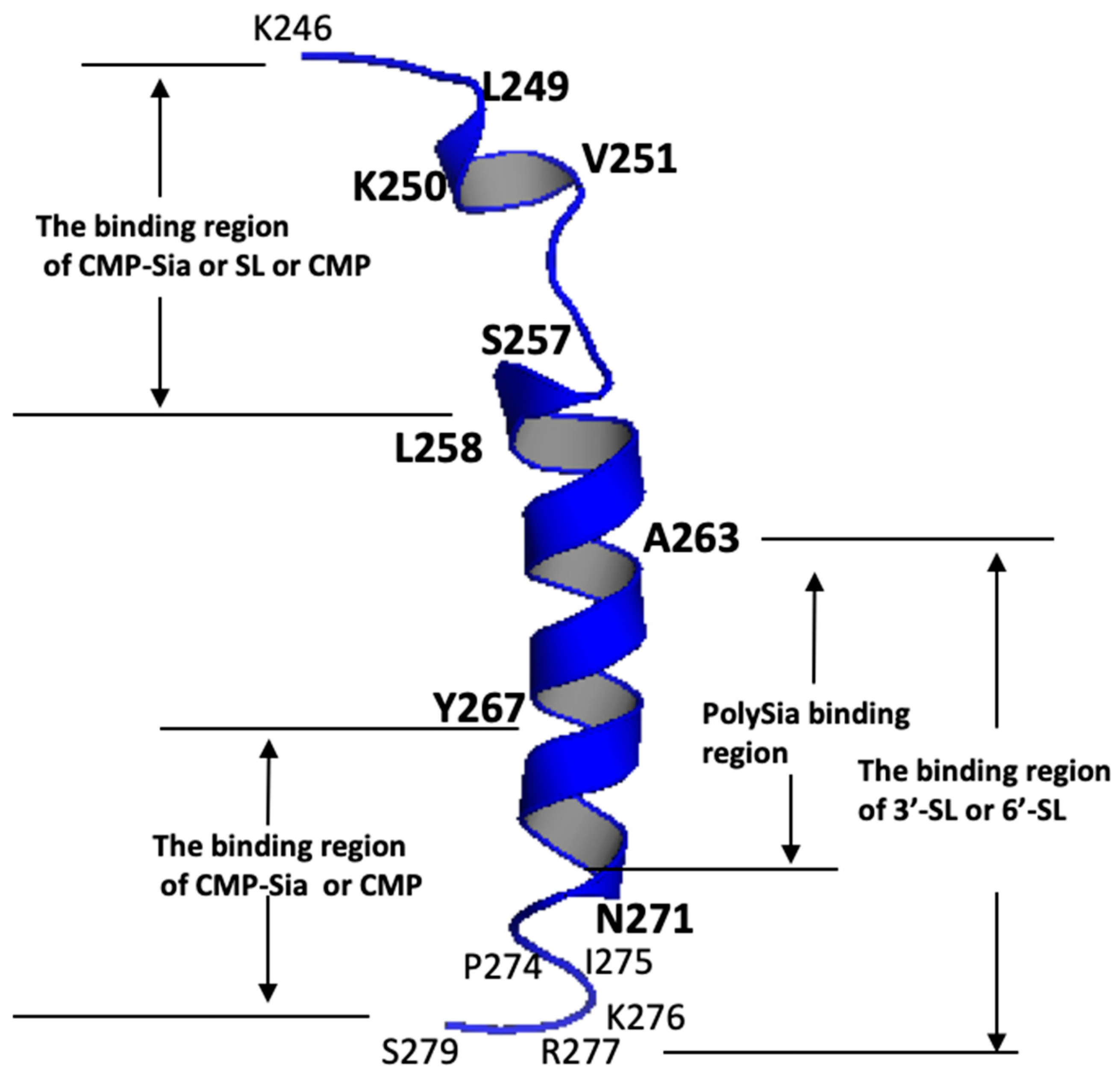
| Ligands Binding to the PSTD | The Main Binding Regions of the Ligands in the PSTD | The Maximum CSPs of Each Binding Region |
|---|---|---|
| CMP-Sia | K246–S257 V267–K276 | 0.070 0.026 |
| polySia | A263–N271 | 0.049 |
| Heparin LMWH | K246–S257 A263–R277 | 0.087 0.072 |
| CMP | K246–S257 A263–R277 | 0.087 0.046 |
| 3′-SL (0.5 mM) | K246–S257 A263–R277 | 0.070 0.043 |
| 6′-SL (1 mM) | K246–S257 A263–R277 | 0.070 0.026 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, B.; Liao, S.-M.; Liang, S.-J.; Li, J.-X.; Liu, X.-H.; Huang, R.-B.; Zhou, G.-P. NMR Studies of the Interactions between Sialyllactoses and the Polysialytransferase Domain for Polysialylation Inhibition. Curr. Issues Mol. Biol. 2024, 46, 5682-5700. https://doi.org/10.3390/cimb46060340
Lu B, Liao S-M, Liang S-J, Li J-X, Liu X-H, Huang R-B, Zhou G-P. NMR Studies of the Interactions between Sialyllactoses and the Polysialytransferase Domain for Polysialylation Inhibition. Current Issues in Molecular Biology. 2024; 46(6):5682-5700. https://doi.org/10.3390/cimb46060340
Chicago/Turabian StyleLu, Bo, Si-Ming Liao, Shi-Jie Liang, Jian-Xiu Li, Xue-Hui Liu, Ri-Bo Huang, and Guo-Ping Zhou. 2024. "NMR Studies of the Interactions between Sialyllactoses and the Polysialytransferase Domain for Polysialylation Inhibition" Current Issues in Molecular Biology 46, no. 6: 5682-5700. https://doi.org/10.3390/cimb46060340
APA StyleLu, B., Liao, S.-M., Liang, S.-J., Li, J.-X., Liu, X.-H., Huang, R.-B., & Zhou, G.-P. (2024). NMR Studies of the Interactions between Sialyllactoses and the Polysialytransferase Domain for Polysialylation Inhibition. Current Issues in Molecular Biology, 46(6), 5682-5700. https://doi.org/10.3390/cimb46060340






