Phenotypic and Genotypic Characterization with MALDI-TOF-MS Based Identification of Staphylococcus spp. Isolated from Mobile Phones with their Antibiotic Susceptibility, Biofilm Formation, and Adhesion Properties
Abstract
:1. Introduction
2. Results
2.1. Staphylococcus spp. Morphotypes on CSA
2.2. Species Identification with MALDI-TOF-MS and Hydrolytic Enzymes Production
2.3. Susceptibility to Antibiotics and Detection of mecA Gene
2.4. Adhesive Properties
2.5. Detection of Biofilm, Exoenzymes, and Haemolysin Related Genes
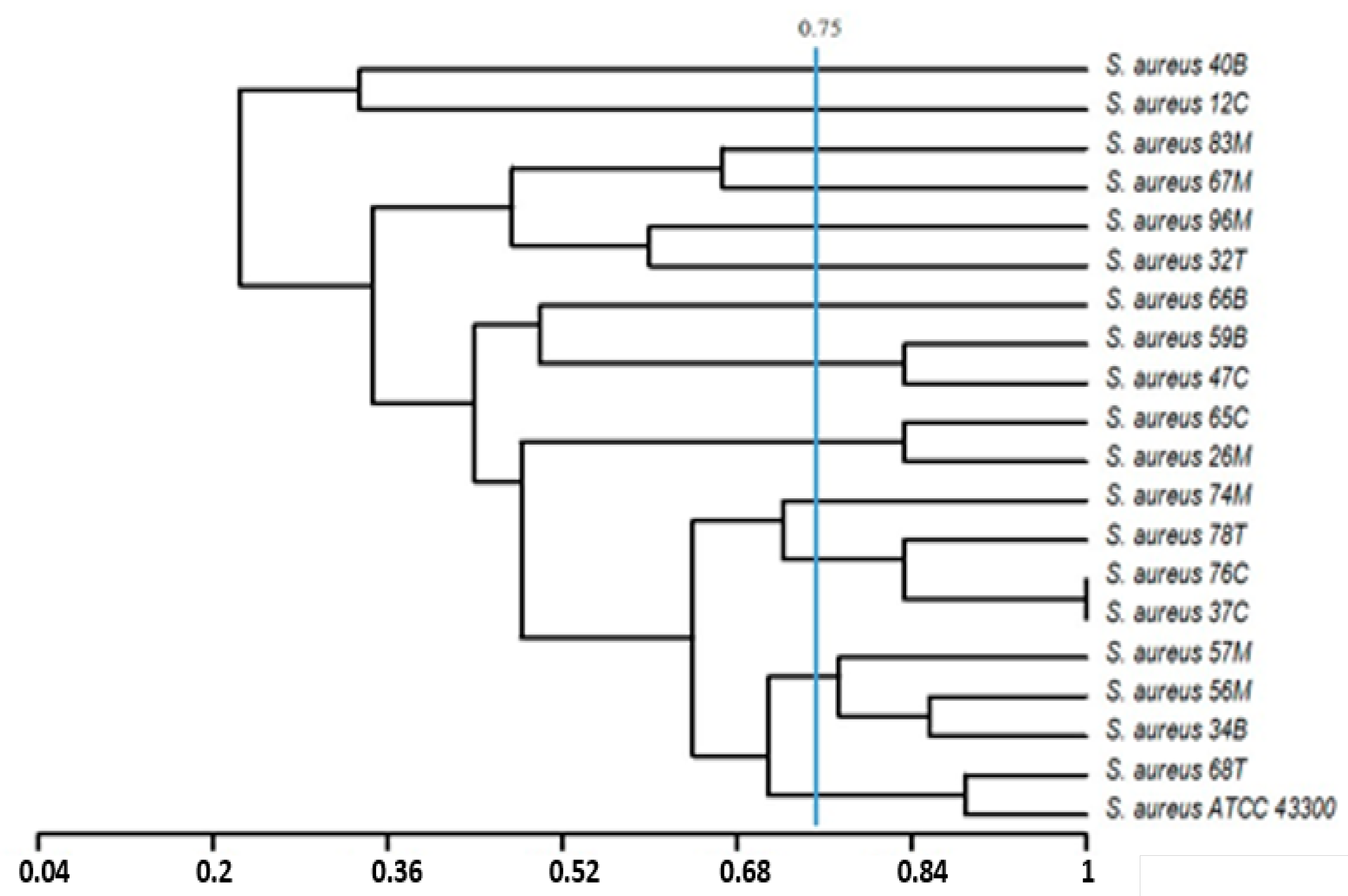
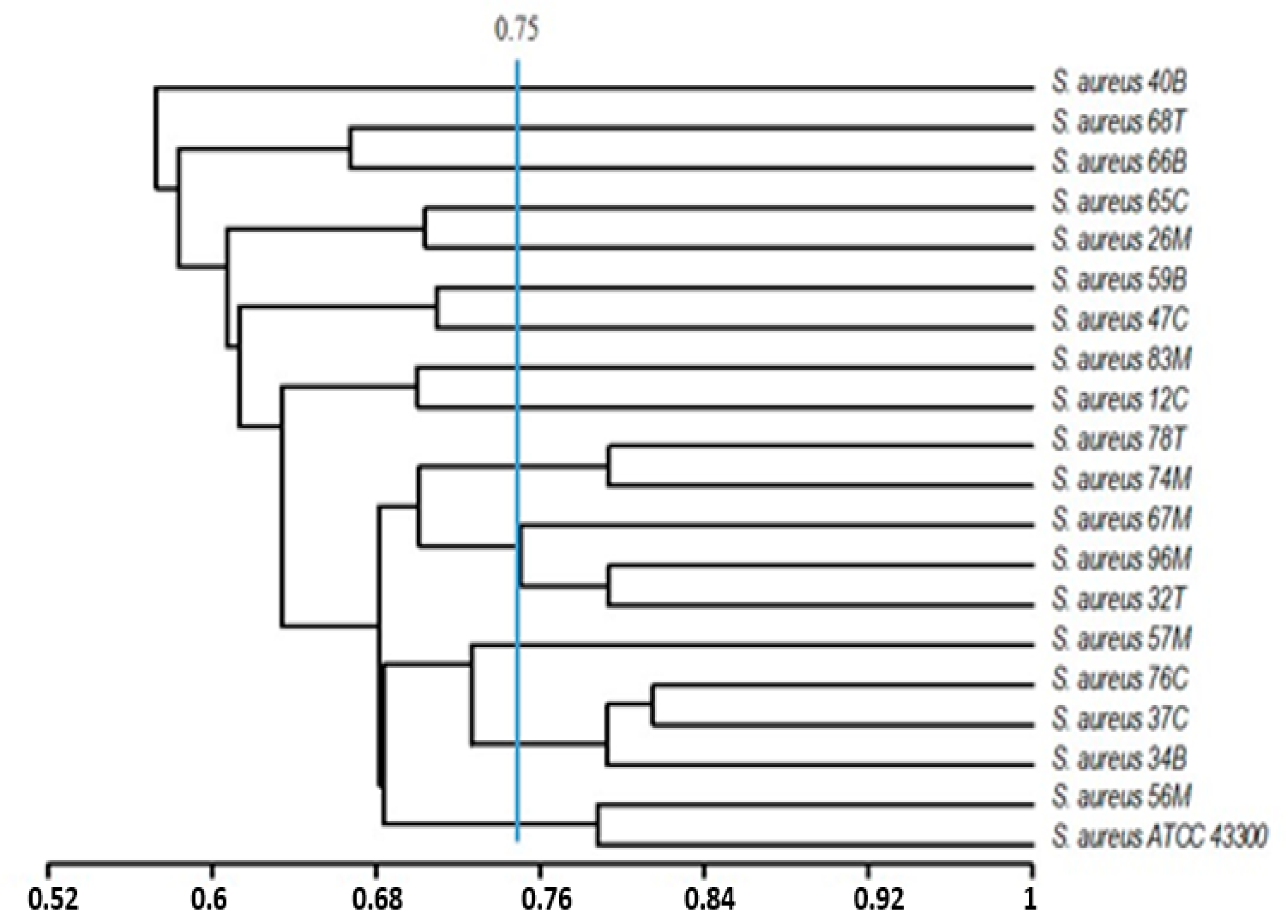
2.6. Statistical Analysis
3. Discussion
4. Materials and Methods
4.1. Sample Collection and Bacteriological Analysis
4.2. Susceptibility to Antibiotics
4.3. Adhesive Properties of the Identified Bacteria
4.4. Detection of Methicillin Resistance, Protease (sspA, sspB), Lipase (geh), α-Hemolysin (hla), and Adhesion Genes (icaA, icaD, cna, fnbA) in Staphylococcus spp. Strains
4.5. Data Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Halayem, S.; Nouira, O.; Bourgou, S.; Bouden, A.; Othman, S.; Halayem, M. The mobile: A new addiction upon adolescents. Tunis. Med. 2010, 88, 593–596. [Google Scholar]
- Schabrun, S.M.; Hoorn, W.V.D.; Moorcroft, A.; Greenland, C.; Hodges, P.W. Texting and Walking: Strategies for Postural Control and Implications for Safety. PLoS ONE 2014, 9, e84312. [Google Scholar] [CrossRef] [Green Version]
- Hamilton, J. Are main lines and mobile phones substitutes or complements? Evidence from Africa. Telecommun. Policy 2003, 27, 109–133. [Google Scholar] [CrossRef]
- Laatar, R.; Kachouri, H.; Borji, R.; Rebai, H.; Sahli, S. The effect of cell phone use on postural balance and mobility in older compared to young adults. Physiol. Behav. 2017, 173, 293–297. [Google Scholar] [CrossRef]
- Ekrakene, T.; Igeleke, C.L. Microorganisms Associated with Public Mobile Phones Along Benin-Sapele express way. J. Appl. Sci. Res. 2007, 3, 2009–2012. [Google Scholar]
- Bhat, S.S.; Hegde, S.K.; Salian, S. Potential of Mobile Phones to Serve as a Reservoir in Spread of Nosocomial Pathogens. Online J. Health Allied Sci. 2011, 10, 14. [Google Scholar]
- Egert, M.; Späth, K.; Weik, K.; Kunzelmann, H.; Horn, C.; Kohl, M.; Blessing, F. Bacteria on smartphone touchscreens in a German university setting and evaluation of two popular cleaning methods using commercially available cleaning products. Folia Microbiol. 2014, 60, 159–164. [Google Scholar] [CrossRef] [PubMed]
- Pierson, D.J. Is your smart phone spreading infection? Int. Med. Alert 2013, 35, 92–93. [Google Scholar]
- Brady, R.R.; Verran, J.; Damani, N.; Gibb, A. Review of mobile communication devices as potential reservoirs of nosocomial pathogens. J. Hosp. Infect. 2009, 71, 295–300. [Google Scholar] [CrossRef]
- Brady, R.; Wasson, A.; Stirling, I.; McAllister, C.; Damani, N. Is your phone bugged? The incidence of bacteria known to cause nosocomial infection on healthcare workers’ mobile phones. J. Hosp. Infect. 2006, 62, 123–125. [Google Scholar] [CrossRef]
- Gil Goldblatt, J.; Krief, I.; Klonsky, T.; Haller, D.; Milloul, V.; Sixsmith, D.M.; Srugo, I.; Potasman, I. Use of Cellular Telephones and Transmission of Pathogens by Medical Staff in New York and Israel. Infect. Control Hosp. Epidemiol. 2007, 28, 500–503. [Google Scholar] [CrossRef] [PubMed]
- Onyango, L.; Alreshidi, M.M. Adaptive Metabolism in Staphylococci: Survival and Persistence in Environmental and Clinical Settings. J. Pathog. 2018, 2018, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Alreshidi, M.M.; Dunstan, R.H.; Onyango, L.A.; Roberts, T.K. “Staphylococcal phenomics: Metabolomic and proteomic responses to environmental stressors”. In Microbial Pathogens and Strategies for Combating Them: Science, Technology and Education; Mendez-Vilas, A., Ed.; Formatex Research Center: Badajoz, Spain, 2013. [Google Scholar]
- Alreshidi, M.M.; Dunstan, R.H.; Macdonald, M.M.; Smith, N.D.; Gottfries, J.; Roberts, T.K.; Gottries, J. Metabolomic and proteomic responses of Staphylococcus aureus to prolonged cold stress. J. Proteom. 2015, 121, 44–55. [Google Scholar] [CrossRef] [PubMed]
- Alreshidi, M.M.; Dunstan, R.H.; Gottfries, J.; Macdonald, M.M.; Crompton, M.J.; Ang, C.-S.; Williamson, N.A.; Roberts, T.K. Changes in the Cytoplasmic Composition of Amino Acids and Proteins Observed in Staphylococcus aureus during Growth under Variable Growth Conditions Representative of the Human Wound Site. PLoS ONE 2016, 11, e0159662. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Onyango, L.; Dunstan, R.H.; Gottfries, J.; Von Eiff, C.; Roberts, T.K. Effect of Low Temperature on Growth and Ultra-Structure of Staphylococcus spp. PLoS ONE 2012, 7, e29031. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vuong, C.; Kocianova, S.; Voyich, J.M.; Yao, Y.; Fischer, E.R.; DeLeo, F.R.; Otto, M. A Crucial Role for Exopolysaccharide Modification in Bacterial Biofilm Formation, Immune Evasion, and Virulence. J. Biol. Chem. 2004, 279, 54881–54886. [Google Scholar] [CrossRef] [Green Version]
- Kanayama, A.K.; Takahashi, H.; Yoshizawa, S.; Tateda, K.; Kaneko, A.; Kobayashi, I. Staphylococcus aureus surface contamination of mobile phones and presence of genetically identical strains on the hands of nursing personnel. Am. J. Infect. Control 2017, 45, 929–931. [Google Scholar] [CrossRef]
- Foster, T.J.; Höök, M. Surface protein adhesins of Staphylococcus aureus. Trends Microbiol. 1998, 6, 484–688. [Google Scholar] [CrossRef]
- Saïd-Salim, B.; Dunman, P.M.; McAleese, F.M.; Macapagal, D.; Murphy, E.; McNamara, P.J.; Arvidson, S.; Foster, T.J.; Projan, S.J.; Kreiswirth, B.N. Global Regulation of Staphylococcus aureus Genes by Rot. J. Bacteriol. 2003, 185, 610–619. [Google Scholar] [CrossRef] [Green Version]
- Lowy, F.D. Staphylococcus aureus infections. N. Engl. J. Med. 1998, 339, 520–532. [Google Scholar] [CrossRef]
- Deresse, D.; Daka, D. Antibiotic-resistant Staphylococcus aureus isolated from mobile phone and hands of Health care workers in the Hawassa referral Hospital, South Ethiopia. J. Microbiol. Antimicrob. 2014, 6, 72–78. [Google Scholar] [CrossRef] [Green Version]
- Krumperman, P.H. Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl. Environ. Microbiol. 1983, 46, 165–170. [Google Scholar] [CrossRef] [Green Version]
- Srikanth, P.; Ezhil, R.; Suchitra, S.; Anandhi, I.; Maheswari, U.; Kalyani, J. The Mobile Phone in a Tropical Setting—Emerging Threat for Infection Control. Int. J. Infect. Dis. 2008, 12, e367. [Google Scholar] [CrossRef] [Green Version]
- Mark, D.; Leonard, C.; Breen, H.; Graydon, R.; O’Gorman, C.; Kirk, S. Mobile phones in clinical practice: Reducing the risk of bacterial contamination. Int. J. Clin. Pract. 2014, 68, 1060–1064. [Google Scholar] [CrossRef] [PubMed]
- Tagoe, D.N.; Gyande, V.K.; Ansah, E.O. Bacterial contamination of mobile phones: When your mobile phone could transmit more than just a call. WebmedCentral 2011, 2, 1–9. [Google Scholar]
- Datta, P.; Rani, H.; Chander, J.; Gupta, V. Bacterial contamination of mobile phones of health care workers. Indian J. Med. Microbiol. 2009, 27, 279. [Google Scholar] [CrossRef]
- Ulger, F.; Essen, S.; Dilek, A.; Yanik, K.; Gunaydin, M.; Leblebicioglu, H. Are we aware how contaminated our mobile phones are with nosocomial pathogens? Ann. Clin. Microb. Antimirob. 2009, 8, 7. [Google Scholar] [CrossRef] [Green Version]
- Badr, R.I.; Badr, H.I.; Ali, N.M. Mobile phones and nosocomial infections. Int. J. Infect. Control 2012, 8, 1–5. [Google Scholar] [CrossRef] [Green Version]
- Lavanya, J.; Rani, N.S.; Jais, M.; Upadhya, A.K. Microbial Contamination of Mobile Phones in a Tertiary Health Care Setting. Int. J. Curr. Microbiol. Appl. Sci. 2016, 5, 508–513. [Google Scholar] [CrossRef] [Green Version]
- Trivedi, H.R.; Desai, K.J.; Trivedi, L.P.; Malek, S.S.; Javdekar, T. Role of Mobile Phone in Spreading Hospital Acquired Infection: A Study in Different Group of Health Care Workers. Natl. J. Integr. Res. Med. 2011, 2, 61–66. [Google Scholar]
- Pal, K.; Chatterjee, M.; Sen, P.; Adhya, S. Cell Phones of Health Care Professionals: A Silent Source of Bacteria. Natl. J. Lab. Med. 2015, 4, 33–38. [Google Scholar]
- Tekerekoǧlu, M.S.; Duman, Y.; Serindag, A.; Cuǧlan, S.S.; Kaysadu, H.; Tunc, E.; Yakupogullari, Y.; Tekerekoglu, M.S. Do mobile phones of patients, companions and visitors carry multidrug-resistant hospital pathogens? Am. J. Infect. Control 2011, 39, 379–381. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Li, T.; Huang, X.; Xie, J.; Xu, Y.; Tu, J.; Qin, Z.; Parsons, C.; Wang, J.; Hu, L.; et al. Virulence gene profiling and molecular characterization of hospital-acquired Staphylococcus aureus isolates associated with bloodstream infection. Diagn. Microbiol. Infect. Dis. 2012, 74, 363–368. [Google Scholar] [CrossRef]
- Vorobieva, V.; Bazhukova, T.; Hanssen, A.M.; Caugant, D.A.; Semenova, N.; Haldorsen, B.C.; Simonsen, G.S.; Sundsfjord, A. Clinical isolates of Staphylococcus aureus from the Arkhangelsk region, Russia: Antimicrobial susceptibility, molecular epidemiology, and distribution of panton-valentine leukocidin genes. APMIS 2008, 116, 877–887. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, M.M.; Araujo, M.; Silva-Carvalho, M.; Beltrame, C.O.; Oliveira, C.; Figueiredo, A.; Oliveira, A. Emergence of clonal complex 5 (CC5) methicillin-resistant Staphylococcus aureus (MRSA) isolates susceptible to trimethoprim-sulfamethoxazole in a Brazilian hospital. Braz. J. Med. Biol. Res. 2012, 45, 637–643. [Google Scholar] [CrossRef] [PubMed]
- Maple, P.A.C.; Hamilton-Miller, J.; Brumfitt, W. World-wide antibiotic resistance in methicillin-resistant Staphylococcus aureus. Lancet 1989, 333, 537–540. [Google Scholar] [CrossRef]
- Nimmo, G.R.; Bell, J.M.; Mitchell, D.; Gosbell, I.; Pearman, J.W.; Turnidge, J.D. Antimicrobial Resistance in Staphylococcus aureus in Australian Teaching Hospitals, 1989–1999. Microb. Drug Resist. 2003, 9, 155–160. [Google Scholar] [CrossRef]
- Tokajian, S.; Haddad, M.; Andraos, R.; Hashwa, F.; Araj, G. Toxins and Antibiotic Resistance in Staphylococcus aureus Isolated from a Major Hospital in Lebanon. ISRN Microbiol. 2011, 2011, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Hamze, M.; Dabboussi, F.; Daher, W.; Izard, D. Antibiotic resistance of Staphylococcus aureus at north Lebanon: Place of the methicillin resistance and comparison of detection methods. Pathol. Biol. 2003, 51, 21–26. [Google Scholar] [CrossRef]
- Bischoff, W.E.; Wallis, M.L.; Tucker, K.B.; Reboussin, B.A.; Sherertz, R.J. Staphylococcus aureus Nasal Carriage in a Student Community Prevalence, Clonal Relationships, and Risk Factors. Infect. Control Hosp. Epidemiol. 2004, 25, 485–491. [Google Scholar] [CrossRef]
- Tambe, N.N.; Pai, C. A Study of microbial flora and MRSA harboured by mobile phones of health care personnel. Int. J. Recent Trends Sci. Technol. 2012, 4, 14–18. [Google Scholar]
- Morubagal, R.R.; Shivappa, S.G.; Mahale, R.P.; Neelambike, S.M. Study of bacterial flora associated with mobile phones of healthcare workers and non-healthcare workers. Iran J. Microbiol. 2017, 9, 143–151. [Google Scholar] [PubMed]
- Safdari, H.; Aryan, E.; Sadeghian, H.; Shams, S.F.; Aganj, M. Frequency of methicillin-resistant Staphylococcus aureus (MRSA) in nose and cellular phone of medical and non-medical personnel of emergency departments of Ghaem hospital in Mashhad city. Clin. Epidemiol. Glob. Health. 2020. [Google Scholar] [CrossRef] [Green Version]
- Wu, P.Z.; Zhu, H.; Thakur, A.; Willcox, M.D. Comparison of potential pathogenic traits of staphylococci that may contribute to corneal ulceration and inflammation. Aust. N.-Z. J. Ophthalmol. 1999, 27, 234–236. [Google Scholar] [CrossRef] [PubMed]
- Barretti, P.; Montelli, A.C.; Batalha, J.E.; Caramori, J.C.; Cunha, M.L. The role of virulence factors in the outcome of staphylococcal peritonitis in CAPD patients. BMC Infect. Dis. 2009, 9, 212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kouidhi, B.; Zmantar, T.; Hentati, H.; Bakhrouf, A. Cell surface hydrophobicity, biofilm formation, adhesives properties and molecular detection of adhesins genes in Staphylococcus aureus associated to dental caries. Microb. Pathog. 2010, 49, 14–22. [Google Scholar] [CrossRef]
- Khoramian, B.; Jabalameli, F.; Niasari-Naslaji, A.; Taherikalani, M.; Emaneini, M. Comparison of virulence factors and biofilm formation among Staphylococcus aureus strains isolated from human and bovine infections. Microb. Pathog. 2015, 88, 73–77. [Google Scholar] [CrossRef]
- Zmantar, T.; Chaieb, K.; Makni, H.; Miladi, H.; Ben Abdallah, F.; Mahdouani, K.; Bakhrouf, A. Detection by PCR of adhesins genes and slime production in clinical Staphylococcus aureus. J. Basic Microbiol. 2008, 48, 308–314. [Google Scholar] [CrossRef]
- Haddad, O.; Merghni, A.; Elargoubi, A.; Rhim, H.; Kadri, Y.; Mastouri, M. Comparative study of virulence factors among methicillin resistant Staphylococcus aureus clinical isolates. BMC Infect. Dis. 2018, 18, 560. [Google Scholar] [CrossRef] [Green Version]
- Pinchuk, I.V.; Beswick, E.J.; Reyes, V.E. Staphylococcal enterotoxins. Toxins (Basel) 2010, 2, 2177–2197. [Google Scholar] [CrossRef] [Green Version]
- Benkerroum, N. Staphylococcal enterotoxins and enterotoxin-like toxins with special reference to dairy products: An overview. Crit. Rev. Food Sci. Nutr. 2017, 58, 1943–1970. [Google Scholar] [CrossRef] [PubMed]
- Mkrtchyan, H.V.; Russell, C.A.; Wang, N.; Cutler, R.R. Could Public Restrooms Be an Environment for Bacterial Resistomes? PLoS ONE 2013, 8, e54223. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhoonderowa, A.; Gookool, S.; Biranjia-Hurdoyal, S.D.; Biranjia-Hurdoyal, S. The Importance of Mobile Phones in the Possible Transmission of Bacterial Infections in the Community. J. Community Health 2014, 39, 965–967. [Google Scholar] [CrossRef] [PubMed]
- Arciola, C.R.; Baldassarri, L.; Montanaro, L. Presence of icaA and icaD Genes and Slime Production in a Collection of Staphylococcal Strains from Catheter-Associated Infections. J. Clin. Microbiol. 2001, 39, 2151–2156. [Google Scholar] [CrossRef] [Green Version]
- Rohde, H.; Knobloch, J.; Horstkotte, M.A.; Mack, D. Correlation of Staphylococcus aureus icaADBC genotype and biofilm expression phenotype. J. Clin. Microbiol. 2001, 39, 4595–4596. [Google Scholar] [CrossRef] [Green Version]
- Arciola, C.R.; Campoccia, D.; Gamberini, S.; Baldassarri, L.; Montanaro, L. Prevalence of cna, fnbA and fnbB adhesin genes among Staphylococcus aureus isolates from orthopedic infections associated to different types of implant. FEMS Microbiol. Lett. 2005, 246, 81–86. [Google Scholar] [CrossRef] [Green Version]
- Thomas, M.G.; Peacock, S.; Daenke, S.; Berendt, A.R.; Young, B.; Johnson, S.; Minoo, B.; Shugarts, D.; Allen, M.; Ramey, R.R.; et al. Adhesion of Staphylococcus aureus to Collagen Is Not a Major Virulence Determinant for Septic Arthritis, Osteomyelitis, or Endocarditis. J. Infect. Dis. 1999, 179, 291–293. [Google Scholar] [CrossRef] [Green Version]
- Peacock, S.J.; Moore, C.E.; Justice, A.; Kantzanou, M.; Story, L.; Mackie, K.; O’Neill, G.; Day, N.P.J. Virulent Combinations of Adhesin and Toxin Genes in Natural Populations of Staphylococcus aureus. Infect. Immun. 2002, 70, 4987–4996. [Google Scholar] [CrossRef] [Green Version]
- Hartman, B.J.; Tomasz, A. Low-affinity penicillin-binding protein associated with β-lactam resistance in Staphylococcus aureus. J. Bacteriol. 1984, 158, 513–516. [Google Scholar] [CrossRef] [Green Version]
- Ubukata, K.; Nonoguchi, R.; Matsuhashi, M.; Konno, M. Expression and inducibility in Staphylococcus aureus of the mecA gene, which encodes a methicillin-resistant S. aureus-specific penicillin-binding protein. J. Bacteriol. 1989, 171, 2882–2885. [Google Scholar] [CrossRef] [Green Version]
- Lim, D.; Strynadka, N.C. Structural basis for the β-lactam resistance of PBP2a from methicillin-resistant Staphylococcus aureus. Nat. Struct. Biol. 2002, 9, 870–876. [Google Scholar] [CrossRef] [PubMed]
- Fuda, C.; Suvorov, M.; Vakulenko, S.B.; Mobashery, S. The Basis for Resistance to β-Lactam Antibiotics by Penicillin-binding Protein 2a of Methicillin-resistant Staphylococcus aureus. J. Biol. Chem. 2004, 279, 40802–40806. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bouchami, O.; Achour, W.; Mekni, M.A.; Rolo, J.; Ben Hassen, A. Antibiotic resistance and molecular characterization of clinical isolates of methicillin-resistant coagulase-negative staphylococci isolated from bacteremic patients in oncohematology. Folia Microbiol. 2011, 56, 122–130. [Google Scholar] [CrossRef] [PubMed]
- Noto, M.J.; Archer, G.L. A Subset of Staphylococcus aureus Strains Harboring Staphylococcal Cassette Chromosome mec (SCCmec) Type IV Is Deficient in CcrAB-Mediated SCCmec Excision. Antimicrob. Agents Chemother. 2006, 50, 2782–2788. [Google Scholar] [CrossRef] [Green Version]
- Harrison, E.M.; Paterson, G.K.; Holden, M.T.G.; Ba, X.; Rolo, J.; Morgan, F.J.E.; Pichon, B.; Kearns, A.; Zadoks, R.N.; Peacock, S.J.; et al. A novel hybrid SCCmec-mecC region in Staphylococcus sciuri. J. Antimicrob. Chemother. 2013, 69, 911–918. [Google Scholar] [CrossRef]
- Szczuka, E.; Krzyminska, S.; Bogucka, N.; Kaznowski, A. Multifactorial mechanisms of the pathogenesis of methicillin-resistant Staphylococcus hominis isolated from bloodstream infections. Antonie Van Leeuwenhoek 2017, 111, 1259–1265. [Google Scholar] [CrossRef] [Green Version]
- Harrison, E.M.; Paterson, G.K.; Holden, M.T.G.; Larsen, J.; Stegger, M.; Larsen, A.R.; Petersen, A.; Skov, R.L.; Christensen, J.M.; Zeuthen, A.B.; et al. Whole genome sequencing identifies zoonotic transmission of MRSA isolates with the novel mecA homologue mecC. EMBO Mol. Med. 2013, 5, 509–515. [Google Scholar] [CrossRef]
- Petersen, A.; Stegger, M.; Heltberg, O.; Christensen, J.; Zeuthen, A.; Knudsen, L.; Urth, T.; Sorum, M.; Schouls, L.; Larsen, J.; et al. Epidemiology of methicillin-resistant Staphylococcus aureus carrying the novel mecC gene in Denmark corroborates a zoonotic reservoir with transmission to humans. Clin. Microbiol. Infect. 2013, 19, E16–E22. [Google Scholar] [CrossRef] [Green Version]
- Snoussi, M.; Noumi, E.; Hajlaoui, H.; Usai, D.; Sechi, L.A.; Zanetti, S.; Bakhrouf, A. High potential of adhesion to abiotic and biotic materials in fish aquaculture facility by Vibrio alginolyticus strains. J. Appl. Microbiol. 2009, 106, 1591–1599. [Google Scholar] [CrossRef]
- Eddouzi, J.; Hofstetter, V.; Groenewald, M.; Manai, M.; Sanglard, D. Characterization of a New Clinical Yeast Species, Candida tunisiensis sp. nov., Isolated from a Strain Collection from Tunisian Hospitals. J. Clin. Microbiol. 2012, 51, 31–39. [Google Scholar] [CrossRef] [Green Version]
- Manjusha, S.; Sarita, G.; Elyas, K.; Chandrasekaran, M. Multiple Antibiotic Resistances of Vibrio Isolates from Coastal and Brackish Water Areas. Am. J. Biochem. Biotechnol. 2005, 1, 201–206. [Google Scholar] [CrossRef]
- Christensen, G.D.; Simpson, W.A.; Bisno, A.L.; Beachey, E.H. Adherence of biofilm producing strains of Staphylococcus epidermidis to smooth surfaces. Infect. Immun. 1982, 37, 318–326. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Freeman, D.J.; Falkiner, F.R.; Keane, C.T. New method for detecting slime production by coagulase negative staphylococci. J. Clin. Pathol. 1989, 42, 872–874. [Google Scholar] [CrossRef] [Green Version]
- Mack, D.; Bartscht, K.; Fischer, C.; Rohde, H.; De Grahl, C.; Dobinsky, S.; Horstkotte, M.A.; Kiel, K.; Knobloch, J.K. Genetic and biochemical analysis of Staphylococcus epidermidis biofilm accumulation. Methods Enzymol. 2001, 336, 215–239. [Google Scholar] [CrossRef] [PubMed]
- Rachid, S.; Ohlsen, K.; Wallner, U.; Hacker, J.; Hecker, M.; Ziebuhr, W. Alternative Transcription Factor ςB Is Involved in Regulation of Biofilm Expression in a Staphylococcus aureus Mucosal Isolate. J. Bacteriol. 2000, 182, 6824–6826. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- da Silva Meira, Q.G.; de Medeiros Barbosa, I.; Alves Aguiar Athayde, A.J.; de Siqueira-Júnior, J.P.; de Souza, E.L. Influence of temperature and surface kind on biofilm formation by Staphylococcus aureus from food-contact surfaces and sensitivity to sanitizers. Food Control 2012, 25, 469–475. [Google Scholar] [CrossRef] [Green Version]
- Geha, D.J.; Uhl, J.R.; Gustaferro, C.A.; Persing, D.H. Multiplex PCR for identification of methicillin-resistant staphylococci in the clinical laboratory. J. Clin. Microbiol. 1994, 32, 1768–1772. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karlsson, A.; Arvidson, S. Variation in Extracellular Protease Production among Clinical Isolates of Staphylococcus aureus Due to Different Levels of Expression of the Protease Repressor sarA. Infect. Immun. 2002, 70, 4239–4246. [Google Scholar] [CrossRef] [Green Version]
- Arciola, C.R.; Borsetti, E.; Collamati, S.; Baldassarri, L.; Montanaro, L. Detection of fibronectin-binding protein genes in staphylococcal strains from peri-prosthesis infections. New Microbiol. 1999, 22, 331–336. [Google Scholar]
- Liang, X.; Ji, Y. Alpha-toxin interferes with integrin-mediated adhesion and internalization of Staphylococcus aureus by epithelial cells. Cell. Microbiol. 2006, 8, 1656–1668. [Google Scholar] [CrossRef]
- Snoussi, M.; Noumi, E.; Usai, D.; Sechi, L.A.; Zanetti, S.; Bakhrouf, A. Distribution of some virulence related-properties of Vibrio alginolyticus strains isolated from Mediterranean seawater (Bay of Khenis, Tunisia): Investigation of eight Vibrio cholerae virulence genes. World J. Microbiol. Biotechnol. 2008, 24, 2133–2141. [Google Scholar] [CrossRef]

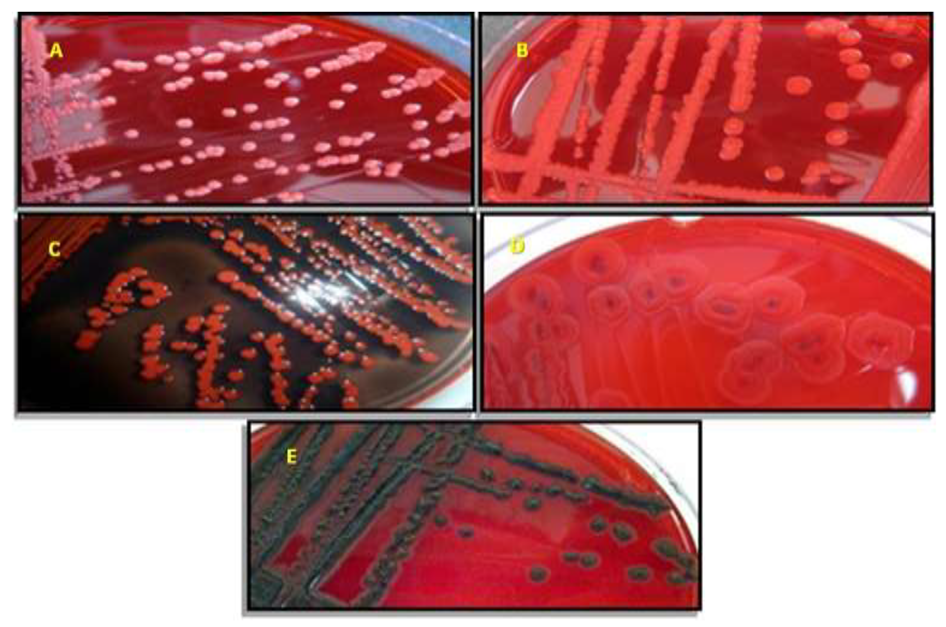

| Organism (No. Tested) | Color of Isolated Colonies |
|---|---|
| Methicillin susceptible (1) | Mauve |
| Methicillin resistant (18) | Mauve |
| S. haemolyticus (2) | Light blue |
| S. warneri (3) | White |
| Strains | MALDI-TOF-MS | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 |
|---|---|---|---|---|---|---|---|---|---|---|
| ATCC 43300 | S. aureus | + | + | + | + | − | + | + | + | β |
| 12C | S. aureus | + | + | + | + | − | + | − | + | β |
| 26M | S. aureus | + | + | + | + | + | + | + | + | β |
| 32T | S. aureus | + | + | + | + | + | − | + | − | α |
| 34B | S. aureus | + | + | + | + | − | − | + | + | α |
| 37C | S. aureus | + | + | + | + | − | − | + | − | α |
| 40B | S. aureus | + | + | + | − | + | + | + | + | α |
| 47C | S. aureus | + | + | + | − | + | − | + | − | β |
| 56M | S. aureus | + | + | + | + | − | − | + | − | α |
| 57 M | S. aureus | + | + | + | + | − | − | + | − | α |
| 59B | S. aureus | + | + | + | + | + | − | + | − | σ |
| 65C | S. aureus | + | + | + | + | + | − | − | − | σ |
| 66B | S. aureus | + | + | + | − | − | − | + | − | α |
| 67M | S. aureus | + | + | + | − | − | + | + | + | β |
| 68T | S. aureus | + | + | + | − | − | + | + | − | β |
| 74M | S. aureus | + | + | + | + | + | + | + | − | α |
| 76C | S. aureus | + | + | + | + | − | − | + | + | β |
| 78T | S. aureus | + | + | + | + | + | − | + | − | σ |
| 83M | S. aureus | + | + | + | + | + | + | + | + | β |
| 96M | S. aureus | + | + | + | + | + | + | + | − | σ |
| 7T | S. warneri | − | + | + | + | − | − | − | − | α |
| 31C | S. warneri | − | + | + | + | − | + | − | − | α |
| 47B | S. warneri | − | + | + | − | − | + | + | − | σ |
| 30C | S. haemolyticus | − | + | + | + | + | + | + | − | α |
| 39B | S. haemolyticus | − | + | + | − | − | − | + | − | α |
| % positivity | 80 | 100 | 100 | 70.83 | 45.83 | 45.83 | 83.33 | 29.16 | 29.16 (β) | |
| Strains | Antibiotics | MARI | mecA | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | |||
| ATCC 43300 | R | S | R | R | R | R | R | S | S | R | S | S | R | R | S | S | R | R | 0.611 | mecA+ |
| 12C | R | S | S | R | R | S | R | S | R | R | R | S | R | R | S | R | R | R | 0.666 | mecA− |
| 26M | R | S | S | S | R | S | R | S | R | R | S | S | R | S | S | S | R | R | 0.444 | mecA+ |
| 32T | R | S | R | S | R | S | R | S | R | R | R | S | R | R | S | S | R | R | 0.611 | mecA− |
| 34B | R | S | R | S | R | S | S | S | R | S | R | S | R | S | S | S | R | S | 0.388 | mecA− |
| 37C | R | S | R | S | R | S | R | S | R | R | R | S | R | S | S | S | R | R | 0.555 | mecA− |
| 40B | R | S | R | S | R | S | S | S | S | S | R | S | R | R | S | S | R | R | 0.444 | mecA− |
| 47C | R | S | R | S | R | S | S | S | R | R | R | S | R | R | S | S | R | R | 0.555 | mecA− |
| 56M | S | S | R | R | R | S | R | S | R | R | S | S | R | R | S | S | R | R | 0.555 | mecA− |
| 57M | R | S | R | R | R | R | S | S | R | R | R | S | R | S | S | R | R | R | 0.666 | mecA− |
| 59B | R | S | R | R | R | R | R | R | R | R | R | S | R | S | S | R | R | R | 0.777 | mecA+ |
| 65C | R | S | S | S | R | S | S | S | S | R | S | S | R | R | S | S | R | R | 0.388 | mecA+ |
| 67M | R | S | R | R | R | S | R | S | R | R | R | S | R | R | S | S | R | R | 0.666 | mecA− |
| 66B | R | S | S | S | R | S | S | R | R | R | R | S | R | S | S | R | R | R | 0.555 | mecA+ |
| 68T | R | R | R | R | - | S | - | S | R | R | R | R | R | S | R | R | R | R | 0.812 | mecA+ |
| 74M | R | S | R | S | R | R | R | S | S | R | S | R | R | R | S | S | R | R | 0.611 | mecA+ |
| 76C | R | S | S | S | R | S | R | S | R | S | R | S | R | R | S | S | R | R | 0.500 | mecA− |
| 78T | R | S | R | S | R | S | R | S | S | R | R | S | R | R | S | S | R | R | 0.555 | mecA+ |
| 83M | R | S | S | S | R | S | R | S | S | R | R | S | R | R | R | S | R | R | 0.555 | mecA− |
| 96M | R | S | R | R | R | R | R | S | R | R | R | S | R | R | S | S | R | R | 0.722 | mecA− |
| 7T | R | R | S | R | R | S | R | S | R | R | R | S | R | R | S | R | R | R | 0.722 | mecA+ |
| 31C | R | S | R | R | R | R | R | R | R | R | R | S | R | S | S | R | R | R | 0.777 | mecA− |
| 47B | R | S | S | S | R | R | - | S | R | R | S | S | R | R | S | S | R | R | 0.529 | mecA− |
| 39B | R | S | R | R | R | S | R | S | R | R | R | R | R | R | S | S | R | R | 0.722 | mecA+ |
| 30C | R | S | S | R | R | S | R | S | R | R | R | S | R | R | S | R | R | R | 0.666 | mecA− |
| Strains | Safranin Assay | Slime Phenotype (CRA) | Biofilm on Polystyrene | Biofilm on Glass | |||
|---|---|---|---|---|---|---|---|
| (OD570) ± SD | Interpretation | (OD570) ± SD | Interpretation | ||||
| ATCC 43300 | +++ | Black | S+ | 1.89 ± 0.13 | H | 1.48 ± 0.15 | H |
| 12C | +++ | Black | S+ | 1.34 ± 0.18 | H | 1.09 ± 0.05 | H |
| 26M | +++ | Red | S− | 1.31 ± 0.15 | H | 0.57 ± 0.1 | M |
| 32T | + | Bordeaux | S− | 0.57 ± 0.4 | M | 0.55 ± 0.04 | M |
| 34B | +++ | Red | S− | 0.64 ± 0 | M | 0.65 ± 0.09 | M |
| 37C | ++ | Red | S− | 0.16 ± 0.35 | M | 0.99 ± 0.01 | M |
| 40B | +++ | Bordeaux | S− | 0.28 ± 0 | M | 1.10 ± 0.16 | H |
| 47C | ++ | Red | S− | 0.64 ± 0 | M | 0.57 ± 0.06 | M |
| 56M | ++ | Red with black center | S+ | 0.21 ± 0 | M | 1.11 ± 0.08 | H |
| 57 M | +++ | Red | S− | 0.74 ± 0 | M | 1.68 ± 0.2 | H |
| 59B | +++ | Pink | S− | 0.16 ± 0 | M | 0.717 ± 0.12 | M |
| 65C | ++ | Pink | S− | 0.15 ± 0 | M | 0.88 ± 0.05 | M |
| 66B | ++ | Red | S− | 2.73 ± 0.56 | H | 0.75 ± 0.2 | M |
| 67M | +++ | Orange | S− | 1.27 ± 0.17 | H | 1.28 ± 0.1 | H |
| 68T | +++ | Red | S− | 2.14 ± 0.64 | H | 0.45 ± 0.09 | M |
| 74M | +++ | Pink | S− | 0.40 ± 0 | M | 0.50 ± 0.09 | M |
| 76C | +++ | Black | S+ | 1.63 ± 0.57 | H | 0.38 ± 0.04 | M |
| 78T | + | Red | S− | 0.47 ± 0 | M | 0.90 ± 0.15 | M |
| 83M | +++ | Red with black center | S+ | 0.85 ± 0.13 | M | 0.32 ± 0.04 | M |
| 96M | +++ | Red | S− | 0.36 ± 0.3 | M | 0.30 ± 0.05 | M |
| 7T | +++ | Red | S− | 0.33 ± 0 | M | 0.87 ± 0.13 | M |
| 31C | +++ | Pink | S− | 0.59 ± 0 | M | 0.46 ± 0.08 | M |
| 47B | ++ | Pink | S− | 0.66 ± 0.12 | M | 0.56 ± 0.03 | M |
| 30C | + | Red | S− | 0.17 ± 0 | M | 0.98 ± 0.07 | M |
| 39B | ++ | Pink | S− | 1.07 ± 0.19 | H | 0.84 ± 0.11 | M |
| Strains | Adhesion | Haemolysins | Exoenymes | |||||
|---|---|---|---|---|---|---|---|---|
| icaA | icaD | cna | fnbA | Hla | geh | sspA | sspB | |
| ATCC 43300 | + | + | + | + | + | + | + | + |
| 12C | − | − | − | + | − | + | − | + |
| 26M | − | − | + | + | + | + | − | + |
| 32T | + | + | − | + | + | + | − | − |
| 34B | + | + | + | + | − | + | − | + |
| 37C | + | + | + | + | − | − | − | + |
| 40B | − | − | − | − | − | − | − | + |
| 47C | − | + | + | − | + | − | + | + |
| 56M | + | + | + | + | + | + | − | + |
| 57M | + | + | − | + | − | + | − | + |
| 59B | − | + | + | − | + | − | + | + |
| 65C | − | − | + | + | + | − | − | + |
| 66B | + | − | − | + | − | − | − | + |
| 67M | + | + | + | − | + | − | − | − |
| 68T | + | + | + | + | + | + | − | + |
| 74M | + | + | + | + | − | − | − | − |
| 76C | + | + | + | + | − | − | − | + |
| 78T | + | + | + | + | − | − | − | + |
| 83M | + | − | − | + | − | − | − | − |
| 96M | + | − | − | + | + | − | − | − |
| 7T | − | + | − | − | − | − | − | − |
| 31C | − | + | − | − | − | − | − | + |
| 47B | − | + | − | + | − | − | − | − |
| 30C | + | + | − | + | − | − | + | − |
| 39B | + | − | + | + | − | + | + | + |
| % positivity | 62.5 | 66.66 | 50.00 | 75.00 | 37.50 | 33.33 | 16.66 | 66.66 |
| p-Value (Univariate Analysis) | Adhesion Related Genes | Haemolysin Gene | Exoenymes Genes | |||||
|---|---|---|---|---|---|---|---|---|
| icaA | icaD | cna | fnbA | Hla | geh | sspA | sspB | |
| Biofilm on glass | 0.489 | 0.783 | 0.783 | 0.489 | 0.581 | 0.945 | 0.836 | 0.581 |
| Biofilm on polystyrene | 0.489 | 0.783 | 0.783 | 0.489 | 0.581 | 0.945 | 0.836 | 0.581 |
| Primer | Primer Sequence | PCR Conditions | Product Size (pb) | Reference |
|---|---|---|---|---|
| mecA-F | 5′-GTA GAA ATG ACT GAA CGT CCG ATAA-3′ | 94 °C for 5 min 30 Cycles (1 min at 94 °C, 1 min at 55 °C, 2 min at 72 °C) 72 °C for 10 min | 310 | [78] |
| mecA-R | 5′-CCAATT CCA CAT TGT TTC GGT CTAA-3′ | |||
| sspA-F | 5′-GAC AAC AGC GAC ACT TGT GA-3′ | 94 °C for 5 min 30 Cycles (30 s at 94 °C, 30 s min at 45 °C, 45 s at 72 °C) 72 °C for 10 min | 292 | [79] |
| sspA-R | 5′-AGT ATC TTT ACC TAC AAC TAC A-3′ | |||
| sspB-F | 5′-TGA AGA AGA TGG CAA AGT TAG-3′ | 94 °C for 5 min 30 Cycles (30 s at 94 °C, 30 s min at 47 °C, 45 s at 72 °C) 72 °C for 10 min | 493 | |
| sspB-R | 5′-TTG AGA TAC ACT TTG TGC AAG-3′ | |||
| geh-F | 5′-GCACAAGCCTCGG-3′ | 94 °C for 5 min 30 Cycles (30 s at 94 °C, 30 s min at 40 °C, 45 s at 72 °C) 72 °C for 10 min | 473 | [20] |
| geh-R | 5′-GACGGGGGTGTAG-3′ | |||
| icaA-F | 5′-ACACTTGCTGGCGCAGTCAA-3′ | 94 °C for 5 min 30 Cycles (30 s at 94 °C, 30 s min at 45 °C, 45 s at 72 °C) 72 °C for 10 min | 188 | [80] |
| icaA-R | 5′-TCTGGAACCAACATCCAACA-3′ | |||
| icaD-F | 5′-ACACTTGCTGGCGCAGTCAA-3′ | 94 °C for 5 min 30 Cycles (30 s at 94 °C, 30 s min at 55 °C, 45 s at 72 °C) 72 °C for 10 min | 198 | |
| icaD-R | 5′-TCTGGAACCAACATCCAACA-3′ | |||
| cna-F | 5′-AAAGCGTTGCCTAGTGGAGA-3′ | 94 °C for 5 min 30 Cycles (30 s at 94 °C, 30 s min at 62 °C, 45 s at 72 °C) 72 °C for 10 min | 192 | [57] |
| cna-R | 5′-AGTGCCTTCCCAAACCTTTT-3′ | |||
| fnbA-F | 5′-GATACAAACCCAGGTGGTGG-3′ | 191 | ||
| fnbA-R | 5′-TGTGCTTGACCATGCTCTTC-3′ | |||
| hla-F | 5′CAACTGATAAAAAAGTAGGCTGGAAAGTGAT-3′ | 94 °C for 5 min 35 Cycles (30 s at 94 °C, 60 s min at 59 °C, 60 s at 72 °C) 72 °C for 10 min | 201 | [81] |
| hla-R | 5′-CTGGTGAAAACCCTGAAGATAATAGAG-3′ |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Noumi, E.; Merghni, A.; Alreshidi, M.; Del Campo, R.; Adnan, M.; Haddad, O.; De Feo, V.; Snoussi, M. Phenotypic and Genotypic Characterization with MALDI-TOF-MS Based Identification of Staphylococcus spp. Isolated from Mobile Phones with their Antibiotic Susceptibility, Biofilm Formation, and Adhesion Properties. Int. J. Environ. Res. Public Health 2020, 17, 3761. https://doi.org/10.3390/ijerph17113761
Noumi E, Merghni A, Alreshidi M, Del Campo R, Adnan M, Haddad O, De Feo V, Snoussi M. Phenotypic and Genotypic Characterization with MALDI-TOF-MS Based Identification of Staphylococcus spp. Isolated from Mobile Phones with their Antibiotic Susceptibility, Biofilm Formation, and Adhesion Properties. International Journal of Environmental Research and Public Health. 2020; 17(11):3761. https://doi.org/10.3390/ijerph17113761
Chicago/Turabian StyleNoumi, Emira, Abderrahmen Merghni, Mousa Alreshidi, Rosa Del Campo, Mohd Adnan, Ons Haddad, Vincenzo De Feo, and Mejdi Snoussi. 2020. "Phenotypic and Genotypic Characterization with MALDI-TOF-MS Based Identification of Staphylococcus spp. Isolated from Mobile Phones with their Antibiotic Susceptibility, Biofilm Formation, and Adhesion Properties" International Journal of Environmental Research and Public Health 17, no. 11: 3761. https://doi.org/10.3390/ijerph17113761









