Altered Gene Transcription in Human Cells Treated with Ludox® Silica Nanoparticles
Abstract
:1. Introduction
2. Experimental Section
2.1. Nanoparticle Characterization
2.2. Cell Line and Treatments
2.3. Assessment of Cytotoxicity and Apoptosis Detection
2.4. Assessment of Microarray Experiments
2.5. Microarray Data Analysis
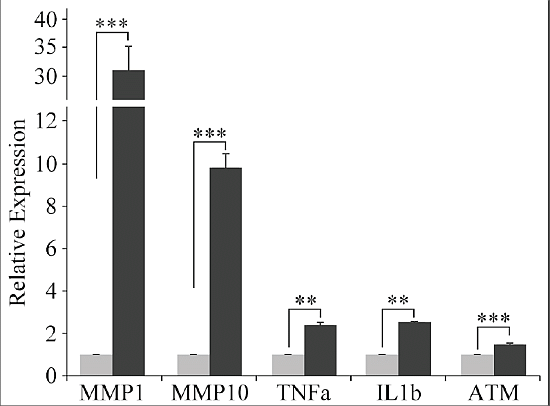
2.6. qRT-PCR Experiments
| Primer Name | Sequence |
|---|---|
| MMP1 forward | AGAGAGCAGCTTCAGTGACA |
| MMP1 reverse | CTTGAGCTGCTTTTCCTCCG |
| MMP10 forward | TTGACCCCAATGCCAGGAT |
| MMP10 reverse | CCCCTATCTCGCCTAGCAAT |
| TNFa forward | AGTGCTGGCAACCACTAAGAA |
| TNFa reverse | AGATGTCAGGGATCAAAGCTG |
| IL1b forward | TACTCACTTAAAGCCCGCCT |
| IL1b reverse | ATGTGGGAGCGAATGACAGA |
| ATM forward | ACTGGCCAGAACTTTCAAGAAC |
| ATM reverse | TGCCCAGAATACTTGTGCTTC |
| GAPDH forward | TCCTCTGACTTCAACAGCGA |
| GAPDH reverse | GGGTCTTACTCCTTGGAGGC |
3. Results and Discussion
3.1. Characterization of Ludox® AS30 and SM30 Nanoparticles

| NP Type | Counterion * | ζ Potential in PBS | DLS Diameter in PBS | Diameter from TEM in PBS | Surface Area * | pH * |
|---|---|---|---|---|---|---|
| SM30 | sodium | −26.3 mV | 14 ± 4 nm | 9 ± 3 nm | 345 m2/g | 10.0 |
| AS30 | ammonium | −25.9 mV | 20 ± 4 nm | 18 ± 3 nm | 230 m2/g | 9.1 |
3.2. Cytoxicity Induced by Ludox® AS30 and SM30 Nanoparticles

3.3. Microarray Analysis: Differentially Expressed Genes

| GO.ID | Term | Count | p Value |
|---|---|---|---|
| GO:0042981 | Regulation of apoptosis | 29 | 2.9 × 10−6 |
| GO:0043067 | Regulation of cell death | 29 | 3.5 × 10−6 |
| GO:0006357 | Regulation of transcription from RNA polymerase II promoter | 25 | 3.8 × 10−5 |
| GO:0006954 | Inflammatory response | 12 | 0.00436 |
| GO:0009611 | Response to wounding | 16 | 0.00507 |
| GO:0006952 | Defense response | 16 | 0.01816 |
| Topological Parameters | SM30 Network |
|---|---|
| Average clustering coefficient | 0.651 |
| Connected components | 32 |
| Avg. number of neighbors | 6.278 |
| Network radius | 1 |
| Network diameter | 11 |
| Network centralization | 0.065 |
| Network density | 0.009 |
| Network heterogeneity | 0.997 |

3.4. Supervised Approach: Pathway Analysis
| Pathway | Set Size | NTk Stat | NTk q-Value |
|---|---|---|---|
| Activation of ATR in response to replication stress | 33 | −5.89 | 0 |
| G2/M Checkpoints | 37 | −5.31 | 0 |
| CDC6 association with the ORC: origin complex | 10 | −4.94 | 0 |
| Activation of the pre-replicative complex | 28 | −4.87 | 0 |
| E2F mediated regulation of DNA replication | 30 | −4.76 | 0 |
| M Phase | 96 | −3.67 | 0 |
| Association of licensing factors with the pre-replicative complex | 14 | −3.09 | 0.012315271 |
| G1/S-Specific Transcription | 17 | −2.65 | 0.031397174 |
| Synthesis of glycosylphosphatidylinositol (GPI) | 15 | −2.51 | 0.036945813 |
| DCC mediated attractive signaling | 11 | 2.37 | 0.048701299 |
| Regulation of Complement cascade | 10 | 2.41 | 0.044994376 |
| Activation of Matrix Metalloproteinases | 21 | 2.41 | 0.044994376 |
| Acyl chain remodelling of PE | 13 | 2.58 | 0.035714286 |
| Activation of BH3-only proteins | 16 | 2.58 | 0.035714286 |
| Signaling by Robo receptor | 24 | 2.65 | 0.031397174 |
| Nucleotide-binding domain, leucine rich repeat containing receptor (NLR) signaling pathways | 44 | 2.65 | 0.031397174 |
| p38MAPK events | 12 | 2.65 | 0.031397174 |
| Acyl chain remodelling of PC | 14 | 2.75 | 0.027472527 |
| Chemokine receptors bind chemokines | 27 | 2.88 | 0.020120724 |
| GAB1 signalosome | 71 | 2.88 | 0.020120724 |
| Antigen Activates B Cell Receptor Leading to Generation of Second Messengers | 18 | 3.09 | 0.012315271 |
| Translocation of GLUT4 to the Plasma Membrane | 47 | 3.09 | 0.012315271 |
| Signalling to RAS | 28 | 3.09 | 0.012315271 |
| Cell junction organization | 66 | 3.09 | 0.012315271 |
| O-linked glycosylation of mucins | 44 | 3.2 | 0 |
| Interleukin-2 signaling | 38 | 3.53 | 0 |
| Signalling to ERKs | 34 | 3.59 | 0 |
| Downstream signal transduction | 120 | 3.9 | 0 |
| Glycerophospholipid biosynthesis | 68 | 4.21 | 0 |
| Cell-Cell communication | 101 | 4.23 | 0 |
| Signaling by ERBB4 | 106 | 4.45 | 0 |
| TRAF6 Mediated Induction of proinflammatory cytokines | 64 | 4.47 | 0 |
| MyD88 cascade initiated on plasma membrane | 73 | 4.56 | 0 |
| Toll Like Receptor 10 (TLR10) Cascade | 73 | 4.56 | 0 |
| Toll Like Receptor 5 (TLR5) Cascade | 73 | 4.56 | 0 |
| TRAF6 mediated induction of NFkB and MAP kinases upon TLR7/8 or 9 activation | 73 | 4.62 | 0 |
| NFkB and MAP kinases activation mediated by TLR4 signaling repertoire | 71 | 4.71 | 0 |
| MyD88-independent cascade | 76 | 4.71 | 0 |
| Toll Like Receptor 3 (TLR3) Cascade | 76 | 4.71 | 0 |
| MyD88 dependent cascade initiated on endosome | 74 | 4.72 | 0 |
| Toll Like Receptor 7/8 (TLR7/8) Cascade | 74 | 4.72 | 0 |
| Toll Like Receptor 4 (TLR4) Cascade | 92 | 4.76 | 0 |
| Toll Receptor Cascades | 105 | 4.79 | 0 |
| Signaling by SCF-KIT | 106 | 4.8 | 0 |
| Activated TLR4 signalling | 88 | 5.09 | 0 |
| MyD88:Mal cascade initiated on plasma membrane | 78 | 5.1 | 0 |
| Toll Like Receptor 2 (TLR2) Cascade | 78 | 5.1 | 0 |
| Toll Like Receptor TLR1:TLR2 Cascade | 78 | 5.1 | 0 |
| Toll Like Receptor TLR6:TLR2 Cascade | 78 | 5.1 | 0 |
| Signaling by Interleukins | 91 | 5.1 | 0 |



4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Aillon, K.L.; Xie, Y.; el-Gendy, N.; Berkland, C.J.; Forrest, M.L. Effects of nanomaterial physicochemical properties on in vivo toxicity. Adv. Drug Deliv. Rev. 2009, 61, 457–466. [Google Scholar] [CrossRef]
- Drobne, D. Nanotoxicology for safe and sustainable nanotechnology. Arh. Hig. Rada Toksikol. 2007, 58, 471–478. [Google Scholar]
- Flörke, O.W.; Graetsch, H.A.; Brunk, F.; Benda, L.; Paschen, S.; Bergna, H.E.; Roberts, W.O.; Welsh, W.A.; Libanati, C.; Ettlinger, M.; et al. (Eds.) Silica. In Ullmann’s Encyclopedia of Industrial Chemistry; Wiley-VCH Verlag GmbH and Co. KGaA: Weinheim, Germany, 2000.
- Sharifi, S.; Behzadi, S.; Laurent, S.; Forrest, M.L.; Stroeve, P.; Mahmoudi, M. Toxicity of nanomaterials. Chem. Soc. Rev. 2012, 41, 2323–2343. [Google Scholar] [CrossRef]
- Kan, H.; London, S.J.; Chen, G.; Zhang, Y.; Song, G.; Zhao, N.; Jiang, L.; Chen, B. Season, sex, age, and education as modifiers of the effects of outdoor air pollution on daily mortality in Shanghai, China: The Public Health and Air Pollution in Asia (PAPA) Study. Environ. Health Perspect. 2008, 116, 1183–1188. [Google Scholar] [CrossRef]
- Goeminne, P.C.; Kicinski, M.; Vermeulen, F.; Fierens, F.; de Boeck, K.; Nemery, B.; Nawrot, T.S.; Dupont, L.J. Impact of air pollution on cystic fibrosis pulmonary exacerbations: A case-crossover analysis. Chest 2013, 143, 946–954. [Google Scholar] [CrossRef]
- Ruckerl, R.; Schneider, A.; Breitner, S.; Cyrys, J.; Peters, A. Health effects of particulate air pollution: A review of epidemiological evidence. Inhal. Toxicol. 2011, 23, 555–592. [Google Scholar] [CrossRef]
- Hampel, R.; Ruckerl, R.; Yli-Tuomi, T.; Breitner, S.; Lanki, T.; Kraus, U.; Cyrys, J.; Belcredi, P.; Bruske, I.; Laitinen, T.M.; et al. Impact of personally measured pollutants on cardiac function. Int. J. Hyg. Environ. Health 2014, 217, 460–464. [Google Scholar] [CrossRef]
- Elder, A.; Gelein, R.; Silva, V.; Feikert, T.; Opanashuk, L.; Carter, J.; Potter, R.; Maynard, A.; Ito, Y.; Finkelstein, J.; et al. Translocation of inhaled ultrafine manganese oxide particles to the central nervous system. Environ. Health Perspect. 2006, 114, 1172–1178. [Google Scholar] [CrossRef]
- Guadagnini, R.; Moreau, K.; Hussain, S.; Marano, F.; Boland, S. Toxicity evaluation of engineered nanoparticles for medical applications using pulmonary epithelial cells. Nanotoxicology 2013. [Google Scholar] [CrossRef]
- Yan, L.; Gu, Z.; Zhao, Y. Chemical mechanisms of the toxicological properties of nanomaterials: Generation of intracellular reactive oxygen species. Chemistry 2013, 8, 2342–2353. [Google Scholar]
- Fubini, B.; Ghiazza, M.; Fenoglio, I. Physico-chemical features of engineered nanoparticles relevant to their toxicity. Nanotoxicology 2010, 4, 347–363. [Google Scholar] [CrossRef]
- Etoc, F.; Lisse, D.; Bellaiche, Y.; Piehler, J.; Coppey, M.; Dahan, M. Subcellular control of Rac-GTPase signalling by magnetogenetic manipulation inside living cells. Nat. Nanotechnol. 2013, 8, 193–198. [Google Scholar] [CrossRef]
- Rothen-Rutishauser, B.; Blank, F.; Muhlfeld, C.; Gehr, P. In vitro models of the human epithelial airway barrier to study the toxic potential of particulate matter. Expert Opin. Drug Metab. Toxicol 2008, 4, 1075–1089. [Google Scholar]
- Sabella, S.; Kote, S.; Pompa, P.P. In vivo nanotoxicology: Toxicoproteomics. Nanotech 2013 2013, 3, 151–154. [Google Scholar]
- Moos, P.J.; Olszewski, K.; Honeggar, M.; Cassidy, P.; Leachman, S.; Woessner, D.; Cutler, N.S.; Veranth, J.M. Responses of human cells to ZnO nanoparticles: A gene transcription study. Metallomics 2011, 3, 1199–1211. [Google Scholar] [CrossRef]
- Balakumaran, A.; Pawelczyk, E.; Ren, J.; Sworder, B.; Chaudhry, A.; Sabatino, M.; Stroncek, D.; Frank, J.A.; Robey, P.G. Superparamagnetic iron oxide nanoparticles labeling of bone marrow stromal (mesenchymal) cells does not affect their “stemness”. PLoS One 2010, 5. [Google Scholar] [CrossRef]
- Tuomela, S.; Autio, R.; Buerki-Thurnherr, T.; Arslan, O.; Kunzmann, A.; Andersson-Willman, B.; Wick, P.; Mathur, S.; Scheynius, A.; Krug, H.F.; et al. Gene expression profiling of immune-competent human cells exposed to engineered zinc oxide or titanium dioxide nanoparticles. PLoS One 2013, 8. [Google Scholar] [CrossRef]
- Busch, W.; Kuhnel, D.; Schirmer, K.; Scholz, S. Tungsten carbide cobalt nanoparticles exert hypoxia-like effects on the gene expression level in human keratinocytes. BMC Genomics 2010, 11. [Google Scholar] [CrossRef]
- Fujita, K.; Horie, M.; Kato, H.; Endoh, S.; Suzuki, M.; Nakamura, A.; Miyauchi, A.; Yamamoto, K.; Kinugasa, S.; Nishio, K.; et al. Effects of ultrafine TiO2 particles on gene expression profile in human keratinocytes without illumination: Involvement of extracellular matrix and cell adhesion. Toxicol. Lett. 2009, 191, 109–117. [Google Scholar] [CrossRef]
- Gras, R.; Almonacid, L.; Ortega, P.; Serramia, M.J.; Gomez, R.; de la Mata, F.J.; Lopez-Fernandez, L.A.; Munoz-Fernandez, M.A. Changes in gene expression pattern of human primary macrophages induced by carbosilane dendrimer 2G-NN16. Pharm. Res. 2009, 26, 577–586. [Google Scholar] [CrossRef]
- Huang, Y.C.; Karoly, E.D.; Dailey, L.A.; Schmitt, M.T.; Silbajoris, R.; Graff, D.W.; Devlin, R.B. Comparison of gene expression profiles induced by coarse, fine, and ultrafine particulate matter. J. Toxicol. Environ. Health Part A 2011, 74, 296–312. [Google Scholar] [CrossRef]
- Karoly, E.D.; Li, Z.; Dailey, L.A.; Hyseni, X.; Huang, Y.C. Up-regulation of tissue factor in human pulmonary artery endothelial cells after ultrafine particle exposure. Environ. Health Perspect. 2007, 115, 535–540. [Google Scholar] [CrossRef]
- Eder, C.; Frankenberger, M.; Stanzel, F.; Seidel, A.; Schramm, K.W.; Ziegler-Heitbrock, L.; Hofer, T.P. Ultrafine carbon particles down-regulate CYP1B1 expression in human monocytes. Part. Fibre Toxicol. 2009, 6. [Google Scholar] [CrossRef]
- Kim, E.Y.; Schulz, R.; Swantek, P.; Kunstman, K.; Malim, M.H.; Wolinsky, S.M. Gold nanoparticle-mediated gene delivery induces widespread changes in the expression of innate immunity genes. Gene Ther. 2012, 19, 347–353. [Google Scholar] [CrossRef]
- Kawata, K.; Osawa, M.; Okabe, S. In vitro toxicity of silver nanoparticles at noncytotoxic doses to HepG2 human hepatoma cells. Environ. Sci. Technol. 2009, 43, 6046–6051. [Google Scholar] [CrossRef]
- Shen, K.; Tseng, G.C. Meta-analysis for pathway enrichment analysis when combining multiple genomic studies. Bioinformatics 2010, 26, 1316–1323. [Google Scholar] [CrossRef]
- Martini, P.; Risso, D.; Sales, G.; Romualdi, C.; Lanfranchi, G.; Cagnin, S. Statistical test of expression pattern (STEPath): A new strategy to integrate gene expression data with genomic information in individual and meta-analysis studies. BMC Bioinform. 2011, 12. [Google Scholar] [CrossRef]
- Martini, P.; Sales, G.; Calura, E.; Brugiolo, M.; Lanfranchi, G.; Romualdi, C.; Cagnin, S. Systems biology approach to the dissection of the complexity of regulatory networks in the S. scrofa cardiocirculatory system. Int. J. Mol. Sci. 2013, 14, 23160–23187. [Google Scholar] [CrossRef]
- Gene Ontology Consortium. Available online: http://www.geneontology.org (accessed on 22 April 2014).
- KEGG: Kyoto Encyclopedia of Genes and Genomes. Available online: http://www.genome.jp/kegg/ (accessed on 22 April 2014).
- Reactome. Available online: http://www.reactome.org/ (accessed on 22 April 2014).
- Fede, C.; Selvestrel, F.; Compagnin, C.; Mognato, M.; Mancin, F.; Reddi, E.; Celotti, L. The toxicity outcome of silica nanoparticles (Ludox(R)) is influenced by testing techniques and treatment modalities. Anal. Bioanal. Chem. 2012, 404, 1789–1802. [Google Scholar] [CrossRef]
- Compagnin, C.; Mognato, M.; Celotti, L.; Canti, G.; Palumbo, G.; Reddi, E. Cell proliferation and cell cycle alterations in oesophageal p53-mutated cancer cells treated with cisplatin in combination with photodynamic therapy. Cell. Prolif. 2010, 43, 262–274. [Google Scholar] [CrossRef]
- Bolstad, B.M.; Irizarry, R.A.; Astrand, M.; Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 2003, 19, 185–193. [Google Scholar] [CrossRef]
- Tusher, V.G.; Tibshirani, R.; Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 2001, 98, 5116–5121. [Google Scholar] [CrossRef]
- Saeed, A.I.; Sharov, V.; White, J.; Li, J.; Liang, W.; Bhagabati, N.; Braisted, J.; Klapa, M.; Currier, T.; Thiagarajan, M.; et al. TM4: A free, open-source system for microarray data management and analysis. BioTechniques 2003, 34, 374–378. [Google Scholar]
- Huang, da W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar]
- Cline, M.S.; Smoot, M.; Cerami, E.; Kuchinsky, A.; Landys, N.; Workman, C.; Christmas, R.; Avila-Campilo, I.; Creech, M.; Gross, B.; et al. Integration of biological networks and gene expression data using Cytoscape. Nat. Protoc. 2007, 2, 2366–2382. [Google Scholar] [CrossRef]
- Agilent Literature Search. Available online: http://apps.cytoscape.org/apps/agilentliteraturesearch (accessed on March 2014).
- Doncheva, N.T.; Assenov, Y.; Domingues, F.S.; Albrecht, M. Topological analysis and interactive visualization of biological networks and protein structures. Nat. Protoc. 2012, 7, 670–685. [Google Scholar] [CrossRef]
- Dong, J.; Horvath, S. Understanding network concepts in modules. BMC Syst. Biol. 2007, 1. [Google Scholar] [CrossRef]
- Tian, L.; Greenberg, S.A.; Kong, S.W.; Altschuler, J.; Kohane, I.S.; Park, P.J. Discovering statistically significant pathways in expression profiling studies. Proc. Natl. Acad. Sci. USA 2005, 102, 13544–13549. [Google Scholar] [CrossRef]
- Martini, P.; Sales, G.; Massa, M.S.; Chiogna, M.; Romualdi, C. Along signal paths: An empirical gene set approach exploiting pathway topology. Nucleic Acids Res. 2013, 41. [Google Scholar] [CrossRef]
- Sales, G.; Calura, E.; Martini, P.; Romualdi, C. Graphite Web: Web tool for gene set analysis exploiting pathway topology. Nucleic Acids Res. 2013, 41, 89–97. [Google Scholar] [CrossRef]
- Martinez-Castanon, G.A.; Nino-Martinez, N.; Martinez-Gutierrez, F.; Martinez-Mendoza, J.R.; Ruiz, F. Synthesis and antibacterial activity of silver nanoparticles with different sizes. J. Nanopart Res. 2008, 10, 1343–1348. [Google Scholar]
- Assenov, Y.; Ramirez, F.; Schelhorn, S.E.; Lengauer, T.; Albrecht, M. Computing topological parameters of biological networks. Bioinformatics 2008, 24, 282–284. [Google Scholar] [CrossRef]
- Carlson, M.R.; Zhang, B.; Fang, Z.; Mischel, P.S.; Horvath, S.; Nelson, S.F. Gene connectivity, function, and sequence conservation: Predictions from modular yeast co-expression networks. BMC Genomics 2006, 7. [Google Scholar] [CrossRef]
- Horvath, S.; Zhang, B.; Carlson, M.; Lu, K.V.; Zhu, S.; Felciano, R.M.; Laurance, M.F.; Zhao, W.; Qi, S.; Chen, Z.; et al. Analysis of oncogenic signaling networks in glioblastoma identifies ASPM as a molecular target. Proc. Natl. Acad. Sci. USA 2006, 103, 17402–17407. [Google Scholar] [CrossRef]
- Carter, S.L.; Brechbuhler, C.M.; Griffin, M.; Bond, A.T. Gene co-expression network topology provides a framework for molecular characterization of cellular state. Bioinformatics 2004, 20, 2242–2250. [Google Scholar]
- Dagouassat, M.; Lanone, S.; Boczkowski, J. Interaction of matrix metalloproteinases with pulmonary pollutants. Eur. Respir. J. 2012, 39, 1021–1032. [Google Scholar] [CrossRef]
- Amara, N.; Bachoual, R.; Desmard, M.; Golda, S.; Guichard, C.; Lanone, S.; Aubier, M.; Ogier-Denis, E.; Boczkowski, J. Diesel exhaust particles induce matrix metalloprotease-1 in human lung epithelial cells via a NADP(H) oxidase/NOX4 redox-dependent mechanism. Am. J. Physiol. Lung Cell. Mol. Physiol. 2007, 293, 170–181. [Google Scholar] [CrossRef]
- Croft, D.; Mundo, A.F.; Haw, R.; Milacic, M.; Weiser, J.; Wu, G.; Caudy, M.; Garapati, P.; Gillespie, M.; Kamdar, M.R.; et al. The Reactome pathway knowledgebase. Nucleic Acids Res. 2014, 42, 472–477. [Google Scholar] [CrossRef]
- D’Eustachio, P. Pathway databases: Making chemical and biological sense of the genomic data flood. Chem. Biol. 2013, 20, 629–635. [Google Scholar]
- Mordes, D.A.; Cortez, D. Activation of ATR and related PIKKs. Cell Cycle 2008, 7, 2809–2812. [Google Scholar] [CrossRef]
- Cuddihy, A.R.; O’Connell, M.J. Cell-cycle responses to DNA damage in G2. Int. Rev. Cytol. 2003, 222, 99–140. [Google Scholar]
- DiPaola, R.S. To arrest or not to G(2)-M Cell-cycle arrest: Commentary re: A.K. Tyagi et al., Silibinin strongly synergizes human prostate carcinoma DU145 cells to doxorubicin-induced growth inhibition, G(2)-M arrest, and apoptosis. Clin. cancer res., 8: 3512–3519, 2002. Clin. Cancer Res. 2002, 8, 3311–3314. [Google Scholar]
- Liu, W.; Jiang, F.; Bi, X.; Zhang, Y.Q. Drosophila FMRP participates in the DNA damage response by regulating G2/M cell cycle checkpoint and apoptosis. Hum. Mol. Genet. 2012, 21, 4655–4668. [Google Scholar] [CrossRef]
- Chen, M.; von Mikecz, A. Formation of nucleoplasmic protein aggregates impairs nuclear function in response to SiO2 nanoparticles. Exp. Cell Res. 2005, 305, 51–62. [Google Scholar] [CrossRef]
- Wan, R.; Mo, Y.; Zhang, X.; Chien, S.; Tollerud, D.J.; Zhang, Q. Matrix metalloproteinase-2 and -9 are induced differently by metal nanoparticles in human monocytes: The role of oxidative stress and protein tyrosine kinase activation. Toxicol. Appl. Pharmacol. 2008, 233, 276–285. [Google Scholar] [CrossRef]
- Corbalan, J.J.; Medina, C.; Jacoby, A.; Malinski, T.; Radomski, M.W. Amorphous silica nanoparticles trigger nitric oxide/peroxynitrite imbalance in human endothelial cells: Inflammatory and cytotoxic effects. Int. J. Nanomed. 2011, 6, 2821–2835. [Google Scholar]
- Choi, M.; Cho, W.S.; Han, B.S.; Cho, M.; Kim, S.Y.; Yi, J.Y.; Ahn, B.; Kim, S.H.; Jeong, J. Transient pulmonary fibrogenic effect induced by intratracheal instillation of ultrafine amorphous silica in A/J mice. Toxicol. Lett. 2008, 182, 97–101. [Google Scholar] [CrossRef]
- Lund, A.K.; Lucero, J.; Lucas, S.; Madden, M.C.; McDonald, J.D.; Seagrave, J.C.; Knuckles, T.L.; Campen, M.J. Vehicular emissions induce vascular MMP-9 expression and activity associated with endothelin-1-mediated pathways. Arterioscler. Thromb. Vasc. Biol. 2009, 29, 511–517. [Google Scholar] [CrossRef]
- Zhang, Q.; Kleeberger, S.R.; Reddy, S.P. DEP-induced fra-1 expression correlates with a distinct activation of AP-1-dependent gene transcription in the lung. Am. J. Physiol. Lung Cell. Mol. Physiol. 2004, 286, L427–L436. [Google Scholar] [CrossRef]
- Ding, M.; Kisin, E.R.; Zhao, J.; Bowman, L.; Lu, Y.; Jiang, B.; Leonard, S.; Vallyathan, V.; Castranova, V.; Murray, A.R.; et al. Size-dependent effects of tungsten carbide-cobalt particles on oxygen radical production and activation of cell signaling pathways in murine epidermal cells. Toxicol. Appl. Pharmacol. 2009, 241, 260–268. [Google Scholar] [CrossRef]
- Vaillant, C.; Meissirel, C.; Mutin, M.; Belin, M.F.; Lund, L.R.; Thomasset, N. MMP-9 deficiency affects axonal outgrowth, migration, and apoptosis in the developing cerebellum. Mol. Cell. Neurosci. 2003, 24, 395–408. [Google Scholar] [CrossRef]
- Bouillet, P.; Strasser, A. BH3-only proteins—Evolutionarily conserved proapoptotic Bcl-2 family members essential for initiating programmed cell death. J. Cell Sci. 2002, 115, 1567–1574. [Google Scholar]
- Paur, H.-R.; Cassee, F.R.; Teeguarden, J.; Fissan, H.; Diabate, S.; Aufderheide, M.; Kreyling, W.G.; Hänninen, O.; Kasper, G.; Riediker, M.; et al. In-vitro cell exposure studies for the assessment of nanoparticle toxicity in the lung—A dialog between aerosol science and biology. J. Aerosol Sci. 2011, 42, 668–692. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Fede, C.; Millino, C.; Pacchioni, B.; Celegato, B.; Compagnin, C.; Martini, P.; Selvestrel, F.; Mancin, F.; Celotti, L.; Lanfranchi, G.; et al. Altered Gene Transcription in Human Cells Treated with Ludox® Silica Nanoparticles. Int. J. Environ. Res. Public Health 2014, 11, 8867-8890. https://doi.org/10.3390/ijerph110908867
Fede C, Millino C, Pacchioni B, Celegato B, Compagnin C, Martini P, Selvestrel F, Mancin F, Celotti L, Lanfranchi G, et al. Altered Gene Transcription in Human Cells Treated with Ludox® Silica Nanoparticles. International Journal of Environmental Research and Public Health. 2014; 11(9):8867-8890. https://doi.org/10.3390/ijerph110908867
Chicago/Turabian StyleFede, Caterina, Caterina Millino, Beniamina Pacchioni, Barbara Celegato, Chiara Compagnin, Paolo Martini, Francesco Selvestrel, Fabrizio Mancin, Lucia Celotti, Gerolamo Lanfranchi, and et al. 2014. "Altered Gene Transcription in Human Cells Treated with Ludox® Silica Nanoparticles" International Journal of Environmental Research and Public Health 11, no. 9: 8867-8890. https://doi.org/10.3390/ijerph110908867