Automatic Text-Mining Approach to Identify Molecular Target Candidates Associated with Metabolic Processes for Myotonic Dystrophy Type 1
Abstract
:1. Introduction
2. Material and Methods
2.1. Search Approach
2.2. Automatic Text-Mining Analysis Using VOSviewer
2.3. DisGeNET Search
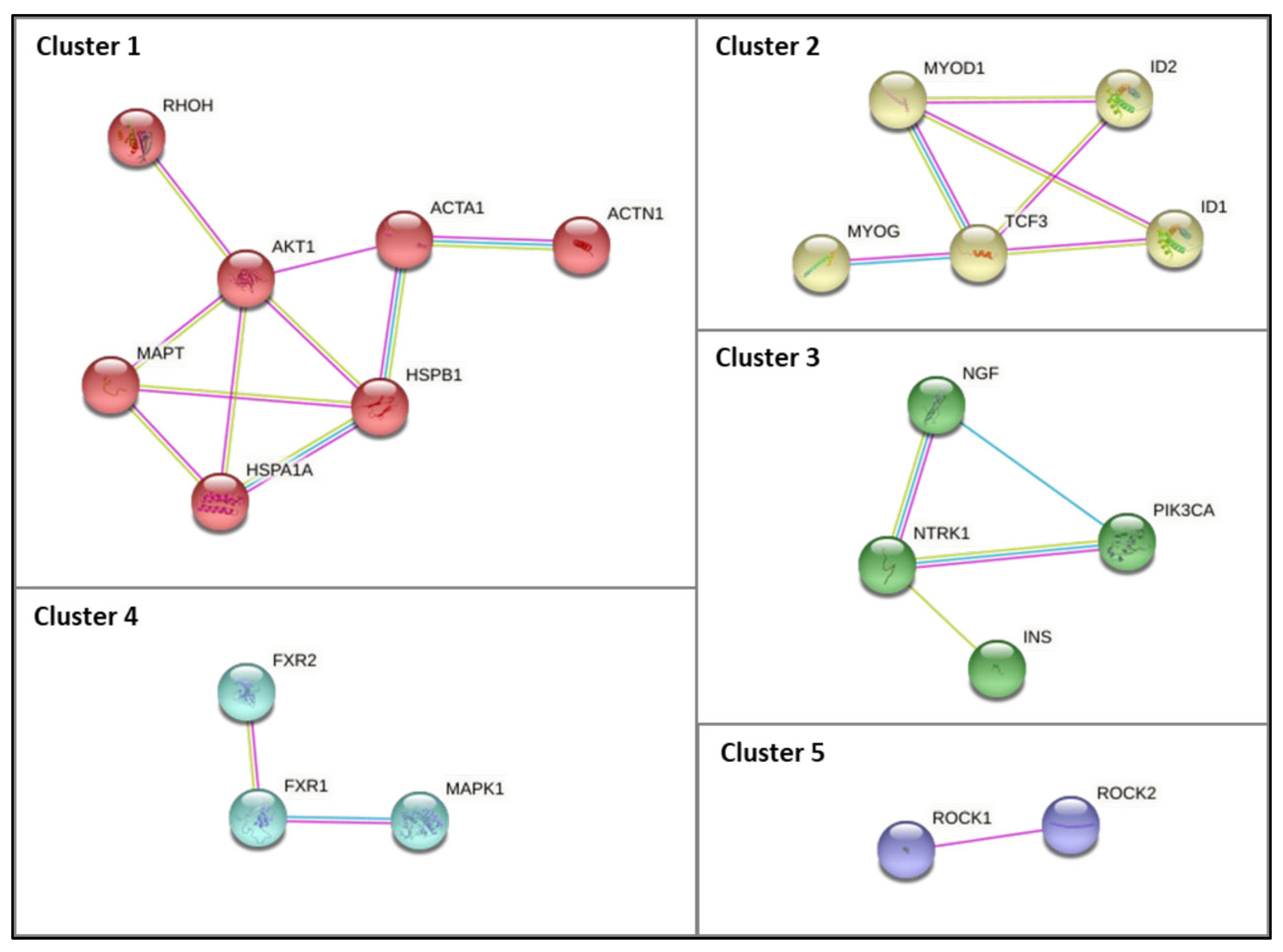
2.4. Protein–Protein Interaction Network Construction
2.5. Gene Ontology Functional and Pathway Enrichment Analyses
3. Results
3.1. Literature Search and Automatic Text-Mining Analysis to Unveil Novel Metabolism Molecular Targets in DM1
3.2. Comparative Analysis of the Novel Identified Metabolism-Associated Molecular Targets in DM1 Using VOSviewer with Molecular Associations Previously Described in DisGeNET
3.3. Characterization and Functional Enrichment Analysis of the Novel Identified Metabolism Associated Molecular Targets in DM1
4. Discussion
4.1. Dysregulation of Biological Processes and Signaling Pathways Associated with DM1
4.1.1. Muscle Function and Associated Pathways
4.1.2. NGF and TRK Signaling Pathways
4.1.3. Insulin Signaling Pathways
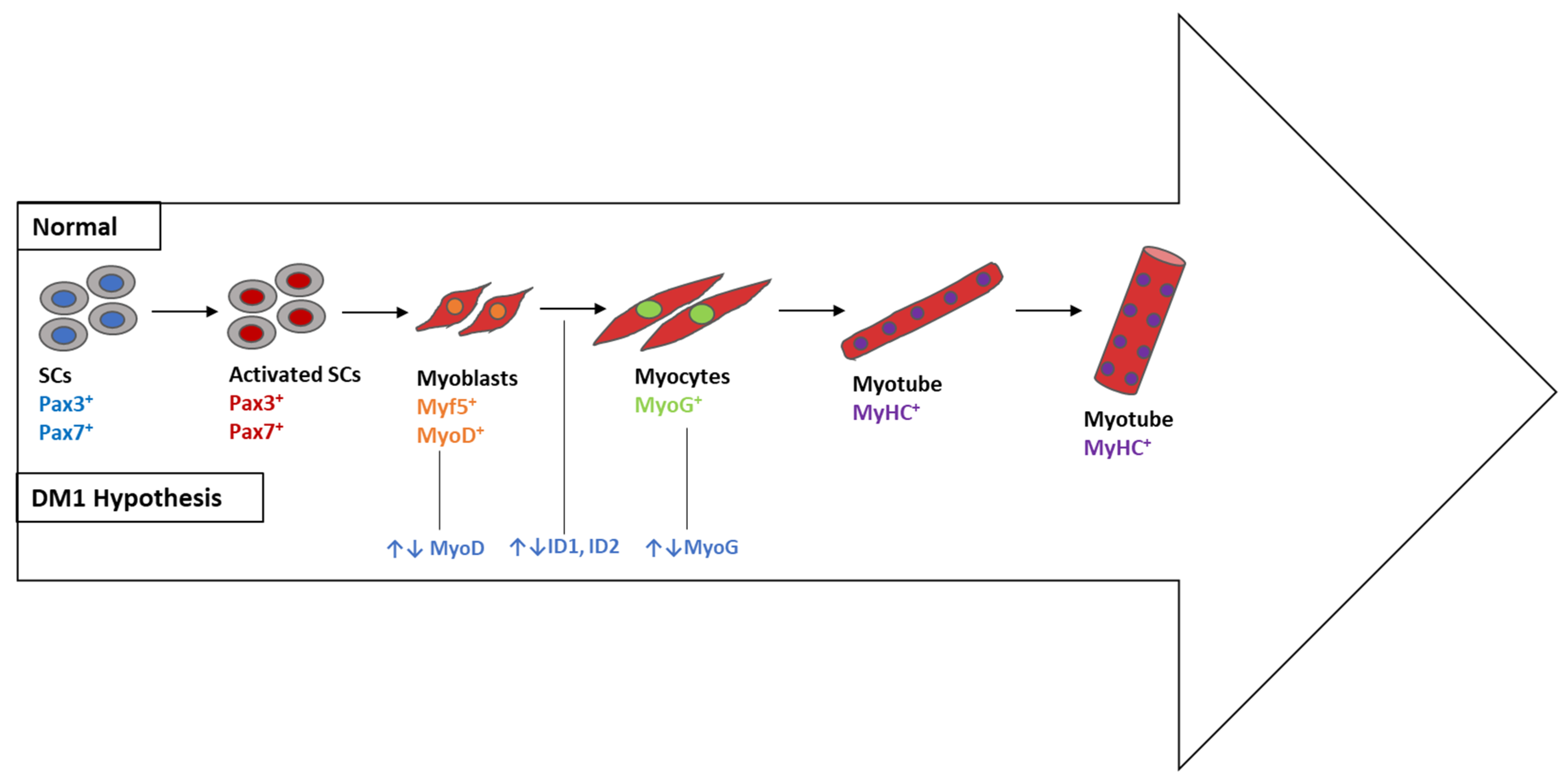
4.1.4. Myogenesis- and Signaling-Associated Pathways
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siciliano, G.; Manca, M.L.; Gennarelli, M.; Angelini, C.; Rocchi, A.; Iudice, A.; Miorin, M.; Mostacciuolo, M.L. Epidemiology of Myotonic Dystrophy in Italy: Re-Apprisal after Genetic Diagnosis. Clin. Genet. 2001, 59, 344–349. [Google Scholar] [CrossRef] [PubMed]
- Johnson, N.E.; Butterfield, R.J.; Mayne, K.; Newcomb, T.; Imburgia, C.; Dunn, D.; Duval, B.; Feldkamp, M.L.; Weiss, R.B. Population-Based Prevalence of Myotonic Dystrophy Type 1 Using Genetic Analysis of Statewide Blood Screening Program. Neurology 2021, 96, e1045–e1053. [Google Scholar] [CrossRef] [PubMed]
- Johnson, N.E.; Aldana, E.Z.; Angeard, N.; Ashizawa, T.; Berggren, K.N.; Marini-Bettolo, C.; Duong, T.; Ekström, A.B.; Sansone, V.; Tian, C.; et al. Consensus-Based Care Recommendations for Congenital and Childhood-Onset Myotonic Dystrophy Type 1. Neurol. Clin. Pract. 2019, 9, 443–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aslanidis, C.; Jansen, G.; Amemiya, C.; Shutler, G.; Mahadevan, M.; Tsilfidis, C.; Chen, C.; Alleman, J.; Wormskamp, N.G.M.; Vooijs, M.; et al. Cloning of the Essential Myotonic Dystrophy Region and Mapping of the Putative Defect. Nature 1992, 355, 548–551. [Google Scholar] [CrossRef]
- Fu, Y.H.; Pizzuti, A.; Fenwick, R.G.; King, J.; Rajnarayan, S.; Dunne, P.W.; Dubel, J.; Nasser, G.A.; Ashizawa, T.; De Jong, P.; et al. An Unstable Triplet Repeat in a Gene Related to Myotonic Muscular Dystrophy. Science 1992, 255, 1256–1258. [Google Scholar] [CrossRef]
- Yum, K.; Wang, E.T.; Kalsotra, A. Myotonic Dystrophy: Disease Repeat Range, Penetrance, Age of Onset, and Relationship between Repeat Size and Phenotypes. Curr. Opin. Genet. Dev. 2017, 44, 30–37. [Google Scholar] [CrossRef]
- De Antonio, M.; Dogan, C.; Hamroun, D.; Mati, M.; Zerrouki, S.; Eymard, B.; Katsahian, S.; Bassez, G. Unravelling the Myotonic Dystrophy Type 1 Clinical Spectrum: A Systematic Registry-Based Study with Implications for Disease Classification. Rev. Neurol. 2016, 172, 572–580. [Google Scholar] [CrossRef]
- López-Otín, C.; Galluzzi, L.; Freije, J.M.P.; Madeo, F.; Kroemer, G. Metabolic Control of Longevity. Cell 2016, 166, 802–821. [Google Scholar] [CrossRef] [Green Version]
- Mattson, M.P.; Arumugam, T.V. Hallmarks of Brain Aging: Adaptive and Pathological Modification by Metabolic States. Cell Metab. 2018, 27, 1176–1199. [Google Scholar] [CrossRef] [Green Version]
- Mateos-Aierdi, A.J.; Goicoechea, M.; Aiastui, A.; Fernández-Torrón, R.; Garcia-Puga, M.; Matheu, A.; de Munain, A. Muscle Wasting in Myotonic Dystrophies: A Model of Premature Aging. Front. Aging Neurosci. 2015, 7, 125. [Google Scholar] [CrossRef]
- Turner, C.; Hilton-Jones, D. Myotonic Dystrophy: Diagnosis, Management and New Therapies. Curr. Opin. Neurol. 2014, 27, 599–606. [Google Scholar] [CrossRef]
- Lee, J.E.; Cooper, T.A. Pathogenic Mechanisms of Myotonic Dystrophy. Biochem. Soc. Trans. 2009, 37, 1281–1286. [Google Scholar] [CrossRef] [PubMed]
- Udd, B.; Krahe, R. The Myotonic Dystrophies: Molecular, Clinical, and Therapeutic Challenges. Lancet Neurol. 2012, 11, 891–905. [Google Scholar] [CrossRef] [PubMed]
- Koutsoulidou, A.; Koutalianos, D.; Georgiou, K.; Kakouri, A.C.; Oulas, A.; Tomazou, M.; Kyriakides, T.C.; Roos, A.; Papadimas, G.K.; Papadopoulos, C.; et al. Serum MiRNAs as Biomarkers for the Rare Types of Muscular Dystrophy. Neuromuscul. Disord. 2022, 32, 332–346. [Google Scholar] [CrossRef] [PubMed]
- Koutsoulidou, A.; Photiades, M.; Kyriakides, T.C.; Georgiou, K.; Prokopi, M.; Kapnisis, K.; Lusakowska, A.; Nearchou, M.; Christou, Y.; Papadimas, G.K.; et al. Identification of Exosomal Muscle-Specific MiRNAs in Serum of Myotonic Dystrophy Patients Relating to Muscle Disease Progress. Hum. Mol. Genet. 2017, 26, 3285–3302. [Google Scholar] [CrossRef] [Green Version]
- Koehorst, E.; Ballester-lopez, A.; Arechavala-gomeza, V.; Martínez-piñeiro, A.; Nogales-gadea, G. The Biomarker Potential of Mirnas in Myotonic Dystrophy Type I. J. Clin. Med. 2020, 9, 3939. [Google Scholar] [CrossRef]
- Pegoraro, V.; Cudia, P.; Baba, A.; Angelini, C. MyomiRNAs and Myostatin as Physical Rehabilitation Biomarkers for Myotonic Dystrophy. Neurol. Sci. 2020, 41, 2953–2960. [Google Scholar] [CrossRef]
- Espinosa-Espinosa, J.; González-Barriga, A.; López-Castel, A.; Artero, R. Deciphering the Complex Molecular Pathogenesis of Myotonic Dystrophy Type 1 through Omics Studies. Int. J. Mol. Sci. 2022, 23, 1441. [Google Scholar] [CrossRef]
- García-Puga, M.; Saenz-Antoñanzas, A.; Fernández-Torrón, R.; de Munain, A.L.; Matheu, A. Myotonic Dystrophy Type 1 Cells Display Impaired Metabolism and Mitochondrial Dysfunction That Are Reversed by Metformin. Aging 2020, 12, 6260–6275. [Google Scholar] [CrossRef]
- García-Puga, M.; Saenz-Antoñanzas, A.; Matheu, A.; de Munain, A. Targeting Myotonic Dystrophy Type 1 with Metformin. Int. J. Mol. Sci. 2022, 23, 2901. [Google Scholar] [CrossRef]
- Burnham, J.F. Scopus Database: A Review. Biomed. Digit. Libr. 2006, 3, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Falagas, M.E.; Pitsouni, E.I.; Malietzis, G.A.; Pappas, G. Comparison of PubMed, Scopus, Web of Science, and Google Scholar: Strengths and Weaknesses. FASEB J. 2008, 22, 338–342. [Google Scholar] [CrossRef] [PubMed]
- Van Eck, N.J.; Waltman, L. Software Survey: VOSviewer, a Computer Program for Bibliometric Mapping. Scientometrics 2010, 84, 523–538. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Callon, M.; Courtial, J.P.; Turner, W.A.; Bauin, S. From Translations to Problematic Networks: An Introduction to Co-Word Analysis. Soc. Sci. Inf. 1983, 22, 191–235. [Google Scholar] [CrossRef]
- Börner, K.; Chen, C.; Boyack, K.W. Visualizing Knowledge Domains. Annu. Rev. Inf. Sci. Technol. 2003, 37, 179–255. [Google Scholar] [CrossRef]
- Bateman, A. UniProt: A Worldwide Hub of Protein Knowledge. Nucleic Acids Res. 2019, 47, D506–D515. [Google Scholar] [CrossRef] [Green Version]
- Piñero, J.; Ramírez-Anguita, J.M.; Saüch-Pitarch, J.; Ronzano, F.; Centeno, E.; Sanz, F.; Furlong, L.I. The DisGeNET Knowledge Platform for Disease Genomics: 2019 Update. Nucleic Acids Res. 2020, 48, D845–D855. [Google Scholar] [CrossRef] [Green Version]
- Piñero, J.; Saüch, J.; Sanz, F.; Furlong, L.I. The DisGeNET Cytoscape App: Exploring and Visualizing Disease Genomics Data. Comput. Struct. Biotechnol. J. 2021, 19, 2960–2967. [Google Scholar] [CrossRef]
- Bardou, P.; Mariette, J.; Escudié, F.; Djemiel, C.; Klopp, C. Jvenn: An Interactive Venn Diagram Viewer. BMC Bioinform. 2014, 15, 293. [Google Scholar] [CrossRef] [Green Version]
- Snel, B.; Lehmann, G.; Bork, P.; Huynen, M.A. String: A Web-Server to Retrieve and Display the Repeatedly Occurring Neighbourhood of a Gene. Nucleic Acids Res. 2000, 28, 3442–3444. [Google Scholar] [CrossRef] [Green Version]
- Reimand, J.; Arak, T.; Adler, P.; Kolberg, L.; Reisberg, S.; Peterson, H.; Vilo, J. G:Profiler—A Web Server for Functional Interpretation of Gene Lists (2016 Update). Nucleic Acids Res. 2016, 44, W83–W89. [Google Scholar] [CrossRef] [PubMed]
- Silla, T.; Schmid, M.; Dou, Y.; Garland, W.; Milek, M.; Imami, K.; Johnsen, D.; Polak, P.; Andersen, J.S.; Selbach, M.; et al. The Human ZC3H3 and RBM26/27 Proteins Are Critical for PAXT-Mediated Nuclear RNA Decay. Nucleic Acids Res. 2020, 48, 2518–2530. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, J.; Tan, L.; Yin, Z.; Zhu, W.; Tao, K.; Wang, G.; Shi, W.; Gao, J. MIR17HG-MiR-18a/19a Axis, Regulated by Interferon Regulatory Factor-1, Promotes Gastric Cancer Metastasis via Wnt/β-Catenin Signalling. Cell Death Dis. 2019, 10, 454. [Google Scholar] [CrossRef] [PubMed]
- Nieuwenhuis, S.; Okkersen, K.; Widomska, J.; Blom, P.; ’t Hoen, P.A.C.; van Engelen, B.; Glennon, J.C. Insulin Signaling as a Key Moderator in Myotonic Dystrophy Type 1. Front. Neurol. 2019, 10, 1229. [Google Scholar] [CrossRef]
- Vihola, A.; Bachinski, L.L.; Sirito, M.; Olufemi, S.-E.; Hajibashi, S.; Baggerly, K.A.; Raheem, O.; Haapasalo, H.; Suominen, T.; Holmlund-Hampf, J.; et al. Differences in Aberrant Expression and Splicing of Sarcomeric Proteins in the Myotonic Dystrophies DM1 and DM2. Acta Neuropathol. 2010, 119, 465–479. [Google Scholar] [CrossRef] [Green Version]
- Pantic, B.; Trevisan, E.; Citta, A.; Rigobello, M.P.; Marin, O.; Bernardi, P.; Salvatori, S.; Rasola, A. Myotonic Dystrophy Protein Kinase (DMPK) Prevents ROS-Induced Cell Death by Assembling a Hexokinase II-Src Complex on the Mitochondrial Surface. Cell Death Dis. 2013, 4, e858. [Google Scholar] [CrossRef] [Green Version]
- Shi, D.-L.; Grifone, R. RNA-Binding Proteins in the Post-Transcriptional Control of Skeletal Muscle Development, Regeneration and Disease. Front. Cell Dev. Biol. 2021, 9, 738978. [Google Scholar] [CrossRef]
- Laing, N.G.; Dye, D.E.; Wallgren-Pettersson, C.; Richard, G.; Monnier, N.; Lillis, S.; Winder, T.L.; Lochmüller, H.; Graziano, C.; Mitrani-Rosenbaum, S.; et al. Mutations and Polymorphisms of the Skeletal Muscle α-Actin Gene (ACTA1). Hum. Mutat. 2009, 30, 1267–1277. [Google Scholar] [CrossRef] [Green Version]
- Velcicky, J.; Bodendorf, U.; Rigollier, P.; Epple, R.; Beisner, D.R.; Guerini, D.; Smith, P.; Liu, B.; Feifel, R.; Wipfli, P.; et al. Discovery of the First Potent, Selective, and Orally Bioavailable Signal Peptide Peptidase-Like 2a (SPPL2a) Inhibitor Displaying Pronounced Immunomodulatory Effects In Vivo. J. Med. Chem. 2018, 61, 865–880. [Google Scholar] [CrossRef] [Green Version]
- Zhang, T.; Cheng, G.; Deng, L.; Yang, Y.; Sun, L.; Chen, P.; He, X.; Su, D.; Bi, N.; Qiu, B. Silence of S1 RNA Binding Domain 1 Represses Cell Growth and Promotes Apoptosis in Human Non-Small Cell Lung Cancer Cells. Transl. Lung Cancer Res. 2019, 8, 760. [Google Scholar] [CrossRef]
- Hintze, S.; Knaier, L.; Limmer, S.; Schoser, B.; Meinke, P. Nuclear Envelope Transmembrane Proteins in Myotonic Dystrophy Type. Front. Physiol. 2018, 9, 1532. [Google Scholar] [CrossRef] [Green Version]
- Rübben, A.; Wahl, R.U.; Eggermann, T.; Dahl, E.; Ortiz-Brüchle, N.; Cacchi, C. Mutation Analysis of Multiple Pilomatricomas in a Patient with Myotonic Dystrophy Type 1 Suggests a DM1-Associated Hypermutation Phenotype. PLoS ONE 2020, 15, e0230003. [Google Scholar] [CrossRef] [Green Version]
- Shi, J.; Wu, X.; Surma, M.; Vemula, S.; Zhang, L.; Yang, Y.; Kapur, R.; Wei, L. Distinct Roles for ROCK1 and ROCK2 in the Regulation of Cell Detachment. Cell Death Dis. 2013, 4, e483. [Google Scholar] [CrossRef] [Green Version]
- Sun, L.; Trausch-Azar, J.S.; Ciechanover, A.; Schwartz, A.L. Ubiquitin-Proteasome-Mediated Degradation, Intracellular Localization, and Protein Synthesis of MyoD and Id1 during Muscle Differentiation. J. Biol. Chem. 2005, 280, 26448–26456. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoshida, H.; Goedert, M. Phosphorylation of Microtubule-Associated Protein Tau by AMPK-Related Kinases. J. Neurochem. 2012, 120, 165–176. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.S.; Bedrin, N.G.; Ades, P.A.; Palmer, B.M.; Toth, M.J. Molecular Determinants of Force Production in Human Skeletal Muscle Fibers: Effects of Myosin Isoform Expression and Cross-Sectional Area. Am. J. Physiol. Cell Physiol. 2015, 308, C473–C484. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smith, J.A.; Curry, E.G.; Blue, R.E.; Roden, C.; Dundon, S.E.R.; Rodríguez-Vargas, A.; Jordan, D.C.; Chen, X.; Lyons, S.M.; Crutchley, J.; et al. FXR1 Splicing Is Important for Muscle Development and Biomolecular Condensates in Muscle Cells. J. Cell Biol. 2020, 219, e201911129. [Google Scholar] [CrossRef] [Green Version]
- Shelton, D.L.; Reichardt, L.F. Studies on the Expression of the Beta Nerve Growth Factor (NGF) Gene in the Central Nervous System: Level and Regional Distribution of NGF MRNA Suggest That NGF Functions as a Trophic Factor for Several Distinct Populations of Neurons. Proc. Natl. Acad. Sci. USA 1986, 83, 2714–2718. [Google Scholar] [CrossRef] [Green Version]
- Harding, H.P.; Lackey, J.G.; Hsu, H.-C.; Zhang, Y.; Deng, J.; Xu, R.-M.; Damha, M.J.; Ron, D. An Intact Unfolded Protein Response in Trpt1 Knockout Mice Reveals Phylogenic Divergence in Pathways for RNA Ligation. RNA 2008, 14, 225–232. [Google Scholar] [CrossRef] [Green Version]
- Wei, X.W.; Liu, X.Q.; Zhang, Y.C.; Qin, C.M.; Lin, Y.; Tian, F.J. TCF3 Regulates Human Endometrial Stromal Cell Proliferation and Migration in RPL. Reproduction 2022, 163, 281–291. [Google Scholar] [CrossRef]
- Xu, F.; Na, L.; Li, Y.; Chen, L. RETRACTED ARTICLE: Roles of the PI3K/AKT/MTOR Signalling Pathways in Neurodegenerative Diseases and Tumours. Cell Biosci. 2020, 10, 54. [Google Scholar] [CrossRef] [Green Version]
- Bourque, J.; Kousnetsov, R.; Hawiger, D. Roles of Hopx in the Differentiation and Functions of Immune Cells. Eur. J. Cell Biol. 2022, 101, 151242. [Google Scholar] [CrossRef] [PubMed]
- Xia, Z.; Zhang, X.; Liu, P.; Zhang, R.; Huang, Z.; Li, D.; Xiao, X.; Wu, M.; Ning, N.; Zhang, Q.; et al. GNA13 Regulates BCL2 Expression and the Sensitivity of GCB-DLBCL Cells to BCL2 Inhibitors in a Palmitoylation-Dependent Manner. Cell Death Dis. 2021, 12, 54. [Google Scholar] [CrossRef]
- Huang, X.; Liu, N.; Xiong, X. ZNF24 Is Upregulated in Prostate Cancer and Facilitates the Epithelial-to-mesenchymal Transition through the Regulation of Twist1. Oncol. Lett. 2020, 19, 3593–3601. [Google Scholar] [CrossRef] [PubMed]
- Aavani, P.; Allen, L.J.S. The Role of CD4 T Cells in Immune System Activation and Viral Reproduction in a Simple Model for HIV Infection. Appl. Math. Model. 2019, 75, 210–222. [Google Scholar] [CrossRef]
- Gobbi, G.; Galli, D.; Carubbi, C.; Neri, L.M.; Masselli, E.; Pozzi, G.; Vitale, M.; Mirandola, P. PKC Proteins and Muscular Dystrophy. J. Funct. Morphol. Kinesiol. 2018, 3, 12. [Google Scholar] [CrossRef] [Green Version]
- Mateus, T.; Martins, F.; Nunes, A.; Herdeiro, M.T.; Rebelo, S. Metabolic Alterations in Myotonic Dystrophy Type 1 and Their Correlation with Lipin. Int. J. Environ. Res. Public. Health 2021, 18, 1794. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.D.; Harley, M.E.; Kemp, A.J.; Wills, J.; Lee, M.; Arends, M.; von Kriegsheim, A.; Behrends, C.; Wilkinson, S. CCPG1 Is a Non-Canonical Autophagy Cargo Receptor Essential for ER-Phagy and Pancreatic ER Proteostasis. Dev. Cell 2018, 44, 217–232. [Google Scholar] [CrossRef] [Green Version]
- Huang, Y.; Xing, Y.; Li, H. Heterozygous Desmin Gene (DES) Mutation Contributes to Familial Dilated Cardiomyopathy. J. Int. Med. Res. 2021, 49, 03000605211006598. [Google Scholar] [CrossRef]
- Mosaddeghzadeh, N.; Ahmadian, M.R. The RHO Family GTPases: Mechanisms of Regulation and Signaling. Cells 2021, 10, 1831. [Google Scholar] [CrossRef]
- Zhao, Z.; Manser, E. Myotonic Dystrophy Kinase-Related Cdc42-Binding Kinases (MRCK), the ROCK-like Effectors of Cdc42 and Rac1. Small GTPases 2015, 6, 81–88. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Gong, H.; Wang, P.; Zhu, Y.; Peng, H.; Cui, Y.; Li, H.; Liu, J.; Wang, Z. The Emerging Role of ISWI Chromatin Remodeling Complexes in Cancer. J. Exp. Clin. Cancer Res. 2021, 40, 346. [Google Scholar] [CrossRef]
- Blum, R.; Dynlacht, B.D. The Role of MyoD1 and Histone Modifications in the Activation of Muscle Enhancers. Epigenetics 2013, 8, 778–784. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shan, Z.; Shakoori, A.; Bodaghi, S.; Goldsmith, P.; Jin, J.; Wiest, J.S. TUSC1, a Putative Tumor Suppressor Gene, Reduces Tumor Cell Growth In Vitro and Tumor Growth In Vivo. PLoS ONE 2013, 8, e66114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hasan, N.; Ray, T.A.; Gregg, R.G. CACNA1S Expression in Mouse Retina: Novel Isoforms and Antibody Cross-Reactivity with GPR179. Vis. Neurosci. 2016, 33, E009. [Google Scholar] [CrossRef] [PubMed]
- Faralli, H.; Dilworth, F.J. Turning on Myogenin in Muscle: A Paradigm for Understanding Mechanisms of Tissue-Specific Gene Expression. Comp. Funct. Genomics 2012, 2012, 836374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bedford, L.; Lowe, J.; Dick, L.R.; Mayer, R.J.; Brownell, J.E. Ubiquitin-like Protein Conjugation and the Ubiquitin–Proteasome System as Drug Targets. Nat. Rev. Drug Discov. 2011, 10, 29–46. [Google Scholar] [CrossRef]
- Kästle, M.; Grune, T. Chapter 4—Interactions of the Proteasomal System with Chaperones: Protein Triage and Protein Quality Control. In The Proteasomal System in Aging and Disease; Grune, T., Ed.; Progress in Molecular Biology and Translational Science; Academic Press: Cambridge, MA, USA, 2012; Volume 109, pp. 113–160. [Google Scholar]
- André, L.M.; Ausems, C.R.M.; Wansink, D.G.; Wieringa, B. Abnormalities in Skeletal Muscle Myogenesis, Growth, and Regeneration in Myotonic Dystrophy. Front. Neurol. 2018, 9, 368. [Google Scholar] [CrossRef] [Green Version]
- Zhu, S.; Kuek, V.; Bennett, S.; Xu, H.; Rosen, V.; Xu, J. Protein Cytl1: Its Role in Chondrogenesis, Cartilage Homeostasis, and Disease. Cell. Mol. Life Sci. 2019, 76, 3515–3523. [Google Scholar] [CrossRef] [Green Version]
- Simoncini, C.; Spadoni, G.; Lai, E.; Santoni, L.; Angelini, C.; Ricci, G.; Siciliano, G. Central Nervous System Involvement as Outcome Measure for Clinical Trials Efficacy in Myotonic Dystrophy Type 1. Front. Neurol. 2020, 11, 624. [Google Scholar] [CrossRef]
- Sen, S.; Dong, M.; Kumar, S. Isoform-Specific Contributions of α-Actinin to Glioma Cell Mechanobiology. PLoS ONE 2009, 4, e8427. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Du, J.; Zhu, Z.; Xu, L.; Chen, X.; Li, X.; Lan, T.; Li, W.; Yuan, K.; Zeng, Y. ARHGEF11 Promotes Proliferation and Epithelial-Mesenchymal Transition of Hepatocellular Carcinoma through Activation of β-Catenin Pathway. Aging 2020, 12, 20235–20253. [Google Scholar] [CrossRef]
- Fusco, N.; Malapelle, U.; Fassan, M.; Marchiò, C.; Buglioni, S.; Zupo, S.; Criscitiello, C.; Vigneri, P.; Dei Tos, A.P.; Maiorano, E.; et al. PIK3CA Mutations as a Molecular Target for Hormone Receptor-Positive, HER2-Negative Metastatic Breast Cancer. Front. Oncol. 2021, 11, 644737. [Google Scholar] [CrossRef]
- Jiang, Y.; Wu, S.-Y.; Chen, Y.-L.; Zhang, Z.-M.; Tao, Y.-F.; Xie, Y.; Liao, X.-M.; Li, X.-L.; Li, G.; Wu, D.; et al. CEBPG Promotes Acute Myeloid Leukemia Progression by Enhancing EIF4EBP1. Cancer Cell Int. 2021, 21, 598. [Google Scholar] [CrossRef] [PubMed]
- Kunitomi, H.; Kobayashi, Y.; Wu, R.-C.; Takeda, T.; Tominaga, E.; Banno, K.; Aoki, D. LAMC1 Is a Prognostic Factor and a Potential Therapeutic Target in Endometrial Cancer. J. Gynecol. Oncol. 2020, 31, e11. [Google Scholar] [CrossRef] [PubMed]
- Zou, D.; Qin, B.; Wang, J.; Shi, Y.; Zhou, P.; Yi, Y.; Liao, J.; Lu, X. AFF2 Is Associated With X-Linked Partial (Focal) Epilepsy with Antecedent Febrile Seizures. Front. Mol. Neurosci. 2022, 15, 795840. [Google Scholar] [CrossRef]
- Souidi, A.; Zmojdzian, M.; Jagla, K. Dissecting Pathogenetic Mechanisms and Therapeutic Strategies in Drosophila Models of Myotonic Dystrophy Type 1. Int. J. Mol. Sci. 2018, 19, 4104. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.H.; Yang, C.K.; Heo, K.; Roeder, R.G.; An, W.; Stallcup, M.R. CCAR1, a Key Regulator of Mediator Complex Recruitment to Nuclear Receptor Transcription Complexes. Mol. Cell 2008, 31, 510–519. [Google Scholar] [CrossRef] [Green Version]
- McAlear, T.S.; Bechstedt, S. The Mitotic Spindle Protein CKAP2 Potently Increases Formation and Stability of Microtubules. eLife 2022, 11, e72202. [Google Scholar] [CrossRef]
- Zhang, Y.; O’Connor, J.P.; Siomi, M.C.; Srinivasan, S.; Dutra, A.; Nussbaum, R.L.; Dreyfuss, G. The Fragile X Mental Retardation Syndrome Protein Interacts with Novel Homologs FXR1 and FXR2. EMBO J. 1995, 14, 5358–5366. [Google Scholar] [CrossRef]
- Takahashi, S.; Andreoletti, G.; Chen, R.; Munehira, Y.; Batra, A.; Afzal, N.A.; Beattie, R.M.; Bernstein, J.A.; Ennis, S.; Snyder, M. De Novo and Rare Mutations in the HSPA1L Heat Shock Gene Associated with Inflammatory Bowel Disease. Genome Med. 2017, 9, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Geuens, T.; De Winter, V.; Rajan, N.; Achsel, T.; Mateiu, L.; Almeida-Souza, L.; Asselbergh, B.; Bouhy, D.; Auer-Grumbach, M.; Bagni, C.; et al. Mutant HSPB1 Causes Loss of Translational Repression by Binding to PCBP1, an RNA Binding Protein with a Possible Role in Neurodegenerative Disease. Acta Neuropathol. Commun. 2017, 5, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Navarro, M.; Valentinis, B.; Belletti, B.; Romano, G.; Reiss, K.; Baserga, R. Regulation of Id2 Gene Expression by the Type 1 IGF Receptor and the Insulin Receptor Substrate-1. Endocrinology 2001, 142, 5149–5157. [Google Scholar] [CrossRef] [PubMed]
- Elmas, M.; Gogus, B.; Solak, M. Understanding What You Have Found: A Family with a Mutation in the LAMA1 Gene with Literature Review. Clin. Med. Insights Case Rep. 2020, 13, 1179547620948666. [Google Scholar] [CrossRef]
- Ran, T.; Chen, Z.; Zhao, L.; Ran, W.; Fan, J.; Hong, S.; Yang, Z. LAMB1 Is Related to the T Stage and Indicates Poor Prognosis in Gastric Cancer. Technol. Cancer Res. Treat. 2021, 20, 15330338211004944. [Google Scholar] [CrossRef]
- Campbell, C.S.; Hombauer, H.; Srivatsan, A.; Bowen, N.; Gries, K.; Desai, A.; Putnam, C.D.; Kolodner, R.D. Mlh2 Is an Accessory Factor for DNA Mismatch Repair in Saccharomyces Cerevisiae. PLoS Genet. 2014, 10, e1004327. [Google Scholar] [CrossRef]
- Franc, D.T.; Muetzel, R.L.; Robinson, P.R.; Rodriguez, C.P.; Dalton, J.C.; Naughton, C.E.; Mueller, B.A.; Wozniak, J.R.; Lim, K.O.; Day, J.W. Cerebral and Muscle MRI Abnormalities in Myotonic Dystrophy. Neuromuscul. Disord. 2012, 22, 483–491. [Google Scholar] [CrossRef] [Green Version]
- Telese, R.; Pagliarani, S.; Lerario, A.; Ciscato, P.; Fagiolari, G.; Cassandrini, D.; Grimoldi, N.; Conte, G.; Cinnante, C.; Santorelli, F.M.; et al. MYH2 Myopathy, a New Case Expands the Clinical and Pathological Spectrum of the Recessive Form. Mol. Genet. Genom. Med. 2020, 8, e1320. [Google Scholar] [CrossRef] [PubMed]
- Amatu, A.; Sartore-Bianchi, A.; Bencardino, K.; Pizzutilo, E.G.; Tosi, F.; Siena, S. Tropomyosin Receptor Kinase (TRK) Biology and the Role of NTRK Gene Fusions in Cancer. Ann. Oncol. 2019, 30, viii5. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.; Su, Z.; Li, J.; Wang, Q.; Meng, G.; Zhang, Y.; Yang, W.; Zhang, J.; Gao, P. Role of RNA-binding Protein 5 in the Diagnosis and Chemotherapeutic Response of Lung Cancer (Review). Oncol. Lett. 2019, 17, 2013–2019. [Google Scholar] [CrossRef]
- Zhao, X.; McKillop-Smith, S.; Müller, B. The Human Histone Gene Expression Regulator HBP/SLBP Is Required for Histone and DNA Synthesis, Cell Cycle Progression and Cell Proliferation in Mitotic Cells. J. Cell Sci. 2004, 117, 6043–6051. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loaiza, N.; Hartgers, M.L.; Reeskamp, L.F.; Balder, J.-W.; Rimbert, A.; Bazioti, V.; Wolters, J.C.; Winkelmeijer, M.; Jansen, H.P.G.; Dallinga-Thie, G.M.; et al. Taking One Step Back in Familial Hypercholesterolemia. Arterioscler. Thromb. Vasc. Biol. 2020, 40, 973–985. [Google Scholar] [CrossRef] [PubMed]
- Kitai, Y.; Iwakami, M.; Saitoh, K.; Togi, S.; Isayama, S.; Sekine, Y.; Muromoto, R.; Kashiwakura, J.; Yoshimura, A.; Oritani, K.; et al. STAP-2 Protein Promotes Prostate Cancer Growth by Enhancing Epidermal Growth Factor Receptor Stabilization. J. Biol. Chem. 2017, 292, 19392–19399. [Google Scholar] [CrossRef] [Green Version]
- Bisikirska, B.C.; Adam, S.J.; Alvarez, M.J.; Rajbhandari, P.; Cox, R.; Lefebvre, C.; Wang, K.; Rieckhof, G.E.; Felsher, D.W.; Califano, A. STK38 Is a Critical Upstream Regulator of MYC’s Oncogenic Activity in Human B-Cell Lymphoma. Oncogene 2013, 32, 5283–5291. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu-Monette, Z.Y.; Medeiros, L.J.; Li, Y.; Orlowski, R.Z.; Andreeff, M.; Bueso-Ramos, C.E.; Greiner, T.C.; McDonnell, T.J.; Young, K.H. Dysfunction of the TP53 Tumor Suppressor Gene in Lymphoid Malignancies. Blood 2012, 119, 3668–3683. [Google Scholar] [CrossRef] [Green Version]
- Rasmussen, M.; Jin, J.-P. Troponin Variants as Markers of Skeletal Muscle Health and Diseases. Front. Physiol. 2021, 12, 1623. [Google Scholar] [CrossRef]
- Bosè, F.; Renna, L.V.; Fossati, B.; Arpa, G.; Labate, V.; Milani, V.; Botta, A.; Micaglio, E.; Meola, G.; Cardani, R. TNNT2 Missplicing in Skeletal Muscle as a Cardiac Biomarker in Myotonic Dystrophy Type 1 but Not in Myotonic Dystrophy Type 2. Front. Neurol. 2019, 10, 992. [Google Scholar] [CrossRef]
- Rhodes, J.D.; Lott, M.C.; Russell, S.L.; Moulton, V.; Sanderson, J.; Wormstone, I.M.; Broadway, D.C. Activation of the Innate Immune Response and Interferon Signalling in Myotonic Dystrophy Type 1 and Type 2 Cataracts. Hum. Mol. Genet. 2012, 21, 852–862. [Google Scholar] [CrossRef] [Green Version]
- Hale, M.A.; Richardson, J.I.; Day, R.C.; McConnell, O.L.; Arboleda, J.; Wang, E.T.; Berglund, J.A. An Engineered RNA Binding Protein with Improved Splicing Regulation. Nucleic Acids Res. 2018, 46, 3152–3168. [Google Scholar] [CrossRef] [Green Version]
- Bae, J.-H.; Jeong, H.-J.; Kim, H.; Leem, Y.-E.; Ryu, D.; Park, S.C.; Lee, Y.-I.; Cho, S.C.; Kang, J.-S. ZNF746/PARIS Overexpression Induces Cellular Senescence through FoxO1/P21 Axis Activation in Myoblasts. Cell Death Dis. 2020, 11, 359. [Google Scholar] [CrossRef]
- Esposito, F.; Cè, E.; Rampichini, S.; Monti, E.; Limonta, E.; Fossati, B.; Meola, G. Electromechanical Delays during a Fatiguing Exercise and Recovery in Patients with Myotonic Dystrophy Type 1. Eur. J. Appl. Physiol. 2017, 117, 551–566. [Google Scholar] [CrossRef]
- Johnson, N.E. Myotonic Muscular Dystrophies. Contin. Lifelong Learn. Neurol. 2019, 25, 1682–1695. [Google Scholar] [CrossRef] [PubMed]
- Kalsotra, A.; Xiao, X.; Ward, A.J.; Castle, J.C.; Johnson, J.M.; Burge, C.B.; Cooper, T.A. A Postnatal Switch of CELF and MBNL Proteins Reprograms Alternative Splicing in the Developing Heart. Proc. Natl. Acad. Sci. USA 2008, 105, 20333–20338. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Overby, S.J.; Cerro-Herreros, E.; Llamusi, B.; Artero, R. RNA-Mediated Therapies in Myotonic Dystrophy. Drug Discov. Today 2018, 23, 2013–2022. [Google Scholar] [CrossRef]
- Ozimski, L.L.; Sabater-Arcis, M.; Bargiela, A.; Artero, R. The Hallmarks of Myotonic Dystrophy Type 1 Muscle Dysfunction. Biol. Rev. 2021, 96, 716–730. [Google Scholar] [CrossRef] [PubMed]
- Cauteruccio, F.; Stamile, C.; Terracina, G.; Ursino, D.; Sappey-Marinier, D. An Automated String-Based Approach to Extracting and Characterizing White Matter Fiber-Bundles. Comput. Biol. Med. 2016, 77, 64–75. [Google Scholar] [CrossRef]
- Thakur, S.S.; Swiderski, K.; Ryall, J.G.; Lynch, G.S. Therapeutic Potential of Heat Shock Protein Induction for Muscular Dystrophy and Other Muscle Wasting Conditions. Philos. Trans. R. Soc. B Biol. Sci. 2017, 373, 20160528. [Google Scholar] [CrossRef] [Green Version]
- Kampinga, H.H.; Hageman, J.; Vos, M.J.; Kubota, H.; Tanguay, R.M.; Bruford, E.A.; Cheetham, M.E.; Chen, B.; Hightower, L.E. Guidelines for the Nomenclature of the Human Heat Shock Proteins. Cell Stress Chaperones 2009, 14, 105–111. [Google Scholar] [CrossRef] [Green Version]
- Senf, S.M.; Dodd, S.L.; Judge, A.R. FOXO Signaling Is Required for Disuse Muscle Atrophy and Is Directly Regulated by Hsp70. Am. J. Physiol. Cell Physiol. 2010, 298, C38–C45. [Google Scholar] [CrossRef] [Green Version]
- Senf, S. Skeletal Muscle Heat Shock Protein 70: Diverse Functions and Therapeutic Potential for Wasting Disorders. Front. Physiol. 2013, 4, 330. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.-B.; Venkatraman, V.; Crowgey, E.L.; Liu, T.; Fu, Z.; Holewinski, R.; Ranek, M.; Kass, D.A.; O’Rourke, B.; Van Eyk, J.E. Protein S-Nitrosylation Controls Glycogen Synthase Kinase 3β Function Independent of Its Phosphorylation State. Circ. Res. 2018, 122, 1517–1531. [Google Scholar] [CrossRef]
- Wei, C.; Stock, L.; Valanejad, L.; Zalewski, Z.A.; Karns, R.; Puymirat, J.; Nelson, D.; Witte, D.; Woodgett, J.; Timchenko, N.A.; et al. Correction of GSK3ß at Young Age Prevents Muscle Pathology in Mice with Myotonic Dystrophy Type 1. FASEB J. 2018, 32, 2073–2085. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grande, V.; Hathazi, D.; O’Connor, E.; Marteau, T.; Schara-Schmidt, U.; Hentschel, A.; Gourdon, G.; Nikolenko, N.; Lochmüller, H.; Roos, A. Dysregulation of GSK3β-Target Proteins in Skin Fibroblasts of Myotonic Dystrophy Type 1 (DM1) Patients. J. Neuromuscul. Dis. 2021, 8, 603–619. [Google Scholar] [CrossRef] [PubMed]
- Mei, W.; Wen-Chin, W.; Lauren, S.; Diana, L.; Ana, M.; Genevieve, G.; Nikolai, T.; Mike, S.; Lubov, T. Correction of Glycogen Synthase Kinase 3β in Myotonic Dystrophy 1 Reduces the Mutant RNA and Improves Postnatal Survival of DMSXL Mice. Mol. Cell. Biol. 2019, 39, e00155-19. [Google Scholar] [CrossRef] [Green Version]
- Mirzoev, T.M.; Sharlo, K.A.; Shenkman, B.S. The Role of GSK-3β in the Regulation of Protein Turnover, Myosin Phenotype, and Oxidative Capacity in Skeletal Muscle under Disuse Conditions. Int. J. Mol. Sci. 2021, 22, 5081. [Google Scholar] [CrossRef] [PubMed]
- Huang, E.J.; Reichardt, L.F. Neurotrophins: Roles in Neuronal Development and Function. Annu. Rev. Neurosci. 2001, 24, 677–736. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Westphalen, C.B.; Krebs, M.G.; Le Tourneau, C.; Sokol, E.S.; Maund, S.L.; Wilson, T.R.; Jin, D.X.; Newberg, J.Y.; Fabrizio, D.; Veronese, L.; et al. Genomic Context of NTRK1/2/3 Fusion-Positive Tumours from a Large Real-World Population. NPJ Precis. Oncol. 2021, 5, 69. [Google Scholar] [CrossRef]
- Kaplan, D.R.; Martin-Zanca, D.; Parada, L.F. Tyrosine Phosphorylation and Tyrosine Kinase Activity of the Trk Proto-Oncogene Product Induced by NGF. Nature 1991, 350, 158–160. [Google Scholar] [CrossRef]
- Arévalo, J.C.; Wu, S.H. Neurotrophin Signaling: Many Exciting Surprises! Cell. Mol. Life Sci. CMLS 2006, 63, 1523–1537. [Google Scholar] [CrossRef]
- Fujisawa, K.; Fujita, A.; Ishizaki, T.; Saito, Y.; Narumiya, S. Identification of the Rho-Binding Domain of P160ROCK, a Rho-Associated Coiled-Coil Containing Protein Kinase. J. Biol. Chem. 1996, 271, 23022–23028. [Google Scholar] [CrossRef] [Green Version]
- Sahai, E.; Ishizaki, T.; Narumiya, S.; Treisman, R. Transformation Mediated by RhoA Requires Activity of ROCK Kinases. Curr. Biol. 1999, 9, 136–145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sahai, E.; Olson, M.F.; Marshall, C.J. Cross-Talk between Ras and Rho Signalling Pathways in Transformation Favours Proliferation and Increased Motility. EMBO J. 2001, 20, 755–766. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ehrenreiter, K.; Piazzolla, D.; Velamoor, V.; Sobczak, I.; Small, J.V.; Takeda, J.; Leung, T.; Baccarini, M. Raf-1 Regulates Rho Signaling and Cell Migration. J. Cell Biol. 2005, 168, 955–964. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amatu, A.; Sartore-Bianchi, A.; Siena, S. NTRK Gene Fusions as Novel Targets of Cancer Therapy across Multiple Tumour Types. ESMO Open 2016, 1, e000023. [Google Scholar] [CrossRef] [Green Version]
- Zou, J.; Lei, T.; Guo, P.; Yu, J.; Xu, Q.; Luo, Y.; Ke, R.; Huang, D. Mechanisms Shaping the Role of ERK1/2 in Cellular Senescence (Review). Mol. Med. Rep. 2018, 19, 759–770. [Google Scholar] [CrossRef] [Green Version]
- Beffy, P.; Del Carratore, R.; Masini, M.; Furling, D.; Puymirat, J.; Masiello, P.; Simili, M. Altered Signal Transduction Pathways and Induction of Autophagy in Human Myotonic Dystrophy Type 1 Myoblasts. Int. J. Biochem. Cell Biol. 2010, 42, 1973–1983. [Google Scholar] [CrossRef]
- Gadalla, S.M.; Lund, M.; Pfeiffer, R.M.; Gørtz, S.; Mueller, C.M.; Moxley, R.T.; Kristinsson, S.Y.; Björkholm, M.; Shebl, F.M.; Hilbert, J.E.; et al. Cancer Risk Among Patients with Myotonic Muscular Dystrophy. JAMA 2011, 306, 2480–2486. [Google Scholar] [CrossRef] [Green Version]
- Win, A.K.; Perattur, P.G.; Pulido, J.S.; Pulido, C.M.; Lindor, N.M. Increased Cancer Risks in Myotonic Dystrophy. Mayo Clin. Proc. 2012, 87, 130–135. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alsaggaf, R.; St. George, D.M.M.; Zhan, M.; Pfeiffer, R.M.; Wang, Y.; Wagner, K.R.; Greene, M.H.; Amr, S.; Gadalla, S.M. Cancer Risk in Myotonic Dystrophy Type I: Evidence of a Role for Disease Severity. JNCI Cancer Spectr. 2018, 2, pky052. [Google Scholar] [CrossRef] [Green Version]
- Roskoski, R. ERK1/2 MAP Kinases: Structure, Function, and Regulation. Pharmacol. Res. 2012, 66, 105–143. [Google Scholar] [CrossRef]
- Mao, Z.; Zhang, W. Role of MTOR in Glucose and Lipid Metabolism. Int. J. Mol. Sci. 2018, 19, 2043. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Renna, L.V.; Bosè, F.; Brigonzi, E.; Fossati, B.; Meola, G.; Cardani, R. Aberrant Insulin Receptor Expression Is Associated with Insulin Resistance and Skeletal Muscle Atrophy in Myotonic Dystrophies. PLoS ONE 2019, 14, e0214254. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chevrel, G.; Hohlfeld, R.; Sendtner, M. The Role of Neurotrophins in Muscle under Physiological and Pathological Conditions. Muscle Nerve 2006, 33, 462–476. [Google Scholar] [CrossRef]
- Deponti, D.; Buono, R.; Catanzaro, G.; De Palma, C.; Longhi, R.; Meneveri, R.; Bresolin, N.; Bassi, M.T.; Cossu, G.; Clementi, E.; et al. The Low-Affinity Receptor for Neurotrophins P75NTR Plays a Key Role for Satellite Cell Function in Muscle Repair Acting via RhoA. Mol. Biol. Cell 2009, 20, 3620–3627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Romagnoli, C.; Iantomasi, T.; Brandi, M.L. Available In Vitro Models for Human Satellite Cells from Skeletal Muscle. Int. J. Mol. Sci. 2021, 22, 13221. [Google Scholar] [CrossRef]
- Esteves de Lima, J.; Relaix, F. Master Regulators of Skeletal Muscle Lineage Development and Pluripotent Stem Cells Differentiation. Cell Regen. 2021, 10, 31. [Google Scholar] [CrossRef] [PubMed]
- Mourikis, P.; Sambasivan, R.; Castel, D.; Rocheteau, P.; Bizzarro, V.; Tajbakhsh, S. A Critical Requirement for Notch Signaling in Maintenance of the Quiescent Skeletal Muscle Stem Cell State. Stem Cells 2012, 30, 243–252. [Google Scholar] [CrossRef]
- Lahmann, I.; Bröhl, D.; Zyrianova, T.; Isomura, A.; Czajkowski, M.T.; Kapoor, V.; Griger, J.; Ruffault, P.-L.; Mademtzoglou, D.; Zammit, P.S.; et al. Oscillations of MyoD and Hes1 Proteins Regulate the Maintenance of Activated Muscle Stem Cells. Genes Dev. 2019, 33, 524–535. [Google Scholar] [CrossRef] [Green Version]
- Borok, M.J.; Mademtzoglou, D.; Relaix, F. Bu-M-P-Ing Iron: How BMP Signaling Regulates Muscle Growth and Regeneration. J. Dev. Biol. 2020, 8, 4. [Google Scholar] [CrossRef] [Green Version]
- Friedrichs, M.; Wirsdöerfer, F.; Flohé, S.B.; Schneider, S.; Wuelling, M.; Vortkamp, A. BMP Signaling Balances Proliferation and Differentiation of Muscle Satellite Cell Descendants. BMC Cell Biol. 2011, 12, 26. [Google Scholar] [CrossRef] [Green Version]
- Ono, Y.; Calhabeu, F.; Morgan, J.E.; Katagiri, T.; Amthor, H.; Zammit, P.S. BMP Signalling Permits Population Expansion by Preventing Premature Myogenic Differentiation in Muscle Satellite Cells. Cell Death Differ. 2010, 18, 222–234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, J.S.L.; Cui, W. Proliferation, Survival and Metabolism: The Role of PI3K/AKT/MTOR Signalling in Pluripotency and Cell Fate Determination. Development 2016, 143, 3050–3060. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huichalaf, C.H.; Sakai, K.; Wang, G.-L.; Timchenko, N.A.; Timchenko, L. Regulation of the Promoter of CUG Triplet Repeat Binding Protein, Cugbp1, during Myogenesis. Gene 2007, 396, 391–402. [Google Scholar] [CrossRef] [PubMed]

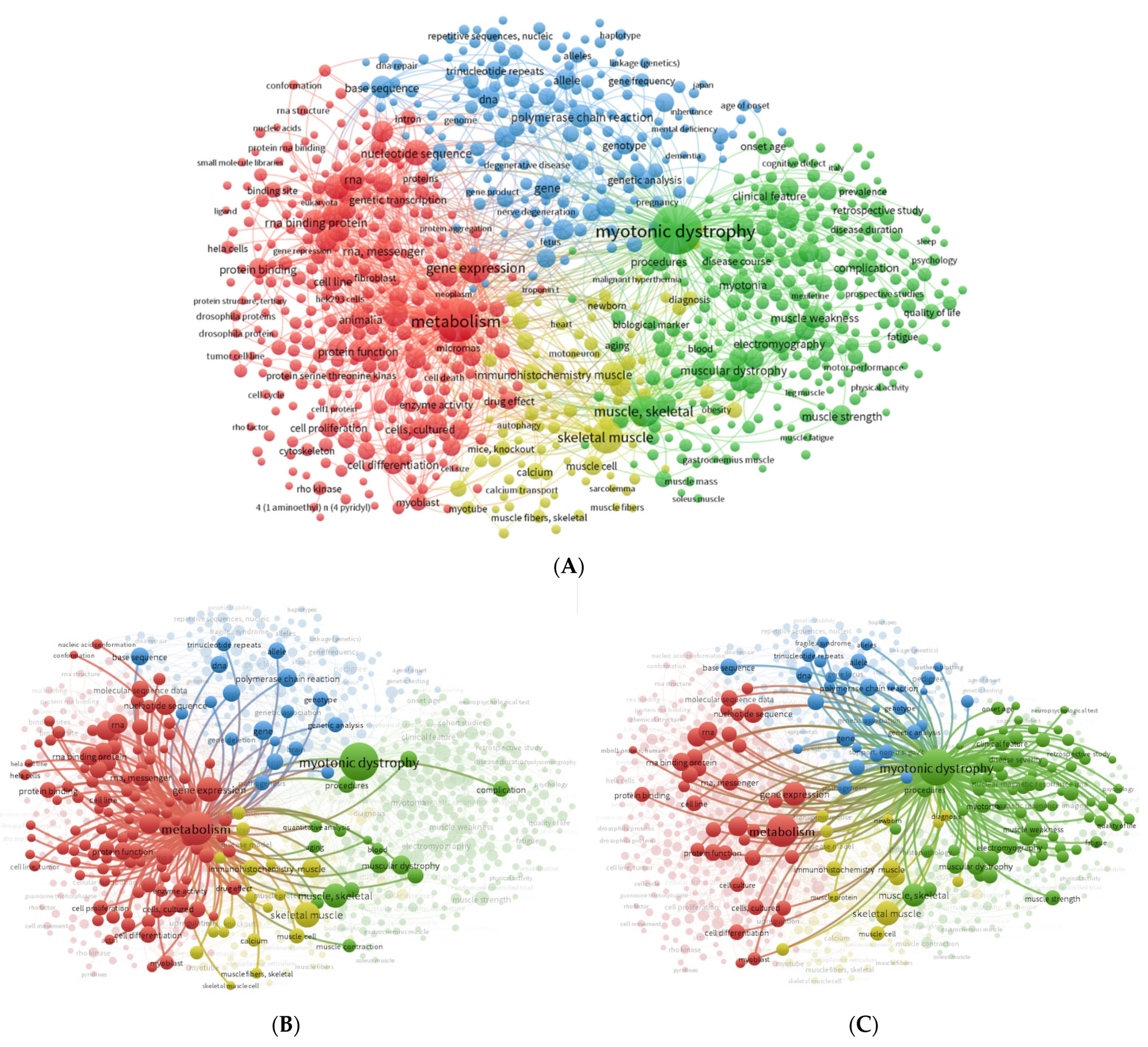
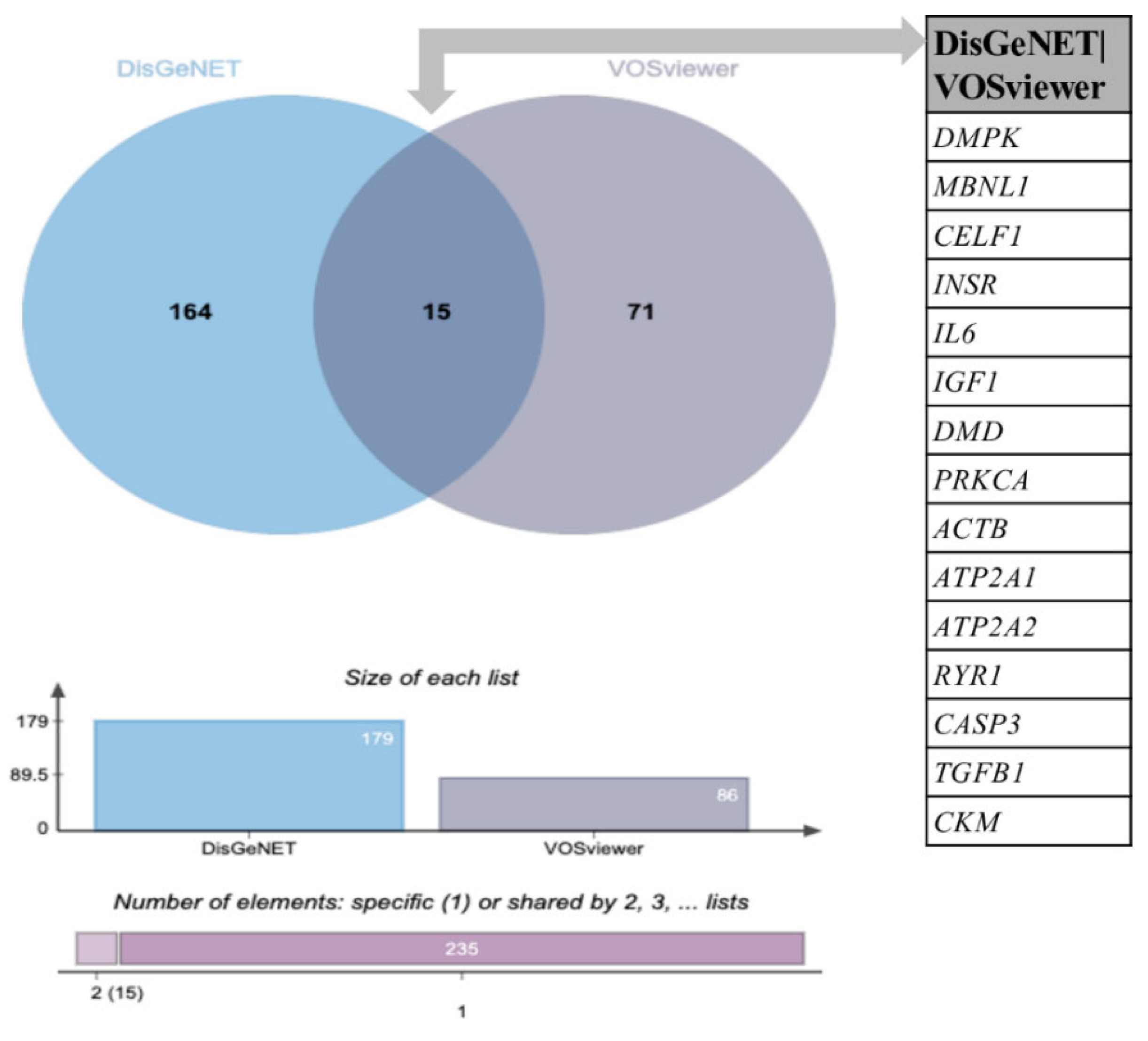
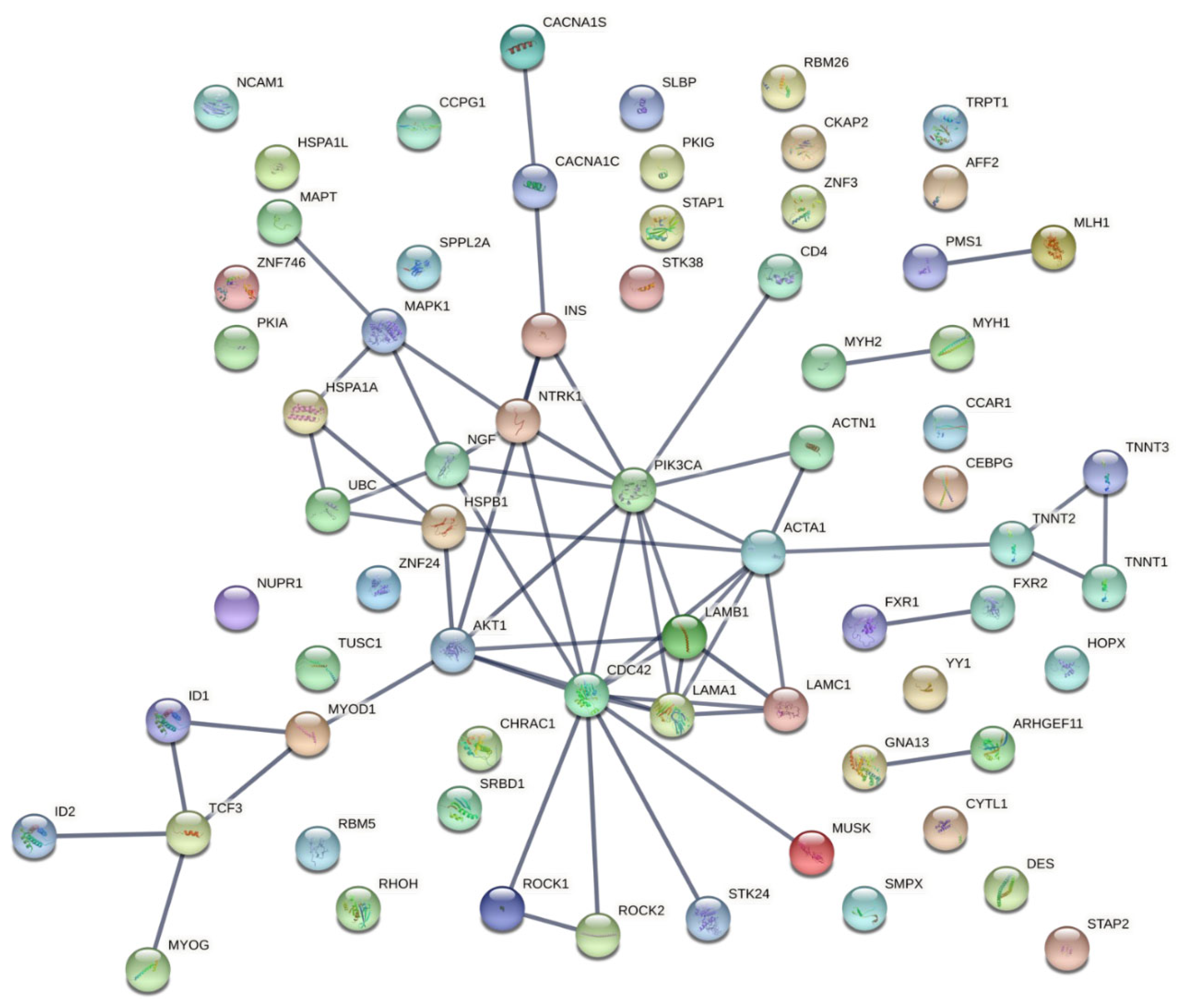
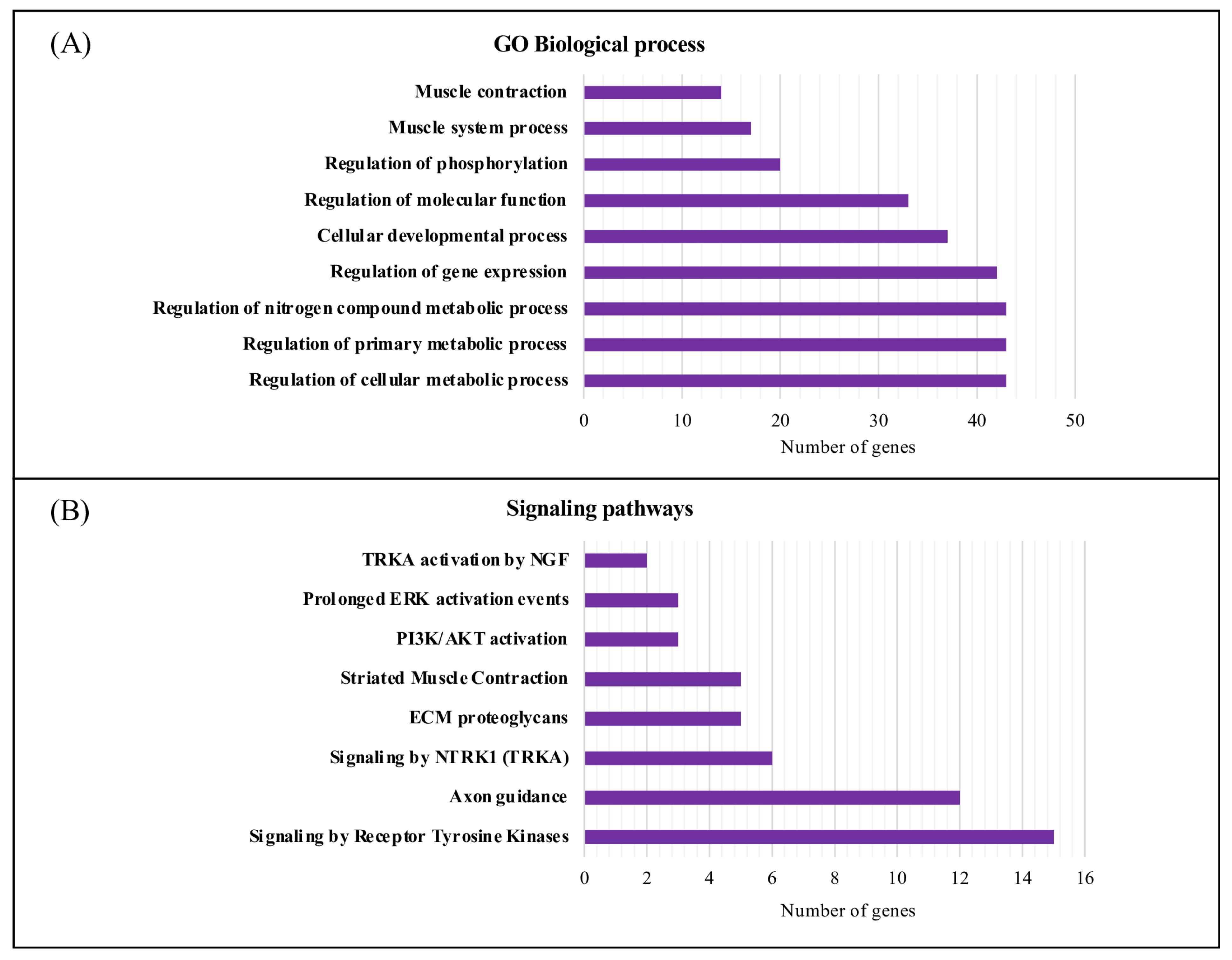

| Rank | Keyword | Total Link Strength |
|---|---|---|
| 1 | Myotonic dystrophy | 29,037 |
| 2 | Metabolism | 28,505 |
| 3 | Gene expression | 15,044 |
| 4 | Skeletal muscle | 14,660 |
| 5 | Muscle, skeletal | 13,750 |
| 6 | RNA-binding proteins | 10,329 |
| UniProt Accession Number | Gene Name | Protein Name | Biological Process |
|---|---|---|---|
| Q09013 | DMPK | Myotonic dystrophy protein kinase | Muscle cell apoptotic processing; intercellular signal transduction; regulation of excitatory postsynaptic membrane potential in skeletal muscle contraction; cellular calcium ion homeostasis |
| Q9NR56 | MBNL1 | Muscleblind Like Splicing Regulator 1 | Neurogenesis; mRNA splicing; mRNA processing; myoblast differentiation |
| Q92879 | CELF1 | CUGBP Elav-Like Family Member 1 | mRNA processing; germ cell development; positive regulation of gene expression and cell death |
| P06213 | INSR | Insulin Receptor | Carbohydrate metabolism; activation and positive regulation of protein kinase activity; glucose homeostasis; insulin receptor signaling pathway |
| P05231 | IL-6 | Interleukin 6 | Acute-phase inflammatory response; cellular response to lipopolysaccharide; glucose homeostasis; negative regulation of lipid storage; positive regulation of MAPK cascade |
| P05019 | IGF-1 | Insulin-like growth factor 1 | Myogenesis; ERK1 and ERK2 cascade; glycolate metabolic process; muscle hypertrophy; myoblast differentiation and proliferation |
| P11532 | DMD | Dystrophin | Myogenesis; actin cytoskeleton organization; muscle cell development; regulation of muscle system process; skeletal muscle tissue development |
| P17252 | PRKCA | Protein Kinase C alpha | Cell adhesion; neurotransmitter; angiogenesis; protein synthesis inhibitor; positive regulation of bone resorption; protein phosphorylation; regulation of mRNA stability |
| P60709 | ACTB | Beta-actin | Blood coagulation; rotamase; cell motility; axongenesis |
| O14983 | ATP2A1 | ATPase Sarcoplasmic/Endoplasmic Reticulum Ca2+ Transporting 1 | Blood coagulation; calcium transport; ATP synthesis; maintenance of mitochondrion location; negative regulation of striated muscle contraction |
| P16615 | ATP2A2 | ATPase Sarcoplasmic/Endoplasmic Reticulum Ca2+ Transporting 2 | Cell adhesion; blood coagulation; translocation; transport; ATP synthesis |
| P21817 | RYR-1 | Ryanodine receptor 1 | Calcium transport; muscle contraction; skeletal muscle fiber development |
| P42574 | CASP3 | Caspase 3 | Protease; apoptosis; DNA damage; proteolysis; protein processing |
| P01137 | TGFB1 | Transforming Growth Factor Beta 1 | ATP synthesis; growth arrest; inflammatory response; DNA replication inhibition; neurodegeneration; MAPK cascade |
| P06732 | CKM | Creatine kinase, M-type | Creatine metabolic process; phosphocreatine biosynthetic process |
| UniProt ID | Gene/miRNA | Protein | Biological Process | Generic Gene Role/Function in DM1 | VOSviewer Occurrences | Reference |
|---|---|---|---|---|---|---|
| Q5T8P6 | RBM26 | RNA-binding protein 26 | mRNA processing | - Protein-coding gene critical for PAXT-mediated nuclear RNA. | 478 | [32] |
| Q75NE6 | MIR17HG | Putative microRNA 17 host gene protein | - | - Exhibits complex links to cancer metastasis. | 253 | [33] |
| P01398 | INS | Insulin | Glucose metabolism, sigma factor, transcription | - INS is linked with the regulation of muscle protein synthesis, through which reduced insulin sensitivity effects occur in lower muscle mass. - Insulin signaling is known to be a significant contributor to DM1. | 196 | [34] |
| P13591 | NCAM1 | Neural cell adhesion molecule 1 | Cell adhesion plasma membrane, virus receptor activity | - Its site of expression/function in the membrane protein - Known to occur more frequently in nuclear clump fibers in DM2 than in DM1. | 188 | [35] |
| Q9Y6E0 | STK24 | Serine/threonine-protein kinase 24 | Protein phosphorylation, signal transduction | - Promotes apoptosis in response to caspase activation and stress stimuli. | 162 | [36] |
| Q9UHP9 | SMPX | Small muscular protein | Striated muscle contraction | - Functions to promote myocyte fusion by increasing the activity of the nuclear factor of activated T cells and MEF2 transcription factors through IGF1 signaling. - Important for muscle fiber organization and distal myopathy. | 145 | [37] |
| P68133 | ACTA1 | Actin, alpha skeletal muscle | Muscle contraction, skeletal muscle fiber adaption and development, skeletal muscle thin filament assembly | - Plays a role in the integrity, motility, and structure of all eukaryotic cells. It is also expressed in skeletal muscle. - Mutations in this gene are likely to cause various kinds of myopathy. - ACTA1 mutations are a notable cause of serious congenital myopathies with no treatment. | 128 | [38] |
| Q8TCT8 | SPPL2A | Signal peptide peptidase-like 2A | Membrane protein ectodomain proteolysis, membrane protein proteolysis | - Plays a key role in the development and purpose of antigen-presenting cells such as dendritic cells. - Shows a druggable pharmacological target, with the potential to provide a novel approach for treating autoimmune diseases by targeting dendritic cells and B cells. | 112 | [39] |
| Q8N5C6 | SRBD1 | S1 RNA-binding domain-containing protein 1 | Translation, nucleobase-containing compound metabolic process | - SRBD1 is an RNA-binding protein usually identified in E.coli. - Participates maintainence of homeostasis, cell growth, induction of apoptosis, and protein synthesis. - SRBD1 is known to be sensitive to early-onset normal tension glaucoma, although the functions of this gene in other fields are unclear. | 112 | [40] |
| O60356 | NUPR1 | Nuclear protein 1 | Detoxification, protein acetylation, skeletal muscle cell differentiation | - Little information about NUPR1 is available in relation to DM1, nuclear envelope alterations are observed in DM1 primary myoblasts. | 101 | [41] |
| P40692 | MLH1 | DNA mismatch repair protein MLH1 | Spermatogenesis, oogenesis, mismatch repair | - Provides instructions to make proteins that play a role in repairing DNA. | 92 | [42] |
| Q13464 | ROCK1 | Rho-associated protein kinase 1 | Regulation of cell adhesion, signal transduction | - Destabilizes the actin cytoskeleton via regulation of myosin light chain 2 (MLC2) phosphorylation. | 89 | [43] |
| P41134 | ID1 | DNA-binding protein inhibitor ID-1 | DNA-directed RNA polymerase transcription, apoptosis, angiogenesis, antibiotic resistance | - Helix–loop–helix protein involved in cellular growth and processes that has no basic region and does not bind to DNA. - ID1 inhibits skeletal muscle but has no direct association with DM1. | 78 | [44] |
| P10636 | MAPT | Microtubule-associated protein tau | rRNA metabolic process, protein homo-oligomerization | - Contributes to microtubule assembly and stability. | 78 | [45] |
| P12882 | MYH1 | Myosin-1 | Muscle contraction | - Myosin changes chemical energy into mechanical energy via the hydrolysis of ATP. | 77 | [46] |
| P51114 | FXR1 | Fragile X mental retardation syndrome-related protein 1 | Apoptotic process, muscle organ development, skeletal muscle organ development | - FXR1 splicing is key for muscle development and biomolecular condensates in muscle cells. | 74 | [47] |
| P01138 | NGF | Beta nerve growth factor | Apoptosis, inflammatory response | - Provides directions for the manufacture of a protein called nerve growth factor beta (NGFβ). - Contributes to the development and survival of nerve cells. | 71 | [48] |
| Q86TN4 | TRPT1 | tRNA 2’-phosphotransferase 1 | tRNA processing, regulation of protein kinase activity | - Predicted to enable tRNA 2’-phosphotransferase activity. - Anticipated to be involved in tRNA splicing through ligation and endonucleolytic cleavage. | 68 | [49] |
| P15923 | TCF3 | Transcription factor E2-alpha | DNA-directed RNA polymerase, transcription, transcription, regulation, apoptosis | - Plays a vital role in embryogenesis. | 66 | [50] |
| P31749 | AKT1 | Protein kinase B | Cell differentiation, glucose metabolic process, protein kinase B signaling, protein phosphorylation | - AKT1 regulates processes including cell survival, metabolism, and angiogenesis. This gene plays a role in various signaling pathways in neurodegenerative diseases. - Impairment of AKT signaling in DM1 pathogenesis causes surges in apoptosis and autophagy, which can be affected in DM1 by AMPK downregulation. | 65 | [51] |
| Q9BPY8 | HOPX | Homeodomain-only protein | DNA-directed RNA polymerase, Transcription, regulation of protein binding | - Involved in the regulation of growth and differentiation of myocytes, stem cells, and lymphocytes. | 61 | [52] |
| Q14344 | GNA13 | Guanine nucleotide-binding protein subunit alpha-13 | Transduction, blood coagulation, differentiation, cell shape | - Contributes to bone homeostasis, angiogenesis, platelet activation, and localization of germinal B cells. | 60 | [53] |
| P17028 | ZNF24 | Zinc finger protein 24 | DNA-directed RNA polymerase, transcription, myelination | - Controls proliferation, differentiation, and migration in many kinds of cells. | 60 | [54] |
| P01730 | CD4 | T-cell surface glycoprotein CD4 | Transduction, cell adhesion, innate immunity | - CD4 (T cells) fights infection. - Crucial role in adaptive immune responses, such as the stimulation of cytotoxic lymphocytes. - HIV particles are reproduced by infected CD4 T cells. | 57 | [55] |
| P28482 | MAPK1 | Mitogen-activated protein kinase 1 | DNA-directed RNA polymerase, cytosine metabolic process, ERK1 and ERK2 cascade, protein phosphorylation | - Involved in muscular dystrophies. - Many researchers have reported the activation of extracellular signal-related kinases (ERKs), c-Jun N-terminal kinases (JNKs), and p-38 MAPK in skeletal and cardiac muscle of Duchenne muscular dystrophies. - ERKs were also found to be deregulated in DM1. - MAPKs are targets of PKC proteins. | 55 | [56,57] |
| Q9ULG6 | CCPG1 | Cell cycle progression protein 1 | Positive regulation of cell cycle, positive regulation of cell population proliferation | - Cell cycle progression gene 1 and has no known physiological role. - This gene is important for ER-phagy and pancreatic ER proteostasis. | 53 | [58] |
| P17661 | DES | Desmin | Cytoskeleton organization, muscle contraction | - A class III intermediate filament used as a marker of myogenic cells. - Found in cardiac and skeletal muscle. | 53 | [59] |
| Q15669 | RHOH | Rho-related GTP-binding protein RhoH | Sigma factor, transcription regulation | - Rho genes are part of the Ras superfamily of GTPases. - Alteration of signal transduction by Rho GTPases is a repeated theme in the flow of human malignancies. | 48 | [60] |
| P60953 | CDC42 | Cell division control protein 42 homolog | Actin cytoskeleton organization, actin filament branching | - Plays an essential role in local F-actin organization by several kinase and non-kinase effector proteins - Cell cycle regulator. | 45 | [61] |
| Q9NRG0 | CHRAC1 | Chromatin accessibility complex protein 1 | Nucleosome assembly and mobilization, regulation of DNA replication | - A potential target for lung cancer. | 45 | [62] |
| P15172 | MYOD1 | Myoblast determination protein 1 | Muscle organ development, positive regulation of muscle cell differentiation, protein phosphorylation | - Encodes a nuclear protein and is known to be involved in muscle regeneration and differentiation. | 45 | [63] |
| Q2TAM9 | TUSC1 | Tumor suppressor candidate gene 1 protein | - | - Known to exhibit tumor-suppressor activity as a candidate tumor suppressor gene. | 44 | [64] |
| Q13698 | CACNA1S | Voltage-dependent L-type calcium channel subunit alpha-1S. | Muscle contraction, calcium ion import and transport | - Delivers directions to make a subunit of a structure called a calcium channel. | 43 | [65] |
| P15173 | MYOG | Myogenin | Positive regulation of muscle atrophy, regulation of myoblast fusion, skeletal muscle cell differentiation | - Plays a role in muscle atrophy, muscle differentiation, and embryonic skeletal fiber muscle differentiation. | 43 | [66] |
| P0CG48 | UBC | Polyubiquitin-C | Protein ubiquitination | - The ubiquitin (Ub) system plays a vital role in protein homeostasis. | 43 | [67] |
| P0DMV8 | HSPA1A | Heat shock 70 kDa protein 1A | ATP metabolic process, cellular response to heat, lysosomal transport, protein refolding and stabilization | - Important role in the protein quality control system. - Ensures the correct folding of proteins and controls the targeting of proteins for upcoming degradation. | 42 | [68] |
| P45378 | TNNT3 | Troponin T, fast skeletal muscle | Muscle contraction, skeletal muscle contraction | - Provides instructions to make a protein called troponin T. - Troponin T is found in skeletal muscles, which are used for movement. - This gene is linked to impaired muscle function. | 42 | [69] |
| Q9NRR1 | CYTL1 | Cytokine-like protein 1 | Signaling receptor binding, extracellular space, cartilage homeostasis | - Features of a secretory protein. - Linked with conditions such as smoking, cardiac fibrosis, and various tumors. | 40 | [70] |
| P61925 | PKIA | cAMP-dependent protein kinase inhibitor alpha | Negative regulation of protein import into the nucleus | - Protein kinase inhibitors are widely referenced in the literature. - However, because the DMPK gene encodes for myotonic dystrophy protein kinase, a serine–threonine kinase with similarity to adenosine monophosphate-dependent protein kinases undergoes self-phosphorylation, in contrast to other membrane proteins. | 40 | [71] |
| P12814 | ACTN1 | Alpha-actinin-1 | Actin cytoskeleton organization, focal adhesion assembly, muscle cell development | - Cytoskeletal proteins known to exhibit non-muscle functions. - Important for glioma cell motility. | 39 | [72] |
| O15085 | ARHGEF11 | Rho guanine nucleotide exchange factor 11 | Striated muscle contraction, G-protein-coupled receptor | - Promotes tumor metastasis in glioblastoma and ovarian cancer. | 38 | [73] |
| P42336 | PIK3CA | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform | Glucose metabolism, response to muscle stretching, positive regulation of smooth muscle cell proliferation | - Produces an enzyme called PI3K, which has been associated with DM1 - Participates in cellular signaling in response to different growth factors. - The PIK3CA gene mainly plays a role in breast cancer and causes mutations in different human malignancies. | 38 | [57,74] |
| P53567 | CEBPG | CCAAT/enhancer-binding protein gamma | mRNA metabolic process, positive regulation of DNA binding and repair | - Potential biomarkers for cancer prognosis. - Plays a role in gastric cancer progression and breast cancer cell migration. | 36 | [75] |
| P11047 | LAMC1 | Laminin subunit gamma 1 | Cell adhesion, tissue development | - Associated with the development and occurrence of tubal cancer and other malignant tumors. - Its molecular mechanism remains unclear. | 36 | [76] |
| P51816 | AFF2 | AF4/FMR2 family member 2 | mRNA processing, RNA splicing, regulation of gene expression | - Encodes a recognized transcriptional activator. - conserved and expressed in the human brain. | ND | [77] |
| Q13936 | CACNA1C | Voltage-dependent L-type calcium channel subunit alpha-1C | Cardiac muscle cell action potential involved in contraction, cardiac conduction | - Encodes calcium channels in heart tissue and its gain-of-purpose mutations in arrhythmias and sudden death. - CACNA1C and GJA1 upregulation may contribute to cardiac impairment observed in DM1 patients. | ND | [78] |
| Q8IX12 | CCAR1 | Cell division cycle and apoptosis regulator protein 1 | DNA-directed RNA polymerase, transcription, sigma factor, transcription, Regulation, mRNA splicing | - Functions as a key proliferation-inducing gene and p35 coactivator. | ND | [79] |
| Q8WWK9 | CKAP2 | Cytoskeleton-associated protein 2 | Apoptotic process, mitotic cytokinesis | - A powerful microtubule growth factor. - Plays important roles as an oncogene and spindle protein and in proliferation. | ND | [80] |
| P51116 | FXR2 | Fragile X mental retardation syndrome-related protein 2 | Regulation of mRNA stability, positive regulation of protein phosphorylation | - FXR2 is an RNA-binding protein known to play a role in fragile X cognitive disability syndrome. | ND | [81] |
| P34931 | HSPA1L | Heat shock 70 kDa protein 1-like | Unfolded protein response, vesicle-mediated transport | - Encodes a 70kDa heat shock protein. - Ensures protein quality control of the cell. | ND | [82] |
| P04792 | HSPB1 | Heat shock protein beta 1 | Regulation of protein phosphorylation, chaperon-mediated protein folding | - Preserves cellular proteostasis during the course of stress conditions. | ND | [83] |
| Q02363 | ID2 | DNA-binding protein inhibitor ID-2 | Developmental protein, regulation of lipid metabolic process, cellular senescence | - Regulator of developmentally associated genes and tumor growth in vitro, as well as in vivo, in Ewing sarcoma tumors. - Involved in cellular growth. | ND | [84] |
| P25391 | LAMA1 | Laminin subunit alpha 1 | Cell adhesion, tissue development, protein phosphorylation | - Encodes the alpha 1 subunits of laminin. - Implicated in various biological processes such as migration, cell adhesion, signaling, differentiation, and metastasis. - Mutations in this gene may be related to Poretti-Boltshauser syndrome. | ND | [85] |
| P07942 | LAMB1 | Laminin subunit beta 1 | Cell adhesion, cell migration, tissue development | - Plays significant roles in different kinds of tumors, including breast cancer, glioblastoma multiforme, prostate cancer, and hepatocellular carcinoma. | ND | [86] |
| P43246 | MLH2 | DNA mismatch repair protein Msh2 | DNA repair, oxidative phosphorylation, mismatch repair | - Delivers instructions for making a protein that play an important role in DNA repair. | ND | [87] |
| O15146 | MUSK | Muscle, skeletal receptor tyrosine-protein kinase | Skeletal muscle acetylcholine-gated channel clustering | - Encodes a muscle-specific tyrosine kinase receptor that plays a role in clustering of the acetylcholine receptor in postsynaptic neuromuscular connections. - Mutations in this gene have been related to congenital myasthenic syndrome. | ND | [88] |
| Q9UKX2 | MYH2 | Myosin-2 | Muscle contraction, muscle filament sliding | - Needed for cytoskeleton organization and muscle contraction. | ND | [89] |
| P04629 | NTRK1 | High affinity nerve growth factor receptor | Aging, nerve growth factor signaling pathway | NTRK plays a role in the growth and normal functioning of the nervous system. However, tropomyosin receptor kinase (trk) is involved in different kinds of cancer. | ND | [90] |
| Q9Y2B9 | PKIG | cAMP-dependent protein kinase inhibitor gamma | Negative regulation of protein import into the nucleus | - Protein kinase inhibitors are broadly referenced in the literature. - However, because the DMPK gene encodes for myotonic dystrophy protein kinase, a serine–threonine kinase with similarity to adenosine monophosphate-dependent protein kinases undergoes self-phosphorylation, in contrast to other membrane proteins. | ND | [71] |
| P52756 | RBM5 | RNA-binding protein 5 | Apoptosis, mRNA splicing | - Acts as a tumor suppressor. - Controls cell growth, cell cycle progression, and apoptosis in cell homeostasis. - RBM5 expression may be a potential curative target for drug-resistant lung cancer. | ND | [91] |
| O75116 | ROCK2 | Rho-associated protein kinase 2 | Actin cytoskeleton organization, positive regulation of MAPK cascade | - Stabilizes actin cytoskeleton via regulation of cofilin phosphorylation. | ND | [43] |
| Q14493 | SLBP | Histone RNA hairpin-binding protein | mRNA processing, mRNA transport | - Essential for the coordinate synthesis of DNA and histone proteins. - Needed for progression through the cell division cycle. | ND | [92] |
| Q9ULZ2 | STAP1 | Signal-transducing adaptor protein 1 | Transduction, positive regulation of gene expression | - Candidate gene related to familial hypercholesterolemia. - STAP1 is observed in immune cells. | ND | [93] |
| Q9UGK3 | STAP2 | Signal-transducing adaptor protein 2 | --- | - Regulates different intercellular signaling pathways and promotes prostate cancer flow via EGFR activation. | ND | [94] |
| Q15208 | STK38 | Serine/threonine-protein kinase 38 | Protein phosphorylation | - Negative regulator of MAPK1/2 signaling. | ND | [95] |
| P04637 | TB53 | Cellular tumor antigen p53 | Cell aging, protein stabilization, regulation of cell cycle | - Acts as a tumor suppressor in various tumor types. - Regulates in DNA repair, autophagy, senescence, and cell cycle arrest. | ND | [96] |
| P13805 | TNNT1 | Troponin T, slow skeletal muscle | Muscle contraction, skeletal muscle contraction | - TNNT1 is known as the slow skeletal troponin T subunit. - Mutations in this gene lead to nemaline myopathy type 5. - Causes most troponin-related skeletal myopathies. | ND | [97] |
| P45379 | TNNT2 | Troponin T, cardiac muscle | Muscle contraction, muscle filament sliding, actin crosslink formation | - It is unclear whether TNNT2 mis-splicing can be considered a specific cardiac biomarker in adult skeletal muscles of DM1 patients, suggesting that alternative splicing of this gene may be useful as a cardiac biomarker. | ND | [98] |
| P25490 | YY1 | Transcriptional repressor protein YY1 | DNA-directed RNA polymerase, transcription, differentiation spermatogenesis, DNA damage | - Involved in regulating the expression of a large number of genes and binds to interferon-beta (IFN-β) promoter to either repress or activate its expression. - It could play a role in modulating the cellular response to dsRNA. | ND | [99] |
| P17036 | ZNF3 | Zinc finger protein 3 (isoform) | Cell differentiation, leukocyte activation | - Serves as a specific RNA-binding domain, which helps MBNL to identify various target mRNAs by binding to a wide area of motifs. | ND | [100] |
| Q6NUN9 | ZNF746 | Zinc finger protein 746 | DNA-directed RNA polymerase, transcription | - Performs the function of promoting the occurrence of hepatocellular carcinoma. - Recognized as a substrate of E3 ligase Parkin, and its accumulation is related to Parkinson’s disease. | ND | [101] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kuntawala, D.H.; Martins, F.; Vitorino, R.; Rebelo, S. Automatic Text-Mining Approach to Identify Molecular Target Candidates Associated with Metabolic Processes for Myotonic Dystrophy Type 1. Int. J. Environ. Res. Public Health 2023, 20, 2283. https://doi.org/10.3390/ijerph20032283
Kuntawala DH, Martins F, Vitorino R, Rebelo S. Automatic Text-Mining Approach to Identify Molecular Target Candidates Associated with Metabolic Processes for Myotonic Dystrophy Type 1. International Journal of Environmental Research and Public Health. 2023; 20(3):2283. https://doi.org/10.3390/ijerph20032283
Chicago/Turabian StyleKuntawala, Dhvani H., Filipa Martins, Rui Vitorino, and Sandra Rebelo. 2023. "Automatic Text-Mining Approach to Identify Molecular Target Candidates Associated with Metabolic Processes for Myotonic Dystrophy Type 1" International Journal of Environmental Research and Public Health 20, no. 3: 2283. https://doi.org/10.3390/ijerph20032283





