Control Strategies of Plastic Biodegradation through Adjusting Additives Ratios Using In Silico Approaches Associated with Proportional Factorial Experimental Design
Abstract
:1. Introduction
2. Materials and Methods
2.1. Biodegradability Assessment of the Plastic Component PS and Its Additives Using Molecular Docking and Molecular Dynamics Simulations
2.2. Proportional Factorial Experimental Design Method for Screening the Proportioning Scheme of Plastic Components Affecting Biodegradability
3. Results and Discussion
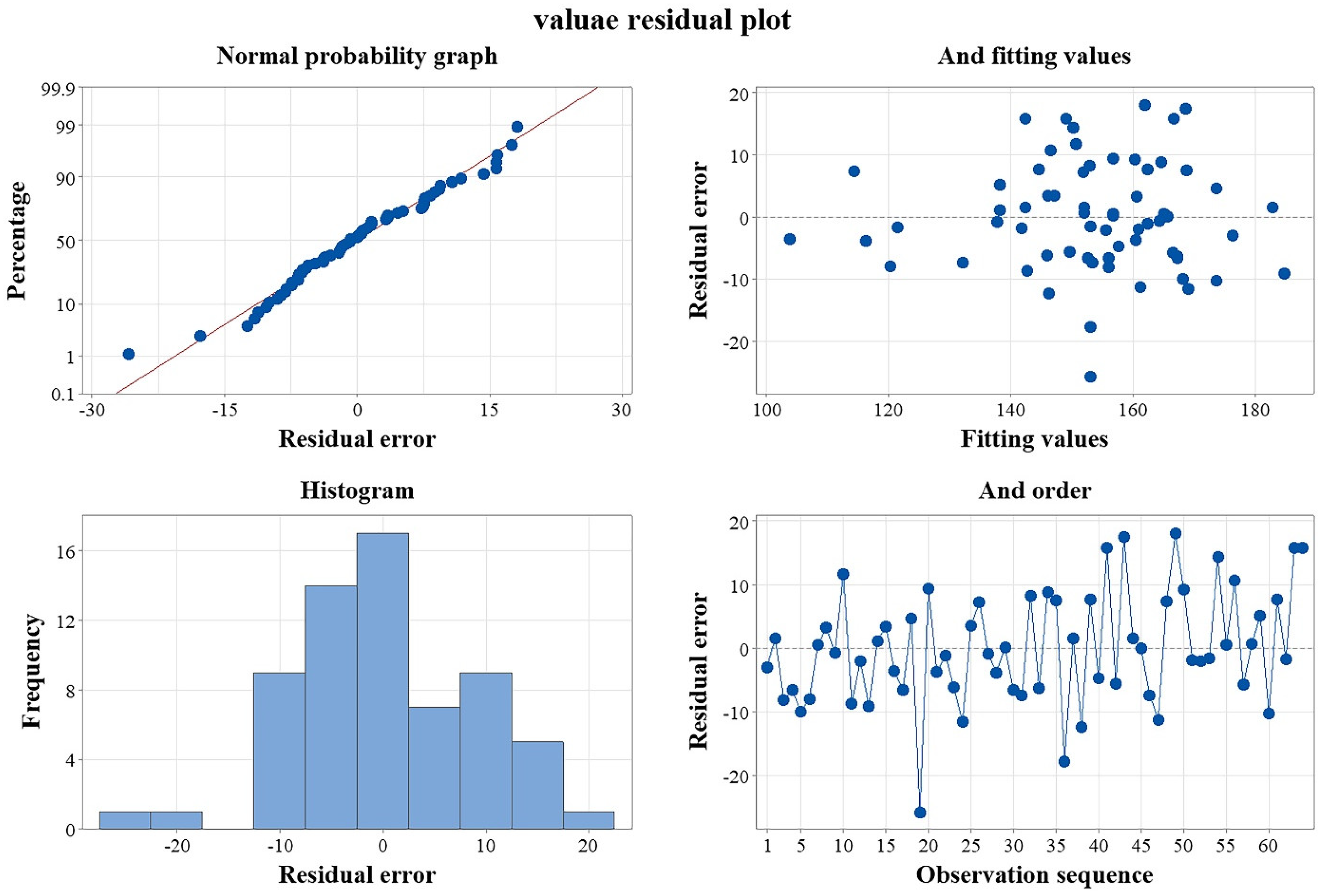
3.1. Biodegradability Features of Plastics Using the Entire Factorial Experimental Design Group’s Distribution Proportion Scheme
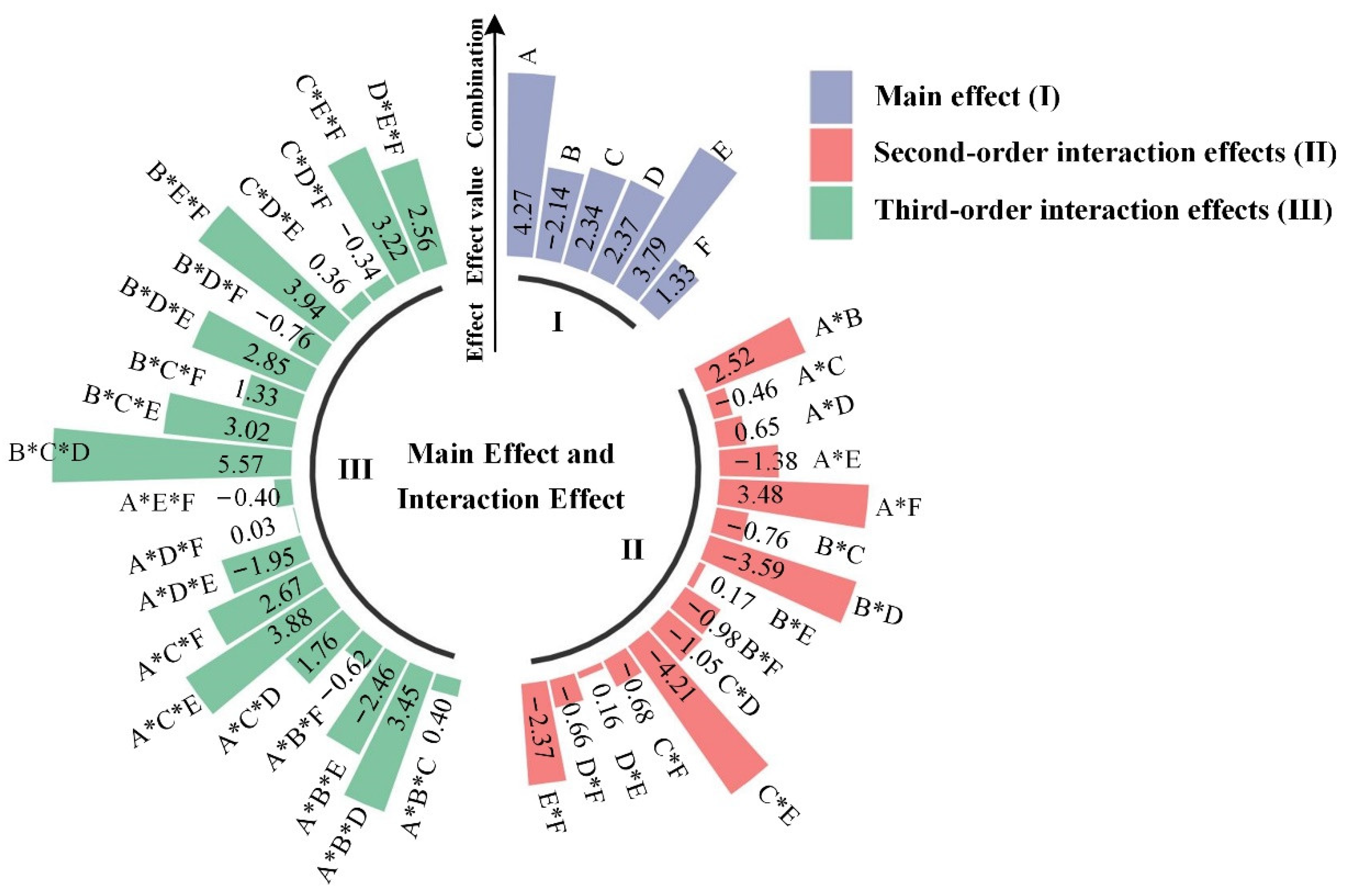
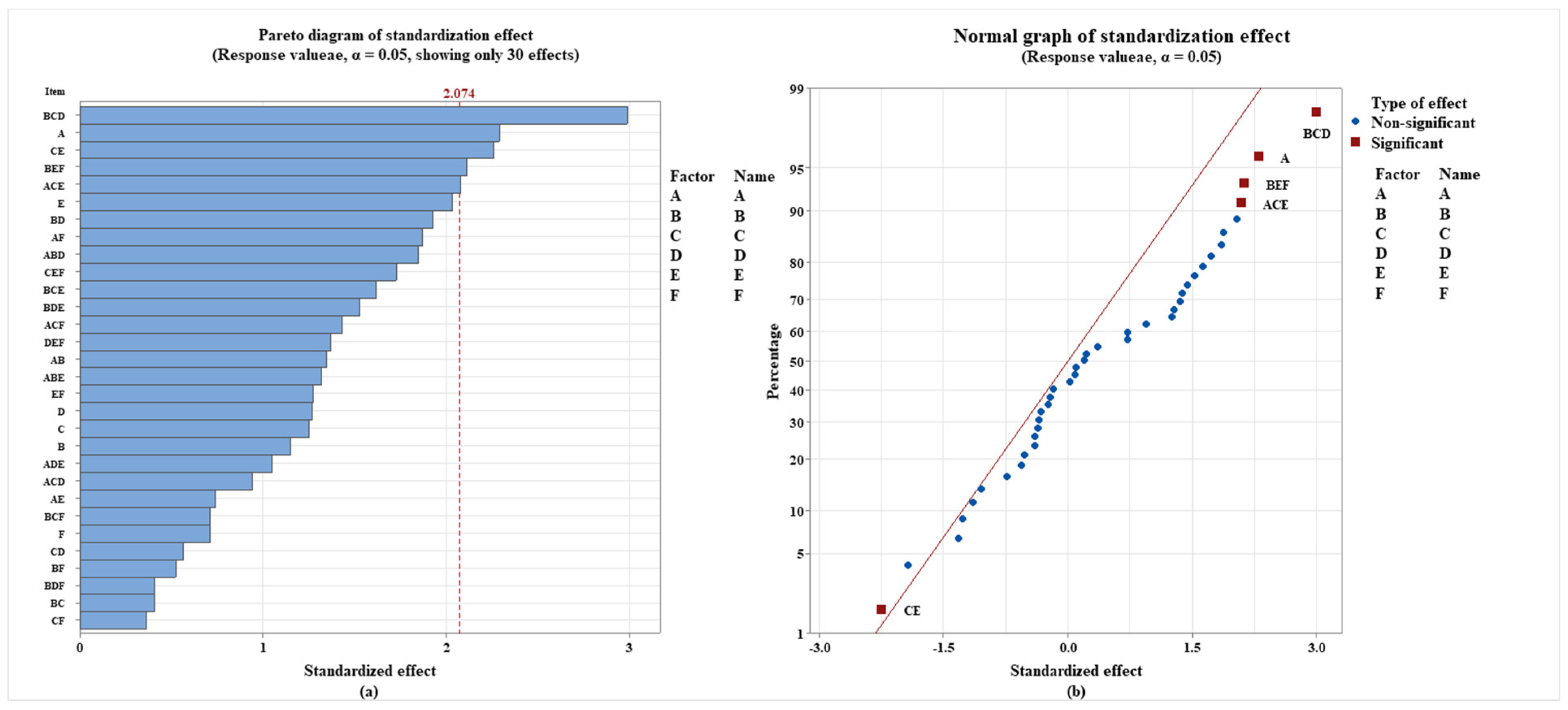
3.2. Analysis of the Primary Impacts of Plastic Additives and Their Interactions on the Biodegradability of Plastics
3.2.1. Biodegradation Characteristics of Plastics under an Additive Combination Scheme Using a Full Factorial Design
3.2.2. The Effect of Plastic Additives on the Biodegradability of Polymers Was Investigated
3.3. Factorial Analysis and Molecular Dynamics Simulation Were Used to Investigate the Mechanism of Plastic Biodegradation
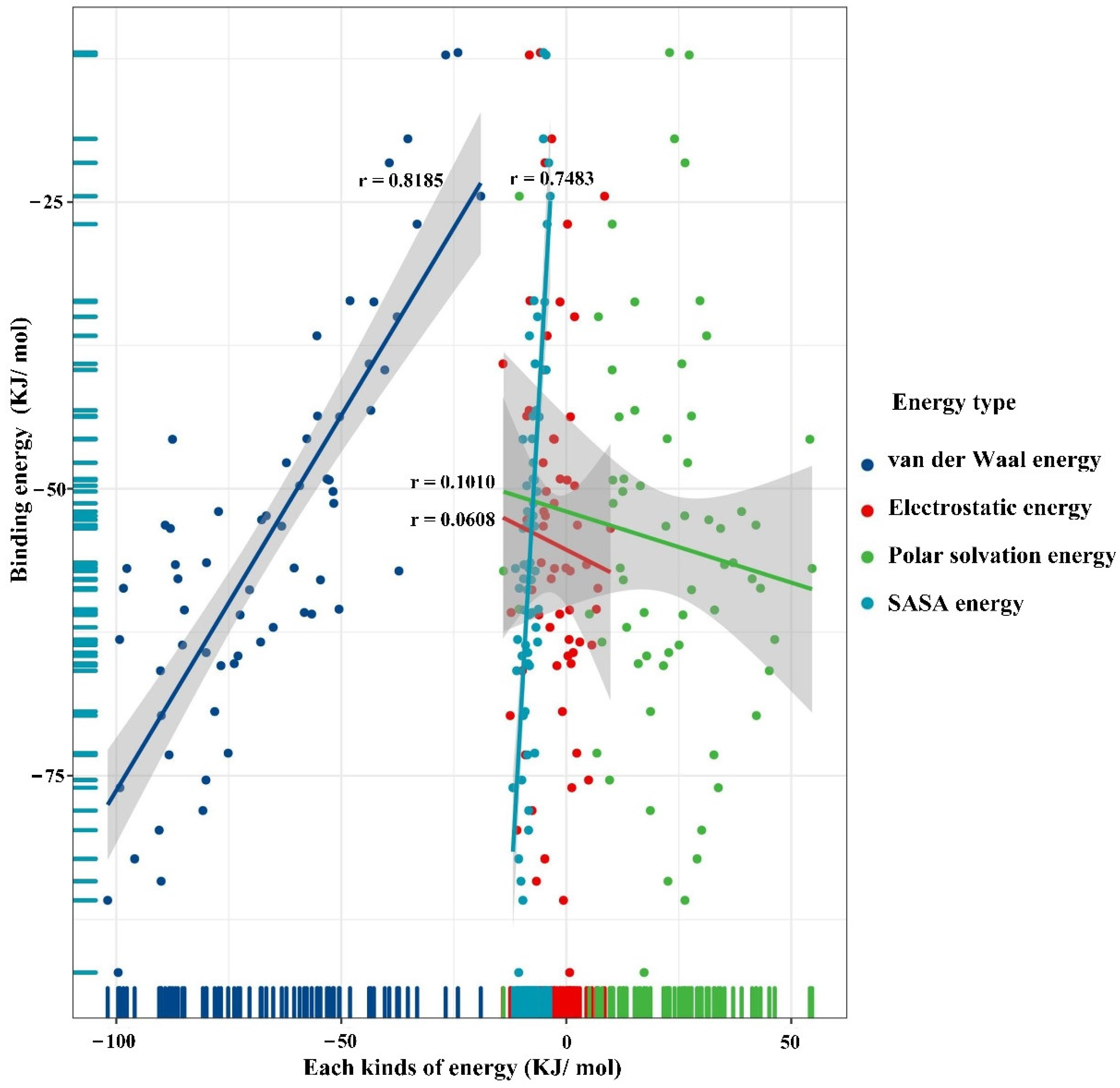
3.3.1. Molecular Dynamics Simulation-Based Mechanism Investigation of the Biodegradation Effect of Polystyrene 5 Polymers
3.3.2. Total Factor and Amino Acid Residue Analysis Were Used to Investigate the Mechanism of Biodegradation of Polystyrene 5 Polymers
3.3.3. Based on Factorial Design Verification, We Examined the Biodegradation Mechanism of the Polystyrene 5 Polymer
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Derraik, J.G.B. The pollution of the marine environment by plastic debris: A review. Mar. Pollut. Bull. 2002, 44, 842–852. [Google Scholar] [CrossRef]
- Laist, D.W. Overview of the biological effects of lost and discarded plastic debris in the marine environment. Mar. Pollut. Bull. 1987, 18, 319–326. [Google Scholar] [CrossRef]
- Heidbreder, L.M.; Bablok, I.; Drews, S.; Menzel, C. Tackling the plastic problem: A review on perceptions, behaviors, and interventions. Sci. Total Environ. 2019, 668, 1077–1093. [Google Scholar] [CrossRef] [PubMed]
- Worm, B.; Lotze, H.K.; Jubinville, I.; Wilcox, C.; Jambeck, J. Plastic as a Persistent Marine Pollutant. Annu. Rev. Environ. Resour. 2017, 42, 1–26. [Google Scholar] [CrossRef]
- Andrady, A.L. Microplastics in the marine environment. Mar. Pollut. Bull. 2011, 62, 1596–1605. [Google Scholar] [CrossRef] [PubMed]
- Ogata, Y.; Takada, H.; Mizukawa, K.; Hirai, H.; Iwasa, S.; Endo, S.; Mato, Y.; Saha, M.; Okuda, K.; Nakashima, A.; et al. International Pellet Watch: Global monitoring of persistent organic pollutants (POPs) in coastal waters. 1. Initial phase data on PCBs, DDTs, and HCHs. Mar. Pollut. Bull. 2009, 58, 1437–1446. [Google Scholar] [CrossRef] [PubMed]
- Doyle, M.J.; Watson, W.; Bowlin, N.M.; Sheavly, S.B. Plastic particles in coastal pelagic ecosystems of the Northeast Pacific ocean. Mar. Environ. Res. 2011, 71, 41–52. [Google Scholar] [CrossRef]
- Lu, Y.; Zhang, Y.; Deng, Y.; Jiang, W.; Zhao, Y.; Geng, J.; Ding, L.; Ren, H.-Q. Uptake and Accumulation of Polystyrene Microplastics in Zebrafish (Danio rerio) and Toxic Effects in Liver. Environ. Sci. Technol. 2016, 50, 4054–4060. [Google Scholar] [CrossRef]
- Wan, Z.; Wang, C.; Zhou, J.; Shen, M.; Wang, X.; Fu, Z.; Jin, Y. Effects of polystyrene microplastics on the composition of the microbiome and metabolism in larval zebrafish. Chemosphere 2018, 217, 646–658. [Google Scholar] [CrossRef]
- Sussarellu, R.; Suquet, M.; Thomas, Y.; Lambert, C.; Fabioux, C.; Pernet, M.E.J.; Le Goïc, N.; Quillien, V.; Mingant, C.; Epelboin, Y.; et al. Oyster reproduction is affected by exposure to polystyrene microplastics. Proc. Natl. Acad. Sci. USA 2016, 113, 2430–2435. [Google Scholar] [CrossRef] [Green Version]
- Yu, Y.; Chen, H.; Hua, X.; Dang, Y.; Han, Y.; Yu, Z.; Chen, X.; Ding, P.; Li, H. Polystyrene microplastics (PS-MPs) toxicity induced oxidative stress and intestinal injury in nematode Caenorhabditis elegans. Sci. Total Environ. 2020, 726, 138679. [Google Scholar] [CrossRef] [PubMed]
- Beiras, R.; Verdejo, E.; Campoy-López, P.; Vidal-Liñán, L. Aquatic toxicity of chemically defined microplastics can be explained by functional additives. J. Hazard. Mater. 2020, 406, 124338. [Google Scholar] [CrossRef] [PubMed]
- Campanale, C.; Massarelli, C.; Savino, I.; Locaputo, V.; Uricchio, V.F. A Detailed Review Study on Potential Effects of Microplastics and Additives of Concern on Human Health. Int. J. Environ. Res. Public Health 2020, 17, 1212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hahladakis, J.N.; Velis, C.A.; Weber, R.; Iacovidou, E.; Purnell, P. An overview of chemical additives present in plastics: Migration, release, fate and environmental impact during their use, disposal and recycling. J. Hazard. Mater. 2018, 344, 179–199. [Google Scholar] [CrossRef]
- Liu, W.; Zhao, Y.; Shi, Z.; Li, Z.; Liang, X. Ecotoxicoproteomic assessment of microplastics and plastic additives in aquatic organisms: A review. Comp. Biochem. Physiol. Part D Genom. Proteom. 2020, 36, 100713. [Google Scholar] [CrossRef]
- Faure, F.; Demars, C.; Wieser, O.; Kunz, M.; de Alencastro, L.F. Plastic pollution in Swiss surface waters: Nature and concentrations, interaction with pollutants. Environ. Chem. 2015, 12, 582. [Google Scholar] [CrossRef]
- Solleiro-Villavicencio, H.; León, C.T.G.; Araiza, V.H.D.R.; Morales-Montor, J. The detrimental effect of microplastics on critical periods of development in the neuroendocrine system. Birth Defects Res. 2020, 112, 1326–1340. [Google Scholar] [CrossRef]
- Wei, G.-L.; Li, D.-Q.; Zhuo, M.-N.; Liao, Y.-S.; Xie, Z.-Y.; Guo, T.-L.; Li, J.-J.; Zhang, S.-Y.; Liang, Z.-Q. Organophosphorus flame retardants and plasticizers: Sources, occurrence, toxicity and human exposure. Environ. Pollut. 2015, 196, 29–46. [Google Scholar] [CrossRef]
- Zhang, H.; Zhou, Q.; Xie, Z.; Zhou, Y.; Tu, C.; Fu, C.; Mi, W.; Ebinghaus, R.; Christie, P.; Luo, Y. Occurrences of organophosphorus esters and phthalates in the microplastics from the coastal beaches in north China. Sci. Total Environ. 2017, 616–617, 1505–1512. [Google Scholar] [CrossRef]
- Browne, M.A.; Niven, S.J.; Galloway, T.S.; Rowland, S.J.; Thompson, R. Microplastic Moves Pollutants and Additives to Worms, Reducing Functions Linked to Health and Biodiversity. Curr. Biol. 2013, 23, 2388–2392. [Google Scholar] [CrossRef] [Green Version]
- Di Renzo, L.; Mascilongo, G.; Berti, M.; Bogdanović, T.; Listeš, E.; Brkljača, M.; Notarstefano, V.; Gioacchini, G.; Giorgini, E.; Olivieri, V.; et al. Potential Impact of Microplastics and Additives on the Health Status of Loggerhead Turtles (Caretta caretta) Stranded Along the Central Adriatic Coast. Water Air Soil Pollut. 2021, 232, 98. [Google Scholar] [CrossRef]
- Nobre, C.R.; Santana, M.F.M.; Maluf, A.; Cortez, F.S.; Cesar, A.; Pereira, C.D.C.; Turra, A. Assessment of microplastic toxicity to embryonic development of the sea urchin Lytechinus variegatus (Echinodermata: Echinoidea). Mar. Pollut. Bull. 2015, 92, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Yanful, E.K.; Bassi, A.S. A Review of Plastic Waste Biodegradation. Crit. Rev. Biotechnol. 2005, 25, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Blanco, I. Lifetime prediction of food and beverage packaging wastes. J. Therm. Anal. 2015, 125, 809–816. [Google Scholar] [CrossRef]
- Blanco, I. Lifetime Prediction of Polymers: To Bet, or Not to Bet-Is This the Question? Materials 2018, 11, 1383. [Google Scholar] [CrossRef] [Green Version]
- Alauddin, M.; Choudhury, I.A.; EI Baradie, M.A.; Hashmi, M.S.J. Plastics and their machining: A review. J. Mater. Processing Technol. 1995, 54, 40–46. [Google Scholar] [CrossRef]
- Allena, A.B.; Hilliard, N.P.; Howard, G.T. Purication and characterization of a soluble polyurethane degrading enzyme from Comamonas acidovorans. Int. Biodeterior. Biodegrad. 1999, 43, 37–41. [Google Scholar] [CrossRef]
- Blanco, I.; Abate, L.; Antonelli, M.L.; Bottino, F. The regression of isothermal thermogravimetric data to evaluate degradation Ea values of polymers: A comparison with literature methods and an evaluation of lifetime predictions reliability. Part II. Polym. Degrad. Stab. 2013, 98, 2291–2296. [Google Scholar] [CrossRef]
- Müller, R.-J.; Kleeberg, I.; Deckwer, W.-D. Biodegradation of polyesters containing aromatic constituents. J. Biotechnol. 2001, 86, 87–95. [Google Scholar] [CrossRef]
- Mor, R.; Sivan, A. Biofilm formation and partial biodegradation of polystyrene by the actinomycete Rhodococcus ruber: Biodegradation of polystyrene. Biodegradation 2008, 19, 851–858. [Google Scholar] [CrossRef]
- Chen, X.; Li, X.; Li, Y. Toxicity inhibition strategy of microplastics to aquatic organisms through molecular docking, molecular dynamics simulation and molecular modification. Ecotoxicol. Environ. Saf. 2021, 226, 112870. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Saika, A.; Shinozaki, Y.; Watanabe, T.; Suzuki, K.; Sameshima-Yamashita, Y.; Fukuoka, T.; Habe, H.; Morita, T.; Kitamoto, H. Degradation profiles of biodegradable plastic films by biodegradable plastic-degrading enzymes from the yeast Pseudozyma antarctica and the fungus Paraphoma sp. B47-9. Polym. Degrad. Stab. 2017, 141, 26–32. [Google Scholar] [CrossRef]
- Kumari, R.; Kumar, R.; Open Source Drug Discovery, C.; Lynn, A. g_mmpbsa—A GROMACS tool for high-throughput MM-PBSA calculations. J. Chem. Inf. Model. 2014, 54, 1951–1962. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Nguyen, P.H.; Pham, K.; Huynh, D.; Le, T.-B.N.; Wang, H.; Ren, P.; Luo, R. Calculating protein-ligand binding affinities with MMPBSA: Method and error analysis. J. Comput. Chem. 2016, 37, 2436–2446. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takamatsu, Y.; Sugiyama, A.; Purqon, A.; Nagao, H.; Nishikawa, K. Binding Free Energy Calculation and Structural Analysis for Antigen-Antibody Complex. AIP Conf. Proc. 2006, 832, 566–569. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, S.; Xu, C.; Lin, M.; Li, Y.; Shen, C.; Liang, Y.; Sun, X.; Wang, D.; Lü, P.; et al. Epitopes prediction for microcystin-LR by molecular docking. Ecotoxicol. Environ. Saf. 2021, 227, 112925. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Gu, W.; Li, Y. Molecular design of 1,3,5,7-TetraCN derivatives with reduced bioconcentration using 3D-QSAR modeling, full factorial design, and molecular docking. J. Mol. Graph. Model. 2018, 84, 197–214. [Google Scholar] [CrossRef]
- Fidiani, E. Modeling of diatomic molecule using the Morse potential and the Verlet algorithm. AIP Conf. Proc. 2016, 1719, 30001. [Google Scholar] [CrossRef]
- Wang, X.; Chu, Z.; Yang, J.; Li, Y. Pentachlorophenol molecule design with lower bioconcentration through 3D-QSAR associated with molecule docking. Environ. Sci. Pollut. Res. 2017, 24, 25114–25125. [Google Scholar] [CrossRef]
- Kragh-Hansen, U.; Chuang, V.T.G.; Otagiri, M. Practical Aspects of the Ligand-Binding and Enzymatic Properties of Human Serum Albumin. Biol. Pharm. Bull. 2002, 25, 695–704. [Google Scholar] [CrossRef] [Green Version]
- Park, G.S.; Saleem, M. Diffusion of additives and plasticizers in poly(vinyl chloride)—I. The 1:2 double disc method for obtaining the diffusion coefficient of additives in polymers. J. Membr. Sci. 1984, 18, 177–185. [Google Scholar] [CrossRef]
- Ho, B.T.; Roberts, T.K.; Lucas, S. An overview on biodegradation of polystyrene and modified polystyrene: The microbial approach. Crit. Rev. Biotechnol. 2017, 38, 308–320. [Google Scholar] [CrossRef] [PubMed]
- Crompton, T. Identification of additives in polyolefins and polystyrenes. Eur. Polym. J. 1968, 4, 473–496. [Google Scholar] [CrossRef]
- Chen, Y.; Zhang, Y.; Zhang, Z. Occurrence, effects, and biodegradation of plastic additives in sludge anaerobic digestion: A review. Environ. Pollut. 2021, 287, 117568. [Google Scholar] [CrossRef]
- Guo, X.; Yao, Y.; Zhao, H.; Chi, C.; Zeng, F.; Qian, F.; Liu, Z.; Huo, L.; Lv, Y. Environmental impacts of functional fillers in polylactide (PLA)-based bottles using life cycle assessment methodology. Sci. Total Environ. 2021, 788, 147852. [Google Scholar] [CrossRef]
- Gu, W.; Zhao, Y.; Li, Q.; Li, Y. Plant-microorganism combined remediation of polychlorinated naphthalenes contaminated soils based on molecular directed transformation and Taguchi experimental design-assisted dynamics simulation. J. Hazard. Mater. 2020, 396, 122753. [Google Scholar] [CrossRef]
- Pablos, M.; Martínez-Fernández, J.; Sánchez, N.; González-Zamora, Á. Temporal and Spatial Comparison of Agricultural Drought Indices from Moderate Resolution Satellite Soil Moisture Data over Northwest Spain. Remote Sens. 2017, 9, 1168. [Google Scholar] [CrossRef] [Green Version]
- Yang, S.; DeMars, M.D., 2nd; Grandner, J.M.; Olson, N.M.; Anzai, Y.; Sherman, D.H.; Houk, K.N. Computational-Based Mechanistic Study and Engineering of Cytochrome P450 MycG for Selective Oxidation of 16-Membered Macrolide Antibiotics. J. Am. Chem. Soc. 2020, 142, 17981–17988. [Google Scholar] [CrossRef]
- Li, X.; Hou, Y.; Li, Q.; Gu, W.; Li, Y. Molecular design of high-efficacy and high drug safety Fluoroquinolones suitable for a variety of aerobic biodegradation bacteria. J. Environ. Manag. 2021, 299, 113628. [Google Scholar] [CrossRef]
- Lv, C.; Zhao, G.; Ning, Y. Interactions between plant proteins/enzymes and other food components, and their effects on food quality. Crit. Rev. Food Sci. Nutr. 2015, 57, 1718–1728. [Google Scholar] [CrossRef]
- Zhang, H.; Zhao, C.; Na, H. Theoretical Design of Biodegradable Phthalic Acid Ester Derivatives in Marine and Freshwater Environments. ChemistryOpen 2020, 9, 1033–1045. [Google Scholar] [CrossRef] [PubMed]
| Compositions | Chemicals | Molecular Formula | CAS Number |
|---|---|---|---|
| Plastic Matrix | Polystyrene | (C8H8)n, n = 5 | 100-42-5 |
| Plasticizers (A) | Diethylhexyl phthalate (1) | C24H38O4 | 117-81-7 |
| Diisononyl phthalate (2) | C26H42O4 | 28553-12-0 | |
| Antioxidants (B) | Acetone diphenylamine (1) | C15H17NO | 68412-48-6 |
| Nonyl phenol (2) | C15H24O | 25154-52-3 | |
| Light stabilizers (C) | 2-(2H-benzotriazol-2-yl)-4,6-di-tert-pentylphenol (1) | C22H29N3O | 25973-55-1 |
| Bumetrizole (2) | C17H18N3OCl | 3896-11-5 | |
| Heat stabilizers (D) | Calcium stearate (1) | C36H70O4Ca | 1592-23-0 |
| Zinc stearate (2) | C36H70O4Zn | 557-05-1 | |
| Flame retardants (E) | Decabromodiphenyl ether (1) | C12Br10O | 1163-19-5 |
| Tetrabromobisphenol A (2) | C15H12Br4O2 | 79-94-7 | |
| Lubricants (F) | Oleamide (1) | C18H35NO | 301-02-0 |
| Stearic acid (2) | C18H36O2 | 57-11-4 | |
| Fillers (G) | Calcium carbonate (1) | CaCO3 | 471-34-1 |
| Calcium sulfate (2) | CaSO4 | 10101-41-4 |
| Compositions | Chemicals | Molecular Formula | CAS Number |
|---|---|---|---|
| Plastic Matrix | Polystyrene | (C8H8)n, n = 5 | 100-42-5 |
| Plasticizers (A) | None (0) | - | - |
| Diisononyl phthalate (1) | C26H42O4 | 28553-12-0 | |
| Antioxidants (B) | None (0) | - | - |
| Acetone diphenylamine (1) | C15H17NO | 68412-48-6 | |
| Light stabilizers (C) | None (0) | - | - |
| Bumetrizole (1) | C17H18N3OCl | 3896-11-5 | |
| Heat stabilizers (D) | None (0) | - | - |
| Zinc stearate (1) | C36H70O4Zn | 557-05-1 | |
| Flame retardants (E) | None (0) | - | - |
| Tetrabromobisphenol A (1) | C15H12Br4O2 | 79-94-7 | |
| Lubricants (F) | None (0) | - | - |
| Oleamide (1) | C18H35NO | 301-02-0 |
| No. | Combination | Binding Energy (kJ/mol) | Change Rate (%) | No. | Combination | Binding Energy (kJ/mol) | Change Rate (%) |
|---|---|---|---|---|---|---|---|
| Blank Group | − | −100.374 | − | Blank Group | − | −100.374 | − |
| 1 | 2122121 | −165.347 | 64.73 | 65 | 1211212 | −127.231 | 26.76 |
| 2 | 1212111 | −159.899 | 59.30 | 66 | 2121112 | −139.953 | 39.43 |
| 3 | 2222111 | −171.493 | 70.85 | 67 | 1222212 | −146.936 | 46.39 |
| 4 | 2211222 | −149.510 | 48.95 | 68 | 1121221 | −110.175 | 9.76 |
| 5 | 1212121 | −149.627 | 49.07 | 69 | 2222212 | −181.704 | 81.03 |
| 6 | 2111211 | −176.630 | 75.97 | 70 | 2211112 | −149.459 | 48.90 |
| 7 | 1212211 | −189.748 | 89.04 | 71 | 2222112 | −182.239 | 81.56 |
| 8 | 1122122 | −115.771 | 15.34 | 72 | 2222222 | −188.257 | 87.56 |
| 9 | 1211121 | −165.639 | 65.02 | 73 | 2222211 | −147.114 | 46.57 |
| 10 | 2211122 | −116.508 | 16.07 | 74 | 2122222 | −118.484 | 18.04 |
| 11 | 2122211 | −167.863 | 67.24 | 75 | 1211111 | −129.242 | 28.76 |
| 12 | 2211211 | −168.668 | 68.04 | 76 | 2212121 | −101.028 | 0.65 |
| 13 | 2211212 | −139.039 | 38.52 | 77 | 2211121 | −139.678 | 39.16 |
| 14 | 2111122 | −133.583 | 33.09 | 78 | 1222121 | −135.564 | 35.06 |
| 15 | 1111122 | −146.259 | 45.71 | 79 | 2121121 | −166.372 | 65.75 |
| 16 | 1121222 | −142.824 | 42.29 | 80 | 1222211 | −168.897 | 68.27 |
| 17 | 2221221 | −149.945 | 49.39 | 81 | 1222222 | −179.741 | 79.07 |
| 18 | 1111211 | −186.012 | 85.32 | 82 | 2121122 | −139.483 | 38.96 |
| 19 | 2111212 | −140.396 | 39.87 | 83 | 1212122 | −153.210 | 52.64 |
| 20 | 1211222 | −122.226 | 21.77 | 84 | 1221212 | −174.317 | 73.67 |
| 21 | 1111112 | −159.221 | 58.63 | 85 | 1122211 | −175.672 | 75.02 |
| 22 | 2211221 | −176.404 | 75.75 | 86 | 2122221 | −164.832 | 64.22 |
| 23 | 1212221 | −162.598 | 61.99 | 87 | 1121122 | −143.276 | 42.74 |
| 24 | 1221112 | −133.649 | 33.15 | 88 | 1111111 | −168.066 | 67.44 |
| 25 | 1211211 | −146.415 | 45.87 | 89 | 2212211 | −145.248 | 44.71 |
| 26 | 2111222 | −104.090 | 3.70 | 90 | 2121212 | −108.537 | 8.13 |
| 27 | 1121212 | −162.534 | 61.93 | 91 | 1222111 | −126.167 | 25.70 |
| 28 | 1121211 | −124.222 | 23.76 | 92 | 1122111 | −162.668 | 62.06 |
| 29 | 2211111 | −185.857 | 85.16 | 93 | 1112211 | −156.718 | 56.13 |
| 30 | 2112211 | −151.042 | 50.48 | 94 | 1111222 | −130.514 | 30.03 |
| 31 | 2122111 | −156.230 | 55.65 | 95 | 1212222 | −185.417 | 84.73 |
| 32 | 2112112 | −144.408 | 43.87 | 96 | 1112122 | −172.772 | 72.13 |
| 33 | 1122212 | −171.231 | 70.59 | 97 | 2111221 | −153.231 | 52.66 |
| 34 | 1111212 | −160.521 | 59.92 | 98 | 1112212 | −137.189 | 36.68 |
| 35 | 1222221 | −80.943 | −19.36 | 99 | 2212111 | −173.947 | 73.30 |
| 36 | 1221222 | −133.329 | 32.83 | 100 | 1212212 | −166.364 | 65.74 |
| 37 | 1222122 | −126.188 | 25.72 | 101 | 1112111 | −179.134 | 78.47 |
| 38 | 2112122 | −143.624 | 43.09 | 102 | 2221112 | −160.316 | 59.72 |
| 39 | 2122122 | −174.420 | 73.77 | 103 | 1121111 | −156.163 | 55.58 |
| 40 | 2111121 | −117.572 | 17.13 | 104 | 1211112 | −162.505 | 61.90 |
| 41 | 2111111 | −176.795 | 76.14 | 105 | 2222122 | −133.804 | 33.31 |
| 42 | 2121222 | −149.533 | 48.98 | 106 | 1121121 | −135.199 | 34.70 |
| 43 | 1222112 | −163.450 | 62.84 | 107 | 1122112 | −187.302 | 86.60 |
| 44 | 2212122 | −140.908 | 40.38 | 108 | 2122112 | −122.819 | 22.36 |
| 45 | 2221212 | −143.580 | 43.05 | 109 | 2222221 | −101.376 | 1.00 |
| 46 | 1221221 | −129.625 | 29.14 | 110 | 1122221 | −169.289 | 68.66 |
| 47 | 2121211 | −175.918 | 75.26 | 111 | 2221122 | −172.425 | 71.78 |
| 48 | 2212221 | −172.734 | 72.09 | 112 | 2222121 | −157.271 | 56.68 |
| 49 | 2112212 | −163.466 | 62.86 | 113 | 2112111 | −159.225 | 58.63 |
| 50 | 1221121 | −171.508 | 70.87 | 114 | 1212112 | −151.288 | 50.72 |
| 51 | 1112221 | −180.381 | 79.71 | 115 | 2212222 | −175.982 | 75.33 |
| 52 | 2112221 | −164.344 | 63.73 | 116 | 2122212 | −192.483 | 91.77 |
| 53 | 2212212 | −155.909 | 55.33 | 117 | 2212112 | −163.769 | 63.16 |
| 54 | 1211122 | −157.855 | 57.27 | 118 | 1122222 | −158.070 | 57.48 |
| 55 | 2112121 | −174.158 | 73.51 | 119 | 1221211 | −160.681 | 60.08 |
| 56 | 1111221 | −120.981 | 20.53 | 120 | 1121112 | −170.986 | 70.35 |
| 57 | 1112121 | −151.498 | 50.93 | 121 | 1112222 | −124.770 | 24.31 |
| 58 | 2221121 | −186.537 | 85.84 | 122 | 2112222 | −163.022 | 62.41 |
| 59 | 2221111 | −106.280 | 5.88 | 123 | 1112112 | −138.551 | 38.03 |
| 60 | 1111121 | −104.089 | 3.70 | 124 | 1211221 | −159.955 | 59.36 |
| 61 | 1122121 | −132.955 | 32.46 | 125 | 1221111 | −168.355 | 67.73 |
| 62 | 2221222 | −84.160 | −16.15 | 126 | 2111112 | −104.542 | 4.15 |
| 63 | 2221211 | −172.479 | 71.84 | 127 | 2121221 | −159.279 | 58.69 |
| 64 | 2121111 | −151.320 | 50.76 | 128 | 1221122 | −161.112 | 60.51 |
| Plastic Composition | Binding Energy (kJ/mol) |
|---|---|
| (C8H8)n, n = 5 | −100.374 |
| Diisononyl phthalate | −183.324 |
| Acetone diphenylamine | −146.232 |
| Bumetrizole | −153.988 |
| Zinc stearate | −143.404 |
| Tetrabromobisphenol A | −164.449 |
| Oleamide | −186.741 |
| Calcium sulfate | −144.624 |
| No. | Combination | Binding Energy (kJ/mol) | Change Rate (%) | No. | Combination | Binding Energy (kJ/mol) | Change Rate (%) |
|---|---|---|---|---|---|---|---|
| No. 116-1 | 000111 | −173.410 | 72.76 | No. 116-33 | 100010 | −161.165 | 60.56 |
| No. 116-2 | 000110 | −184.535 | 83.85 | No. 116-34 | 001001 | −173.560 | 72.91 |
| No. 116-3 | 001110 | −148.113 | 47.56 | No. 116-35 | 110001 | −176.420 | 75.76 |
| No. 116-4 | 010110 | −146.021 | 45.48 | No. 116-36 | 110011 | −135.375 | 34.87 |
| No. 116-5 | 101011 | −158.323 | 57.73 | No. 116-37 | 010111 | −153.645 | 53.07 |
| No. 116-6 | 010100 | −112.337 | 11.92 | No. 116-38 | 011010 | −133.981 | 33.48 |
| No. 116-7 | 100111 | −165.617 | 65.00 | No. 116-39 | 000100 | −152.368 | 51.80 |
| No. 116-8 | 001010 | −163.996 | 63.38 | No. 116-40 | 110000 | −153.090 | 52.52 |
| No. 116-9 | 000101 | −163.734 | 63.12 | No. 116-41 | 101000 | −158.225 | 57.64 |
| No. 116-10 | 110111 | −162.456 | 61.85 | No. 116-42 | 011100 | −144.108 | 43.57 |
| No. 116-11 | 111000 | −134.082 | 33.58 | No. 116-43 | 101101 | −186.196 | 85.50 |
| No. 116-12 | 101111 | −159.050 | 58.46 | No. 116-44 | 001111 | −144.039 | 43.50 |
| No. 116-13 | 111111 | −175.752 | 75.10 | No. 116-45 | 101010 | −165.794 | 65.18 |
| No. 116-14 | 010000 | −139.483 | 38.96 | No. 116-46 | 110101 | −146.067 | 45.52 |
| No. 116-15 | 010001 | −150.620 | 50.06 | No. 116-47 | 111110 | −150.120 | 49.56 |
| No. 116-16 | 000000 | −100.374 | - | No. 116-48 | 010101 | −121.953 | 21.50 |
| No. 116-17 | 100110 | −160.875 | 60.28 | No. 116-49 | 111011 | −180.120 | 79.45 |
| No. 116-18 | 001100 | −178.409 | 77.74 | No. 116-50 | 110010 | −169.788 | 69.16 |
| No. 116-19 | 101100 | −127.301 | 26.83 | No. 116-51 | 111010 | −140.016 | 39.49 |
| No. 116-20 | 011000 | −166.234 | 65.61 | No. 116-52 | 111001 | −153.551 | 52.98 |
| No. 116-21 | 001101 | −156.825 | 56.24 | No. 116-53 | 011111 | −151.653 | 51.09 |
| No. 116-22 | 111101 | −161.370 | 60.77 | No. 116-54 | 011110 | −164.654 | 64.04 |
| No. 116-23 | 011001 | −139.879 | 39.36 | No. 116-55 | 110100 | −157.329 | 56.74 |
| No. 116-24 | 101001 | −157.547 | 56.96 | No. 116-56 | 100100 | −157.288 | 56.70 |
| No. 116-25 | 011011 | −149.624 | 49.07 | No. 116-57 | 001000 | −160.944 | 60.34 |
| No. 116-26 | 100001 | −159.186 | 58.59 | No. 116-58 | 110110 | −152.739 | 52.17 |
| No. 116-27 | 001011 | −137.040 | 36.53 | No. 116-59 | 000011 | −143.545 | 43.01 |
| No. 116-28 | 100000 | −112.544 | 12.12 | No. 116-60 | 100101 | −163.517 | 62.91 |
| No. 116-29 | 000010 | −156.999 | 56.41 | No. 116-61 | 010010 | −170.14 | 69.51 |
| No. 116-30 | 010011 | −149.525 | 48.97 | No. 116-62 | 011101 | −119.86 | 19.41 |
| No. 116-31 | 000001 | −124.866 | 24.40 | No. 116-63 | 101110 | −164.943 | 64.33 |
| No. 116-32 | 100011 | −161.298 | 60.70 | No. 116-64 | 111100 | −182.603 | 81.92 |
| Contribution Rate | Percentage (%) |
|---|---|
| Main effect | 20.68 |
| Second-order interaction effects | 25.16 |
| third-order interaction effects | 54.16 |
| Total | 100 |
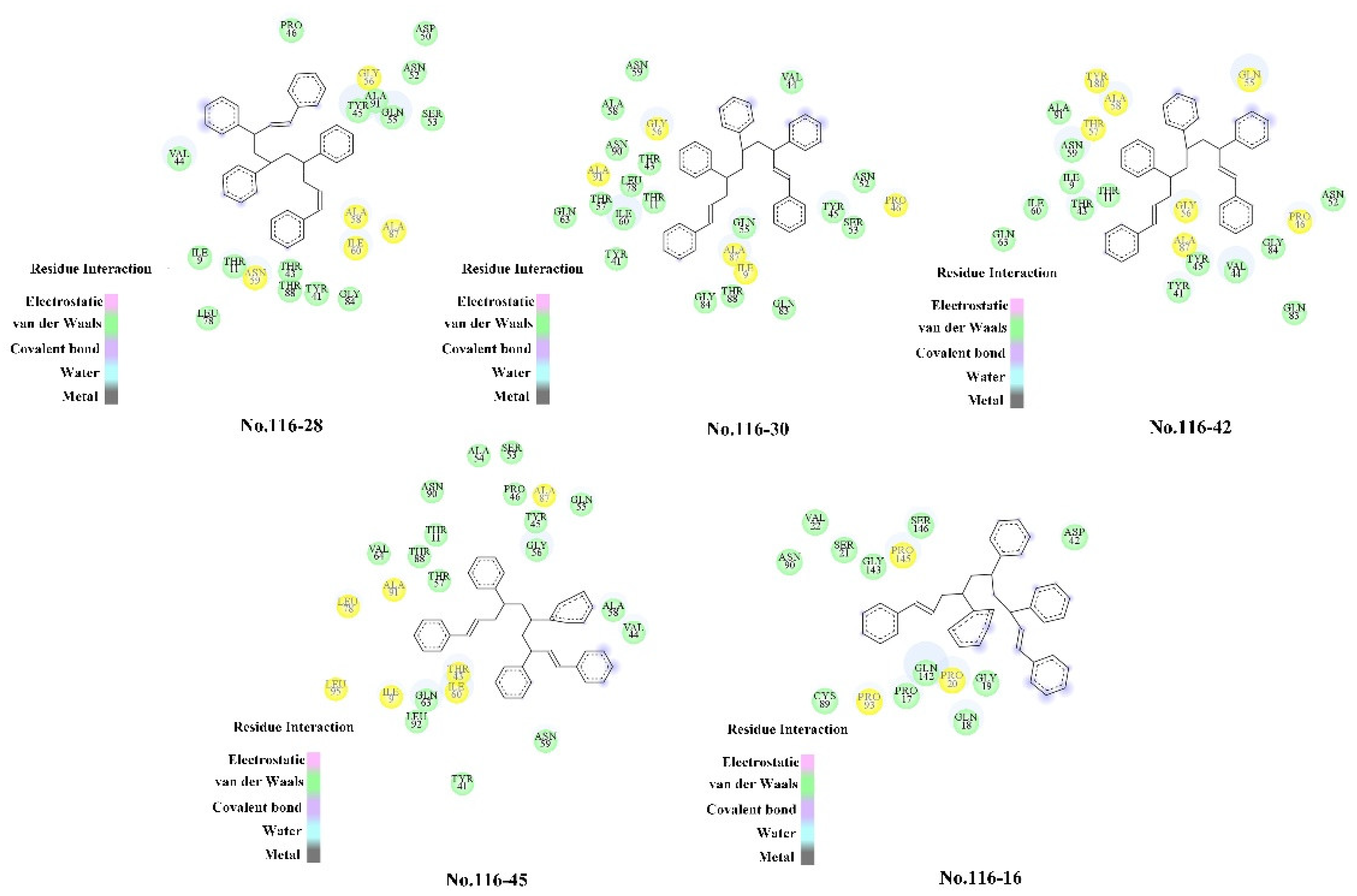
| Groups | Combination | No Bonded Interaction | Interaction Amino Acids | Number |
|---|---|---|---|---|
| No. 116–16 | None | Mixed Pi/Alkyl Hydrophobic | PRO145, PRO93, PRO20 | 4 |
| No. 116–42 | B*C*D | Pi-Alkyl Hydrophobic Mixed Pi/Alkyl Hydrophobic | ALA87, PRO46, GLN55, GLY56, THR57, ALA58, TYR180 | 3 4 |
| No. 116–28 | A | Hydrogen Bonds (no classical) Pi-Alkyl Hydrophobic | GLY56, ALA58, ALA87, ILE60, ASN59 | 1 |
| 5 | ||||
| No. 116–30 | B*E*F | Hydrogen Bonds Mixed Pi/Alkyl Hydrophobic | GLY56, ALA91, ALA87, ILE9, PRO46 | 1 |
| 5 | ||||
| No. 116–45 | A*C*E | Mixed Pi/Alkyl Hydrophobic | ALA87, ALA91, LEU78, LEU95, ILE9, THR43, ILE60 | 8 |
| Groups | PAE Substitutes | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| DBP-CHO | DBP-COOH | DBP-OH | DINP-NH2 | DINP-NO2 | ||||||
| Change Rate (%) | Change Rate (%) | Change Rate (%) | Change Rate (%) | Change Rate (%) | ||||||
| No. 116–28 (A) | −181.290 | 80.61% | −123.126 | 22.67% | −132.944 | 32.45% | −151.873 | 51.31% | −167.071 | 66.45% |
| No. 116–40 (A*B) | −159.726 | 59.13% | −157.740 | 57.15% | −175.469 | 74.82% | −139.738 | 39.22% | −156.185 | 55.60% |
| No. 116–26 (A*F) | −186.207 | 85.51% | −164.083 | 63.47% | −166.870 | 66.25% | −155.496 | 54.92% | −146.213 | 45.67% |
| No. 116–45 (A*C*E) | −153.546 | 52.97% | −138.699 | 38.18% | −163.177 | 62.57% | −134.371 | 33.87% | −161.306 | 60.70% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Hou, Y.; Zhao, W.; Na, H. Control Strategies of Plastic Biodegradation through Adjusting Additives Ratios Using In Silico Approaches Associated with Proportional Factorial Experimental Design. Int. J. Environ. Res. Public Health 2022, 19, 5670. https://doi.org/10.3390/ijerph19095670
Zhang H, Hou Y, Zhao W, Na H. Control Strategies of Plastic Biodegradation through Adjusting Additives Ratios Using In Silico Approaches Associated with Proportional Factorial Experimental Design. International Journal of Environmental Research and Public Health. 2022; 19(9):5670. https://doi.org/10.3390/ijerph19095670
Chicago/Turabian StyleZhang, Haigang, Yilin Hou, Wenjin Zhao, and Hui Na. 2022. "Control Strategies of Plastic Biodegradation through Adjusting Additives Ratios Using In Silico Approaches Associated with Proportional Factorial Experimental Design" International Journal of Environmental Research and Public Health 19, no. 9: 5670. https://doi.org/10.3390/ijerph19095670






