A Rapid and Sensitive Europium Nanoparticle-Based Lateral Flow Immunoassay Combined with Recombinase Polymerase Amplification for Simultaneous Detection of Three Food-Borne Pathogens
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and DNA Template Preparation
2.2. Primer Design
2.3. Multiplex Reaction Protocols for RPA
2.4. Preparation of EuNP Probes
2.5. Preparation of EuNP-Based LFIA and CG–LFIA
2.6. Multiplex EuNP-Based LFIA–RPA Assay
2.7. Optimization of the Multiplex EuNP-Based LFIA–RPA Assay
2.8. Sensitivity and Specificity of the Multiplex EuNP-Based LFIA–RPA Assay
2.9. Stability of the Multiplex EuNP-Based LFIA–RPA Assay
2.10. Analysis of Spiked Samples
2.11. Evaluation of Field Food Samples
2.12. Data Analysis
3. Results
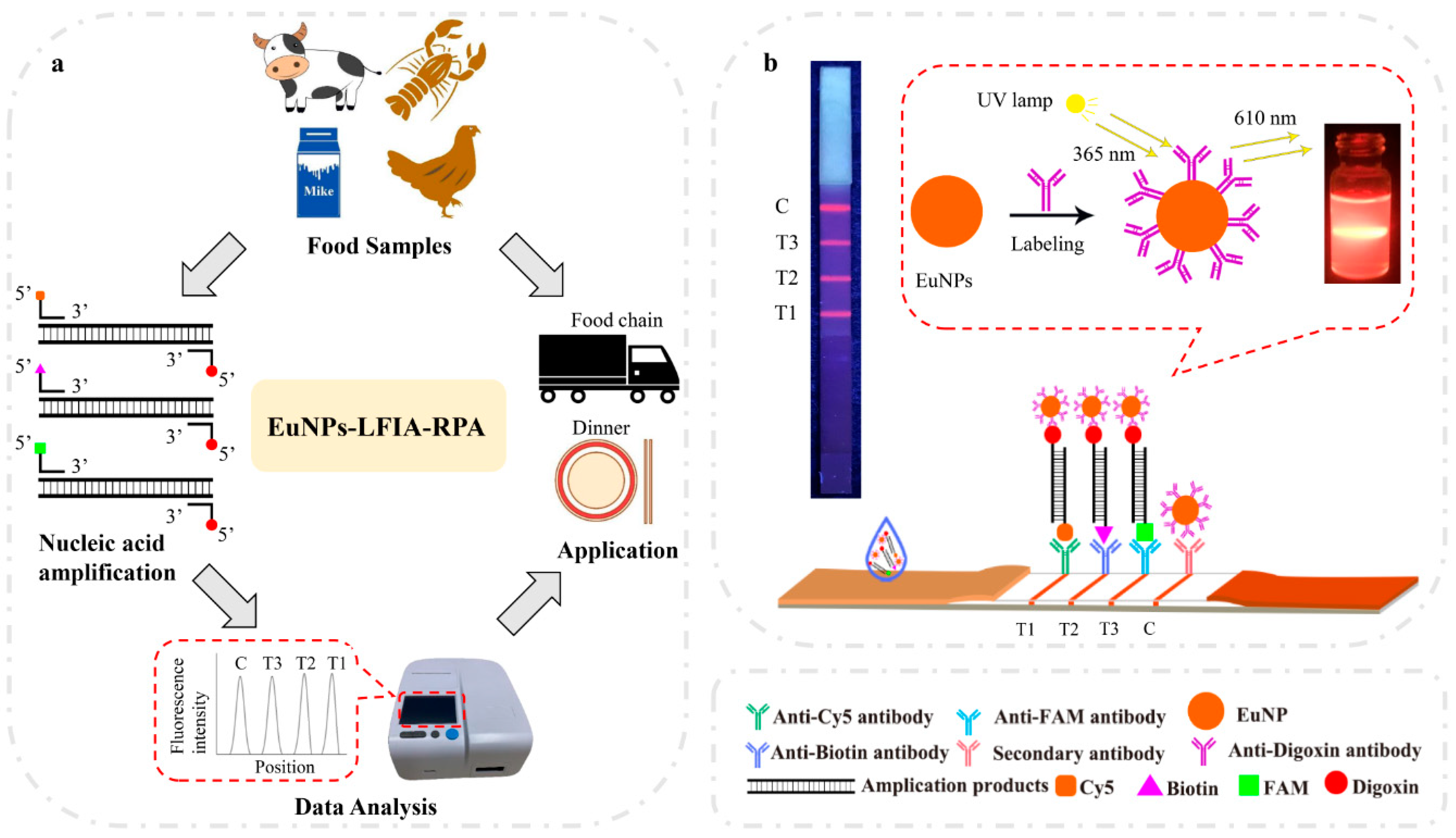
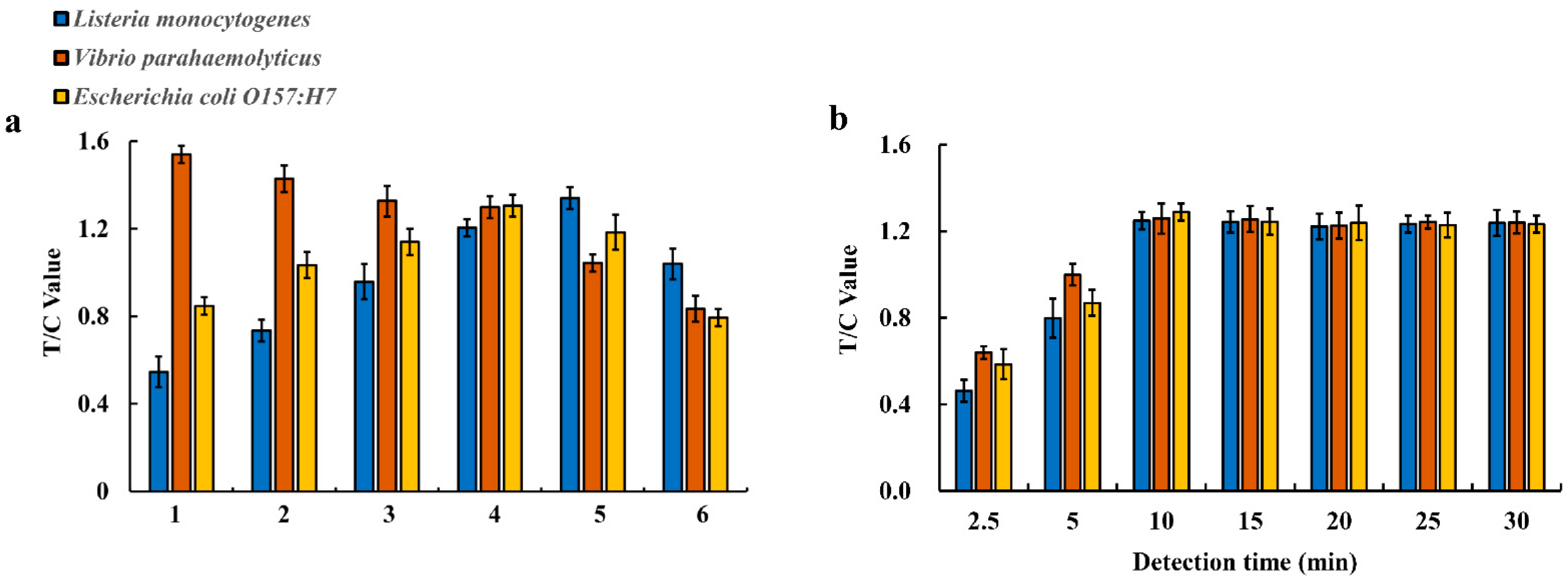
3.1. Establishment and Optimization of Multiplex EuNP-Based LFIA–RPA Assay
3.2. Sensitivity of the Multiplex EuNP-Based LFIA–RPA
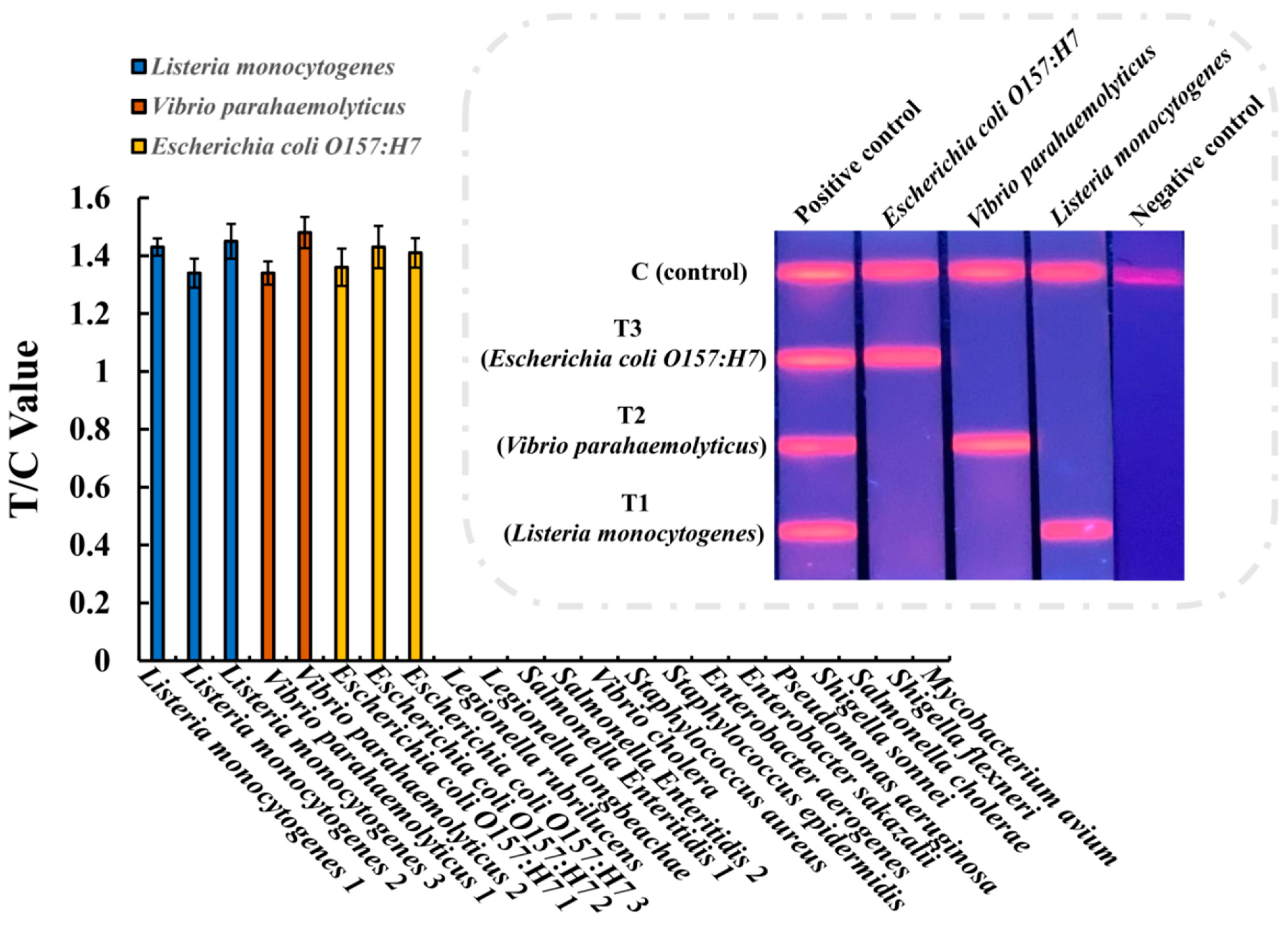
3.3. Specificity of the Multiplex EuNP-Based LFIA–RPA
3.4. Stability of the Multiplex EuNP-Based LFIA–RPA
3.5. Application of the Multiplex EuNP-Based LFIA–RPA to Spiked Samples
3.6. Results of Field Food Samples
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Bhardwaj, N.; Bhardwaj, S.K.; Nayak, M.K.; Mehta, J.; Kim, K.H.; Deep, A. Fluorescent nanobiosensors for the targeted detection of foodborne bacteria. Trends Anal. Chem. 2017, 97, 120–135. [Google Scholar] [CrossRef]
- Liu, H.B.; Du, X.J.; Zang, Y.X.; Li, P.; Wang, S. SERS-based lateral flow strip biosensor for simultaneous detection of Listeria monocytogenes and Salmonella enterica serotype enteritidis. J. Agric. Food Chem. 2017, 65, 10290–10299. [Google Scholar] [CrossRef]
- Ceccarelli, D.; Hasan, N.A.; Huq, A.; Colwell, R.R. Distribution and dynamics of epidemic and pandemic V. parahaemolyticus virulence factors. Front. Cell Infect. Microbiol. 2013, 3, 97. [Google Scholar] [CrossRef] [Green Version]
- Heiman, K.E.; Mody, R.K.; Johnson, S.D.; Griffin, P.M.; Gould, L.H. Escherichia coli O157 outbreaks in the United States, 2003–2012. Emerg. Infect. Dis. 2015, 21, 1293–1301. [Google Scholar] [CrossRef]
- Dun, X.J.; Zang, Y.X.; Liu, H.B.; Li, P.; Wang, S. Recombinase polymerase amplification combined with Lateral Flow Strip for Listeria monocytogenes detection in food. J. Food Sci. 2018, 83, 1041–1047. [Google Scholar]
- Azinheiro, S.; Carvalho, J.; Prado, M.; Garrodo-Maestu, A. Application of Recombinase Polymerase Amplification with Lateral Flow for a Naked-Eye Detection of Listeria monocytogenes on Food Processing Surfaces. Foods 2020, 9, 1249. [Google Scholar] [CrossRef] [PubMed]
- Briet, A.; Helsens, N.; Delannoy, S.; Debuiche, S.; Brisabois, A.; Midelet, G.; Granier, S.A. NDM-1-producing Vibrio parahaemolyticus isolated from imported seafood. J. Antimicrob. Chemother. 2018, 73, 2578–2579. [Google Scholar] [CrossRef]
- Pan, J.Y.; Ye, Z.C.; Cheng, Z.Y.; Peng, X.J.; Wen, L.Y.; Zhao, F.K. Systematic analysis of the lysine acetylome in Vibrio parahemolyticus. J. Proteome Res. 2014, 13, 3294–3302. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, S.; Li, J.; Chen, M.L.; He, M.; Wang, Y.Y.; Pang, B. Application of digital PCR and next generation sequencing in the etiology investigation of a foodborne disease outbreak caused by Vibrio parahaemolyticus. Food Microbiol. 2019, 84, 103233. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Gonzalez-Escalona, N.; Haendiges, J.; Myers, R.A.; Ferguson, J.; Stiles, T.; Whistler, C.A. Sequence type 631 Vibrio parahaemolyticus, an emerging foodborne pathogen in North America. J. Clin. Microbiol. 2017, 55, 645–648. [Google Scholar] [CrossRef] [Green Version]
- Han, J.J.; Zhang, L.; Hu, L.M.; Xing, K.Y.; Lu, X.F.; Huang, Y.J.; Zhang, J.W.; Lai, W.H.; Chen, T. Nanozyme-based lateral flow assay for the sensitive detection of Escherichia coli O157:H7 in milk. J. Dairy Sci. 2018, 101, 5770–5779. [Google Scholar] [CrossRef] [Green Version]
- Fach, P.; Perelle, S.; Grout, J.; Dilasser, J. Comparison of different PCR tests for detecting shiga toxin-producing Escherichia coli O157 and development of an ELISAPCR assay for specific identification of the bacteria. J. Microbiol. Methods 2003, 55, 383–392. [Google Scholar] [CrossRef]
- Thamizhvanan, S.; Sivakumar, S.; Kumar, S.S.; Kumar, D.V.; Suryakodi, S.; Balaji, K.; Rajkumar, T.; Vimal, S.; Majeed, S.A.; Taju, G.; et al. Multiple infections caused by white spot syndrome virus and enterocytozoon hepatopenaei in pond-reared penaeus vannamei in India and multiplex PCR for their simultaneous detection. J. Fish. Dis. 2019, 42, 447–454. [Google Scholar] [CrossRef] [PubMed]
- Suwancharoen, D.; Kulchim, C.; Chirathaworn, C.; Yoshida, S. Development of a novel primer combination to detect pathogenic Leptospira by loop-mediated isothermal amplification. J. Microbiol. Methods 2012, 91, 171–173. [Google Scholar] [CrossRef] [PubMed]
- Euler, M.; Wang, Y.; Otto, P.; Tomaso, H.; Escudero, R.; Anda, P.; Weidmann, M. Recombinase polymerase amplification assay for rapid detection of Francisella tularensis. J. Clin. Microbiol. 2012, 50, 2234–2238. [Google Scholar] [CrossRef] [Green Version]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L.; Armes, N.A. DNA detection using recombination proteins. PLoS Biol. 2006, 4, e204. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Fang, J.H.; Wang, Y.; He, H.Z.; Dai, M.Y.; Lin, W.; Su, W.; Zhang, M.Z. Isothermal method of a recombinase polymerase amplification assay for the detection of most common high-risk human papillomavirus type 16 and type 18 DNA. Clin. Lab. 2017, 63, 27–38. [Google Scholar] [CrossRef]
- Li, J.; Macdonald, J.; von Stetten, F. Review: A comprehensive summary of a decade development of the recombinase polymerase amplification. Analyst 2018, 144, 31–67. [Google Scholar] [CrossRef] [Green Version]
- Jaroenram, W.; Kiatpathomchai, W.; Flegel, T.W. Rapid and sensitive detection of white spot syndrome virus by loop-mediated isothermal amplification combined with a lateral flow dipstick. Mol. Cell Probe 2009, 23, 65–70. [Google Scholar] [CrossRef]
- Wang, Q.; Long, M.Y.; Lv, G.Y.; Xin, S.P.; Han, X.G.; Jiang, W. Lanthanide-labeled fluorescent-nanoparticle immunochromatographic strips enable rapid and quantitative detection of Escherichia coli O157:H7 in food samples. Food Control 2019, 109, 106894. [Google Scholar] [CrossRef]
- Yu, L.S.L.; Reed, S.A.; Golden, M.H. Time-resolved fluorescence immunoassay (TRFIA) for the detection of Escherichia coli O157:H7 in apple cider. J. Microbiol. Methods 2002, 49, 63–68. [Google Scholar] [CrossRef]
- Lin, L.Z.; Zheng, Q.W.; Lin, J.F.; Yuk, H.G.; Guo, L.Q. Immuno- and nucleic acid-based current technique for Salmonella detection in food. Eur. Food Res. Technol. 2020, 246, 373–395. [Google Scholar] [CrossRef]
- Zhou, Y.L.; Xia, X.H.; Xu, Y.; Ke, W.; Yang, W.; Li, Q.G. Application of europium (iii) chelates-bonded silica nanoparticle in time-resolved immunofluorometric detection assay for human thyroid stimulating hormone. Anal. Chim. Acta 2012, 722, 95–99. [Google Scholar] [CrossRef]
- Luo, K.; Hu, L.M.; Guo, Q.; Wu, C.H.; Wu, S.S.; Liu, D.F.; Xiong, Y.H.; Lai, W.H. Comparison of 4 label-based immunochromatographic assays for the detection of Escherichia coli O157:H7 in milk. Comp. Study 2017, 100, 5176–5187. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Li, J.L.; Chen, K.; Yu, X.P.; Sun, C.X.; Zhang, M.Z. Multiplex recombinase polymerase amplification assay for the simultaneous detection of three foodborne pathogens in seafood. Foods 2020, 9, 278. [Google Scholar] [CrossRef] [Green Version]
- Xun, Y.; Ma, B.; Chen, E.J.; Yu, X.P.; Ye, Z.H.; Sun, C.X.; Zhang, M.Z. Dual fluorescent immunochromatographic assay for simultaneous quantitative detection of citrinin and zearalenone in corn samples. Food Chem. 2021, 336, 127713. [Google Scholar]
- Davis, C. Enumeration of probiotic strains: Review of culturedependent and alternative techniques to quantify viable bacteria. J. Microbiol. Methods 2014, 103, 9–17. [Google Scholar] [CrossRef] [Green Version]
- Fraisse, A.; Coudray-Meunier, C.; Martin-Latil, S.; Hennechart-Collette, C.; Delannoy, S.; Fach, P.; Perelle, S. Digital RT-PCR method for hepatitis A virus and norovirus quantification in soft berries. Int. J. Food Microbiol. 2017, 243, 36–45. [Google Scholar] [CrossRef]
- Moriconi, M.; Acke, E.; Petrelli, D.; Preziuso, S. Multiplex PCRbased identification of Streptococcus canis, Streptococcus zooepidemicus and Streptococcus dysgalactiae subspecies from dogs. Comp. Immunol. Microbiol. Infect. Dis. 2017, 50, 48–53. [Google Scholar] [CrossRef] [PubMed]
- Nagaraj, S.; Ramlal, S.; Kingston, J.; Batra, H.V. Development of IgY based sandwich ELISA for the detection of staphylococcal enterotoxin G (SEG), an egc toxin. Int. J. Food Microbiol. 2016, 237, 136–141. [Google Scholar] [CrossRef] [PubMed]
- Brindha, J.; Chanda, K.; Balamurali, M.M. Biosensors for pathogens. Environ. Chem. Lett. 2018, 16, 1325–1337. [Google Scholar]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, e63. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deng, H.M.; Gao, Z.Q. Bioanalytical applications of isothermal nucleic acid amplification techniques. Anal. Chim. Acta 2015, 853, 30–45. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.G.; Xu, Y.C.; Yan, H.; Zhu, Y.Z.; Wang, L.; Zhang, Y.; Lu, Y.; Xing, W.L. Sensitive and rapid detection of pathogenic bacteria from urine samples using multiplex recombinase polymerase amplification. Lab Chip 2018, 18, 2441–2452. [Google Scholar] [CrossRef] [PubMed]
- Tominaga, T. Rapid detection of coliform bacteria using a lateral flow test strip assay. J. Microbiol. Methods 2019, 160, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Song, C.M.; Liu, J.X.; Li, J.W.; Liu, Q. Dual FITC lateral flow immunoassay for sensitive detection of Escherichia coli O157:H7 in food samples. Biosens. Bioelectron. 2016, 85, 734–739. [Google Scholar] [CrossRef]

| Species | ID of Strains | Results of EuNP-Based LFIA–RPA |
|---|---|---|
| Listeria monocytogenes | ATCC19111 | + |
| Listeria monocytogenes | ATCC19115 | + |
| Listeria monocytogenes | ATCC7644 | + |
| Vibrio parahaemolyticus | ATCC17802 | + |
| Vibrio parahaemolyticus | ATCC33847 | + |
| Escherichia coli O157:H7 | GIMCC1.201 | + |
| Escherichia coli O157:H7 | ECO-071 * | + |
| Escherichia coli O157:H7 | ATCC35150 | + |
| Legionella rubrilucens | ATCC35304 | − |
| Legionella longbeachae | ATCC33462 | − |
| Salmonella Enteritidis | GIMCC1.345 | − |
| Salmonella Enteritidis | ATCC13076 | − |
| Vibrio cholera | GIMCC1.449 | − |
| Staphylococcus aureus | ATCC25923 | − |
| Staphylococcus epidermidis | SE-001 * | − |
| Enterobacter aerogenes | CICC10293 | − |
| Enterobacter sakazalii | ATCC21550 | − |
| Pseudomonas aeruginosa | GIMCC1.843 | − |
| Shigella sonnei | GIMCC1.424 | − |
| Salmonella cholerae | CICC21494 | − |
| Shigella flexneri | CICC10865 | − |
| Mycobacterium avium | CMCC93026 | − |
| Target Name | Primer Name | Sequence (5′–3′) | Modifications | Amplification Size |
|---|---|---|---|---|
| Listeria monocytogenes (hlyA) | RPA-FP | CGATCACTCTGGAGGATACGTTGCTCAATT | 5′-Cy5 | 154 bp |
| RPA-RP | TTACCAGGCAAATAGATGGACGATGTGAAA | 5′-Digoxin | ||
| Vibrio parahaemolyticus (toxR) | RPA-FP | TTTGTTTGGCGTGAGCAAGGTTTTGAGGTG | 5′-Biotin | 230 bp |
| RPA-RP | GCAGAGGCGTCATTGTTATCAGAAGCAGGT | 5′-Digoxin | ||
| Escherichia coli O157:H7 (rfbE) | RPA-FP | TATCTGCAAGGTGATTCCTTGATGGTCTCA | 5′-FAM | 176 bp |
| RPA-RP | AGGCCAGTTACCATCCTCAGCTATAGGGTG | 5′- Digoxin |
| Sample (n = 8 Each) | Listeria monocytogenes | Vibrio parahaemolyticus | Escherichia coli O157:H7 | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Spiked Concentration (CFU/mL) | Recovery of EuNP-Based LFIA (%) | Recovery of CG–LFIA (%) | Spiked Concentration (CFU/mL) | Recovery of EuNP-Based LFIA (%) | Recovery of CG–LFIA (%) | Spiked Concentration (CFU/mL) | Recovery of EuNP-Based LFIA (%) | Recovery of CG–LFIA (%) | |
| Beef | 9.0 × 105 | 105.5 | 104.2 | 7.0 × 105 | 110.3 | 105.7 | 4.0 × 105 | 113.5 | 111.1 |
| 9.0 × 104 | 103.3 | 103.8 | 7.0 × 104 | 110.0 | 103.2 | 4.0 × 104 | 113.2 | 110.4 | |
| 9.0 × 103 | 101.7 | 101.5 | 7.0 × 103 | 108.5 | 101.4 | 4.0 × 103 | 112.4 | 107.5 | |
| 9.0 × 102 | 99.5 | 98.9 | 7.0 × 102 | 103.7 | 100.6 | 4.0 × 102 | 107.5 | 101.8 | |
| 9.0 × 101 | 93.1 | 93.7 | 7.0 × 101 | 99.2 | 94.1 | 4.0 × 101 | 99.6 | 96.8 | |
| 9.0 × 100 | 90.9 | / | 7.0 × 100 | 92.4 | / | 4.0 × 100 | 93.6 | / | |
| Milk | 9.0 × 105 | 114.2 | 110.4 | 7.0 × 105 | 113.6 | 103.2 | 4.0 × 105 | 110.5 | 106.2 |
| 9.0 × 104 | 114.0 | 110.1 | 7.0 × 104 | 112.8 | 103.6 | 4.0 × 104 | 109.4 | 106.0 | |
| 9.0 × 103 | 112.9 | 108.3 | 7.0 × 103 | 111.2 | 101.5 | 4.0 × 103 | 108.8 | 104.4 | |
| 9.0 × 102 | 110.4 | 104.3 | 7.0 × 102 | 112.0 | 99.5 | 4.0 × 102 | 109.1 | 102.7 | |
| 9.0 × 101 | 97.9 | 96.1 | 7.0 × 101 | 98.6 | 94.7 | 4.0 × 101 | 94.5 | 94.6 | |
| 9.0 × 100 | 91.3 | / | 7.0 × 100 | 93.7 | / | 4.0 × 100 | 93.3 | / | |
| Chicken breast | 9.0 × 105 | 112.1 | 106.4 | 7.0 × 105 | 111.4 | 113.2 | 4.0 × 105 | 114.1 | 112.8 |
| 9.0 × 104 | 111.4 | 104.8 | 7.0 × 104 | 109.8 | 113.0 | 4.0 × 104 | 113.8 | 110.4 | |
| 9.0 × 103 | 110.5 | 105.1 | 7.0 × 103 | 108.4 | 112.4 | 4.0 × 103 | 113.5 | 111.1 | |
| 9.0 × 102 | 108.1 | 103.2 | 7.0 × 102 | 106.4 | 104.7 | 4.0 × 102 | 106.5 | 106.4 | |
| 9.0 × 101 | 93.8 | 100.3 | 7.0 × 101 | 96.3 | 96.4 | 4.0 × 101 | 96.8 | 99.9 | |
| 9.0 × 100 | 92.4 | / | 7.0 × 100 | 94.1 | / | 4.0 × 100 | 93.9 | / | |
| Shrimp | 9.0 × 105 | 110.8 | 109.3 | 7.0 × 105 | 113.8 | 106.3 | 4.0 × 105 | 112.0 | 104.2 |
| 9.0 × 104 | 109.5 | 107.7 | 7.0 × 104 | 113.2 | 104.9 | 4.0 × 104 | 111.5 | 103.9 | |
| 9.0 × 103 | 106.6 | 105.4 | 7.0 × 103 | 112.6 | 103.3 | 4.0 × 103 | 111.8 | 103.3 | |
| 9.0 × 102 | 104.8 | 102.1 | 7.0 × 102 | 109.0 | 101.1 | 4.0 × 102 | 107.3 | 99.5 | |
| 9.0 × 101 | 97.2 | 93.2 | 7.0 × 101 | 98.1 | 93.1 | 4.0 × 101 | 98.1 | 98.5 | |
| 9.0 × 100 | 94.2 | / | 7.0 × 100 | 93.4 | / | 4.0 × 100 | 95.2 | / | |
| Samples | Listeria monocytogenes | Vibrio parahaemolyticus | Escherichia coli O157:H7 | ||||||
|---|---|---|---|---|---|---|---|---|---|
| EuNP-Based LFIA–RPA | CG–LFIA–RPA | BAM | EuNP-Based LFIA–RPA | CG–LFIA–RPA | BAM | EuNP-Based LFIA–RPA | CG–LFIA–RPA | BAM | |
| BF-1 | − | − | − | − | − | − | − | − | − |
| BF-2 | − | − | − | − | − | − | − | − | − |
| BF-3 | − | − | − | − | − | − | − | − | − |
| BF-4 | − | − | − | − | − | − | − | − | − |
| MK-1 | − | − | − | − | − | − | − | − | − |
| MK-2 | − | − | − | − | − | − | − | − | − |
| MK-3 | − | − | − | − | − | − | − | − | − |
| MK-4 | − | − | − | − | − | − | − | − | − |
| CB-1 | − | − | − | − | − | − | − | − | − |
| CB-2 | − | − | − | − | − | − | − | − | − |
| CB-3 | − | − | − | − | − | − | − | − | − |
| CB-4 | − | − | − | − | − | − | − | − | − |
| SP-1 | − | − | − | − | − | − | − | − | − |
| SP-2 | − | − | − | − | − | − | − | − | − |
| SP-3 | − | − | − | − | − | − | − | − | − |
| SP-4 | − | − | − | − | − | − | − | − | − |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, K.; Ma, B.; Li, J.; Chen, E.; Xu, Y.; Yu, X.; Sun, C.; Zhang, M. A Rapid and Sensitive Europium Nanoparticle-Based Lateral Flow Immunoassay Combined with Recombinase Polymerase Amplification for Simultaneous Detection of Three Food-Borne Pathogens. Int. J. Environ. Res. Public Health 2021, 18, 4574. https://doi.org/10.3390/ijerph18094574
Chen K, Ma B, Li J, Chen E, Xu Y, Yu X, Sun C, Zhang M. A Rapid and Sensitive Europium Nanoparticle-Based Lateral Flow Immunoassay Combined with Recombinase Polymerase Amplification for Simultaneous Detection of Three Food-Borne Pathogens. International Journal of Environmental Research and Public Health. 2021; 18(9):4574. https://doi.org/10.3390/ijerph18094574
Chicago/Turabian StyleChen, Kai, Biao Ma, Jiali Li, Erjing Chen, Ying Xu, Xiaoping Yu, Chuanxin Sun, and Mingzhou Zhang. 2021. "A Rapid and Sensitive Europium Nanoparticle-Based Lateral Flow Immunoassay Combined with Recombinase Polymerase Amplification for Simultaneous Detection of Three Food-Borne Pathogens" International Journal of Environmental Research and Public Health 18, no. 9: 4574. https://doi.org/10.3390/ijerph18094574






