Klebsiella and Enterobacter Isolated from Mangrove Wetland Soils in Thailand and Their Application in Biological Decolorization of Textile Reactive Dyes
Abstract
1. Introduction
2. Materials and Methods
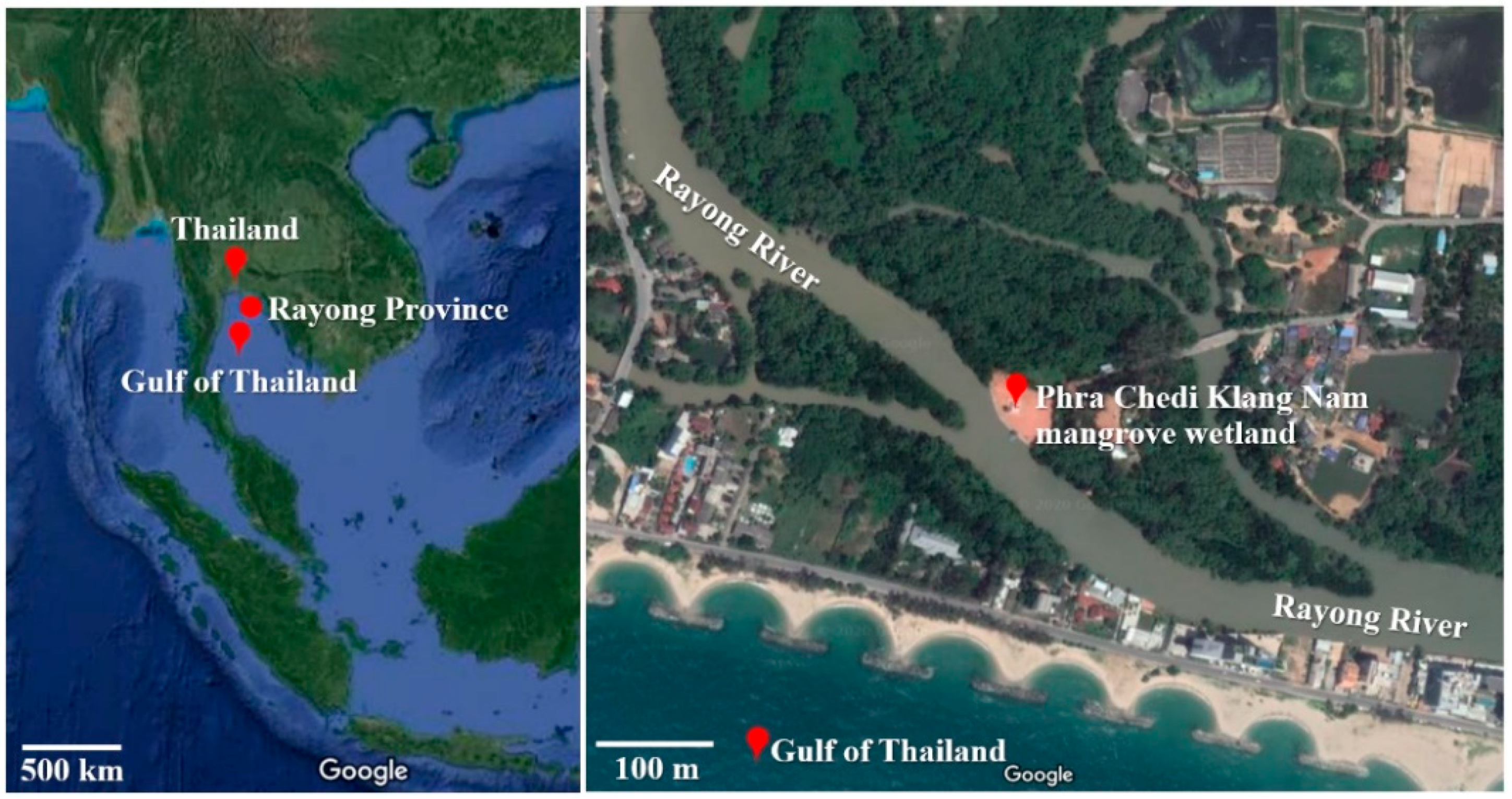
2.1. Study Area
2.2. Sampling of Mangrove Wetland Soils
2.3. Enrichment and Isolation of Bacteria from Mangrove Wetland Soils
2.4. Screening of Dye-Decolorizing Bacteria by Rapid RBBR Method
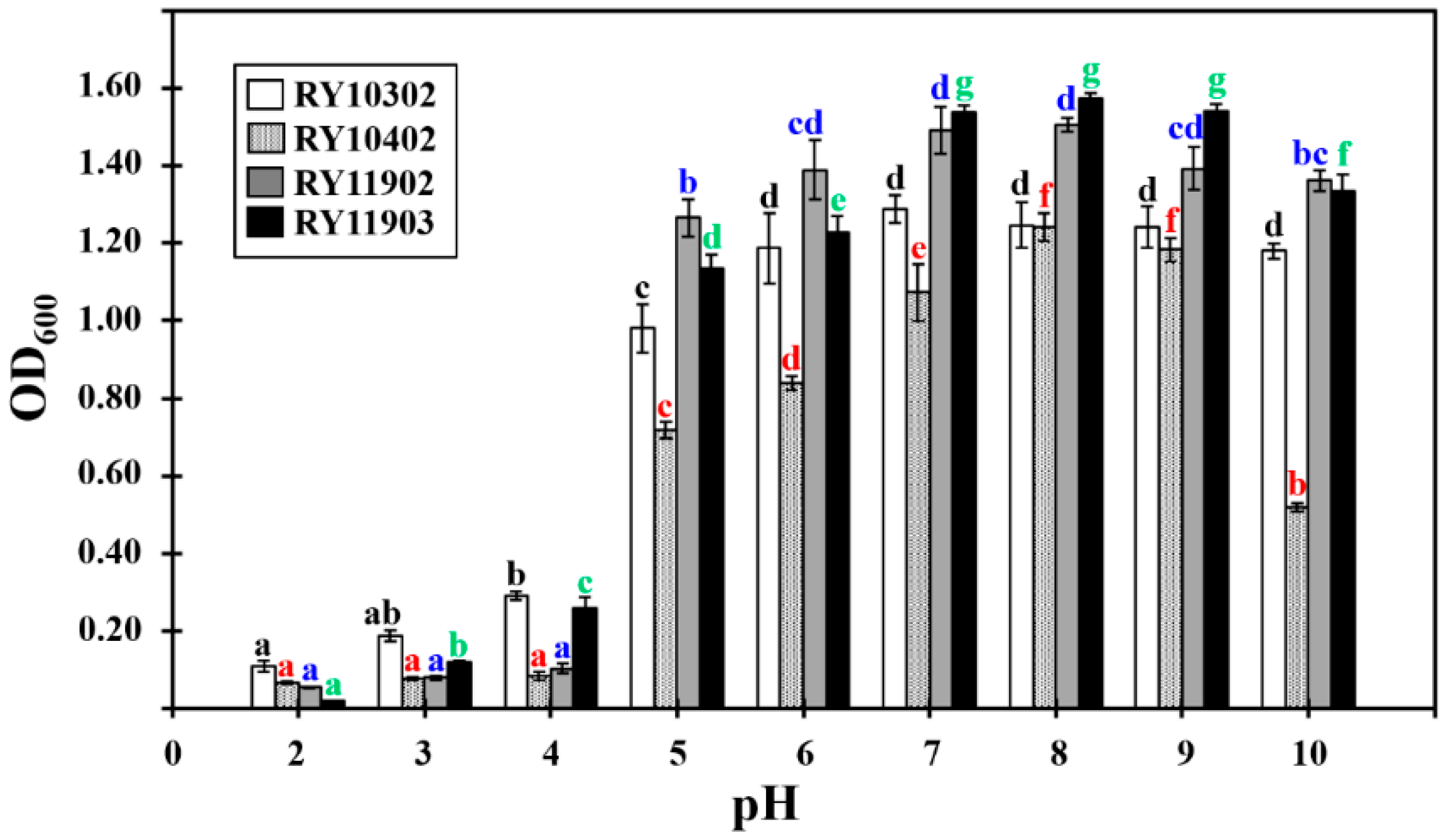
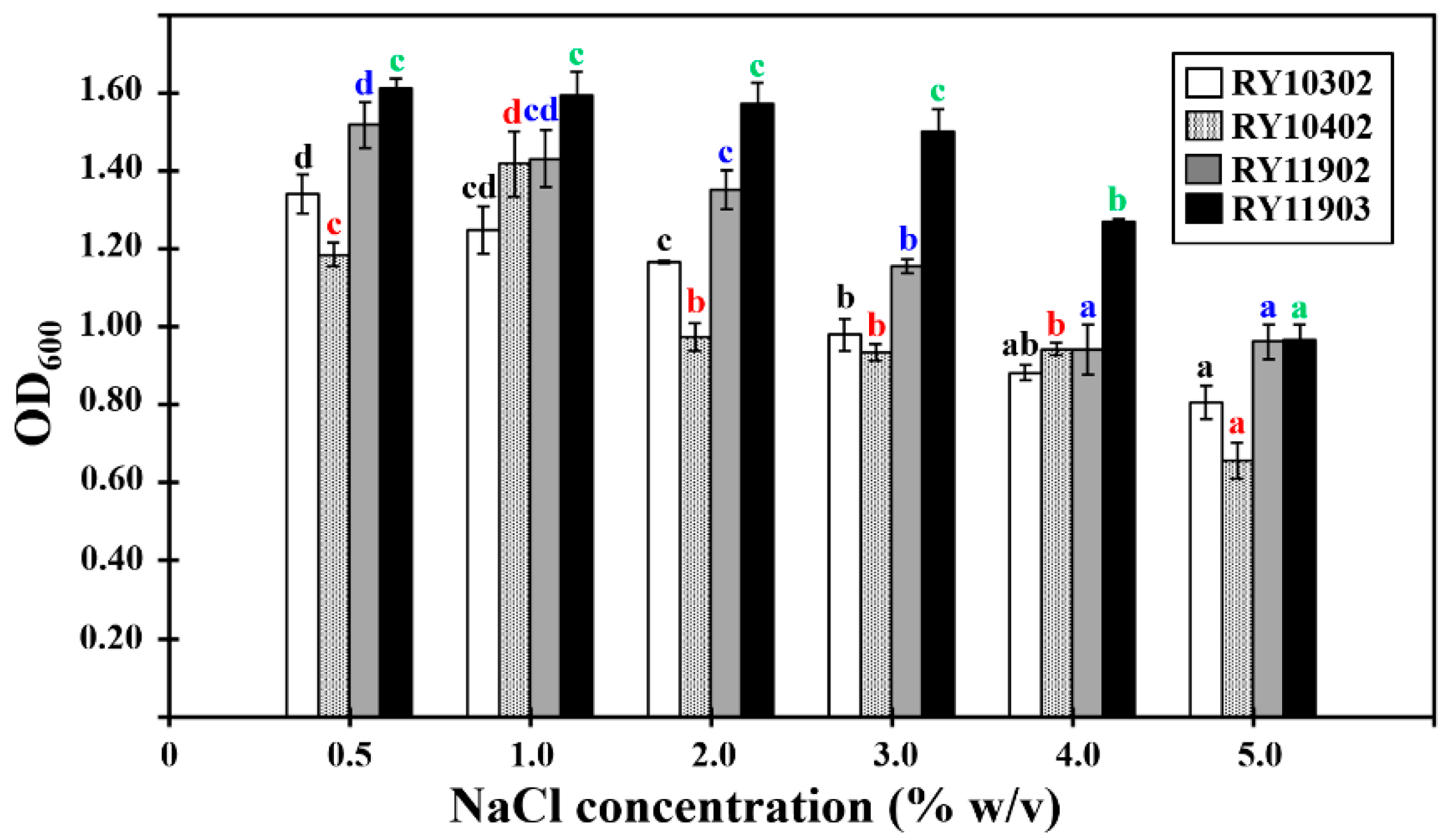
2.5. Growth Characterization of Dye-Decolorizing Bacteria
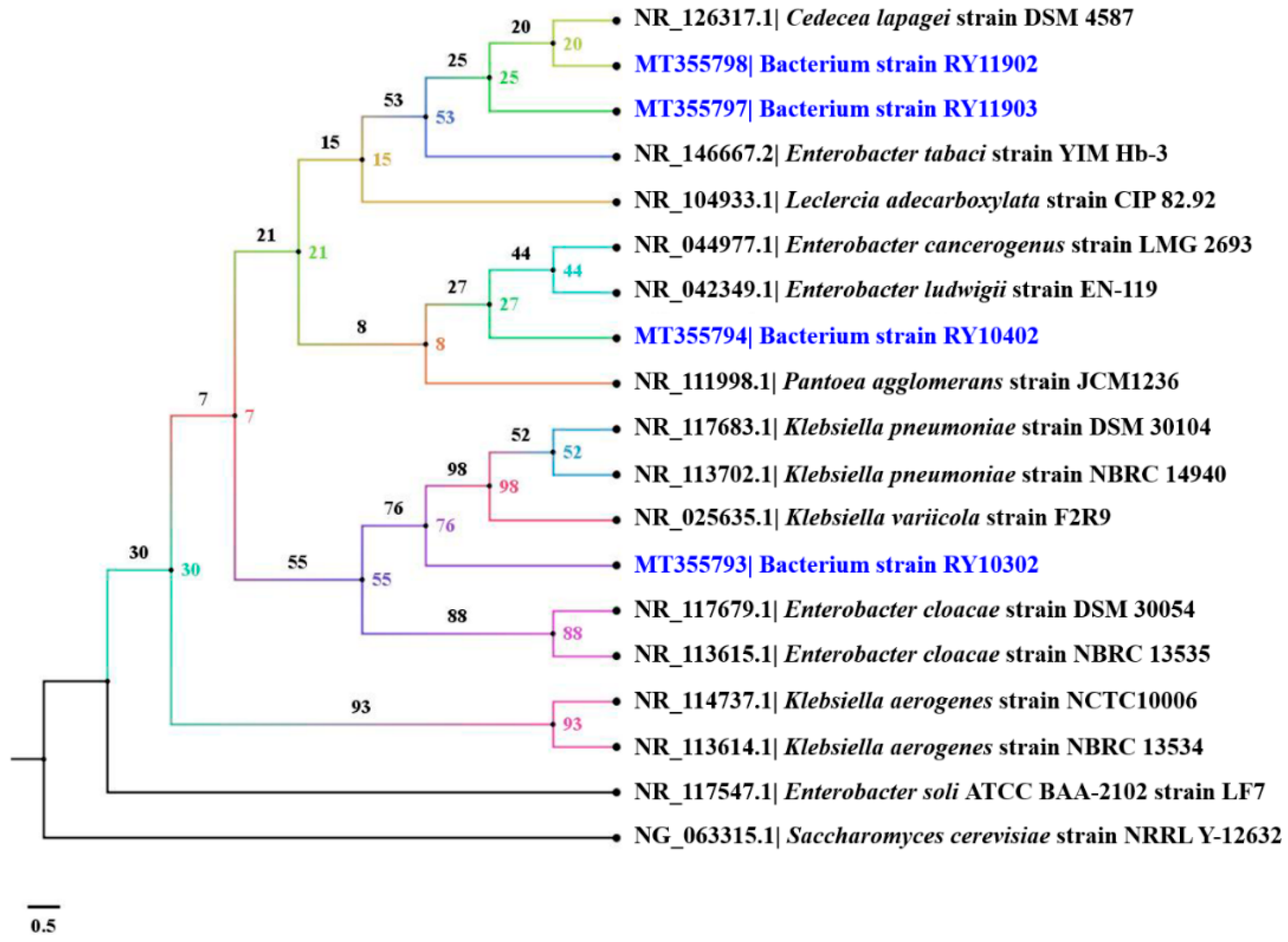
2.6. Identification of Dye-Decolorizing Bacteria
2.7. Preparation of Crude Peroxidases and Laccase
2.8. Determination of Bacterial Peroxidase and Laccase Activities
2.9. Dye Decolorization Efficiency of Bacterial Monoculture and Mixed Culture
2.10. Data Analysis
3. Results and Discussion
3.1. Sampling of Mangrove Soils
3.2. Enrichment and Isolation of Bacteria from Mangrove Soil
3.3. Screening of Dye-Decolorizing Bacteria by Rapid RBBR Method
3.4. Growth Characterization of Dye-Decolorizing Bacteria
3.5. Genotypic Identification of Dye-Decolorizing Bacteria
3.6. Determination of Bacterial Peroxidase and Laccase Activities
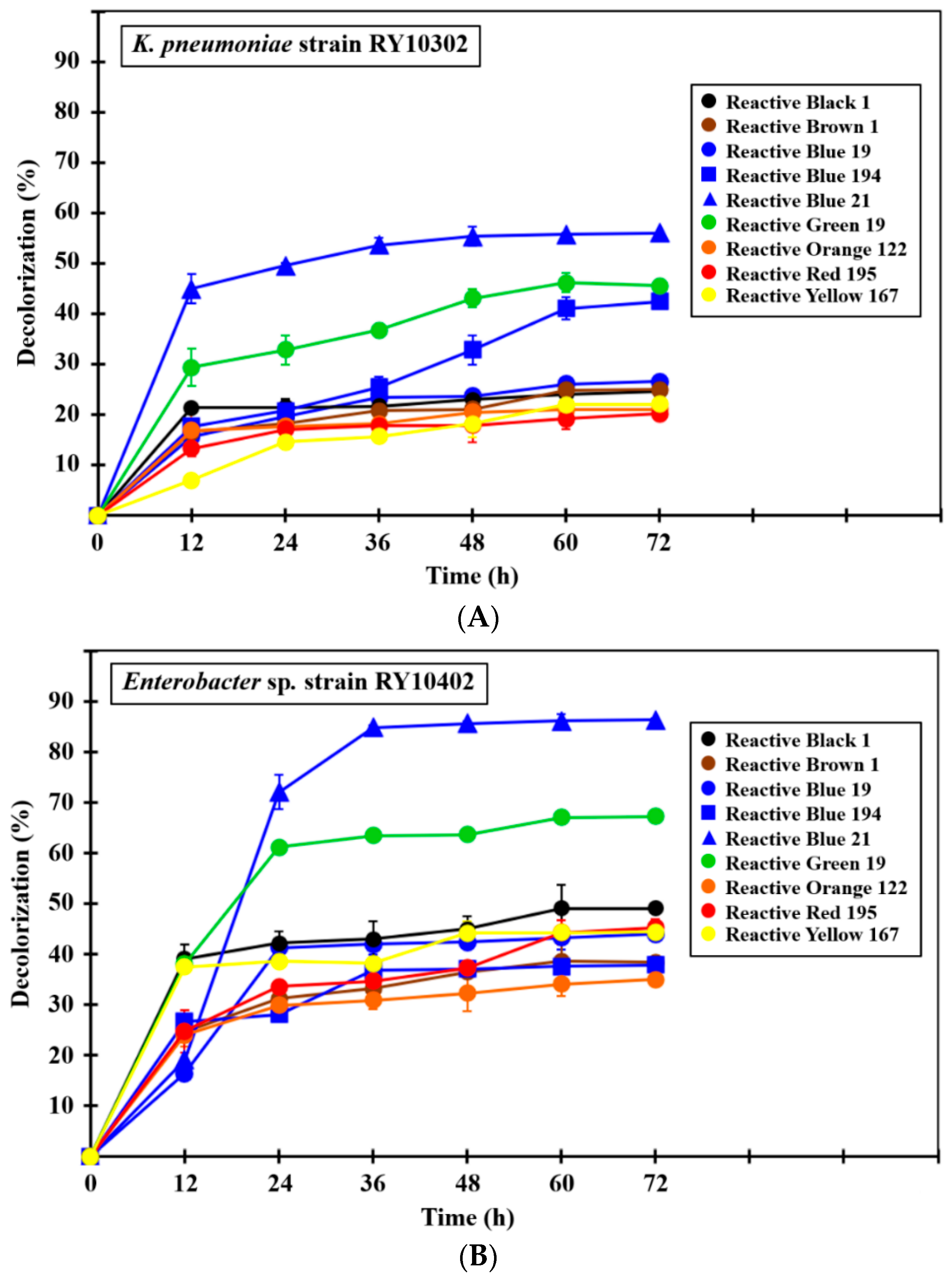
3.7. Dye Decolorization Efficiency of Bacterial Monoculture and Mixed Culture
4. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Chakraborty, J.N. Fundamentals and Practices in Colouration of Textiles; Woodhead Publishing: New Delhi, India, 2010; pp. 57–75. [Google Scholar]
- Khatri, A.; Peerzada, M.H.; Mohsin, M.; White, M. A review on developments in dyeing cotton fabrics with reactive dyes for reducing effluent pollution. J. Clean. Prod. 2015, 87, 50–57. [Google Scholar] [CrossRef]
- Ekanayake, E.M.M.S.; Manage, P.M. Decolorization of CI Direct Blue 201 textile dye by native bacteria. Int. J. Multidiscip. Stud. 2017, 4, 48–57. [Google Scholar] [CrossRef]
- Saratale, R.G.; Saratale, G.D.; Chang, J.S.; Govindwar, S.P. Decolorization and biodegradation of reactive dyes and dye wastewater by a developed bacterial consortium. Biodegradation 2010, 21, 999–1015. [Google Scholar] [CrossRef] [PubMed]
- Bayar, S.; Erdogan, M. Removal of COD and color from Reactive Red 45 azo dye wastewater using Fenton and Fenton-like oxidation processes: Kinetic studies. Appl. Ecol. Environ. Res. 2019, 17, 1517–1529. [Google Scholar] [CrossRef]
- Jegan, J.; Praveen, S.; Bhagavathi Pushpa, T.; Gokulan, R. Sorption kinetics and isotherm studies of cationic dyes using groundnut (Arachis hypogaea) shell derived biochar a low-cost adsorbent. Appl. Ecol. Environ. Res. 2020, 18, 1925–1939. [Google Scholar] [CrossRef]
- Imran, M.; Crowley, D.E.; Khalid, A.; Hussain, S.; Mumtaz, M.W.; Arshad, M. Microbial biotechnology for decolorization of textile wastewaters. Rev. Environ. Sci. Biotechnol. 2015, 14, 73–92. [Google Scholar] [CrossRef]
- Gupta, V.K.; Khamparia, S.; Tyagi, I.; Jaspal, D.; Malviya, A. Decolorization of mixture of dyes: A critical review. Glob. J. Environ. Sci. Manag. 2015, 1, 71–94. [Google Scholar]
- Singh, R.P.; Singh, P.K.; Singh, R.L. Bacterial decolorization of textile azo dye Acid Orange by Staphylococcus hominis RMLRT03. Toxicol. Int. 2014, 21, 160–166. [Google Scholar] [CrossRef]
- Luo, X.; Liang, C.; Hu, Y. Comparison of different enhanced coagulation methods for Azo Dye removal from wastewater. Sustainability 2019, 11, 4760. [Google Scholar] [CrossRef]
- Saranraj, P.; Manigandan, M. Enzymes involved in bacterial decolourization and degradation of textile azo dyes. Indo-Asian J. Multidiscip. Res. 2018, 4, 1369–1376. [Google Scholar]
- Leme, D.M.; de Rodrigues Oliveira, G.A.; Meireles, G.; Brito, L.B.; Rodrigues, L.B.; de Palma Oliveira, D. Eco-and genotoxicological assessments of two reactive textile dyes. J. Toxicol. Environ. Health Part A 2015, 78, 287–300. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.C.; Corrêa, A.D.; Amorim, M.T.S.P.; Parpot, P.; Torres, J.A.; Chagas, P.M.B. Decolorization of the phthalocyanine dye reactive blue 21 by turnip peroxidase and assessment of its oxidation products. J. Mol. Catal. B Enzym. 2012, 77, 9–14. [Google Scholar] [CrossRef]
- Chantarasiri, A.; Boontanom, P. Decolorization of synthetic dyes by ligninolytic Lysinibacillus sphaericus JD1103 isolated from Thai wetland ecosystems. AACL Bioflux 2017, 10, 814–819. [Google Scholar]
- Karim, M.E.; Dhar, K.; Hossain, M.T. Decolorization of textile reactive dyes by bacterial monoculture and consortium screened from textile dyeing effluent. J. Genet. Eng. Biotechnol. 2018, 16, 375–380. [Google Scholar] [CrossRef] [PubMed]
- Roy, D.C.; Biswas, S.K.; Saha, A.K.; Sikdar, B.; Rahman, M.; Roy, A.K.; Prodhan, Z.H.; Tang, S.S. Biodegradation of Crystal Violet dye by bacteria isolated from textile industry effluents. Peer J. 2018, 6, e5015. [Google Scholar] [CrossRef]
- Ghodake, G.S.; Kalme, S.D.; Jadhav, J.P.; Govindwar, S.P. Purification and partial characterization of lignin peroxidase from Acinetobacter calcoaceticus NCIM 2890 and its application in decolorization of textile dyes. Appl. Biochem. Biotechnol. 2009, 152, 6–14. [Google Scholar] [CrossRef]
- Bandounas, L.; Wierckx, N.J.P.; de Winde, J.H.; Ruijssenaars, H.J. Isolation and characterization of novel bacterial strains exhibiting ligninolytic potential. BMC Biotechnol. 2011, 11, 94. [Google Scholar] [CrossRef]
- Chantarasiri, A.; Boontanom, P.; Nuiplot, N. Isolation and characterization of Lysinibacillus sphaericus BR2308 from coastal wetland in Thailand for the biodegradation of lignin. AACL Bioflux 2017, 10, 200–209. [Google Scholar]
- Mitsch, W.J.; Gosselink, J.G. Wetlands, 5th ed.; Wiley: New Jersey, NJ, USA, 2015; pp. 27–44. [Google Scholar]
- Chantarasiri, A. Aquatic Bacillus cereus JD0404 isolated from the muddy sediments of mangrove swamps in Thailand and characterization of its cellulolytic activity. Egypt J. Aquat. Res. 2015, 41, 257–264. [Google Scholar] [CrossRef]
- Sandilyan, S.; Kathiresan, K. Decline of mangroves-a threat of heavy metal poisoning in Asia. Ocean. Coast. Manag. 2014, 102, 161–168. [Google Scholar] [CrossRef]
- Kaewtubtim, P.; Meeinkuirt, W.; Seepom, S.; Pichtel, J. Heavy metal phytoremediation potential of plant species in a mangrove ecosystem in Pattani Bay, Thailand. Appl. Ecol. Environ. Res. 2016, 14, 367–382. [Google Scholar] [CrossRef]
- Abejón, R.; Pérez-Acebo, H.; Clavijo, L. Alternatives for chemical and biochemical lignin valorization: Hot topics from a bibliometric analysis of the research published during the 2000–2016 period. Processes 2018, 6, 98. [Google Scholar] [CrossRef]
- Datta, R.; Kelkar, A.; Baraniya, D.; Molaei, A.; Moulick, A.; Meena, R.S.; Formanek, P. Enzymatic degradation of lignin in soil: A review. Sustainability 2017, 9, 1163. [Google Scholar] [CrossRef]
- Mun, S.P.; Jahan, M.S.; Al-Maruf, A.; Chowdhury, D.A.N. Chemical characterization of six mangrove species in Bangladesh. Wood Sci. Technol. 2011, 45, 281–288. [Google Scholar] [CrossRef]
- Srinivasan, G.P.; Sikkanthar, A.; Elamaran, A.; Delma, C.R.; Subramaniyan, K.; Somasundaram, S.T. Biodegradation of carcinogenic textile azo dyes using bacterial isolates of mangrove sediment. J. Coast. Life Med. 2014, 2, 154–162. [Google Scholar]
- Fang, Z.; Li, T.; Wang, Q.; Zhang, X.; Peng, H.; Fang, W.; Hong, Y.; Ge, H.; Xiao, Y. A bacterial laccase from marine microbial metagenome exhibiting chloride tolerance and dye decolorization ability. Appl. Microbiol. Biotechnol. 2011, 89, 1103–1110. [Google Scholar] [CrossRef]
- Chantarasiri, A. Enrichment and identification of phenanthrene-degrading bacteria isolated from the oil-stained engine sediment in the mangrove swamps of Thailand. Appl. Sci. Eng. Prog 2020, in press. [Google Scholar] [CrossRef]
- Chang, Y.C.; Choi, D.; Takamizawa, K.; Kikuchi, S. Isolation of Bacillus sp. strains capable of decomposing alkali lignin and their application in combination with lactic acid bacteria for enhancing cellulase performance. Bioresour. Technol. 2014, 152, 429–436. [Google Scholar] [CrossRef]
- Ferbiyanto, A.; Rusmana, I.; Raffiudin, R. Characterization and identification of cellulolytic bacteria from gut of worker Macrotermes gilvus. Hayati 2015, 22, 197–200. [Google Scholar] [CrossRef]
- Boontanom, P.; Chantarasiri, A. Short communication: Diversity of culturable epiphytic bacteria isolated from seagrass (Halodule uninervis) in Thailand and their preliminary antibacterial activity. Biodiversitas 2020, 21, 2907–2913. [Google Scholar] [CrossRef]
- Chantarasiri, A. Diversity of cellulolytic bacteria isolated from a freshwater wetland reserve in Thailand and their cellulolytic activity. Appl. Ecol. Environ. Res. 2020, 18, 5965–5983. [Google Scholar]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Gouy, M.; Guindon, S.; Gascuel, O. SeaView version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 2010, 27, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Gascuel, O. BIONJ: An improved version of the NJ algorithm based on a simple model of sequence data. Mol. Biol. Evol. 1997, 14, 685–695. [Google Scholar] [CrossRef] [PubMed]
- Cerboneschi, M.; Corsi, M.; Bianchini, R.; Bonanni, M.; Tegli, S. Decolorization of acid and basic dyes: Understanding the metabolic degradation and cell-induced adsorption/precipitation by Escherichia coli. Appl. Microbiol. Biotechnol. 2015, 99, 8235–8245. [Google Scholar] [CrossRef]
- Pumijumnong, N. Mangrove forests in Thailand. In Mangrove Ecosystems of Asia; Faridah-Hanum, I., Ed.; Springer Science + Business Media: New York, NY, USA, 2014; pp. 61–79. [Google Scholar]
- Srisunont, C.; Jaiyen, T.; Tenrung, M.; Likitchaikul, M. Nutrient accumulation by litterfall in mangrove forest at Klong Khone, Thailand. Thammasat Int. J. Sci. Tech. 2017, 22, 9–18. [Google Scholar]
- Yaseen, D.A.; Scholz, M. Textile dye wastewater characteristics and constituents of synthetic effluents: A critical review. Int. J. Environ. Sci. Technol. 2019, 16, 1193–1226. [Google Scholar] [CrossRef]
- Upadhye, V.B.; Joshi, S.S. Advances in wastewater treatment-a review. Int. J. Chem. Sci. Appl. 2012, 3, 264–268. [Google Scholar]
- Zhang, W.; Liu, W.; Zhang, J.; Zhao, H.; Zhang, Y.; Quan, X.; Jin, Y. Characterisation of acute toxicity, genotoxicity and oxidative stress posed by textile effluent on zebrafish. J. Environ. Sci. 2012, 24, 2019–2027. [Google Scholar] [CrossRef]
- Ghaly, A.; Ananthashankar, R.; Alhattab, M.; Ramakrishnan, V. Production, characterization and treatment of textile effluents: A critical review. J. Chem. Eng. Proc. Technol. 2014, 5, 1–18. [Google Scholar]
- Barati, A.; Ghaderpour, A.; Chew, L.L.; Bong, C.W.; Thong, K.L.; Chong, V.C.; Chai, L.C. Isolation and characterization of aquatic-borne Klebsiella pneumoniae from tropical estuaries in Malaysia. Int. J. Environ. Res. Public Health 2016, 13, 426. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Li, L.; Perseke, M.; Huang, Y.; Wei, J.; Qin, Q. Isolation and characterization of a Klebsiella pneumoniae strain from mangrove sediment for efficient biosynthesis of 1,3-propanediol. Sci. Bull. 2015, 60, 511–521. [Google Scholar] [CrossRef][Green Version]
- Cosgrove, S.E.; Kaye, K.S.; Eliopoulous, G.M.; Carmeli, Y. Health and economic outcomes of the emergence of third-generation cephalosporin resistance in Enterobacter species. Arch. Intern. Med. 2002, 162, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Cooney, S.; O’ Brien, S.; Iversen, C.; Fanning, S. Bacteria: Other pathogenic Enterobacteriaceae-Enterobacter and other genera. In Encyclopedia of Food Safety; Motarjemi, Y., Ed.; Academic Press: Cambridge, MA, USA, 2014; Volume 1, pp. 433–441. [Google Scholar]
- Duan, Y.Q.; Zhou, X.K.; Di-Yan, L.; Li, Q.Q.; Dang, L.Z.; Zhang, Y.G.; Qiu, L.H.; Nimaichand, S.; Li, W.J. Enterobacter tabaci sp. nov., a novel member of the genus Enterobacter isolated from a tobacco stem. Antonie Van Leeuwenhoek 2015, 108, 1161–1169. [Google Scholar] [CrossRef]
- Vazquez, P.; Holguin, G.; Puente, M.E.; Lopez-Cortes, A.; Bashan, Y. Phosphate-solubilizing microorganisms associated with the rhizosphere of mangroves in a semiarid coastal lagoon. Biol. Fertil. Soils. 2000, 30, 460–468. [Google Scholar] [CrossRef]
- Castro, R.A.; Quecine, M.C.; Lacava, P.T.; Batista, B.D.; Luvizotto, D.M.; Marcon, J.; Ferreira, A.; Melo, I.S.; Azevedo, J.L. Isolation and enzyme bioprospection of endophytic bacteria associated with plants of Brazilian mangrove ecosystem. Springer Plus 2014, 3, 382. [Google Scholar] [CrossRef]
- Chen, C.; Li, T. Bacterial dye-decolorizing peroxidases: Biochemical properties and biotechnological opportunities. Phys. Sci. Rev. 2016, 1, 20160051. [Google Scholar] [CrossRef]
- Wong, D.W.S. Structure and action mechanism of ligninolytic enzymes. Appl. Biochem. Biotechnol. 2009, 157, 174–209. [Google Scholar] [CrossRef]
- Lambertz, C.; Ece, S.; Fischer, R.; Commandeur, U. Progress and obstacles in the production and application of recombinant lignin-degrading peroxidases. Bioengineered 2016, 7, 145–154. [Google Scholar] [CrossRef]
- Mester, T.; Tien, M. Oxidation mechanism of ligninolytic enzymes involved in the degradation of environmental pollutants. Int. Biodeterior. Biodegrad. 2000, 46, 51–59. [Google Scholar] [CrossRef]
- Niladevi, K.N.; Prema, P. Mangrove actinomyces as the source of ligninolytic enzymes. Actinomycetologica 2005, 19, 40–47. [Google Scholar] [CrossRef]
- De Gonzalo, G.; Colpa, D.I.; Habib, M.H.M.; Fraaije, M.W. Bacterial enzymes involved in lignin degradation. J. Biotechnol. 2016, 236, 110–119. [Google Scholar] [CrossRef] [PubMed]
- Bilal, M.; Asgher, M.; Parra-Saldivar, R.; Hu, H.; Wang, W.; Zhang, X.; Iqbal, H.M.N. Immobilized ligninolytic enzymes: An innovative and environmental responsive technology to tackle dye-based industrial pollutants-a review. Sci. Total Environ. 2017, 576, 646–659. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.; Chandra, R. Syntrophic co-culture of Bacillus subtilis and Klebsiella pneumonia for degradation of kraft lignin discharged from rayon grade pulp industry. J. Environ. Sci. 2015, 33, 229–238. [Google Scholar] [CrossRef]
- Liu, Y.; Huang, L.; Guo, W.; Jia, L.; Fu, Y.; Gui, S.; Lu, F. Cloning, expression, and characterization of a thermostable and pH-stable laccase from Klebsiella pneumoniae and its application to dye decolorization. Process. Biochem. 2017, 53, 125–134. [Google Scholar] [CrossRef]
- Xu, Z.; Qin, L.; Cai, M.; Hua, W.; Jin, M. Biodegradation of kraft lignin by newly isolated Klebsiella pneumoniae, Pseudomonas putida, and Ochrobactrum tritici strains. Environ. Sci. Pollut. Res. 2018, 25, 14171–14181. [Google Scholar] [CrossRef]
- Gaur, N.; Narasimhulu, K.; Setty, Y.P. Extraction of ligninolytic enzymes from novel Klebsiella pneumoniae strains and its application in wastewater treatment. Appl. Water Sci. 2018, 8, 111. [Google Scholar] [CrossRef]
- DeAngelis, K.M.; D’Haeseleer, P.; Chivian, D.; Fortney, J.L.; Khudyakov, J.; Simmons, B.; Woo, H.; Arkin, A.P.; Davenport, K.W.; Goodwin, L.; et al. Complete genome sequence of “Enterobacter lignolyticus” SCF1. Stand. Genom. Sci. 2011, 5, 69–85. [Google Scholar] [CrossRef]
- Odeniyi, O.A.; Unuofin, J.O.; Adebayo-Tayo, B.C.; Wakil, S.M.; Onilude, A.A. Production characteristics, activity patterns and biodecolourisation applications of thermostable laccases from Corynebacterium efficiens and Enterobacter ludwigii. J. Sci. Ind. Res. 2017, 76, 562–569. [Google Scholar]
- Kumar, V.; Chandra, R. Characterisation of manganese peroxidase and laccase producing bacteria capable for degradation of sucrose glutamic acid-Maillard reaction products at different nutritional and environmental conditions. World J. Microbiol. Biotechnol. 2018, 34, 32. [Google Scholar] [CrossRef]
- Ozer, A.; Rakici, E.; Bektas, K.I.; Canakci, S.; Belduz, A.O. Isolation of lignin-degrading bacteria from different sources and testing of their ligninolytic activities. J. Apitherapy Nat. 2019, 2, 30–45. [Google Scholar]
- Riyadi, F.A.; Tahir, A.A.; Yusof, N.; Sabri, N.S.A.; Noor, M.J.M.M.; Akhir, F.N.M.D.; Othman, N.; Zakaria, Z.; Hara, H. Enzymatic and genetic characterization of lignin depolymerization by Streptomyces sp. S6 isolated from a tropical environment. Sci. Rep. 2020, 10, 7813. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Chai, L.; Tang, C.; Yang, Z.; Zheng, Y.; Chen, Y.; Jing, Q. Biochemical investigation of kraft lignin degradation by Pandoraea sp. B-6 isolated from bamboo slips. Bioprocess. Biosyst. Eng. 2013, 36, 1957–1965. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.H.; Chai, L.Y.; Zhu, Y.H.; Yang, Z.H.; Zheng, Y.; Zhang, H. Biodegradation of kraft lignin by a bacterial strain Comamonas sp. B-9 isolated from eroded bamboo slips. J. Appl. Microbiol. 2012, 112, 900–906. [Google Scholar] [CrossRef] [PubMed]
- Chai, L.Y.; Chen, Y.H.; Tang, C.J.; Yang, Z.H.; Zheng, Y.; Shi, Y. Depolymerization and decolorization of kraft lignin by bacterium Comamonas sp. B-9. Appl. Microbiol. Biotechnol. 2014, 98, 1907–1912. [Google Scholar] [CrossRef] [PubMed]
- Asgher, M.; Bhatti, H.N.; Shah, S.A.H.; Asad, M.J.; Legge, R.L. Decolorization potential of mixed microbial consortia for reactive and disperse textile dyestuffs. Biodegradation 2007, 18, 311–316. [Google Scholar] [CrossRef]
- Sghaier, I.; Ouertani, R.; Mahjoubi, M.; El-Hidri, D.; Hassen, W.; Chamkhi, A.; Chouchane, H.; Jaouani, A.; Cherif, A.; Neifar, M. Application of a mixture design to optimize textile azo-dye decolorization using a bacterial consortium. Biom Biostat. Int. J. 2019, 8, 58–63. [Google Scholar] [CrossRef]
- Chattaraj, S.; Johnson, J.; Madamwar, D. Biotransformation of mixture of dyes by enriched bacterial consortium ASD. Desalin. Water Treat. 2016, 57, 21585–21597. [Google Scholar] [CrossRef]
- Balapure, K.; Aghera, P.; Bhatt, N.; Madamwar, D. Community synergism: Degradation of triazine dye Reactive Black 1 by mixed bacterial cultures KND_PR under microaerophilic and aerobic conditions. Environ. Process. 2019, 6, 713–739. [Google Scholar] [CrossRef]
- Jadhav, P.U.; Bholay, A.D.; Shindikar, M. Bacterial lignin peroxidase in biobleaching of lignin-mimicking indicator dyes. Int. J. Pure Appl. Biosci. 2016, 4, 84–92. [Google Scholar] [CrossRef]
- Gottlieb, A.; Shaw, C.; Smith, A.; Wheatley, A.; Forsythe, S. The toxicity of textile reactive dyes after hydrolysis and decolourisation. J. Biotechnol. 2003, 101, 49–56. [Google Scholar] [CrossRef]
- Saparrat, N.; Carlos, M.; Hammer, E. Decolorization of synthetic dyes by the deuteromycete Pestalotiopsis guepinii CLPS no. 786 strain. J. Basic Microbiol. 2006, 46, 28–33. [Google Scholar] [CrossRef] [PubMed]
- Falade, A.; Mabinya, L.; Okoh, A.; Nwodo, U. Peroxidases produced by new ligninolytic Bacillus strains isolated from marsh and grassland decolourized anthraquinone and azo dyes. Pol. J. Environ. Stud. 2019, 28, 3163–3172. [Google Scholar] [CrossRef]
- Saratale, R.G.; Saratale, G.D.; Chang, J.S.; Govindwar, S.P. Ecofriendly degradation of sulfonated diazo dye C.I. Reactive Green 19A using Micrococcus glutamicus NCIM-2168. Bioresour. Technol. 2009, 100, 3897–3905. [Google Scholar] [CrossRef] [PubMed]
- Meerbergen, K.; Willems, K.A.; Dewil, R.; Impe, J.V.; Appels, L.; Lievens, B. Isolation and screening of bacterial isolates from wastewater treatment plants to decolorize azo dyes. J. Biosci. Bioeng. 2018, 125, 448–456. [Google Scholar] [CrossRef] [PubMed]
- Sahasrabudhe, M.; Pathade, G. Decolourization and degradation of C.I. Reactive Red 195 by Georgenia sp. CC-NMPT-T3. Indian J. Exp. Biol. 2012, 50, 290–299. [Google Scholar]
- Mate, M.S.; Pathade, G. Biodegradation of C.I. Reactive Red 195 by Enterococcus faecalis strain YZ66. World J. Microbiol. Biotechnol. 2012, 28, 815–826. [Google Scholar] [CrossRef]
- Sarnaik, S.; Kanekar, P. Biodegradation of methyl violet by Pseudomonas mendocina MCM B-402. Appl. Microbiol. Biotechnol. 1999, 52, 1–4. [Google Scholar] [CrossRef]



| Textile Reactive Dye | Class of Textile Reactive Dye | λmax (nm) |
|---|---|---|
| C.I. Reactive Black 1 | Azo | 606 |
| C.I. Reactive Brown 1 | Azo | 489 |
| C.I. Reactive Blue 19 | Anthraquinone | 592 |
| C.I. Reactive Blue 194 | Azo | 594 |
| C.I. Reactive Blue 21 | Phthalocyanine | 670 |
| C.I. Reactive Green 19 | Azo | 636 |
| C.I. Reactive Orange 122 | Azo | 489 |
| C.I. Reactive Red 195 | Azo | 544 |
| C.I. Reactive Yellow 167 | Azo | 408 |
| Pigmentation (Percentage) | Shape (Percentage) | Margin (Percentage) | Elevation (Percentage) | ||||
|---|---|---|---|---|---|---|---|
| White | 49.47 | Circular | 94.22 | Entire | 71.57 | Raised | 42.63 |
| Yellow | 23.16 | Filamentous | 3.68 | Erose | 12.11 | Convex | 42.11 |
| Red | 10.00 | Irregular | 1.58 | Undulate | 11.58 | Umbonate | 10.52 |
| Colorless | 17.37 | Punctiform | 0.52 | Filamentous | 4.74 | Flat | 4.74 |
| Total | 100.00 | Total | 100.00 | Total | 100.00 | Total | 100.00 |
| Bacterial Isolate | Close Bacteria | GenBank Accession Number (References) | Query Cover (%) | Identity (%) * | GenBank Accession Number (Deposited) |
|---|---|---|---|---|---|
| RY10302 | Klebsiella pneumoniae | NR_117683.1 | 97 | 99.02 | MT355793 |
| strain DSM 30104 | |||||
| RY10402 | Enterobacter tabaci | NR_146667.2 | 98 | 97.38 | MT355794 |
| strain YIM Hb-3 | |||||
| RY11902 | Enterobacter tabaci | NR_146667.2 | 99 | 97.91 | MT355798 |
| strain YIM Hb-3 | |||||
| RY11903 | Enterobacter tabaci | NR_146667.2 | 100 | 98.36 | MT355797 |
| strain YIM Hb-3 |
| Bacterial Strain | Enzymatic Activities (U/mL) | ||
|---|---|---|---|
| Lignin Peroxidase | Manganese Peroxidase | Laccase | |
| K. pneumoniae | No activity | No activity | 0.45 ± 0.03 a (p < 0.001) |
| strain RY10302 | |||
| Enterobacter sp. | 0.13 ± 0.02 a (p < 0.001) | No activity | 0.81 ± 0.02 c (p < 0.001) |
| strain RY10402 | |||
| Enterobacter sp. | 0.62 ± 0.03 b (p < 0.001) | No activity | 0.66 ± 0.02 b (p < 0.001) |
| strain RY11902 | |||
| Enterobacter sp. | 0.86 ± 0.06 c (p < 0.001) | No activity | 1.47 ± 0.03 d (p < 0.001) |
| strain RY11903 | |||
| Bacterial Strain | Decolorization of Textile Reactive Dye (%) at 72 h of Incubation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Black 1 | Brown 1 | Blue 19 | Blue 194 | Blue 21 | Green 19 | Orange 122 | Red 195 | Yellow 167 | |
| K. pneumoniae | 24.65 ± 0.74 b | 24.96 ± 0.26 b | 26.51 ± 0.44 b | 42.43 ± 0.35 b | 55.97 ± 0.05 b | 45.49 ± 0.60 b | 21.00 ± 0.01 b | 20.10 ± 0.00 b | 22.00 ± 0.01 b |
| strain | |||||||||
| RY10302 | |||||||||
| Enterobacter sp. | 49.00 ± 0.00 c | 38.30 ± 0.26 c | 43.94 ± 0.07 d | 37.83 ± 0.29 b | 86.23 ± 0.32 d | 67.23 ± 0.40 c | 34.96 ± 1.12 c | 45.23 ± 1.71 c | 44.29 ± 0.66 c |
| strain | |||||||||
| RY10402 | |||||||||
| Enterobacter sp. | 89.52 ± 1.41 e | 66.27 ± 4.70 d | 37.27 ± 3.96 c | 92.45 ± 0.40 d | 88.67 ± 0.42 e | 91.09 ± 0.68 e | 54.50 ± 2.17 e | 60.27 ± 2.18d | 40.86 ± 3.71 c |
| strain | |||||||||
| RY11902 | |||||||||
| Enterobacter sp. strain | 73.85 ± 3.18 d | 72.32 ± 1.71 d | 60.34 ± 0.94 e | 69.91 ± 5.24 c | 82.70 ± 0.76 c | 81.35 ± 0.84 d | 40.38 ± 2.02 d | 64.22 ± 2.48 d | 50.17 ± 2.12 d |
| RY11903 | |||||||||
| Mixed | 94.20 ± 0.83 f | 83.03 ± 2.36 e | 58.25 ± 1.86 e | 86.81 ± 1.19 d | 88.32 ± 0.67 de | 92.05 ± 1.78 e | 63.01 ± 1.10 f | 84.32 ± 1.03 e | 68.26 ± 1.37 e |
| culture | |||||||||
| E. coli | 4.67 ± 0.22 a | 12.38 ± 0.44a | 16.56 ± 2.91 a | 5.92 ± 0.81 a | 13.37 ± 1.70 a | 14.53 ± 0.36 a | 15.10 ± 0.94 a | 11.07 ± 0.81 a | 14.36 ± 1.07 a |
| strain | |||||||||
| TISTR 073 | |||||||||
| Reactive Dye | Decolorizing Bacteria | Source of Isolation | Dye Concentration /Decolorization Percentage | Conditions for Decolorization | References |
|---|---|---|---|---|---|
| C.I. Reactive Black 1 | Mixed bacterial culture | Dye contaminated soils, India | 100 mg/L, | 37 °C, 12 h | [73] |
| (Lysinibacillus sp. strains BAB-4931 and BAB-4935, Raoultella sp. strain BAB-4932, Enterococcus sp. strain BAB-4933 and Citrobacter sp. strain BAB-4934) | 65.0% | ||||
| Mixed bacterial culture | Mangrove wetland soils, Thailand | 100 mg/L, | pH 8.0, 30 °C, 0.5% (w/v) NaCl, 72 h | This study | |
| 94.2% | |||||
| C.I. Reactive Blue 19 (RBBR) | Bacillus sp. | Mountain soils, Japan | 100 mg/L, | pH 6.0, 30 °C, | [30] |
| strain CS-1 | 100.0% | 72 h | |||
| Bacillus sp. | Mountain soils, Japan | 100 mg/L, | pH 6.0, 30 °C, | [30] | |
| strain CS-2 | 95.0% | 72 h | |||
| Lysinibacillus sphaericus strain BR2308 | Coastal wetland soils, Thailand | 100 mg/L, | pH 6.0, 30 °C, | [19] | |
| 58.1% | 72 h | ||||
| L. sphaericus | Wetland and estuary soils, Thailand | 50 mg/L, | pH 6.0, 28 °C, | [14] | |
| strain JD1103 | 50.0% | 72 h | |||
| Bacillus sp. | Marsh and grassland samples, South Africa | 100 mg/L, | pH 6.0, 25 °C, | [77] | |
| strain NWODO-3 | 72.1% | 30 min | |||
| Enterobacter sp. | Mangrove wetland soils, Thailand | 100 mg/L, | pH 9.0, 30 °C, 3% (w/v) NaCl, 72 h | This study | |
| strain RY11903 | 60.3% | ||||
| C.I. Reactive Green 19 | Micrococcus glutamicus strain NCIM-2168 | National Chemical Laboratory, India | 50 mg/L, | pH 8.0, 37 °C, | [78] |
| 100.0% | 42 h | ||||
| Acinetobacter sp. | Activated sludges, Belgium | 100 mg/L, | 30 °C, 72 h | [79] | |
| strain ST16.16/164 | 93.4% | ||||
| Mixed bacterial culture | Mangrove wetland soils, Thailand | 100 mg/L, | pH 8.0, 30 °C, 0.5% (w/v) NaCl, 72 h | This study | |
| 92.1% | |||||
| C.I. Reactive Red 195 | Georgenia sp. | India | 50 mg/L, | pH 7.0, 40 °C, 5 h | [80] |
| strain CC-NMPT-T3 | 95.9% | ||||
| Enterococcus faecalis | Dye contaminated soils, India | 50 mg/L, | pH 5.0, 40 °C, | [81] | |
| strain YZ66 | 99.5% | 1.5 h | |||
| Mixed bacterial culture | Mangrove wetland soils, Thailand | 100 mg/L, | pH 8.0, 30 °C, 0.5% (w/v) NaCl, 72 h | This study | |
| 84.3% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chantarasiri, A. Klebsiella and Enterobacter Isolated from Mangrove Wetland Soils in Thailand and Their Application in Biological Decolorization of Textile Reactive Dyes. Int. J. Environ. Res. Public Health 2020, 17, 7531. https://doi.org/10.3390/ijerph17207531
Chantarasiri A. Klebsiella and Enterobacter Isolated from Mangrove Wetland Soils in Thailand and Their Application in Biological Decolorization of Textile Reactive Dyes. International Journal of Environmental Research and Public Health. 2020; 17(20):7531. https://doi.org/10.3390/ijerph17207531
Chicago/Turabian StyleChantarasiri, Aiya. 2020. "Klebsiella and Enterobacter Isolated from Mangrove Wetland Soils in Thailand and Their Application in Biological Decolorization of Textile Reactive Dyes" International Journal of Environmental Research and Public Health 17, no. 20: 7531. https://doi.org/10.3390/ijerph17207531
APA StyleChantarasiri, A. (2020). Klebsiella and Enterobacter Isolated from Mangrove Wetland Soils in Thailand and Their Application in Biological Decolorization of Textile Reactive Dyes. International Journal of Environmental Research and Public Health, 17(20), 7531. https://doi.org/10.3390/ijerph17207531





