Review on Long Non-Coding RNAs as Biomarkers and Potentially Therapeutic Targets for Bacterial Infections
Abstract
1. Introduction
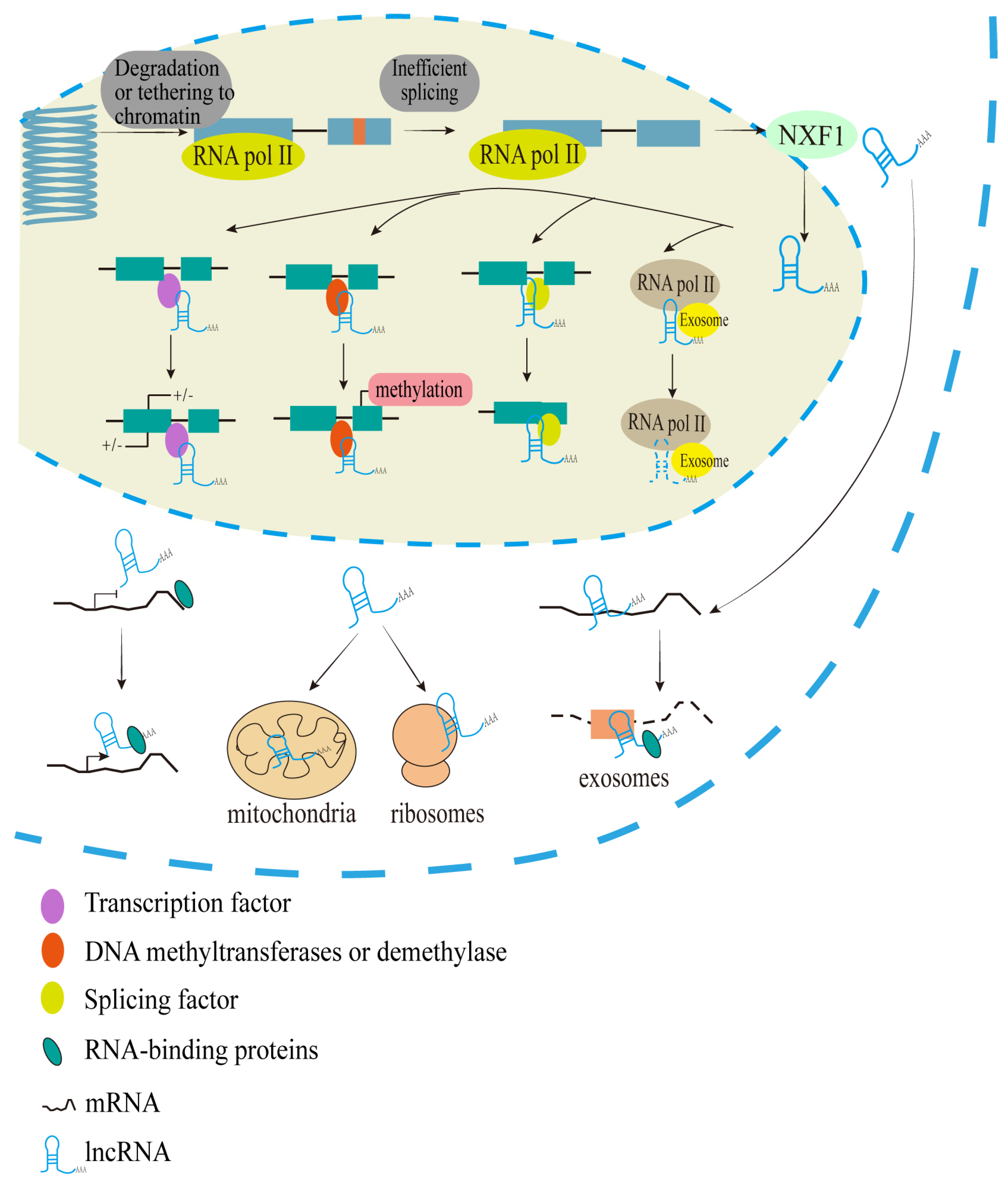
2. Biogenesis, Location, and Molecular Regulation of lncRNAs
3. LncRNAs as Biomarkers for the Occurrence and Development of Bacterial Invasion
4. LncRNAs as Probable Therapeutic Targets for Bacterial Infections
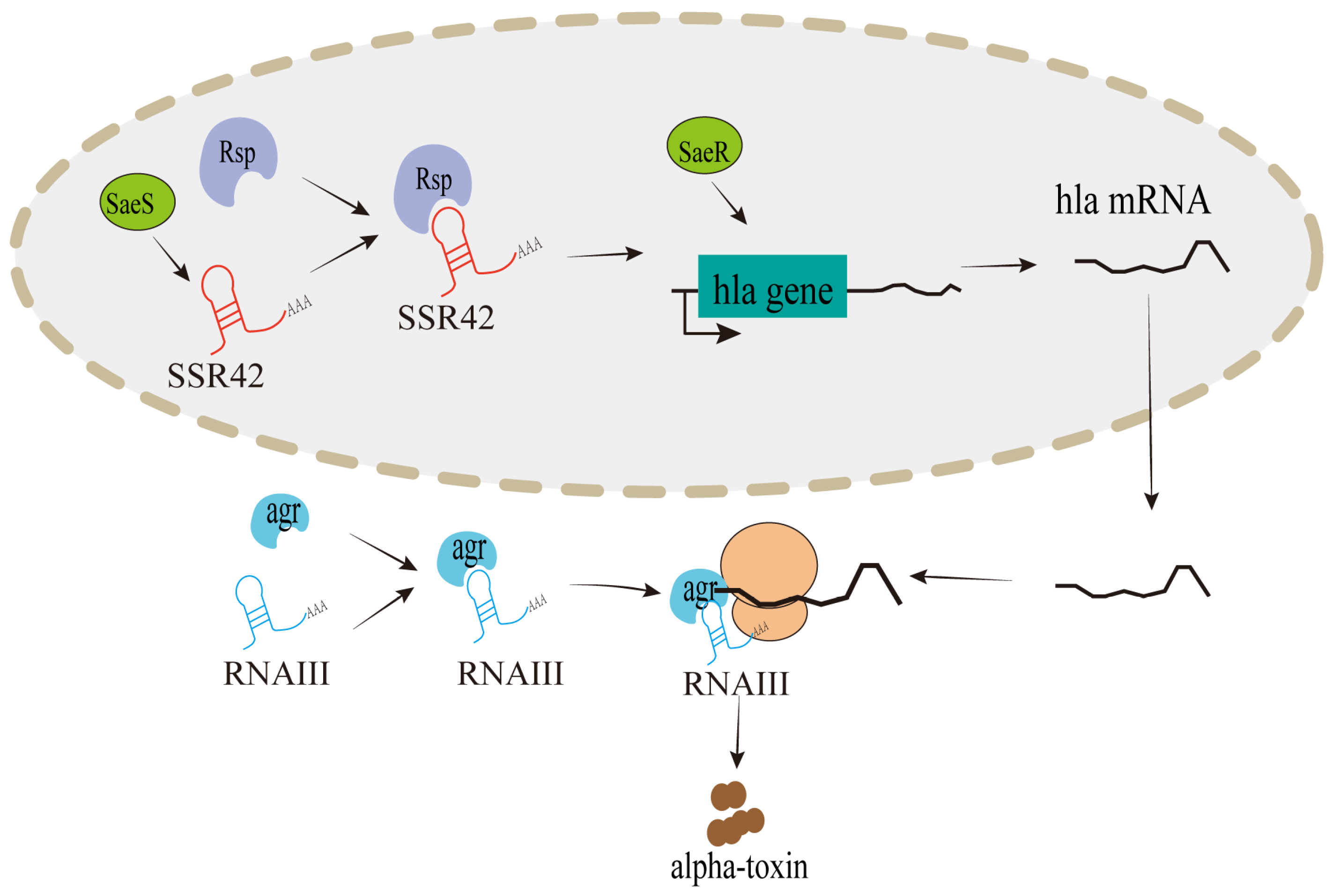
4.1. Targeted Regulation of Pathogenicity by lncRNAs in Bacterial Infections
4.2. Targeted Regulation in Cell Lesions Induced by Bacterial Stimulation
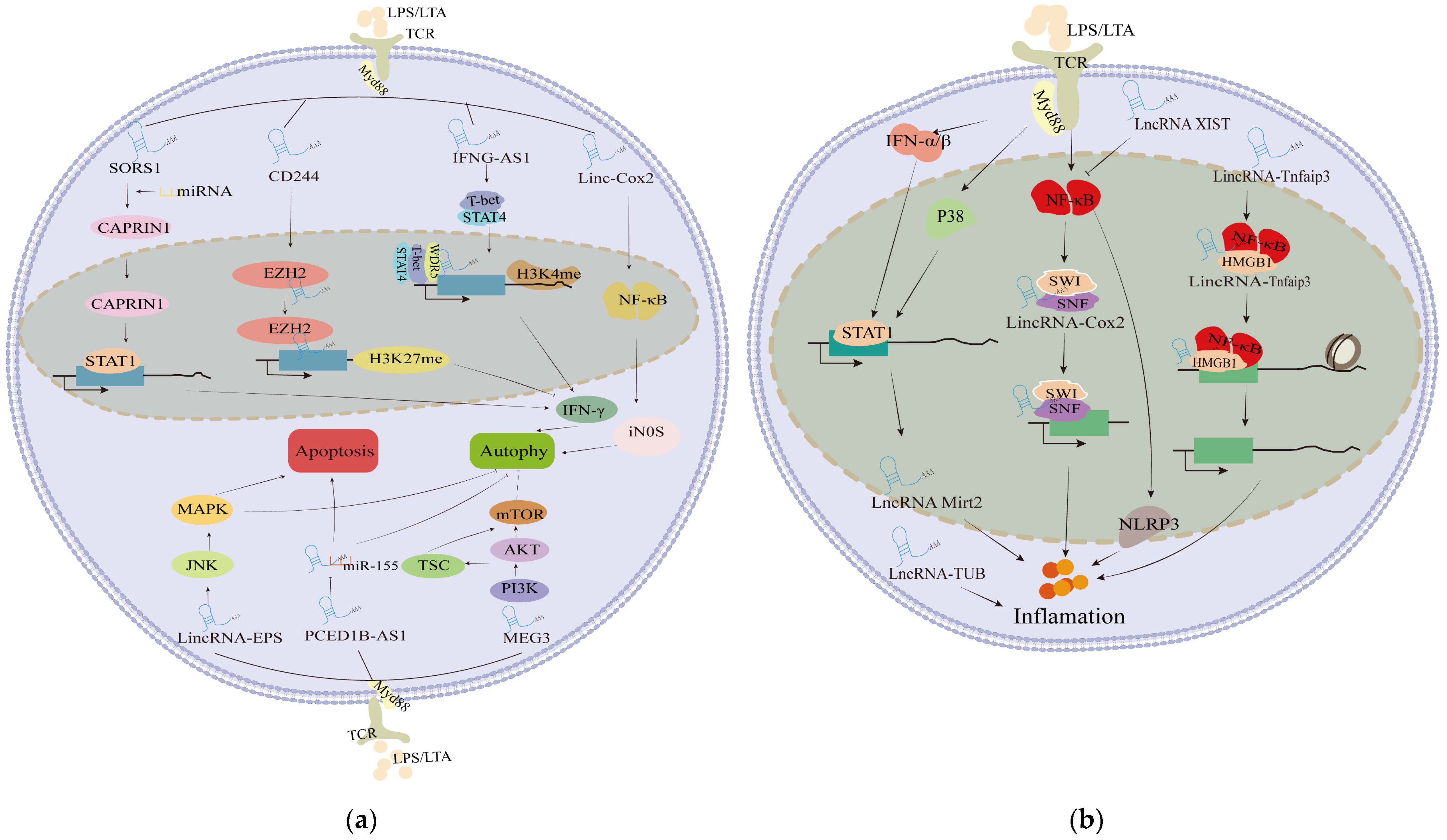
4.3. Targeted Mediation of lncRNAs Inflammation Process Induced by Bacterial Infections
4.4. Targeted Modulation of lncRNAs in the Immune System of the Antibacterial Parade

| Pathogen | LncRNA | Regulation | Target Cells | Mechanisms | References |
|---|---|---|---|---|---|
| S. aureus | SSR42 | Up | Encoded by bacteria | Involves in S. aureus hemolysis induced by antibiotics | [50] |
| LINC00847 | Up | Renal cancer cells | Arrests cell cycle and results in activation of apoptosis through regulatory lncRNA and tst gene | [89] | |
| RNAIII | Up | Encoded by bacteria | Regulates staphylococcal alpha-toxin at the translational level through base pairing with HLA mRNA | [90] | |
| lnc-AFTR | Down | S. aureus-induced MAC-T cells and mastitis tissues | Inhibits apoptosis and inflammation by blocking FAS mRNA translation | [91] | |
| L. monocytogenes | SROS1 | Down | Macrophage | Degraded by upregulated miR-1 to release RBP CAPRIN1, thus improving the stabilization of Stat1 mRNA | [71] |
| lincRNA-EPS | Down | Macrophage | Interacts with hnRNPL to repress immune-responsive gene-1 expression | [72] | |
| lincRNA-Cox2 | Up | Macrophage | Regulates the assembly of NF-κB subunits to the SWI/SNF complex, thereby promoting late inflammatory gene transcription | [92,110] | |
| Morrbid | Up | Eosinophils, Macrophage | Controls apoptosis of highly inflammatory cells such as eosinophils by regulating the expression of the pro-apoptotic factor Bcl2l11 | [107] | |
| AS-IL1α | Up | Macrophage | RNA polymerase II recruitment to the IL-1α promotor | [79] | |
| Mtb | lncRNA CD244 | Up | CD8+ T-cells | Recruits Ezh2 to mediate the trimethylation of H3K27 in IFN-γ/TNF-α locus | [68] |
| NEAT1 | Up | Monocyte, Macrophage | Regulates IL-6 expression | [69] | |
| PCED1B-AS1 | Down | Macrophage | As a miR-155 sponge, inhibiting macrophage apoptosis by regulating FOXO3/Rheb | [94] | |
| lincRNA-Cox2 | Up | Macrophage | Regulates the inflammatory response of TB infection macrophages through Stat3 and NF-κB signaling pathways | [97] | |
| HOTAIR | Up (H37Ra-infected) Down (H37Rv-infected) | Macrophage | Likely regulates the expression of DUSP4 and SATB1 by recruiting EZH2 during infection | [111] | |
| Salmonella | NEAT1 | Up | HeLa cells | Alters expression of host immune genes | [61] |
| IFNG-AS1 | Up | Macrophages and murine tissues | Interacts with WDR5 to promote histone methylation at the IFN-γ locus | [82] | |
| M. smegmatis | MEG3 | Up | Macrophage | Inhibits the expression of TGF-β by forming RNA-DNA triplex structures | [70] |
| M. bovis BCG | MEG3 | Down | Macrophage | IFN-γ-induced autophagy | [81] |
| Pulmonary tuberculosis; M. bovis BCG | lincRNA-EPS | Down | Monocyte, Macrophage | Regulation of apoptosis and autophagy | [80] |
| E. coli or S. aureus | lncRNA-TUB | Up | MAC-T cells | Regulates the morphology, proliferation, migration, and β-casein secretion of mammary epithelial cells | [39] |
| lncRNA-XIST | Up | MAC-T cells | Inhibits E. coli or S. aureus-induced NF-κB phosphorylation and the production of NLRP3 inflammasome | [95] | |
| E. coli | lncRNA-Mirt2 | Up | Macrophage | Inhibits the K63-ubiquitination of TRAF6 and thus alleviates inflammatory responses after TLR4 activation and balances macrophage polarization | [96] |
| LPS stimulation | lincRNA-Tnfaip3 | Up | Macrophage | Acts as a coactivator of NF-κB for the transcription of inflammatory genes through modulation of epigenetic chromatin remodeling | [98] |
| lncRNA-MRF | Up | Monocyte | Increases monocyte recruitment by upregulating the expression of monocyte chemotactic protein | [103] | |
| H. pylori | lnc-SGK1 | Up | T cells in human gastric cancer | Enhances SGK1 transcription in cis to induce TH2 and TH17 differentiation and inhibit TH1 differentiation | [52] |
| lnc-GNAT1 | Down | Gastric cancer cells | Impedes cancer cell migration through Wnt/β-catenin pathway protein expression inhibition | [53] | |
| ZFAS1 | Up | Colorectal cancer tissues | Interacts with CDK1 to be involved in p53-dependent cell cycle control and apoptosis | [77] | |
| P. aeruginosa | MEG3-4 | Down | Macrophage, Epithelial cells | Bindings to miRNA-138, thereby releasing interleukin-1β to augment inflammatory responses | [112] |
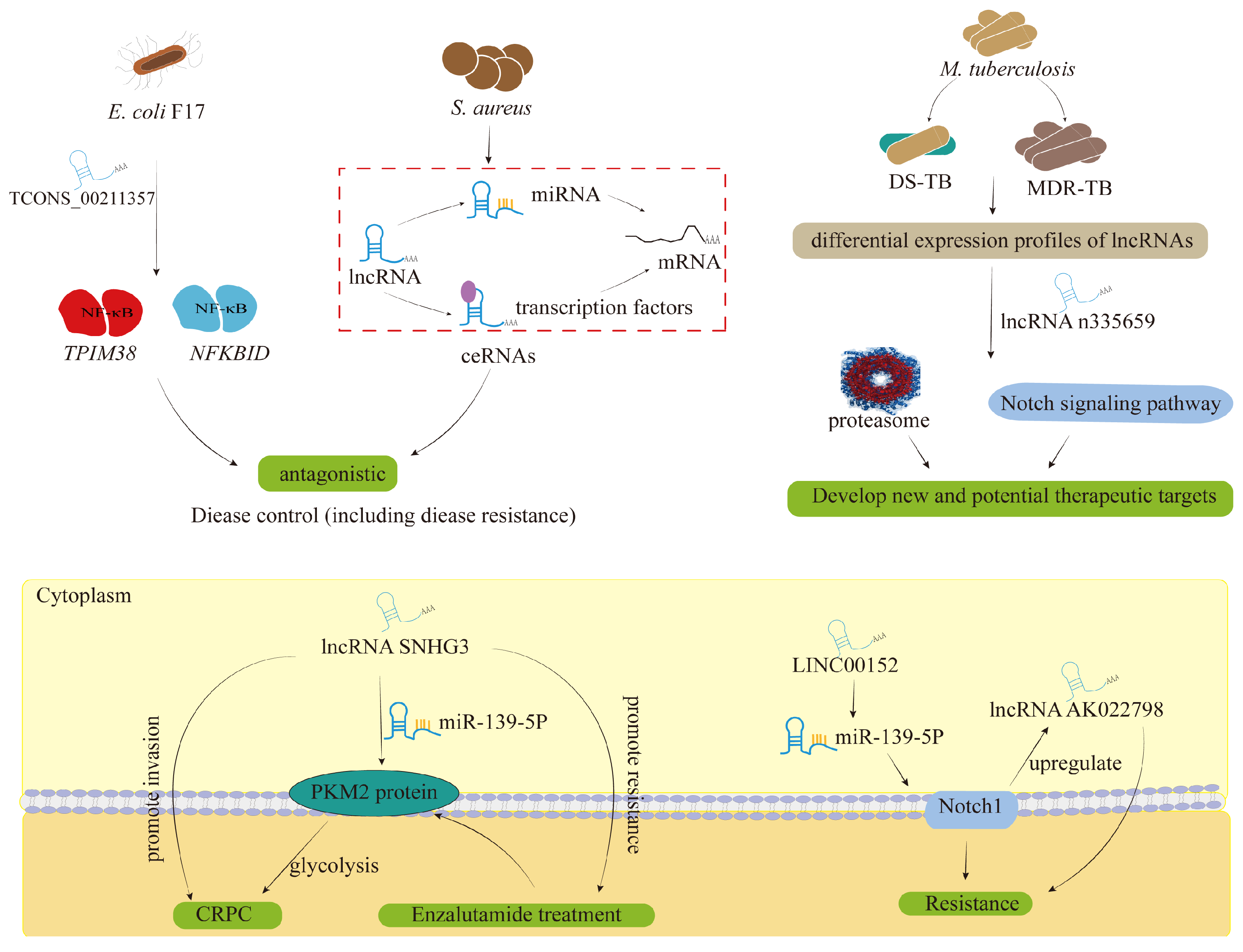
4.5. Could lncRNAs Be Targeted as a Promising Solution to the Current Antibiotic Resistance?
5. Conclusions and Future Outlook
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- van Elsland, D.; Neefjes, J. Bacterial infections and cancer. EMBO Rep. 2018, 19, e46632. [Google Scholar] [CrossRef] [PubMed]
- Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- Mancuso, G.; Midiri, A.; Gerace, E.; Biondo, C. Bacterial Antibiotic Resistance: The Most Critical Pathogens. Pathogens 2021, 10, 1310. [Google Scholar] [CrossRef] [PubMed]
- Piewngam, P.; Chiou, J.; Chatterjee, P.; Otto, M. Alternative approaches to treat bacterial infections: Targeting quorum-sensing. Expert. Rev. Anti-Infect. Ther. 2020, 18, 499–510. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Chen, X. Current Promising Strategies against Antibiotic-Resistant Bacterial Infections. Antibiotics 2022, 12, 67. [Google Scholar] [CrossRef] [PubMed]
- Alaoui Mdarhri, H.; Benmessaoud, R.; Yacoubi, H.; Seffar, L.; Guennouni Assimi, H.; Hamam, M.; Boussettine, R.; Filali-Ansari, N.; Lahlou, F.A.; Diawara, I.; et al. Alternatives Therapeutic Approaches to Conventional Antibiotics: Advantages, Limitations and Potential Application in Medicine. Antibiotics 2022, 11, 1826. [Google Scholar] [CrossRef] [PubMed]
- Kalelkar, P.P.; Riddick, M.; García, A.J. Biomaterial-based antimicrobial therapies for the treatment of bacterial infections. Nat. Rev. Mater. 2022, 7, 39–54. [Google Scholar] [CrossRef]
- Xuan, R.; Zhao, X.; Li, Q.; Zhao, Y.; Wang, Y.; Du, S.; Duan, Q.; Guo, Y.; Ji, Z.; Chao, T.; et al. Characterization of long noncoding RNA in nonlactating goat mammary glands reveals their regulatory role in mammary cell involution and remodeling. Int. J. Biol. Macromol. 2022, 222, 2158–2175. [Google Scholar] [CrossRef] [PubMed]
- Dykes, I.M.; Emanueli, C. Transcriptional and Post-transcriptional Gene Regulation by Long Non-coding RNA. Genom. Proteom. Bioinform. 2017, 15, 177–186. [Google Scholar] [CrossRef] [PubMed]
- DiStefano, J.K. The Emerging Role of Long Noncoding RNAs in Human Disease. Methods Mol. Biol. 2018, 1706, 91–110. [Google Scholar] [CrossRef]
- Luo, T.; Liu, P.; Wang, X.Y.; Li, L.Z.; Zhao, L.P.; Huang, J.; Li, Y.M.; Ou, J.L.; Peng, X.Q. Effect of the autism-associated lncRNA Shank2-AS on architecture and growth of neurons. J. Cell Biochem. 2019, 120, 1754–1762. [Google Scholar] [CrossRef] [PubMed]
- Agliano, F.; Rathinam, V.A.; Medvedev, A.E.; Vanaja, S.K.; Vella, A.T. Long Noncoding RNAs in Host-Pathogen Interactions. Trends Immunol. 2019, 40, 492–510. [Google Scholar] [CrossRef] [PubMed]
- Naorem, L.D.; Prakash, V.S.; Muthaiyan, M.; Venkatesan, A. Comprehensive analysis of dysregulated lncRNAs and their competing endogenous RNA network in triple-negative breast cancer. Int. J. Biol. Macromol. 2020, 145, 429–436. [Google Scholar] [CrossRef] [PubMed]
- Schmerer, N.; Schulte, L.N. Long noncoding RNAs in bacterial infection. Wiley Interdiscip. Rev. RNA 2021, 12, e1664. [Google Scholar] [CrossRef]
- Jiang, W.; Qu, Y.; Yang, Q.; Ma, X.; Meng, Q.; Xu, J.; Liu, X.; Wang, S. D-lnc: A comprehensive database and analytical platform to dissect the modification of drugs on lncRNA expression. RNA Biol. 2019, 16, 1586–1591. [Google Scholar] [CrossRef] [PubMed]
- Zur Bruegge, J.; Einspanier, R.; Sharbati, S. A Long Journey Ahead: Long Non-coding RNAs in Bacterial Infections. Front. Cell. Infect. Microbiol. 2017, 7, 95. [Google Scholar] [CrossRef] [PubMed]
- Zumla, A.; Rao, M.; Wallis, R.S.; Kaufmann, S.H.; Rustomjee, R.; Mwaba, P.; Vilaplana, C.; Yeboah-Manu, D.; Chakaya, J.; Ippolito, G.; et al. Host-directed therapies for infectious diseases: Current status, recent progress, and future prospects. Lancet Infect. Dis. 2016, 16, e47–e63. [Google Scholar] [CrossRef] [PubMed]
- Akerman, I.; Tu, Z.; Beucher, A.; Rolando, D.M.Y.; Sauty-Colace, C.; Benazra, M.; Nakic, N.; Yang, J.; Wang, H.; Pasquali, L.; et al. Human Pancreatic β Cell lncRNAs Control Cell-Specific Regulatory Networks. Cell Metab. 2017, 25, 400–411. [Google Scholar] [CrossRef] [PubMed]
- Morán, I.; Akerman, I.; van de Bunt, M.; Xie, R.; Benazra, M.; Nammo, T.; Arnes, L.; Nakić, N.; García-Hurtado, J.; Rodríguez-Seguí, S.; et al. Human β cell transcriptome analysis uncovers lncRNAs that are tissue-specific, dynamically regulated, and abnormally expressed in type 2 diabetes. Cell Metab. 2012, 16, 435–448. [Google Scholar] [CrossRef]
- Dahariya, S.; Paddibhatla, I.; Kumar, S.; Raghuwanshi, S.; Pallepati, A.; Gutti, R.K. Long non-coding RNA: Classification, biogenesis and functions in blood cells. Mol. Immunol. 2019, 112, 82–92. [Google Scholar] [CrossRef]
- Wu, H.; Yang, L.; Chen, L.L. The Diversity of Long Noncoding RNAs and Their Generation. Trends Genet. 2017, 33, 540–552. [Google Scholar] [CrossRef] [PubMed]
- Silveira, G.O.; Coelho, H.S.; Amaral, M.S.; Verjovski-Almeida, S. Long non-coding RNAs as possible therapeutic targets in protozoa, and in Schistosoma and other helminths. Parasitol. Res. 2022, 121, 1091–1115. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, C.; Sharma, S.; Meghwanshi, K.K.; Patel, S.; Mehta, P.; Shukla, N.; Do, D.N.; Rajpurohit, S.; Suravajhala, P.; Shukla, J.N. Long Non-Coding RNAs in Insects. Animals 2021, 11, 1118. [Google Scholar] [CrossRef] [PubMed]
- Statello, L.; Guo, C.J.; Chen, L.L.; Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 2021, 22, 96–118. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Zhang, Y.H.; Pan, X.; Liu, M.; Wang, S.; Huang, T.; Cai, Y.D. Tissue Expression Difference between mRNAs and lncRNAs. Int. J. Mol. Sci. 2018, 19, 3416. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.J.; Ma, X.K.; Xing, Y.H.; Zheng, C.C.; Xu, Y.F.; Shan, L.; Zhang, J.; Wang, S.; Wang, Y.; Carmichael, G.G.; et al. Distinct Processing of lncRNAs Contributes to Non-conserved Functions in Stem Cells. Cell 2020, 181, 621–636.e622. [Google Scholar] [CrossRef] [PubMed]
- Arunima, A.; van Schaik, E.J.; Samuel, J.E. The emerging roles of long non-coding RNA in host immune response and intracellular bacterial infections. Front. Cell. Infect. Microbiol. 2023, 13, 1160198. [Google Scholar] [CrossRef] [PubMed]
- Takakuwa, H.; Yamazaki, T.; Souquere, S.; Adachi, S.; Yoshino, H.; Fujiwara, N.; Yamamoto, T.; Natsume, T.; Nakagawa, S.; Pierron, G.; et al. Shell protein composition specified by the lncRNA NEAT1 domains dictates the formation of paraspeckles as distinct membraneless organelles. Nat. Cell Biol. 2023, 25, 1664–1675. [Google Scholar] [CrossRef] [PubMed]
- Ninomiya, K.; Adachi, S.; Natsume, T.; Iwakiri, J.; Terai, G.; Asai, K.; Hirose, T. LncRNA-dependent nuclear stress bodies promote intron retention through SR protein phosphorylation. Embo J. 2020, 39, e102729. [Google Scholar] [CrossRef]
- Zhao, M.Y.; Wang, G.Q.; Wang, N.N.; Yu, Q.Y.; Liu, R.L.; Shi, W.Q. The long-non-coding RNA NEAT1 is a novel target for Alzheimer’s disease progression via miR-124/BACE1 axis. Neurol. Res. 2019, 41, 489–497. [Google Scholar] [CrossRef]
- Xie, S.P.; Zhou, F.; Li, J.; Duan, S.J. NEAT1 regulates MPP(+)-induced neuronal injury by targeting miR-124 in neuroblastoma cells. Neurosci. Lett. 2019, 708, 134340. [Google Scholar] [CrossRef]
- Seki, T.; Yamagata, H.; Uchida, S.; Chen, C.; Kobayashi, A.; Kobayashi, M.; Harada, K.; Matsuo, K.; Watanabe, Y.; Nakagawa, S. Altered expression of long noncoding RNAs in patients with major depressive disorder. J. Psychiatr. Res. 2019, 117, 92–99. [Google Scholar] [CrossRef]
- Bridges, M.C.; Daulagala, A.C.; Kourtidis, A. LNCcation: LncRNA localization and function. J. Cell Biol. 2021, 220, e202009045. [Google Scholar] [CrossRef]
- Hadjicharalambous, M.R.; Lindsay, M.A. Long Non-Coding RNAs and the Innate Immune Response. Noncoding RNA 2019, 5, 34. [Google Scholar] [CrossRef]
- Bonetti, A.; Agostini, F.; Suzuki, A.M.; Hashimoto, K.; Pascarella, G.; Gimenez, J.; Roos, L.; Nash, A.J.; Ghilotti, M.; Cameron, C.J.F.; et al. RADICL-seq identifies general and cell type–specific principles of genome-wide RNA-chromatin interactions. Nat. Commun. 2020, 11, 1018. [Google Scholar] [CrossRef]
- Schertzer, M.D.; Braceros, K.C.A.; Starmer, J.; Cherney, R.E.; Lee, D.M.; Salazar, G.; Justice, M.; Bischoff, S.R.; Cowley, D.O.; Ariel, P.; et al. lncRNA-Induced Spread of Polycomb Controlled by Genome Architecture, RNA Abundance, and CpG Island DNA. Mol. Cell 2019, 75, 523–537.e510. [Google Scholar] [CrossRef]
- Saldaña-Meyer, R.; Rodriguez-Hernaez, J.; Escobar, T.; Nishana, M.; Jácome-López, K.; Nora, E.P.; Bruneau, B.G.; Tsirigos, A.; Furlan-Magaril, M.; Skok, J.; et al. RNA Interactions Are Essential for CTCF-Mediated Genome Organization. Mol. Cell 2019, 76, 412–422.e415. [Google Scholar] [CrossRef]
- Wen, Y.; Chen, H.; Luo, F.; Zhou, H.; Li, Z. Roles of long noncoding RNAs in bacterial infection. Life Sci. 2020, 263, 118579. [Google Scholar] [CrossRef]
- Wang, H.; Wang, X.; Li, X.; Wang, Q.; Qing, S.; Zhang, Y.; Gao, M.Q. A novel long non-coding RNA regulates the immune response in MAC-T cells and contributes to bovine mastitis. FEBS J. 2019, 286, 1780–1795. [Google Scholar] [CrossRef]
- Shen, Y.; Gong, L.; Xu, F.; Wang, S.; Liu, H.; Wang, Y.; Hu, L.; Zhu, L. Insight into the lncRNA-mRNA Co-Expression Profile and ceRNA Network in Lipopolysaccharide-Induced Acute Lung Injury. Curr. Issues Mol. Biol. 2023, 45, 6170–6189. [Google Scholar] [CrossRef]
- Yap, K.; Mukhina, S.; Zhang, G.; Tan, J.S.C.; Ong, H.S.; Makeyev, E.V. A Short Tandem Repeat-Enriched RNA Assembles a Nuclear Compartment to Control Alternative Splicing and Promote Cell Survival. Mol. Cell 2018, 72, 525–540.e513. [Google Scholar] [CrossRef]
- Grelet, S.; Link, L.A.; Howley, B.; Obellianne, C.; Palanisamy, V.; Gangaraju, V.K.; Diehl, J.A.; Howe, P.H. A regulated PNUTS mRNA to lncRNA splice switch mediates EMT and tumour progression. Nat. Cell Biol. 2017, 19, 1105–1115. [Google Scholar] [CrossRef]
- Tichon, A.; Perry, R.B.; Stojic, L.; Ulitsky, I. SAM68 is required for regulation of Pumilio by the NORAD long noncoding RNA. Genes. Dev. 2018, 32, 70–78. [Google Scholar] [CrossRef]
- Li, D.; Wang, X.; Miao, H.; Liu, H.; Pang, M.; Guo, H.; Ge, M.; Glass, S.E.; Emmrich, S.; Ji, S.; et al. The lncRNA MIR99AHG directs alternative splicing of SMARCA1 by PTBP1 to enable invadopodia formation in colorectal cancer cells. Sci. Signal 2023, 16, eadh4210. [Google Scholar] [CrossRef]
- Chen, X.; Du, J.; Zhu, Y.; Zhang, C.; Sun, H. Comprehensive analysis of lncRNA and mRNA expression profiles in macrophages activated by Actinidia eriantha polysaccharide. Int. J. Biol. Macromol. 2019, 136, 980–993. [Google Scholar] [CrossRef]
- Yan, Z.; Wang, P.; Yang, Q.; Gao, X.; Gun, S.; Huang, X. Change in Long Non-Coding RNA Expression Profile Related to the Antagonistic Effect of Clostridium perfringens Type C on Piglet Spleen. Curr. Issues Mol. Biol. 2023, 45, 2309–2325. [Google Scholar] [CrossRef]
- Lin, C.; Zhu, Y.; Hao, Z.; Xu, H.; Li, T.; Yang, J.; Chen, X.; Chen, Y.; Guo, A.; Hu, C. Genome-Wide Analysis of LncRNA in Bovine Mammary Epithelial Cell Injuries Induced by Escherichia Coli and Staphylococcus Aureus. Int. J. Mol. Sci. 2021, 22, 9719. [Google Scholar] [CrossRef]
- Chen, Y.; Jing, H.; Chen, M.; Liang, W.; Yang, J.; Deng, G.; Guo, M. Transcriptional Profiling of Exosomes Derived from Staphylococcus aureus-Infected Bovine Mammary Epithelial Cell Line MAC-T by RNA-Seq Analysis. Oxid. Med. Cell. Longevity 2021, 2021, 8460355. [Google Scholar] [CrossRef]
- Wang, X.; Su, F.; Yu, X.; Geng, N.; Li, L.; Wang, R.; Zhang, M.; Liu, J.; Liu, Y.; Han, B. RNA-Seq Whole Transcriptome Analysis of Bovine Mammary Epithelial Cells in Response to Intracellular Staphylococcus aureus. Front. Vet. Sci. 2020, 7, 642. [Google Scholar] [CrossRef]
- Horn, J.; Klepsch, M.; Manger, M.; Wolz, C.; Rudel, T.; Fraunholz, M. Long Noncoding RNA SSR42 Controls Staphylococcus aureus Alpha-Toxin Transcription in Response to Environmental Stimuli. J. Bacteriol. 2018, 200, e00252-18. [Google Scholar] [CrossRef]
- Mi, S.; Tang, Y.; Dari, G.; Shi, Y.; Zhang, J.; Zhang, H.; Liu, X.; Liu, Y.; Tahir, U.; Yu, Y. Transcriptome sequencing analysis for the identification of stable lncRNAs associated with bovine Staphylococcus aureus mastitis. J. Anim. Sci. Biotechnol. 2021, 12, 120. [Google Scholar] [CrossRef]
- Yao, Y.; Jiang, Q.; Jiang, L.; Wu, J.; Zhang, Q.; Wang, J.; Feng, H.; Zang, P. Lnc-SGK1 induced by Helicobacter pylori infection and highsalt diet promote Th2 and Th17 differentiation in human gastric cancer by SGK1/Jun B signaling. Oncotarget 2016, 7, 20549–20560. [Google Scholar] [CrossRef]
- Liu, L.; Shuai, T.; Li, B.; Zhu, L.; Li, X. Long non-coding RNA lnc-GNAT1-1 inhibits gastric cancer cell proliferation and invasion through the Wnt/β-catenin pathway in Helicobacter pylori infection. Mol. Med. Rep. 2018, 18, 4009–4015. [Google Scholar] [CrossRef]
- Bai, H.; Wu, Q.; Hu, X.; Wu, T.; Song, J.; Liu, T.; Meng, Z.; Lv, M.; Lu, X.; Chen, X.; et al. Clinical significance of lnc-AC145676.2.1-6 and lnc-TGS1-1 and their variants in western Chinese tuberculosis patients. Int. J. Infect. Dis. 2019, 84, 8–14. [Google Scholar] [CrossRef]
- He, J.; Ou, Q.; Liu, C.; Shi, L.; Zhao, C.; Xu, Y.; Kong, S.K.; Loo, J.; Li, B.; Gu, D. Differential expression of long non-coding RNAs in patients with tuberculosis infection. Tuberculosis 2017, 107, 73–79. [Google Scholar] [CrossRef]
- Fathizadeh, H.; Hayat, S.M.G.; Dao, S.; Ganbarov, K.; Tanomand, A.; Asgharzadeh, M.; Kafil, H.S. Long non-coding RNA molecules in tuberculosis. Int. J. Biol. Macromol. 2020, 156, 340–346. [Google Scholar] [CrossRef]
- Liu, L.; Yu, Z.; Ma, Q.; Yu, J.; Gong, Z.; Deng, G.; Wu, X. LncRNA NR_003508 Suppresses Mycobacterium tuberculosis-Induced Programmed Necrosis via Sponging miR-346-3p to Regulate RIPK1. Int. J. Mol. Sci. 2023, 24, 8016. [Google Scholar] [CrossRef]
- Xia, J.; Liu, Y.; Ma, Y.; Yang, F.; Ruan, Y.; Xu, J.F.; Pi, J. Advances of Long Non-Coding RNAs as Potential Biomarkers for Tuberculosis: New Hope for Diagnosis? Pharmaceutics 2023, 15, 2096. [Google Scholar] [CrossRef]
- Fang, Y.; Zhao, J.; Wang, X.; Wang, X.; Wang, L.; Liu, L.; Liu, J.; Gao, M.; Yuan, C. Identification of differentially expressed lncRNAs as potential plasma biomarkers for active tuberculosis. Tuberculosis 2021, 128, 102065. [Google Scholar] [CrossRef]
- Hu, X.; Liao, S.; Bai, H.; Gupta, S.; Zhou, Y.; Zhou, J.; Jiao, L.; Wu, L.; Wang, M.; Chen, X.; et al. Long Noncoding RNA and Predictive Model To Improve Diagnosis of Clinically Diagnosed Pulmonary Tuberculosis. J. Clin. Microbiol. 2020, 58, e01973-19. [Google Scholar] [CrossRef]
- Imamura, K.; Takaya, A.; Ishida, Y.I.; Fukuoka, Y.; Taya, T.; Nakaki, R.; Kakeda, M.; Imamachi, N.; Sato, A.; Yamada, T.; et al. Diminished nuclear RNA decay upon Salmonella infection upregulates antibacterial noncoding RNAs. Embo J. 2018, 37, e97723. [Google Scholar] [CrossRef]
- Aznaourova, M.; Janga, H.; Sefried, S.; Kaufmann, A.; Dorna, J.; Volkers, S.M.; Georg, P.; Lechner, M.; Hoppe, J.; Dökel, S.; et al. Noncoding RNA MaIL1 is an integral component of the TLR4-TRIF pathway. Proc. Natl. Acad. Sci. USA 2020, 117, 9042–9053. [Google Scholar] [CrossRef]
- Agliano, F.; Fitzgerald, K.A.; Vella, A.T.; Rathinam, V.A.; Medvedev, A.E. Long Non-coding RNA LincRNA-EPS Inhibits Host Defense Against Listeria monocytogenes Infection. Front. Cell. Infect. Microbiol. 2020, 9, 481. [Google Scholar] [CrossRef]
- Deng, X.; Guo, J.; Sun, Z.; Liu, L.; Zhao, T.; Li, J.; Tang, G.; Zhang, H.; Wang, W.; Cao, S.; et al. Brucella-Induced Downregulation of lncRNA Gm28309 Triggers Macrophages Inflammatory Response Through the miR-3068-5p/NF-κB Pathway. Front. Immunol. 2020, 11, 581517. [Google Scholar] [CrossRef]
- Balloy, V.; Koshy, R.; Perra, L.; Corvol, H.; Chignard, M.; Guillot, L.; Scaria, V. Bronchial Epithelial Cells from Cystic Fibrosis Patients Express a Specific Long Non-coding RNA Signature upon Pseudomonas aeruginosa Infection. Front. Cell. Infect. Microbiol. 2017, 7, 218. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.; Luo, F.; Zhao, L.; Su, S.; Lei, W.; Liu, Y.; Shi, K.; Li, Z. Long Non-Coding RNA FGD5-AS1 Induced by Chlamydia trachomatis Infection Inhibits Apoptosis via Wnt/β-Catenin Signaling Pathway. Front. Cell. Infect. Microbiol. 2021, 11, 701352. [Google Scholar] [CrossRef]
- Fu, Y.; Xu, X.; Xue, J.; Duan, W.; Yi, Z. Deregulated lncRNAs in B Cells from Patients with Active Tuberculosis. PLoS ONE 2017, 12, e0170712. [Google Scholar] [CrossRef]
- Wang, Y.; Zhong, H.; Xie, X.; Chen, C.Y.; Huang, D.; Shen, L.; Zhang, H.; Chen, Z.W.; Zeng, G. Long noncoding RNA derived from CD244 signaling epigenetically controls CD8+ T-cell immune responses in tuberculosis infection. Proc. Natl. Acad. Sci. USA 2015, 112, E3883–E3892. [Google Scholar] [CrossRef]
- Huang, S.; Huang, Z.; Luo, Q.; Qing, C. The Expression of lncRNA NEAT1 in Human Tuberculosis and Its Antituberculosis Effect. BioMed Res. Int. 2018, 2018, 9529072. [Google Scholar] [CrossRef]
- Sharbati, S.; Ravon, F.; Einspanier, R.; Zur Bruegge, J. Mycobacterium smegmatis But Not Mycobacterium avium subsp. hominissuis Causes Increased Expression of the Long Non-Coding RNA MEG3 in THP-1-Derived Human Macrophages and Associated Decrease of TGF-β. Microorganisms 2019, 7, 63. [Google Scholar] [CrossRef]
- Xu, H.; Jiang, Y.; Xu, X.; Su, X.; Liu, Y.; Ma, Y.; Zhao, Y.; Shen, Z.; Huang, B.; Cao, X. Inducible degradation of lncRNA Sros1 promotes IFN-γ-mediated activation of innate immune responses by stabilizing Stat1 mRNA. Nat. Immunol. 2019, 20, 1621–1630. [Google Scholar] [CrossRef]
- Atianand, M.K.; Hu, W.; Satpathy, A.T.; Shen, Y.; Ricci, E.P.; Alvarez-Dominguez, J.R.; Bhatta, A.; Schattgen, S.A.; McGowan, J.D.; Blin, J.; et al. A Long Noncoding RNA lincRNA-EPS Acts as a Transcriptional Brake to Restrain Inflammation. Cell 2016, 165, 1672–1685. [Google Scholar] [CrossRef]
- Gheitasi, R.; Jourghasemi, S.; Pakzad, I.; Hosseinpour Sarmadi, V.; Samieipour, Y.; Sekawi, Z.; Azizi Jalilian, F. A potential marker in brucellosis, long non coding RNA IFNG-AS1. Mol. Biol. Rep. 2019, 46, 6495–6500. [Google Scholar] [CrossRef] [PubMed]
- Luo, F.; Wen, Y.; Zhao, L.; Su, S.; Zhao, Y.; Lei, W.; Li, Z. Chlamydia trachomatis induces lncRNA MIAT upregulation to regulate mitochondria-mediated host cell apoptosis and chlamydial development. J. Cell Mol. Med. 2022, 26, 163–177. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.; Chen, H.; Luo, F.; Zhao, L.; Shu, M.; Su, S.; Zhao, Y.; Huang, Q.; Li, Z. Chlamydia trachomatis Plasmid Protein pORF5 Up-Regulates ZFAS1 to Promote Host Cell Survival via MAPK/p38 Pathway. Front. Microbiol. 2020, 11, 593295. [Google Scholar] [CrossRef]
- Wang, X.; Wang, H.; Zhang, R.; Li, D.; Gao, M.Q. LRRC75A antisense lncRNA1 knockout attenuates inflammatory responses of bovine mammary epithelial cells. Int. J. Biol. Sci. 2020, 16, 251–263. [Google Scholar] [CrossRef]
- Xie, S.; Ge, Q.; Wang, X.; Sun, X.; Kang, Y. Long non-coding RNA ZFAS1 sponges miR-484 to promote cell proliferation and invasion in colorectal cancer. Cell Cycle 2018, 17, 154–161. [Google Scholar] [CrossRef]
- Carpenter, S.; Aiello, D.; Atianand, M.K.; Ricci, E.P.; Gandhi, P.; Hall, L.L.; Byron, M.; Monks, B.; Henry-Bezy, M.; Lawrence, J.B.; et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 2013, 341, 789–792. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.; Atianand, M.; Jiang, Z.; Carpenter, S.; Aiello, D.; Elling, R.; Fitzgerald, K.A.; Caffrey, D.R. Cutting Edge: A Natural Antisense Transcript, AS-IL1α, Controls Inducible Transcription of the Proinflammatory Cytokine IL-1α. J. Immunol. 2015, 195, 1359–1363. [Google Scholar] [CrossRef]
- Ke, Z.; Lu, J.; Zhu, J.; Yang, Z.; Jin, Z.; Yuan, L. Down-regulation of lincRNA-EPS regulates apoptosis and autophagy in BCG-infected RAW264.7 macrophages via JNK/MAPK signaling pathway. Infect., Genet. Evol. 2020, 77, 104077. [Google Scholar] [CrossRef]
- Pawar, K.; Hanisch, C.; Palma Vera, S.E.; Einspanier, R.; Sharbati, S. Down regulated lncRNA MEG3 eliminates mycobacteria in macrophages via autophagy. Sci. Rep. 2016, 6, 19416. [Google Scholar] [CrossRef] [PubMed]
- Gomez, J.A.; Wapinski, O.L.; Yang, Y.W.; Bureau, J.F.; Gopinath, S.; Monack, D.M.; Chang, H.Y.; Brahic, M.; Kirkegaard, K. The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-γ locus. Cell 2013, 152, 743–754. [Google Scholar] [CrossRef]
- Nappi, F. Non-Coding RNA-Targeted Therapy: A State-of-the-Art Review. Int. J. Mol. Sci. 2024, 25, 3630. [Google Scholar] [CrossRef] [PubMed]
- Emami Meybodi, S.M.; Soleimani, N.; Yari, A.; Javadifar, A.; Tollabi, M.; Karimi, B.; Emami Meybodi, M.; Seyedhossaini, S.; Brouki Milan, P.; Dehghani Firoozabadi, A. Circulatory long noncoding RNAs (circulatory-LNC-RNAs) as novel biomarkers and therapeutic targets in cardiovascular diseases: Implications for cardiovascular diseases complications. Int. J. Biol. Macromol. 2023, 225, 1049–1071. [Google Scholar] [CrossRef]
- Abd El Fattah, Y.K.; Abulsoud, A.I.; AbdelHamid, S.G.; Hamdy, N.M. Interactome battling of lncRNA CCDC144NL-AS1: Its role in the emergence and ferocity of cancer and beyond. Int. J. Biol. Macromol. 2022, 222, 1676–1687. [Google Scholar] [CrossRef]
- Wang, Y.; Zeng, M.; Xia, L.; Valerie Olovo, C.; Su, Z.; Zhang, Y. Bacterial strategies for immune systems-Role of the type VI secretion system. Int. Immunopharmacol. 2023, 114, 109550. [Google Scholar] [CrossRef] [PubMed]
- Quereda, J.J.; Cossart, P. Regulating Bacterial Virulence with RNA. Annu. Rev. Microbiol. 2017, 71, 263–280. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Qin, C.; Zhang, X.; Zhu, Y.; Li, A.; Wang, M.; Tang, Y.; Kreiswirth, B.N.; Chen, L.; Zhang, H.; et al. The tst gene associated Staphylococcus aureus pathogenicity island facilitates its pathogenesis by promoting the secretion of inflammatory cytokines and inducing immune suppression. Microb. Pathog. 2020, 138, 103797. [Google Scholar] [CrossRef]
- Safarpour-Dehkordi, M.; Doosti, A.; Jami, M.S. Integrative Analysis of lncRNAs in Kidney Cancer to Discover A New lncRNA (LINC00847) as A Therapeutic Target for Staphylococcal Enterotoxin tst Gene. Cell J. 2020, 22, 101–109. [Google Scholar] [CrossRef]
- Todd, O.A.; Noverr, M.C.; Peters, B.M. Candida albicans Impacts Staphylococcus aureus Alpha-Toxin Production via Extracellular Alkalinization. mSphere 2019, 4, e00780-19. [Google Scholar] [CrossRef]
- Chen, Y.; Yang, J.; Huang, Z.; Jing, H.; Yin, B.; Guo, S.; Deng, G.; Guo, M. Exosomal lnc-AFTR as a novel translation regulator of FAS ameliorates Staphylococcus aureus-induced mastitis. Biofactors 2022, 48, 148–163. [Google Scholar] [CrossRef] [PubMed]
- Elling, R.; Robinson, E.K.; Shapleigh, B.; Liapis, S.C.; Covarrubias, S.; Katzman, S.; Groff, A.F.; Jiang, Z.; Agarwal, S.; Motwani, M.; et al. Genetic Models Reveal cis and trans Immune-Regulatory Activities for lincRNA-Cox2. Cell Rep. 2018, 25, 1511–1524.e1516. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; He, Y.; Moqbel, S.A.A.; Zhou, X.; Wu, L.; Bao, J. SIRT3 ameliorates osteoarthritis via regulating chondrocyte autophagy and apoptosis through the PI3K/Akt/mTOR pathway. Int. J. Biol. Macromol. 2021, 175, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Cui, J.; Niu, W.; Huang, J.; Feng, T.; Sun, B.; Yao, H. Long non-coding PCED1B-AS1 regulates macrophage apoptosis and autophagy by sponging miR-155 in active tuberculosis. Biochem. Biophys. Res. Commun. 2019, 509, 803–809. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Pei, Y.; Wang, X.; Feng, J.; Zhang, Y.; Gao, M.Q. LncRNA XIST mediates bovine mammary epithelial cell inflammatory response via NF-kappaB/NLRP3 inflammasome pathway. Cell Prolif. 2019, 52, e12525. [Google Scholar] [CrossRef]
- Du, M.; Yuan, L.; Tan, X.; Huang, D.; Wang, X.; Zheng, Z.; Mao, X.; Li, X.; Yang, L.; Huang, K.; et al. The LPS-inducible lncRNA Mirt2 is a negative regulator of inflammation. Nat. Commun. 2017, 8, 2049. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Gao, C.; Zhao, L.; Zhang, Y. Inflammatory response is modulated by lincRNACox2 via the NF-κB pathway in macrophages infected by Mycobacterium tuberculosis. Mol. Med. Rep. 2020, 21, 2513–2521. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Ming, Z.; Gong, A.Y.; Wang, Y.; Chen, X.; Hu, G.; Zhou, R.; Shibata, A.; Swanson, P.C.; Chen, X.M. A long noncoding RNA, lincRNA-Tnfaip3, acts as a coregulator of NF-κB to modulate inflammatory gene transcription in mouse macrophages. FASEB J. 2017, 31, 1215–1225. [Google Scholar] [CrossRef]
- Zhou, J.; Xu, J.; Zhang, L.; Liu, S.; Ma, Y.; Wen, X.; Hao, J.; Li, Z.; Ni, Y.; Li, X.; et al. Combined Single-Cell Profiling of lncRNAs and Functional Screening Reveals that H19 Is Pivotal for Embryonic Hematopoietic Stem Cell Development. Cell Stem Cell 2019, 24, 285–298.e285. [Google Scholar] [CrossRef]
- Luo, M.; Jeong, M.; Sun, D.; Park, H.J.; Rodriguez, B.A.; Xia, Z.; Yang, L.; Zhang, X.; Sheng, K.; Darlington, G.J.; et al. Long non-coding RNAs control hematopoietic stem cell function. Cell Stem Cell 2015, 16, 426–438. [Google Scholar] [CrossRef]
- Zhuang, L.; Tian, J.; Zhang, X.; Wang, H.; Huang, C. Lnc-DC regulates cellular turnover and the HBV-induced immune response by TLR9/STAT3 signaling in dendritic cells. Cell. Mol. Biol. Lett. 2018, 23, 43. [Google Scholar] [CrossRef]
- Wang, P.; Xue, Y.; Han, Y.; Lin, L.; Wu, C.; Xu, S.; Jiang, Z.; Xu, J.; Liu, Q.; Cao, X. The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science 2014, 344, 310–313. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Xie, Z.; Zhang, Z.; Li, M.; Ye, G.; Yu, W.; Li, J.; Ye, F.; Su, Z.; Che, Y.; et al. LncRNA MRF drives the regulatory function on monocyte recruitment and polarization through HNRNPD-MCP1 axis in mesenchymal stem cells. J. Biomed. Sci. 2022, 29, 73. [Google Scholar] [CrossRef] [PubMed]
- Jiang, R.; Tang, J.; Chen, Y.; Deng, L.; Ji, J.; Xie, Y.; Wang, K.; Jia, W.; Chu, W.M.; Sun, B. The long noncoding RNA lnc-EGFR stimulates T-regulatory cells differentiation thus promoting hepatocellular carcinoma immune evasion. Nat. Commun. 2017, 8, 15129. [Google Scholar] [CrossRef]
- Yu, L.; Zhang, L.; Jiang, Z.; Yu, B. Decreasing lncRNA PVT1 causes Treg/Th17 imbalance via NOTCH signaling in immune thrombocytopenia. Hematology 2021, 26, 734–740. [Google Scholar] [CrossRef] [PubMed]
- Ranzani, V.; Rossetti, G.; Panzeri, I.; Arrigoni, A.; Bonnal, R.J.; Curti, S.; Gruarin, P.; Provasi, E.; Sugliano, E.; Marconi, M.; et al. The long intergenic noncoding RNA landscape of human lymphocytes highlights the regulation of T cell differentiation by linc-MAF-4. Nat. Immunol. 2015, 16, 318–325. [Google Scholar] [CrossRef]
- Kotzin, J.J.; Spencer, S.P.; McCright, S.J.; Kumar, D.B.U.; Collet, M.A.; Mowel, W.K.; Elliott, E.N.; Uyar, A.; Makiya, M.A.; Dunagin, M.C.; et al. The long non-coding RNA Morrbid regulates Bim and short-lived myeloid cell lifespan. Nature 2016, 537, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Wang, T.; Jiang, C.; Li, S.; Zhou, H.; Li, R. Relish-facilitated lncRNA-CR11538 suppresses Drosophila Imd immune response and maintains immune homeostasis via decoying Relish away from antimicrobial peptide promoters. Dev. Comp. Immunol. 2024, 151, 105098. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Li, S.; Wu, S.; Jin, P.; Ma, F. LncRNA-CR11538 Decoys Dif/Dorsal to Reduce Antimicrobial Peptide Products for Restoring Drosophila Toll Immunity Homeostasis. Int. J. Mol. Sci. 2021, 22, 10117. [Google Scholar] [CrossRef]
- Hu, G.; Gong, A.Y.; Wang, Y.; Ma, S.; Chen, X.; Chen, J.; Su, C.J.; Shibata, A.; Strauss-Soukup, J.K.; Drescher, K.M.; et al. LincRNA-Cox2 Promotes Late Inflammatory Gene Transcription in Macrophages through Modulating SWI/SNF-Mediated Chromatin Remodeling. J. Immunol. 2016, 196, 2799–2808. [Google Scholar] [CrossRef]
- Subuddhi, A.; Kumar, M.; Majumder, D.; Sarkar, A.; Ghosh, Z.; Vasudevan, M.; Kundu, M.; Basu, J. Unraveling the role of H3K4 trimethylation and lncRNA HOTAIR in SATB1 and DUSP4-dependent survival of virulent Mycobacterium tuberculosis in macrophages. Tuberculosis 2020, 120, 101897. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Fang, L.; Pu, Q.; Bu, H.; Zhu, P.; Chen, Z.; Yu, M.; Li, X.; Weiland, T.; Bansal, A.; et al. MEG3-4 is a miRNA decoy that regulates IL-1β abundance to initiate and then limit inflammation to prevent sepsis during lung infection. Sci. Signal 2018, 11, eaao2387. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Lv, X.; Zhang, W.; Hu, T.; Cao, X.; Ren, Z.; Getachew, T.; Mwacharo, J.M.; Haile, A.; Sun, W. Insights Into Long Non-Coding RNA and mRNA Expression in the Jejunum of Lambs Challenged With Escherichia coli F17. Front. Vet. Sci. 2022, 9, 819917. [Google Scholar] [CrossRef] [PubMed]
- Jin, C.; Bao, J.; Wang, Y.; Chen, W.; Wu, T.; Wang, L.; Lv, X.; Gao, W.; Wang, B.; Zhu, G.; et al. Changes in long non-coding RNA expression profiles related to the antagonistic effects of Escherichia coli F17 on lamb spleens. Sci. Rep. 2018, 8, 16514. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Xu, R.; Zhang, X.; Wang, Q.; Pang, J.; Zhang, X.; Chang, X.; Zhang, Y. Identifying differentially expressed long non-coding RNAs in PBMCs in response to the infection of multidrug-resistant tuberculosis. Infect. Drug Resist. 2018, 11, 945–959. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Gao, S.; Chen, C.; Li, H.; Wang, S.; Yu, Y.; Ming, L. Screening and identification of differentially expressed long non-coding RNAs in multidrug-resistant tuberculosis. PeerJ 2022, 10, e12776. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Chen, X.; Wang, X.a.; Li, H.; Zhu, Y.; Li, X.; Xiao, Z.; Zi, T.; Qin, X.; Zhao, Y.; et al. Glycolysis related lncRNA SNHG3 / miR-139-5p / PKM2 axis promotes castration-resistant prostate cancer (CRPC) development and enzalutamide resistance. Int. J. Biol. Macromol. 2024, 260, 129635. [Google Scholar] [CrossRef]
- Hashemi, M.; Hasani, S.; Hajimazdarany, S.; Mirmazloomi, S.R.; Makvandy, S.; Zabihi, A.; Goldoost, Y.; Gholinia, N.; Kakavand, A.; Tavakolpournegari, A.; et al. Non-coding RNAs targeting notch signaling pathway in cancer: From proliferation to cancer therapy resistance. Int. J. Biol. Macromol. 2022, 222, 1151–1167. [Google Scholar] [CrossRef]
- Jin, C.; Lagoudas, G.K.; Zhao, C.; Bullman, S.; Bhutkar, A.; Hu, B.; Ameh, S.; Sandel, D.; Liang, X.S.; Mazzilli, S.; et al. Commensal Microbiota Promote Lung Cancer Development via γδ T Cells. Cell 2019, 176, 998–1013.e1016. [Google Scholar] [CrossRef]
- Chen, Y.; Li, Z.; Chen, X.; Zhang, S. Long non-coding RNAs: From disease code to drug role. Acta Pharm. Sin. B 2021, 11, 340–354. [Google Scholar] [CrossRef]
| Pathogen | LncRNA | Expression | Role | References |
|---|---|---|---|---|
| Brucella spp. | Gm28309 | Down | Shows potential as a biomarker in the disease progression for brucellosis | [64] |
| IFNG-AS1 | Up | Enhances the expression of INF-γ in the immune response of brucellosis patients, revealing its potential in screening, diagnosis, or treatment of brucellosis | [73] | |
| C. trachomatis | FGD5-AS1 | Up | Upregulated FGD5-AS1 provides new ideas for screening potential intervention targets for C. trachomati | [66] |
| lncRNA-IRF1 | Up | MIAT and IRF1 are involved in apoptosis resistance in persistent infection | [74] | |
| MIAT | Up | [74] | ||
| ZEB1-AS1 | Up | A persistent C. trachomatis infection up-regulated lncRNA that could enhance persistent infection-induced anti-apoptosis | [74] | |
| ZFAS1 | Up | Plays a role in the anti-apoptotic process of C. trachomatis | [75] | |
| E. coli | LRRC75A | Down | Knockout of LRRC75A-AS1 attenuated the E. coli-induced inflammatory responses | [76] |
| H. pylori | lncRNA GNAT1-1 | Down | Regulation of invasion and migration of gastric cancer cells | [53] |
| ZFAS1 | Up | Upregulation of lncRNA ZFAS1 predicts poor prognosis and promotes invasion and metastasis in colorectal cancer | [77] | |
| L. monocytogenes | lincRNA-EPS | Down | Downregulation of lincRNA-EPS leads to an increased production of proinflammatory cytokines and iNOS in splenic and liver macrophages and dendritic cells | [63] |
| lncRNA-SROS1 | Down | Shows potential as biomarkers in the disease progression | [71] | |
| lincRNA-Cox2 | Up | [78] | ||
| AS-IL1α | Up | An important regulator of IL-1α transcription during the innate immune response, and has the potential to be used as a diagnostic marker | [79] | |
| M. bovis | lincRNA-Cox2 | Up | Shows potential as biomarkers in the disease progression | [78] |
| lincRNA-EPS | Down | [80] | ||
| MEG3 | Down | Shows potential as a biomarker in the systematic early diagnosis | [81] | |
| M. smegmatis | MEG3 | Up | As a novel mediator of host cell response during mycobacterial infections | [70] |
| M. tuberculosis | XLOC_012582 | Up | These deregulated lncRNAs in Mtb-infected cells can be introduced as promising molecular biomarkers for prognosis and/or diagnosis | [56] |
| PCED1B-AS1 | Up | [56] | ||
| lncRNA-CD244 | Up | A major regulator of CD8 T-cell immune response during in vivo Mtb infection and has the potential to be used as a diagnostic marker | [68] | |
| NEAT1 | Down | Tuberculosis patients had significantly increased the expression of NEAT1, and NEAT1 might serve as a potential indicator for patient prognosis of tuberculosis | [69] | |
| S. aureus | TNK2-AS1 | Up | TNK2-AS1 could be regarded as stable markers associated with bovine S. aureus mastitis | [51] |
| S. typhimurium | NEAT1_2 | Up | Shows potential as biomarkers in the systematic early diagnosis | [61] |
| IFNG-AS1 | Up | [82] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shi, L.; Han, X.; Liu, F.; Long, J.; Jin, Y.; Chen, S.; Duan, G.; Yang, H. Review on Long Non-Coding RNAs as Biomarkers and Potentially Therapeutic Targets for Bacterial Infections. Curr. Issues Mol. Biol. 2024, 46, 7558-7576. https://doi.org/10.3390/cimb46070449
Shi L, Han X, Liu F, Long J, Jin Y, Chen S, Duan G, Yang H. Review on Long Non-Coding RNAs as Biomarkers and Potentially Therapeutic Targets for Bacterial Infections. Current Issues in Molecular Biology. 2024; 46(7):7558-7576. https://doi.org/10.3390/cimb46070449
Chicago/Turabian StyleShi, Liqin, Xueya Han, Fang Liu, Jinzhao Long, Yuefei Jin, Shuaiyin Chen, Guangcai Duan, and Haiyan Yang. 2024. "Review on Long Non-Coding RNAs as Biomarkers and Potentially Therapeutic Targets for Bacterial Infections" Current Issues in Molecular Biology 46, no. 7: 7558-7576. https://doi.org/10.3390/cimb46070449
APA StyleShi, L., Han, X., Liu, F., Long, J., Jin, Y., Chen, S., Duan, G., & Yang, H. (2024). Review on Long Non-Coding RNAs as Biomarkers and Potentially Therapeutic Targets for Bacterial Infections. Current Issues in Molecular Biology, 46(7), 7558-7576. https://doi.org/10.3390/cimb46070449









