Characterization and Degradation Pathways of Microbacterium resistens MZT7, A Novel 17β-Estradiol-Degrading Bacterium
Abstract
:1. Introduction
2. Materials and Methods
2.1. Chemicals and Cultures Medium
2.2. Enrichment, Isolation, and Identification of E2 Degrading Bacteria
2.3. Genome Sequencing
2.4. Biodegradation Characteristics of E2 by Strain MZT7
2.5. Determination of Residual E2
2.6. Extraction and Detection of E2 and Its Metabolites
3. Results and Discussion
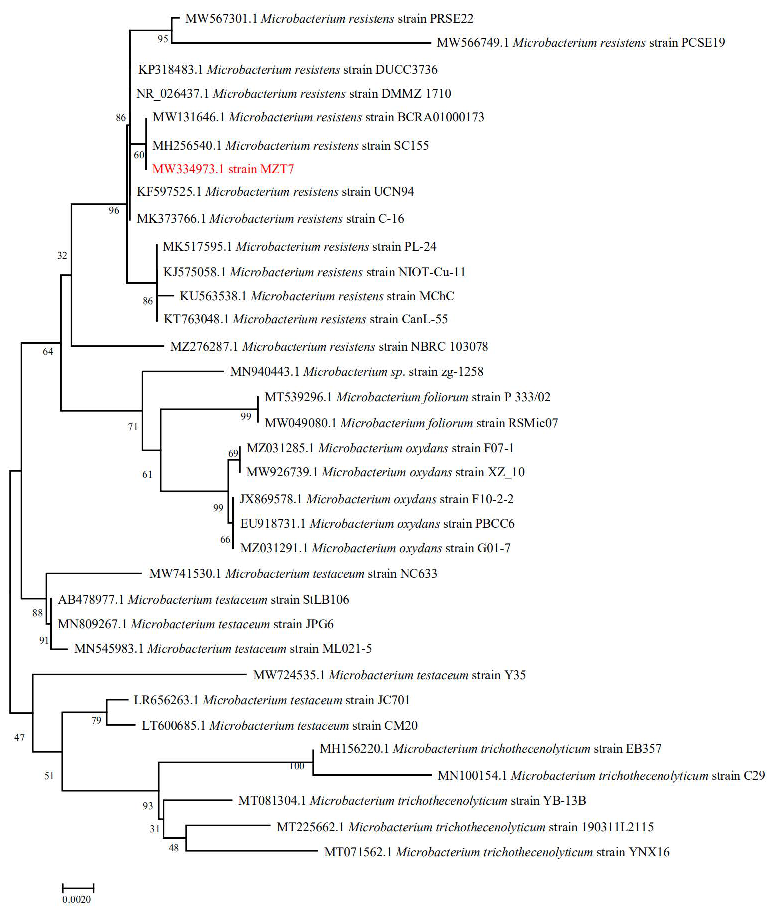
3.1. Isolation and Identification of Strain MZT7
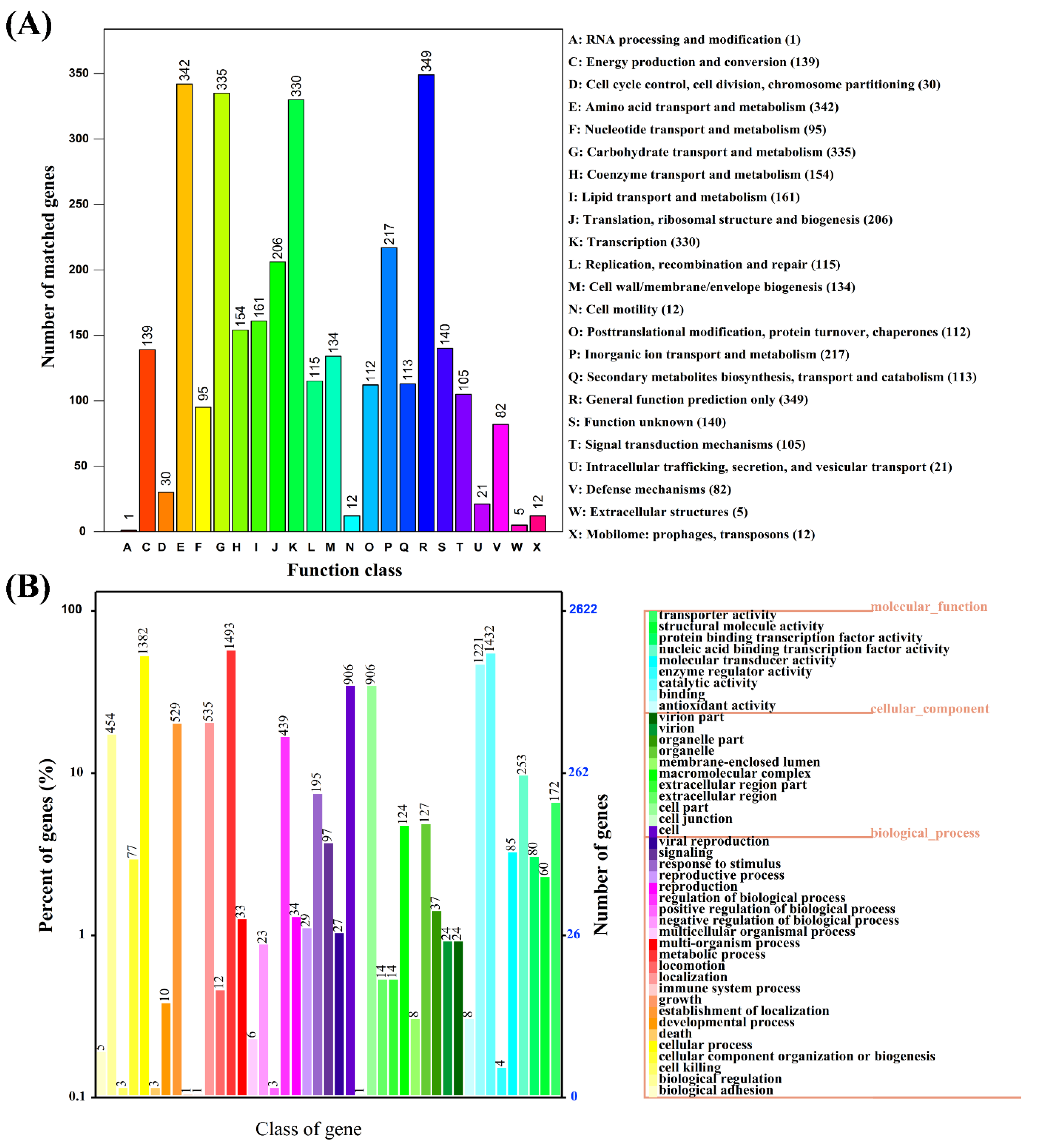
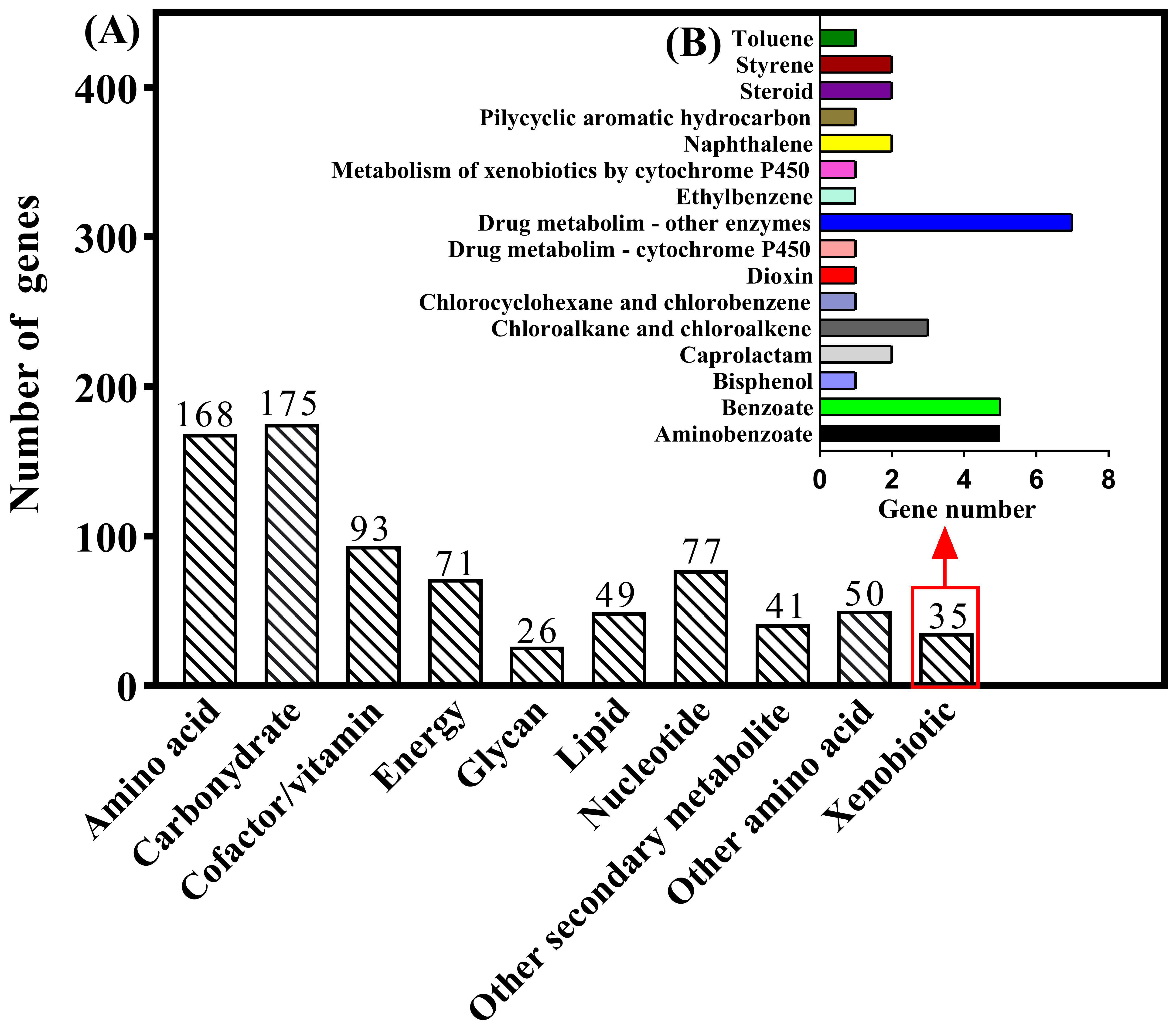
3.2. Genome Characteristics and Database Analysis Related to E2 Degradation
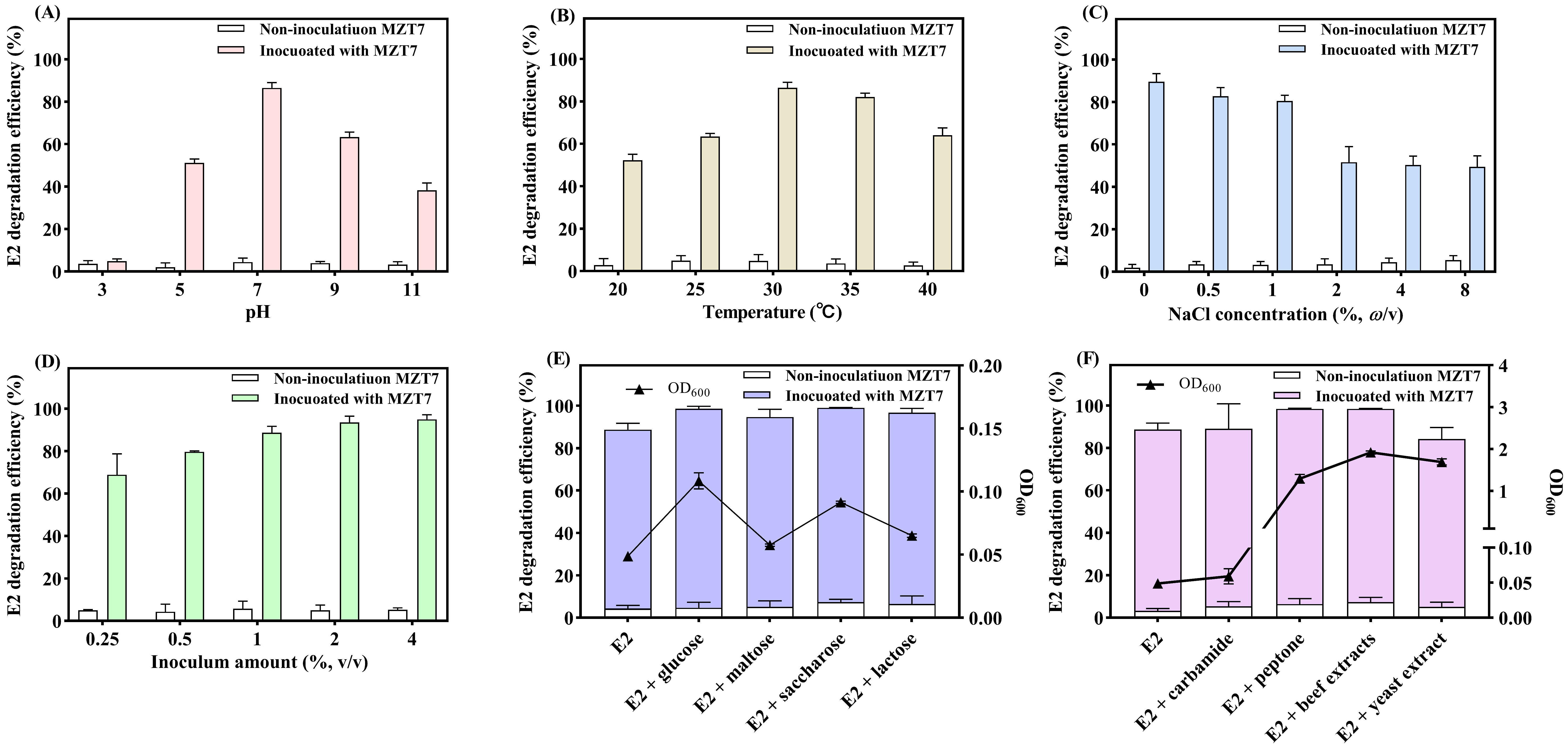
3.3. Effect of pH, Temperature, Salinity, and Inoculum Amount on E2 Biodegradation
3.4. Effect of Carbon Source and Nitrogen Source on E2 Biodegradation
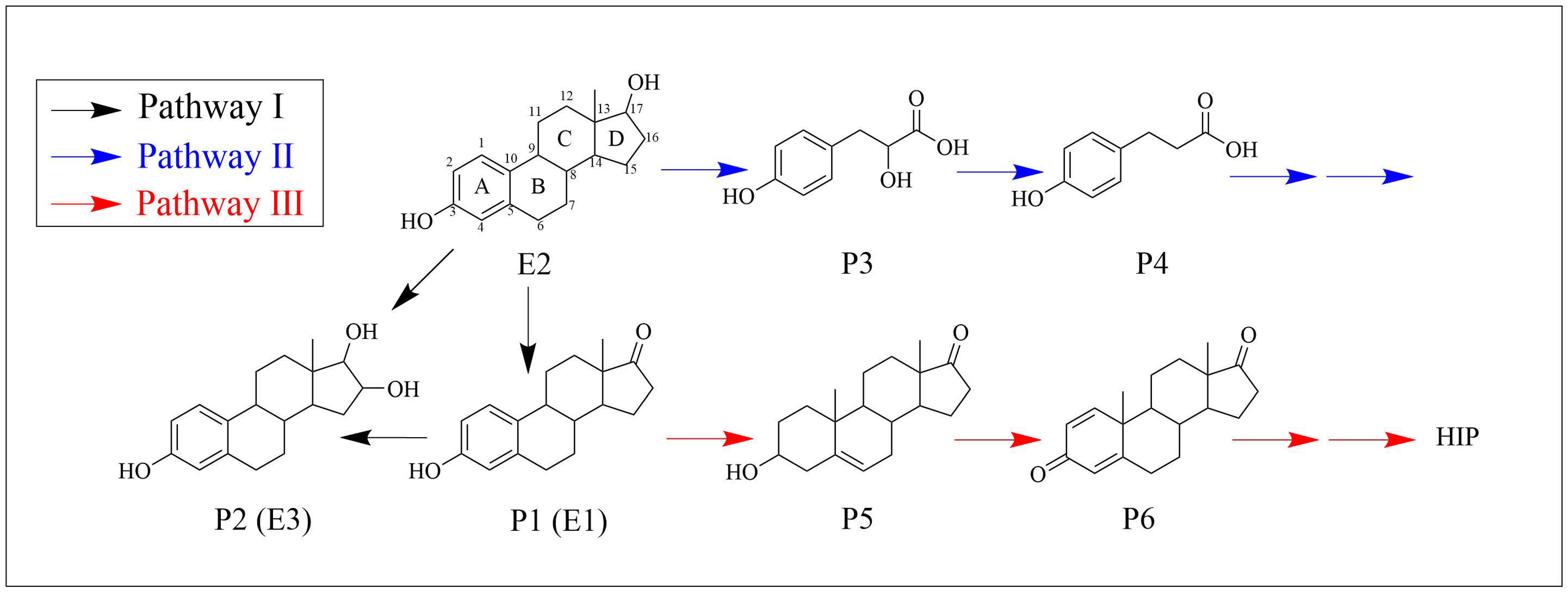
3.5. Biodegradation Products and Metabolic Pathways
3.6. Genome Annotation Reveals Biodegradation Characteristics and Pathways
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hamid, H.; Eskicioglu, C. Fate of estrogenic hormones in wastewater and sludge treatment: A review of properties and analytical detection techniques in sludge matrix. Water Res. 2012, 46, 5813–5833. [Google Scholar] [CrossRef]
- Budeli, P.; Ekwanzala, M.D.; Unuofin, J.O.; Momba, M.N.B. Endocrine disruptive estrogens in wastewater: Revisiting bacterial degradation and zymoremediation. Environ. Technol. Innov. 2021, 21, 101248. [Google Scholar] [CrossRef]
- Lucas, S.; Jones, D. Biodegradation of estrone and 17 β-estradiol in grassland soils amended with animal wastes. Soil Biol. Biochem. 2006, 38, 2803–2815. [Google Scholar] [CrossRef]
- Adeel, M.; Song, X.; Wang, Y.; Francis, D.; Yang, Y. Environmental impact of estrogens on human, animal and plant life: A critical review. Environ. Int. 2017, 99, 107–119. [Google Scholar] [CrossRef] [PubMed]
- Bilal, M.; Iqbal, H.M.N. Persistence and impact of steroidal estrogens on the environment and their laccase-assisted removal. Sci. Total Environ. 2019, 690, 447–459. [Google Scholar] [CrossRef]
- Samavat, H.; Kurzer, M.S. Estrogen metabolism and breast cancer. Cancer Lett. 2015, 356, 231–243. [Google Scholar] [CrossRef]
- Shull, J.D.; Dennison, K.L.; Chack, A.C.; Trentham-Dietz, A. Rat models of 17beta-estradiol-induced mammary cancer reveal novel insights into breast cancer etiology and prevention. Physiol Genom. 2018, 50, 215–234. [Google Scholar] [CrossRef]
- Yang, L.; Lin, L.; Weng, S.; Feng, Z.; Luan, T. Sexually disrupting effects of nonylphenol and diethylstilbestrol on male silver carp (Carassius auratus) in aquatic microcosms. Ecotoxicol. Environ. Saf. 2008, 71, 400–411. [Google Scholar] [CrossRef]
- Jobling, S.; Williams, R.; Johnson, A.; Taylor, A.; Gross-Sorokin, M.; Nolan, M.; Tyler, C.R.; van Aerle, R.; Santos, E.; Brighty, G. Predicted exposures to steroid estrogens in U.K. rivers correlate with widespread sexual disruption in wild fish populations. Environ. Health Perspect. 2006, 114 (Suppl. S1), 32–39. [Google Scholar] [CrossRef]
- Wang, H.-P.; Gao, Z.; Beres, B.; Ottobre, J.; Wallat, G.; Tiu, L.; Rapp, D.; O’Bryant, P.; Yao, H. Effects of estradiol-17β on survival, growth performance, sex reversal and gonadal structure of bluegill sunfish Lepomis macrochirus. Aquaculture 2008, 285, 216–223. [Google Scholar] [CrossRef]
- Zhang, Q.; Xue, C.; Owens, G.; Chen, Z. Isolation and identification of 17β-estradiol degrading bacteria and its degradation pathway. J. Hazard. Mater. 2021, 423, 127185. [Google Scholar] [CrossRef]
- Wang, P.; Zheng, D.; Peng, W.; Wang, Y.; Wang, X.; Xiong, W.; Liang, R. Characterization of 17beta-hydroxysteroid dehydrogenase and regulators involved in estrogen degradation in Pseudomonas putida SJTE-1. Appl. Microbiol. Biotechnol. 2019, 103, 2413–2425. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.; Yin, C.; Peng, W.; Deng, Z.; Lin, S.; Liang, R. Characterization of an 17beta-estradiol-degrading bacterium Stenotrophomonas maltophilia SJTL3 tolerant to adverse environmental factors. Appl. Microbiol. Biotechnol. 2020, 104, 1291–1305. [Google Scholar] [CrossRef]
- Kurisu, F.; Ogura, M.; Saitoh, S.; Yamazoe, A.; Yagi, O. Degradation of natural estrogen and identification of the metabolites produced by soil isolates of Rhodococcus sp. and Sphingomonas sp. J. Biosci. Bioeng. 2010, 109, 576–582. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Qin, D.; Sun, Q.; Zhang, F.; Liu, H.; Yu, C.P. Removal of environmental estrogens by bacterial cell immobilization technique. Chemosphere 2016, 144, 607–614. [Google Scholar] [CrossRef]
- Qiu, Q.; Wang, P.; Kang, H.; Wang, Y.; Tian, K.; Huo, H. Genomic Analysis of a New Estrogen-Degrading Bacterial Strain, Acinetobacter sp. DSSKY-A-001. Int. J. Genomics 2019, 2019, 2804134. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.; Yin, C.; Wang, Y.; Lin, S.; Deng, Z.; Liang, R. Characterization of an efficient estrogen-degrading bacterium Stenotrophomonas maltophilia SJTH1 in saline-, alkaline-, heavy metal-contained environments or solid soil and identification of four 17beta-estradiol-oxidizing dehydrogenases. J. Hazard. Mater. 2020, 385, 121616. [Google Scholar] [CrossRef]
- Nakai, S.; Yamamura, A.; Tanaka, S.; Shi, J.; Nishikawa, M.; Nakashimada, Y.; Hosomi, M. Pathway of 17β-estradiol degradation by Nitrosomonas europaea and reduction in 17β-estradiol-derived estrogenic activity. Environ. Chem. Lett. 2011, 9, 1–6. [Google Scholar] [CrossRef]
- Ye, X.; Wang, H.; Kan, J.; Li, J.; Huang, T.; Xiong, G.; Hu, Z. A novel 17beta-hydroxysteroid dehydrogenase in Rhodococcus sp. P14 for transforming 17beta-estradiol to estrone. Chem. Biol. Interact. 2017, 276, 105–112. [Google Scholar] [CrossRef]
- Yu, C.-P.; Roh, H.; Chu, K.-H. 17β-Estradiol-Degrading Bacteria Isolated from Activated Sludge. Environ. Sci. Technol. 2007, 41, 486–492. [Google Scholar] [CrossRef]
- Li, S.; Sun, K.; Yan, X.; Lu, C.; Waigi, M.G.; Liu, J.; Ling, W. Identification of novel catabolic genes involved in 17beta-estradiol degradation by Novosphingobium sp. ES2-1. Environ. Microbiol. 2021, 23, 2550–2563. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, T.H.; Lee, T.H.; Chuang, M.R.; Wang, P.H.; Meng, M.; Horinouchi, M.; Hayashi, T.; Chen, Y.L.; Chiang, Y.R. Identification of essential beta-oxidation genes and corresponding metabolites for oestrogen degradation by actinobacteria. Microb. Biotechnol. 2022, 15, 949–966. [Google Scholar] [CrossRef] [PubMed]
- Ibero, J.; Galan, B.; Rivero-Buceta, V.; Garcia, J.L. Unraveling the 17beta-Estradiol Degradation Pathway in Novosphingobium tardaugens NBRC 16725. Front. Microbiol. 2020, 11, 588300. [Google Scholar] [CrossRef] [PubMed]
- Ge, H.; Yang, L.; Li, B.; Feng, Y.; Wang, S.; Zheng, Y.; Feng, L.; Liu, Y.; Du, Z.; Zhang, L. A Comparative Study on the Biodegradation of 17β-Estradiol by Candida utilis CU-2 and Lactobacillus casei LC-1. Front. Energy Res. 2021, 9, 661850. [Google Scholar] [CrossRef]
- Chen, Y.L.; Yu, C.P.; Lee, T.H.; Goh, K.S.; Chu, K.H.; Wang, P.H.; Ismail, W.; Shih, C.J.; Chiang, Y.R. Biochemical Mechanisms and Catabolic Enzymes Involved in Bacterial Estrogen Degradation Pathways. Cell Chem. Biol. 2017, 24, 712–724.e7. [Google Scholar] [CrossRef]
- Kube, M.; Chernikova, T.N.; Al-Ramahi, Y.; Beloqui, A.; Lopez-Cortez, N.; Guazzaroni, M.E.; Heipieper, H.J.; Klages, S.; Kotsyurbenko, O.R.; Langer, I.; et al. Genome sequence and functional genomic analysis of the oil-degrading bacterium Oleispira antarctica. Nat. Commun. 2013, 4, 2156. [Google Scholar] [CrossRef]
- Lee, Y.K.; Kim, H.W.; Liu, C.L.; Lee, H.K. A simple method for DNA extraction from marine bacteria that produce extracellular materials. J. Microbiol. Methods 2003, 52, 245–250. [Google Scholar] [CrossRef]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Liu, N.; Shi, Y.E.; Li, J.; Zhu, M.; Zhang, T. Isolation and characterization of a new highly effective 17beta-estradiol-degrading Gordonia sp. strain R9. 3 Biotech 2020, 10, 174. [Google Scholar] [CrossRef]
- Hyndman, D.; Bauman, D.R.; Heredia, V.V.; Penning, T.M. The aldo-keto reductase superfamily homepage. Chem.-Biol. Interact. 2003, 143–144, 621–631. [Google Scholar] [CrossRef]
- Baldanta, S.; Navarro Llorens, J.M.; Guevara, G. Further Studies on the 3-Ketosteroid 9alpha-Hydroxylase of Rhodococcus ruber Chol-4, a Rieske Oxygenase of the Steroid Degradation Pathway. Microorganisms 2021, 9, 1171. [Google Scholar] [CrossRef] [PubMed]
- Rohman, A.; van Oosterwijk, N.; Thunnissen, A.M.; Dijkstra, B.W. Crystal structure and site-directed mutagenesis of 3-ketosteroid Delta1-dehydrogenase from Rhodococcus erythropolis SQ1 explain its catalytic mechanism. J. Biol. Chem. 2013, 288, 35559–35568. [Google Scholar] [CrossRef]
- Wong, J.W.C.; Lai, K.M.; Wan, C.K.; Ma, K.K.; Fang, M. Isolation and optimi zation of PAH-degradative bacteria from contaminated soil for PAHs bioreme diation. Water Air Soil Pollut. Water Air Soil Pollut 2002, 139, 1–13. [Google Scholar] [CrossRef]
- Zhang, H.; Mu, W.; Hou, Z.; Wu, X.; Zhao, W.; Zhang, X.; Pan, H.; Zhang, S. Biodegradation of nicosulfuron by the bacterium Serratia marcescens N80. J. Environ. Sci. Health B 2012, 47, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Zhao, Y.; Ding, H.; Lin, X.; Zheng, H. Co-metabolic degradation of bensulfuron-methyl in laboratory, conditions. J. Hazard. Mater. 2008, 158, 208–214. [Google Scholar] [CrossRef]
- Long, Z.; Wang, X.; Wang, Y.; Dai, H.; Li, C.; Xue, Y.; Deng, Y.; Zhang, H.; Yu, Y.; Fang, H. Characterization of a novel carbendazim-degrading strain Rhodococcus sp. CX-1 revealed by genome and transcriptome analyses. Sci. Total Environ. 2021, 754, 142137. [Google Scholar] [CrossRef]
- Zhang, Q.; Xie, X.; Xu, D.; Hong, R.; Wu, J.; Zeng, X.; Liu, N.; Liu, J. Accelerated azo dye biodegradation and detoxification by Pseudomonas aeruginosa DDMZ1-2 via fructose co-metabolism. Environ. Technol. Innov. 2021, 24, 101878. [Google Scholar] [CrossRef]
- Peng, X.; Jia, X. Optimization of parameters for anaerobic co-metabolic degradation of TBBPA. Bioresour Technol. 2013, 148, 386–393. [Google Scholar] [CrossRef]
- Qi, W.; Long, J.; Feng, C.; Feng, Y.; Cheng, D.; Liu, Y.; Xue, J.; Li, Z. Fe(3+) enhanced degradation of oxytetracycline in water by pseudomonas. Water Res. 2019, 160, 361–370. [Google Scholar] [CrossRef]
- Tang, Y.; Zhang, Y.; Yan, N.; Liu, R.; Rittmann, B.E. The role of electron donors generated from UV photolysis for accelerating pyridine biodegradation. Biotechnol. Bioeng. 2015, 112, 1792–1800. [Google Scholar] [CrossRef]
- Bai, Q.; Yang, L.; Li, R.; Chen, B.; Zhang, L.; Zhang, Y.; Rittmann, B.E. Accelerating Quinoline Biodegradation and Oxidation with Endogenous Electron Donors. Environ. Sci. Technol. 2015, 49, 11536–11542. [Google Scholar] [CrossRef] [PubMed]
- Dytczak, M.A.; Londry, K.L.; Oleszkiewicz, J.A. Biotransformation of Estrogens in Nitrifying Activated Sludge Under Aerobic and Alternating Anoxic/Aerobic Conditions. Water Environ. Res. 2008, 80, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Lai, K.M.; Scrimshaw, M.D.; Lester, J.N. Biotransformation and bioconcentration of steroid estrogens by Chlorella vulgaris. Appl. Environ. Microbiol. 2002, 68, 859–864. [Google Scholar] [CrossRef] [PubMed]
- Combalbert, S.; Hernandez-Raquet, G. Occurrence, fate, and biodegradation of estrogens in sewage and manure. Appl. Microbiol. Biotechnol. 2010, 86, 1671–1692. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.H.; Chen, Y.L.; Wei, S.T.; Wu, K.; Lee, T.H.; Wu, T.Y.; Chiang, Y.R. Retroconversion of estrogens into androgens by bacteria via a cobalamin-mediated methylation. Proc. Natl. Acad. Sci. USA 2020, 117, 1395–1403. [Google Scholar] [CrossRef]
- Wu, K.; Lee, T.H.; Chen, Y.L.; Wang, Y.S.; Wang, P.H.; Yu, C.P.; Chu, K.H.; Chiang, Y.R. Metabolites Involved in Aerobic Degradation of the A and B Rings of Estrogen. Appl. Environ. Microbiol. 2019, 85, e02223-18. [Google Scholar] [CrossRef]
- Xu, J.; Zhang, L.; Hou, J.; Wang, X.; Liu, H.; Zheng, D.; Liang, R. iTRAQ-based quantitative proteomic analysis of the global response to 17beta-estradiol in estrogen-degradation strain Pseudomonas putida SJTE-1. Sci. Rep. 2017, 7, 41682. [Google Scholar] [CrossRef]
- Jiang, W.; Hou, Y.; Inouye, M. CspA, the major cold-shock protein of Escherichia coli, is an RNA chaperone. J. Biol. Chem. 1997, 272, 196–202. [Google Scholar] [CrossRef]
- Vahdani, F.; Ghafouri, H.; Sarikhan, S.; Khodarahmi, R. Molecular cloning, expression, and functional characterization of 70-kDa heat shock protein, DnaK, from Bacillus halodurans. Int. J. Biol. Macromol. 2019, 137, 151–159. [Google Scholar] [CrossRef]
- Johnston, D.M.; Miot, M.; Hoskins, J.R.; Wickner, S.; Doyle, S.M. Substrate Discrimination by ClpB and Hsp104. Front. Mol. Biosci. 2017, 4, 36. [Google Scholar] [CrossRef] [Green Version]
- Shin, D.H.; Choi, I.G.; Busso, D.; Jancarik, J.; Yokota, H.; Kim, R.; Kim, S.H. Structure of OsmC from Escherichia coli: A salt-shock-induced protein. Acta Crystallogr. D Biol. Crystallogr. 2004, 60, 903–911. [Google Scholar] [CrossRef]
- Ibero, J.; Galan, B.; Diaz, E.; Garcia, J.L. Testosterone Degradative Pathway of Novosphingobium tardaugens. Genes 2019, 10, 871. [Google Scholar] [CrossRef] [PubMed]
- Labrie, F.; Luu-The, V.; Lin, S.X.; Labrie, C.; Simard, J.; Breton, R.; Belanger, A. The key role of 17β-hydroxysteroid dehydrogenases in sex steroid biology. Steroids 1997, 62, 148–158. [Google Scholar] [CrossRef]
- Wang, P.; Zheng, D.; Wang, Y.; Liang, R. One 3-oxoacyl-(acyl-Carrier-protein) reductase functions as 17beta-hydroxysteroid dehydrogenase in the estrogen-degrading Pseudomonas putida SJTE-1. Biochem. Biophys. Res. Commun. 2018, 505, 910–916. [Google Scholar] [CrossRef] [PubMed]

| ID | Formula | RT a (min) | m/z b | Exact Mass |
|---|---|---|---|---|
| E2 | C18H24O2 | 12.52 | 273.2530 | 272.1776 |
| P1 (E1) | C18H22O2 | 13.07 | 271.1686 | 270.162 |
| P2 (E3) | C18H24O3 | 9.12 | 288.1493 | 288.1725 |
| P3 | C9H10O4 | 13.03 | 182.0579 | 182.0579 |
| P4 | C9H10O | 1.22 | 167.0126 | 166.063 |
| P5 | C19H28O2 | 13.49 | 269.2105 | 288.2089 |
| P6 | C19H24O2 | 7.81 | 284.1853 | 284.1776 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hao, P.; Wu, S.; Zhang, X.; Gou, C.; Wang, Y.; Wang, L.; Zhu, Y.; Basang, W.; Gao, Y. Characterization and Degradation Pathways of Microbacterium resistens MZT7, A Novel 17β-Estradiol-Degrading Bacterium. Int. J. Environ. Res. Public Health 2022, 19, 11097. https://doi.org/10.3390/ijerph191711097
Hao P, Wu S, Zhang X, Gou C, Wang Y, Wang L, Zhu Y, Basang W, Gao Y. Characterization and Degradation Pathways of Microbacterium resistens MZT7, A Novel 17β-Estradiol-Degrading Bacterium. International Journal of Environmental Research and Public Health. 2022; 19(17):11097. https://doi.org/10.3390/ijerph191711097
Chicago/Turabian StyleHao, Peng, Sicheng Wu, Xiqing Zhang, Changlong Gou, Yuqiong Wang, Lixia Wang, Yanbin Zhu, Wangdui Basang, and Yunhang Gao. 2022. "Characterization and Degradation Pathways of Microbacterium resistens MZT7, A Novel 17β-Estradiol-Degrading Bacterium" International Journal of Environmental Research and Public Health 19, no. 17: 11097. https://doi.org/10.3390/ijerph191711097







