Figure 1.
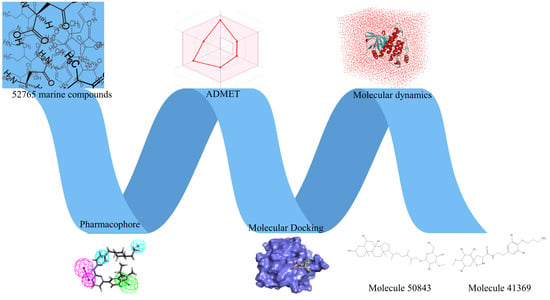
Workflow of this study: marine compound database construction, a pharmacophore, ADMET, molecular docking, and molecular dynamics.
Figure 1.
Workflow of this study: marine compound database construction, a pharmacophore, ADMET, molecular docking, and molecular dynamics.
Figure 2.
Comparison of CDK4/6 protein structures and key residues. (A) Schematic representation of the superimposed CDK4 and CDK6 structures. The CDK4 (PDB: 2W96) structure is shown in blue and the CDK6 (PDB: 5L2S) structure is shown in yellow. (B) Comparison of the amino acid sequences of CDK4 (PDB: 2W96) and CDK6 (PDB: 5L2S). Residues defining the active site boundary are highlighted in red for the PDB structure of CDK4 and in yellow for the PDB structure of CDK6.
Figure 2.
Comparison of CDK4/6 protein structures and key residues. (A) Schematic representation of the superimposed CDK4 and CDK6 structures. The CDK4 (PDB: 2W96) structure is shown in blue and the CDK6 (PDB: 5L2S) structure is shown in yellow. (B) Comparison of the amino acid sequences of CDK4 (PDB: 2W96) and CDK6 (PDB: 5L2S). Residues defining the active site boundary are highlighted in red for the PDB structure of CDK4 and in yellow for the PDB structure of CDK6.
Figure 3.
Comparison of CDK4 and CDK6 protein A-chain residue sequences and conservativeness. Higher scoring levels indicate higher conservativeness of residues. In this case, the yellow residues (which do not contain the active residues of the proteins) could not be classified by the conservativeness grade due to their low frequency of occurrence in the database.
Figure 3.
Comparison of CDK4 and CDK6 protein A-chain residue sequences and conservativeness. Higher scoring levels indicate higher conservativeness of residues. In this case, the yellow residues (which do not contain the active residues of the proteins) could not be classified by the conservativeness grade due to their low frequency of occurrence in the database.
Figure 4.
The pharmacophore model and receiver operating characteristic (ROC) curve validation. Hydrophobic group features are shown as blue spheres, hydrogen bond acceptor features are shown as purple spheres and hydrogen bond donor features are shown as green spheres. (A) Pharmacophore model 01–05. (B) Pharmacophore model 06–10. (C) Coincidence effect drawing of Abemaciclib and pharmacophore 09. (D) ROC curve.
Figure 4.
The pharmacophore model and receiver operating characteristic (ROC) curve validation. Hydrophobic group features are shown as blue spheres, hydrogen bond acceptor features are shown as purple spheres and hydrogen bond donor features are shown as green spheres. (A) Pharmacophore model 01–05. (B) Pharmacophore model 06–10. (C) Coincidence effect drawing of Abemaciclib and pharmacophore 09. (D) ROC curve.
Figure 5.
Analysis of binding mode between compound 41369 and CDK4/6. (A) A 2D interaction schematic of compound 41369 with CDK4. Hydrogen bonds are shown as green dashed lines, hydrophobicity is in red lines. (B) A 3D binding mode of compound 41369 with CDK4. Hydrogen bonds are shown as red dashed lines, while compound 41369 is shown in golden yellow. (C) A 2D interaction schematic of compound 41369 with CDK6. Hydrogen bonds are shown as green dashed lines, hydrophobicity is in red lines. (D) A 3D binding mode of compound 41369 with CDK6. Hydrogen bonds are shown as red dashed lines, while compound 50843 is shown in golden yellow.
Figure 5.
Analysis of binding mode between compound 41369 and CDK4/6. (A) A 2D interaction schematic of compound 41369 with CDK4. Hydrogen bonds are shown as green dashed lines, hydrophobicity is in red lines. (B) A 3D binding mode of compound 41369 with CDK4. Hydrogen bonds are shown as red dashed lines, while compound 41369 is shown in golden yellow. (C) A 2D interaction schematic of compound 41369 with CDK6. Hydrogen bonds are shown as green dashed lines, hydrophobicity is in red lines. (D) A 3D binding mode of compound 41369 with CDK6. Hydrogen bonds are shown as red dashed lines, while compound 50843 is shown in golden yellow.
Figure 6.
Analysis of binding mode between compound 50843 and CDK4/6. (A) A 2D interaction schematic of compound 50843 with CDK4. Hydrogen bonds are shown as green dashed lines, hydrophobicity is in red lines. (B) A 3D binding mode of compound 50843 with CDK4. Hydrogen bonds are shown as red dashed lines, while compound 50843 is shown in blue. (C) A 2D interaction schematic of compound 50843 with CDK6. Hydrogen bonds are shown as green dashed lines, hydrophobicity is in red lines. (D) A 3D binding mode of compound 50843 with CDK6. Hydrogen bonds are shown as red dashed lines, while compound 50843 is shown in blue.
Figure 6.
Analysis of binding mode between compound 50843 and CDK4/6. (A) A 2D interaction schematic of compound 50843 with CDK4. Hydrogen bonds are shown as green dashed lines, hydrophobicity is in red lines. (B) A 3D binding mode of compound 50843 with CDK4. Hydrogen bonds are shown as red dashed lines, while compound 50843 is shown in blue. (C) A 2D interaction schematic of compound 50843 with CDK6. Hydrogen bonds are shown as green dashed lines, hydrophobicity is in red lines. (D) A 3D binding mode of compound 50843 with CDK6. Hydrogen bonds are shown as red dashed lines, while compound 50843 is shown in blue.
Figure 7.
RMSD and RMSF plots of compound 50843 and 41369 with CDK4/6. (A) The RMSD of complexes. CDK4 and compound 41369 is shown as black lines, CDK4 and compound 50843 is in the red line, CDK6 and compound 41369 is shown as blue lines, CDK6 and compound 50843 is in the green line. (B) The RMSD of ligands. Ligand 50843 of complex CDK4 is shown as black lines, Ligand 41369 of complex CDK4 is shown as red lines, Ligand 50843 of complex CDK6 is shown as blue lines, Ligand 41369 of complex CDK6 is shown as green lines. (C) The RMSF of complexes and CDK4. CDK4 is shown as black lines, compound 41369 is in the red line and compound 50843 is in the blue line. (D) The RMSF of complexes and CDK6. CDK6 is shown as black lines, compound 41369 is in the red line and compound 50843 is in the blue line.
Figure 7.
RMSD and RMSF plots of compound 50843 and 41369 with CDK4/6. (A) The RMSD of complexes. CDK4 and compound 41369 is shown as black lines, CDK4 and compound 50843 is in the red line, CDK6 and compound 41369 is shown as blue lines, CDK6 and compound 50843 is in the green line. (B) The RMSD of ligands. Ligand 50843 of complex CDK4 is shown as black lines, Ligand 41369 of complex CDK4 is shown as red lines, Ligand 50843 of complex CDK6 is shown as blue lines, Ligand 41369 of complex CDK6 is shown as green lines. (C) The RMSF of complexes and CDK4. CDK4 is shown as black lines, compound 41369 is in the red line and compound 50843 is in the blue line. (D) The RMSF of complexes and CDK6. CDK6 is shown as black lines, compound 41369 is in the red line and compound 50843 is in the blue line.
Figure 8.
The hydrogen bond of CDK4/6 with compound 41369 and 50843. (A) Compound 41369 (magenta) with CDK4. (B) Compound 50843 (black) with CDK4. (C) Compound 41369 (magenta) with CDK6. (D) Compound 50843 (black) with CDK6.
Figure 8.
The hydrogen bond of CDK4/6 with compound 41369 and 50843. (A) Compound 41369 (magenta) with CDK4. (B) Compound 50843 (black) with CDK4. (C) Compound 41369 (magenta) with CDK6. (D) Compound 50843 (black) with CDK6.
Figure 9.
Solvent accessible surface area (SASA) and radius of gyration (Rg) of 100 ns simulation process. (A) SASA plots of compound 50843 (blue) and 41369 (red) with CDK4 (black). (B) SASA plots of compound 50843 (blue) and 41369 (red) with CDK6 (black). (C) Rg plots of compound 50843 (blue) and 41369 (red) with CDK4 (black). (D) Rg plots of compound 50843 (blue) and 41369 (red) with CDK6 (black).
Figure 9.
Solvent accessible surface area (SASA) and radius of gyration (Rg) of 100 ns simulation process. (A) SASA plots of compound 50843 (blue) and 41369 (red) with CDK4 (black). (B) SASA plots of compound 50843 (blue) and 41369 (red) with CDK6 (black). (C) Rg plots of compound 50843 (blue) and 41369 (red) with CDK4 (black). (D) Rg plots of compound 50843 (blue) and 41369 (red) with CDK6 (black).
Figure 10.
The total energy of CDK4/6 with molecule 41369 and 50843. (A) The total energy of compound 41369 (black) and 50843 (red) with CDK4. (B) The total energy of compound 41369 (black) and 50843 (red) with CDK6.
Figure 10.
The total energy of CDK4/6 with molecule 41369 and 50843. (A) The total energy of compound 41369 (black) and 50843 (red) with CDK4. (B) The total energy of compound 41369 (black) and 50843 (red) with CDK6.
Figure 11.
Residue decomposition diagram of binding energy. (A) Compound 50843 with CDK4 (black). (B) Compound 50843 with CDK6 (magenta).
Figure 11.
Residue decomposition diagram of binding energy. (A) Compound 50843 with CDK4 (black). (B) Compound 50843 with CDK6 (magenta).
Table 1.
The characteristic composition, the number of true/false positive and negative molecules, and the sensitivity of 10 pharmacophore models were constructed. Feature “H” stands for hydrophobic group, while feature “A”, “D” stand for hydrogen bond acceptor and hydrogen bond donor, respectively.
Table 1.
The characteristic composition, the number of true/false positive and negative molecules, and the sensitivity of 10 pharmacophore models were constructed. Feature “H” stands for hydrophobic group, while feature “A”, “D” stand for hydrogen bond acceptor and hydrogen bond donor, respectively.
| Pharmacophore | Features | Ranking Score | True Positives | True Negatives | False Positives | False Negatives | Sensitivity |
|---|
| Phar01 | HHHDA | 55.473 | 3 | 8 | 4 | 4 | 0.42857 |
| Phar02 | HHHDA | 55.280 | 4 | 8 | 4 | 3 | 0.57143 |
| Phar03 | HHDA | 51.607 | 4 | 11 | 1 | 3 | 0.57143 |
| Phar04 | HHHD | 50.761 | 4 | 10 | 2 | 3 | 0.57143 |
| Phar05 | HHDA | 50.129 | 5 | 10 | 2 | 2 | 0.71429 |
| Phar06 | HHHD | 49.714 | 4 | 10 | 2 | 3 | 0.57143 |
| Phar07 | HHDA | 48.862 | 3 | 8 | 4 | 4 | 0.42857 |
| Phar08 | HHDA | 48.828 | 2 | 9 | 3 | 5 | 0.28571 |
| Phar09 | HHDA | 48.726 | 6 | 10 | 2 | 1 | 0.85714 |
| Phar10 | HHDA | 48.442 | 6 | 9 | 3 | 1 | 0.85714 |
Table 2.
Twenty molecules‘ water solubility, intestinal absorption, hepatotoxicity and CYP2D6 enzyme inhibition descriptor properties.
Table 2.
Twenty molecules‘ water solubility, intestinal absorption, hepatotoxicity and CYP2D6 enzyme inhibition descriptor properties.
| Name | Solubility | Absorption Level | Hepatotoxic | CYP2D6 Inhibit |
|---|
| Molecule17227 | 3 | 3 | −9.93282 | −10.6775 |
| Molecule35962 | 3 | 2 | −10.2467 | −9.79215 |
| Molecule35945 | 3 | 2 | −10.3996 | −9.79215 |
| Molecule50853 | 3 | 3 | −9.27768 | −11.7400 |
| Molecule5999 | 3 | 3 | −12.0390 | −11.4830 |
| Molecule20551 | 4 | 2 | −7.04826 | −5.29830 |
| Molecule7211 | 3 | 3 | −11.8022 | −10.2513 |
| Molecule5996 | 3 | 3 | −12.0390 | −11.4830 |
| Molecule23671 | 3 | 3 | −4.67926 | −11.4629 |
| Molecule9567 | 3 | 3 | −27.0804 | −9.75238 |
| Molecule41369 | 3 | 1 | −13.8276 | −0.02226 |
| Molecule6045 | 3 | 3 | −19.2113 | −10.6996 |
| Molecule33567 | 3 | 1 | −9.88709 | −4.57087 |
| Molecule50843 | 4 | 3 | −13.7281 | −8.90702 |
| Molecule6049 | 3 | 3 | −19.2113 | −10.6996 |
| Molecule36157 | 3 | 1 | −4.28935 | −4.45112 |
| Molecule6028 | 3 | 3 | −19.2113 | −10.6996 |
| Molecule22564 | 3 | 2 | −7.74774 | −7.53197 |
| Molecule18748 | 3 | 3 | −5.33843 | −8.68489 |
| Molecule6243 | 3 | 3 | −11.4714 | −9.68009 |
Table 3.
Libdock scores of 20 selected molecules and positive control Abemaciclib with CDK4/6.
Table 4.
Binding energy of binding for the compound 50843 complexed with CDK4/6.
Table 4.
Binding energy of binding for the compound 50843 complexed with CDK4/6.
| Pharmacophore | Features | Ranking Score |
|---|
| Van der Waal energy (kJ/mol) | −192.855 ± 90.101 | −254.799 ± 51.499 |
| Electrostatic energy (kJ/mol) | −84.560 ± 49.773 | −59.732 ± 23.528 |
| Polar solvation energy (kJ/mol) | 139.559 ± 111.556 | 121.507 ± 47.040 |
| SASA energy (kJ/mol) | −16.800 ± 9.461 | −19.058 ± 4.394 |
| Binding energy(kJ/mol) | −154.655 ± 39.178 | −212.082 ± 42.561 |
Table 5.
Synthetic accessibility score parameters (SA score) for molecule 41369 and molecule 50843.
Table 5.
Synthetic accessibility score parameters (SA score) for molecule 41369 and molecule 50843.
| Molecule | SA Score |
|---|
| 41369 | 5.226 |
| 50843 | 5.517 |
| Abemaciclib | 3.415 |
Table 6.
Predicted inhibitory activity of two alternative compounds against tumor cell lines.
Table 6.
Predicted inhibitory activity of two alternative compounds against tumor cell lines.
| Compound | Pa | Pi | Cell Line | Tissue | Tumor Type |
|---|
| 41369 | 0.498 | 0.028 | MDA-MB-231 | Breast | Adenocarcinoma |
| 50843 | 0.625 | 0.014 | HL-60 | Hematopoietic and lymphoid tissue | Leukemia |
| Abemaciclib | 0.750 | 0.004 | LoVo | Colon | Adenocarcinoma |