Development of a 3-D Physical Dynamics Monitoring System Using OCM with DVC for Quantification of Sprouting Endothelial Cells Interacting with a Collagen Matrix
Abstract
:1. Introduction
2. Materials and Methods
2.1. Imaging System
2.1.1. Gabor-Domain Optical Coherence Microscope
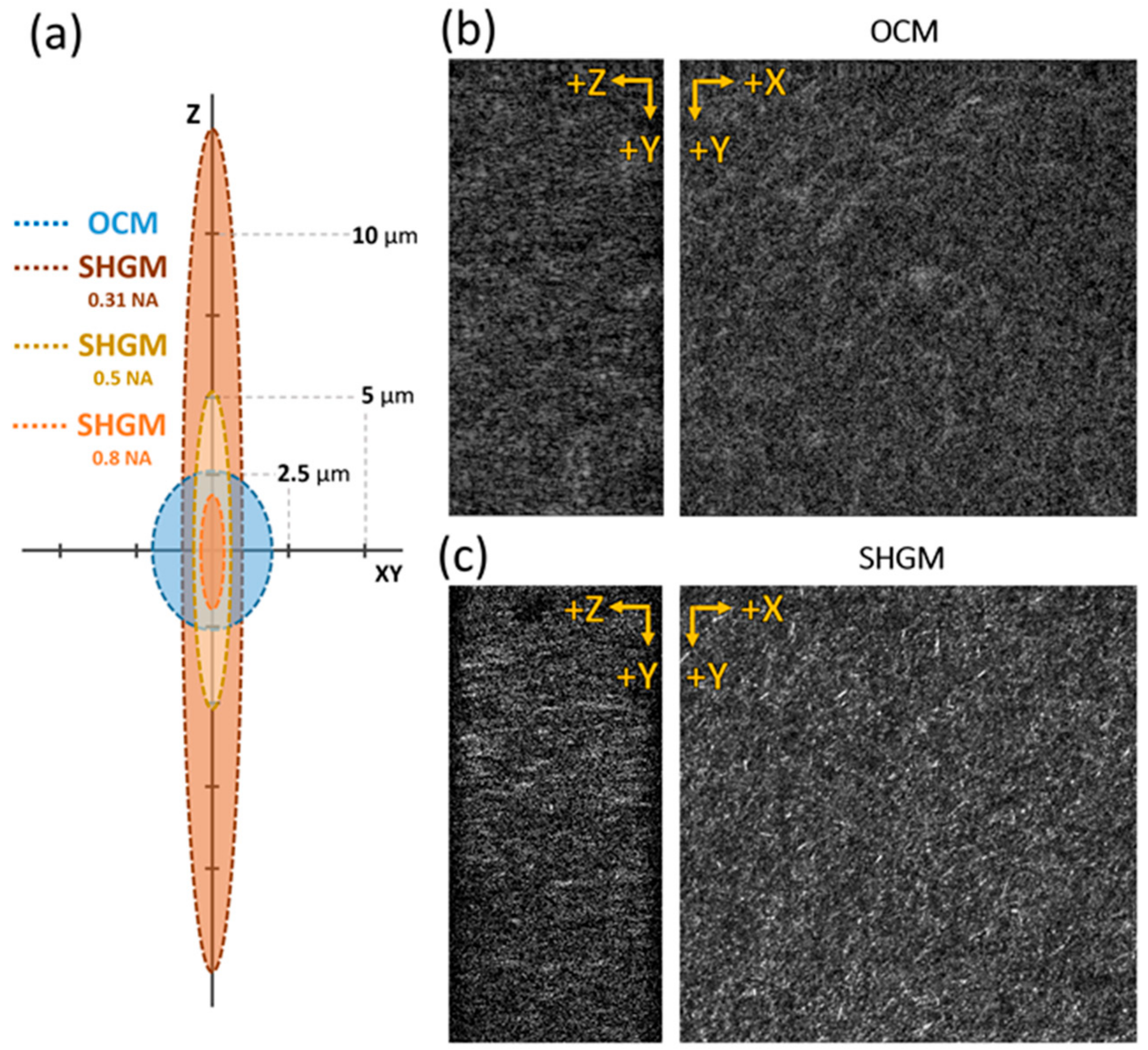
2.1.2. Second-Harmonic Generation Microscope
2.2. Image Acquisition Process
2.2.1. SHGM Image Acquisition
2.2.2. OCM Image Acquisition and Postprocessing
2.3. Sample Preparation
2.3.1. Microfluidic Chip Fabrication
2.3.2. Collagen Type I Gel Preparation
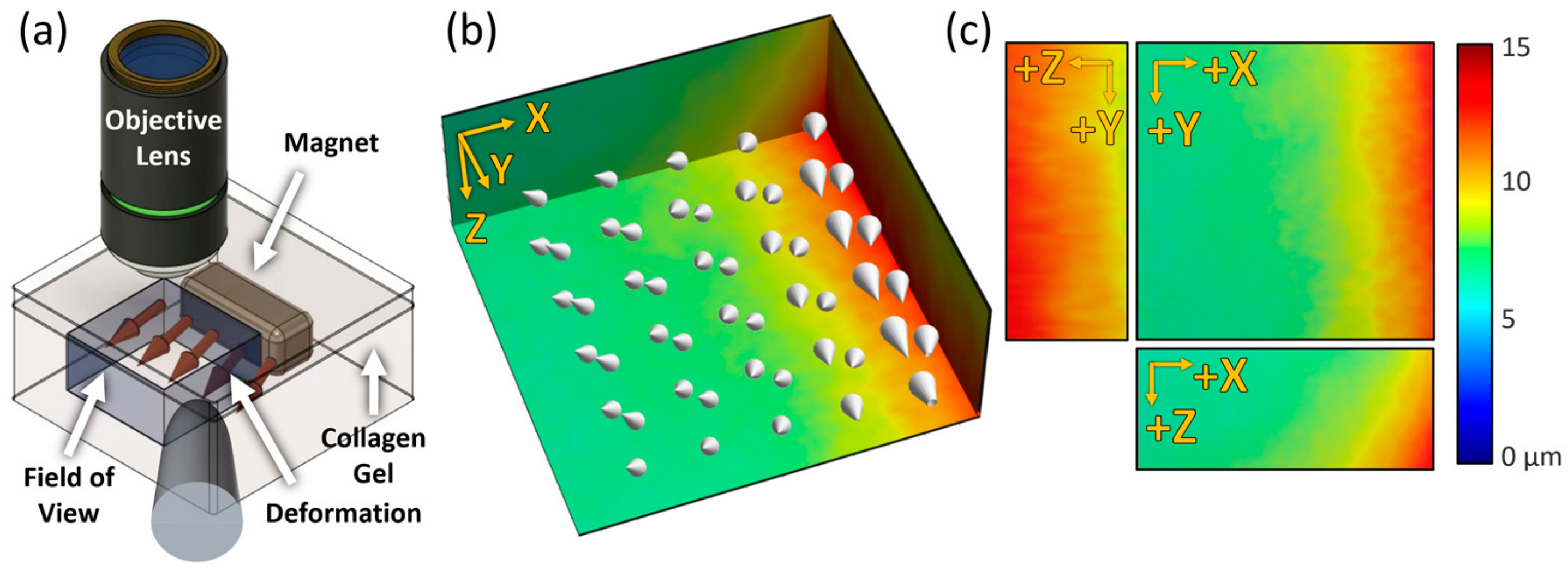
2.3.3. Generating Strain with Nd Magnet and Steel Rod
2.3.4. HUVEC Preparation and Imaging
2.4. DVC Analysis
2.4.1. Error Floor Evaluation
2.4.2. Drift Correction and Calculation of the Incremental Displacement
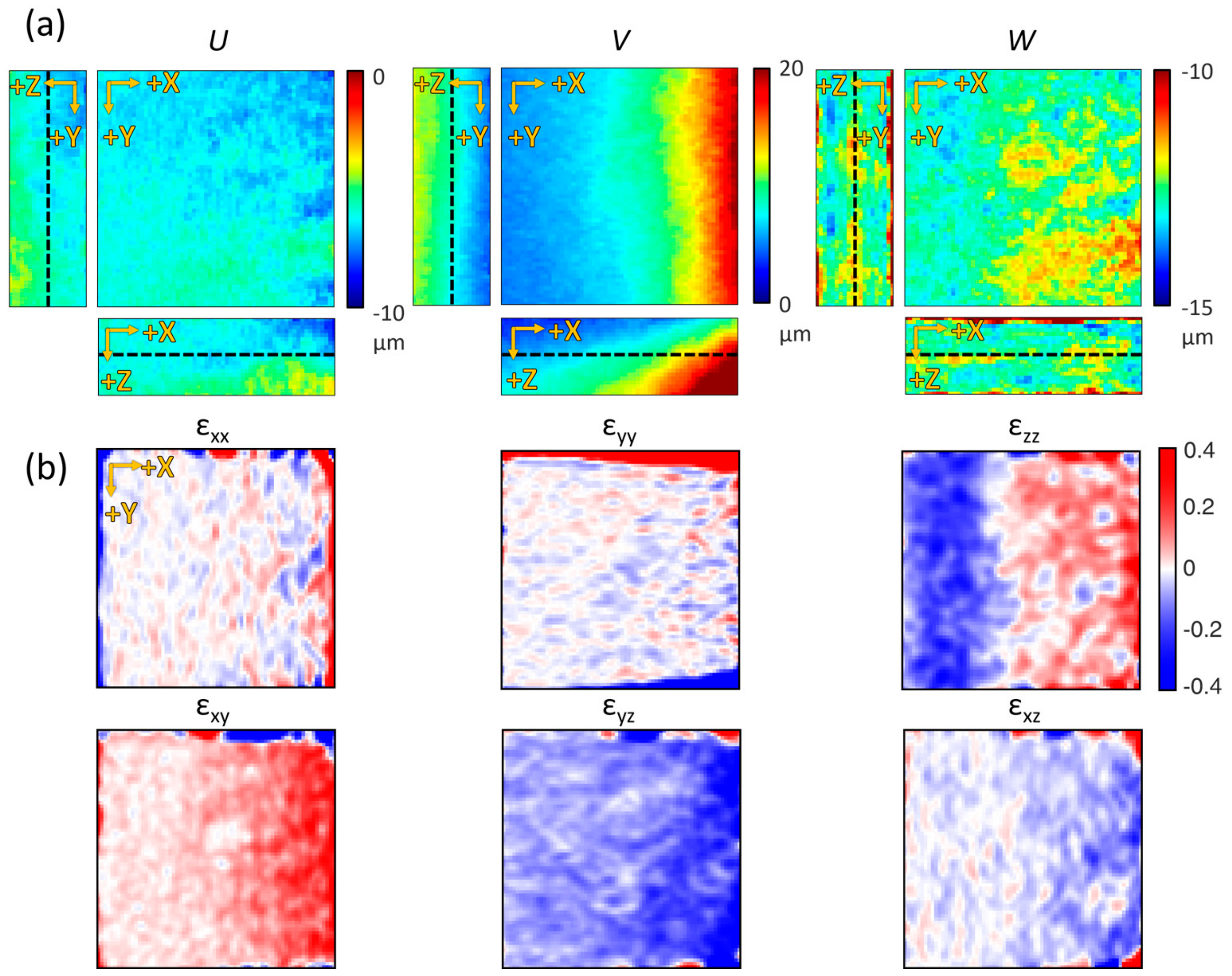
2.4.3. Strain Calculation
3. Results
3.1. GD-OCM System Configuration and Performance Verification
3.2. Optimization of the Efficient Sampling Rate for Detecting Collagen Gel Structural Deformation
3.3. Quantification and Visualization of Collagen Gel Deformation and Its Strain
3.4. Observation of the ECM-Mediated Physical Dynamics of Sprouting Vascular Endothelial Cell and Distant Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rozario, T.; DeSimone, D.W. The extracellular matrix in development and morphogenesis: A dynamic view. Dev. Biol. 2009, 341, 126–140. [Google Scholar] [CrossRef] [Green Version]
- Midwood, K.S.; Williams, L.V.; Schwarzbauer, J.E. Tissue repair and the dynamics of the extracellular matrix. Int. J. Biochem. Cell Biol. 2004, 36, 1031–1037. [Google Scholar] [CrossRef]
- E Discher, D.; Janmey, P.; Wang, Y.-L. Tissue Cells Feel and Respond to the Stiffness of Their Substrate. Science 2005, 310, 1139–1143. [Google Scholar] [CrossRef] [Green Version]
- Kalluri, R.; Zeisberg, M. Fibroblasts in cancer. Nat. Rev. Cancer 2006, 6, 392–401. [Google Scholar] [CrossRef]
- Koch, T.M.; Münster, S.; Bonakdar, N.; Butler, J.P.; Fabry, B. 3D Traction Forces in Cancer Cell Invasion. PLoS ONE 2012, 7, e33476. [Google Scholar] [CrossRef] [Green Version]
- Riching, K.M.; Cox, B.L.; Salick, M.R.; Pehlke, C.; Riching, A.; Ponik, S.M.; Bass, B.R.; Crone, W.C.; Jiang, Y.; Weaver, A.M.; et al. 3D Collagen Alignment Limits Protrusions to Enhance Breast Cancer Cell Persistence. Biophys. J. 2014, 107, 2546–2558. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Egeblad, M.; Rasch, M.G.; Weaver, V.M. Dynamic interplay between the collagen scaffold and tumor evolution. Curr. Opin. Cell Biol. 2010, 22, 697–706. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jabłońska-Trypuć, A.; Matejczyk, M.; Rosochacki, S. Matrix metalloproteinases (MMPs), the main extracellular matrix (ECM) enzymes in collagen degradation, as a target for anticancer drugs. J. Enzym. Inhib. Med. Chem. 2016, 31, 177–183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hur, S.S.; Zhao, Y.; Li, Y.-S.; Botvinick, E.; Chien, S. Live Cells Exert 3-Dimensional Traction Forces on Their Substrata. Cell. Mol. Bioeng. 2009, 2, 425–436. [Google Scholar] [CrossRef] [Green Version]
- Toyjanova, J.; Bar-Kochba, E.; López-Fagundo, C.; Reichner, J.S.; Hoffman-Kim, D.; Franck, C. High Resolution, Large Deformation 3D Traction Force Microscopy. PLoS ONE 2014, 9, e90976. [Google Scholar] [CrossRef]
- Yang, Y.; Kaufman, L.J. Rheology and Confocal Reflectance Microscopy as Probes of Mechanical Properties and Structure during Collagen and Collagen/Hyaluronan Self-Assembly. Biophys. J. 2009, 96, 1566–1585. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brightman, A.; Rajwa, B.; Sturgis, J.; McCallister, M.; Robinson, J.; Voytik-Harbin, S. Time-lapse confocal reflection microscopy of collagen fibrillogenesis and extracellular matrix assembly in vitro. Biopolym. Orig. Res. Biomol. 2000, 54, 222–234. [Google Scholar] [CrossRef]
- Huisken, J. Optical Sectioning Deep Inside Live Embryos by Selective Plane Illumination Microscopy. Science 2004, 305, 1007–1009. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huisken, J.; Stainier, D.Y. Selective plane illumination microscopy techniques in developmental biology. Development 2009, 136, 1963–1975. [Google Scholar] [CrossRef] [Green Version]
- Huang, D.; Swanson, E.A.; Lin, C.P.; Schuman, J.S.; Stinson, W.G.; Chang, W.; Hee, M.R.; Flotte, T.; Gregory, K.; Puliafito, C.A. Optical coherence tomography. Science 1991, 254, 1178–1181. [Google Scholar] [CrossRef] [Green Version]
- Izatt, J.A.; Hee, M.R.; Owen, G.M.; Swanson, E.A.; Fujimoto, J.G. Optical coherence microscopy in scattering media. Opt. Lett. 1994, 19, 590. [Google Scholar] [CrossRef]
- Klein, T.; Huber, R. High-speed OCT light sources and systems [Invited]. Biomed. Opt. Express 2017, 8, 828–859. [Google Scholar] [CrossRef]
- Drexler, W. Ultrahigh-resolution optical coherence tomography. J. Biomed. Opt. 2004, 9, 47. [Google Scholar] [CrossRef]
- Aguirre, A.D.; Zhou, C.; Lee, H.-C.; Ahsen, O.O.; Fujimoto, J.G. Optical Coherence Microscopy. In Optical Coherence Tomography; Drexler, W., Fujimoto, J.G., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 865–911. [Google Scholar]
- Huang, Y.; Wang, S.; Guo, Q.; Kessel, S.; Rubinoff, I.; Chan, L.L.-Y.; Li, P.; Liu, Y.; Qiu, J.; Zhou, C. Optical Coherence Tomography Detects Necrotic Regions and Volumetrically Quantifies Multicellular Tumor Spheroids. Cancer Res. 2017, 77, 6011–6020. [Google Scholar] [CrossRef] [Green Version]
- Mulligan, J.A.; Bordeleau, F.; Reinhart-King, C.A.; Adie, S.G. Measurement of dynamic cell-induced 3D displacement fields in vitro for traction force optical coherence microscopy. Biomed. Opt. Express 2017, 8, 1152–1171. [Google Scholar] [CrossRef] [Green Version]
- Mulligan, J.A.; Feng, X.; Adie, S.G. Quantitative reconstruction of time-varying 3D cell forces with traction force optical coherence microscopy. Sci. Rep. 2019, 9, 4086. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tan, W.; Oldenburg, A.L.; Norman, J.J.; Desai, T.A.; A Boppart, S. Optical coherence tomography of cell dynamics in three-dimensional tissue models. Opt. Express 2006, 14, 7159–7171. [Google Scholar] [CrossRef] [PubMed]
- Mateo, J.L.; Fernández-Caballero, A. Finding out general tendencies in speckle noise reduction in ultrasound images. Expert Syst. Appl. 2009, 36, 7786–7797. [Google Scholar] [CrossRef]
- Yamaguchi, I. A laser-speckle strain gauge. J. Phys. E Sci. Instrum. 1981, 14, 1270–1273. [Google Scholar] [CrossRef]
- Zaitsev, V.Y.; A Matveev, L.; Matveyev, A.L.; Gelikonov, G.V.; Gelikonov, V.M. A model for simulating speckle-pattern evolution based on close to reality procedures used in spectral-domain OCT. Laser Phys. Lett. 2014, 11, 105601. [Google Scholar] [CrossRef] [Green Version]
- Schmitt, J.M.; Xiang, S.H.; Yung, K.M. Speckle in Optical Coherence Tomography. J. Biomed. Opt. 1999, 4, 95. [Google Scholar] [CrossRef]
- Bay, B.K.; Smith, T.S.; Fyhrie, D.P.; Saad, M. Digital volume correlation: Three-dimensional strain mapping using X-ray tomography. Exp. Mech. 1999, 39, 217–226. [Google Scholar] [CrossRef]
- Hall, M.S.; Alisafaei, F.; Ban, E.; Feng, X.; Hui, C.-Y.; Shenoy, V.B.; Wu, M. Fibrous nonlinear elasticity enables positive mechanical feedback between cells and ECMs. Proc. Natl. Acad. Sci. USA 2016, 113, 14043–14048. [Google Scholar] [CrossRef] [Green Version]
- Münster, S.; Jawerth, L.M.; Leslie, B.A.; Weitz, J.I.; Fabry, B.; Weitz, D.A. Strain history dependence of the nonlinear stress response of fibrin and collagen networks. Proc. Natl. Acad. Sci. USA 2013, 110, 12197–12202. [Google Scholar] [CrossRef] [Green Version]
- Liang, L.; Jones, C.; Chen, S.; Sun, B.; Jiao, Y. Heterogeneous force network in 3D cellularized collagen networks. Phys. Biol. 2016, 13, 066001. [Google Scholar] [CrossRef]
- Genge, K.-; Blocki, A.; Franke, R.-P.; Krüger-Genge, A.; Jung, F. Vascular Endothelial Cell Biology: An Update. Int. J. Mol. Sci. 2019, 20, 4411. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ucuzian, A.A.; Greisler, H.P. In Vitro Models of Angiogenesis. World J. Surg. 2007, 31, 654–663. [Google Scholar] [CrossRef]
- Hoang, M.V.; Senger, D.R.; Guan, J.-L. In Vivo and In Vitro Models of Mammalian Angiogenesis. Cell Migration 2004, 294, 269–286. [Google Scholar] [CrossRef]
- Vailhé, B.; Vittet, D.; Feige, J.-J.; Vailh, B. In Vitro Models of Vasculogenesis and Angiogenesis. Lab. Investig. 2001, 81, 439–452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daley, W.P.; Peters, S.B.; Larsen, M. Extracellular matrix dynamics in development and regenerative medicine. J. Cell Sci. 2008, 121, 255–264. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edgar, L.T.; Underwood, C.J.; Guilkey, J.E.; Hoying, J.B.; Weiss, J.A. Extracellular Matrix Density Regulates the Rate of Neovessel Growth and Branching in Sprouting Angiogenesis. PLoS ONE 2014, 9, e85178. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Nadiarynkh, O.; Plotnikov, S.; Campagnola, P. Second harmonic generation microscopy for quantitative analysis of collagen fibrillar structure. Nat. Protoc. 2012, 7, 654–669. [Google Scholar] [CrossRef]
- Lee, S.-W.; Jeong, H.-W.; Kim, B.-M.; Ahn, Y.-C.; Jung, W.; Chen, Z. Optimization for Axial Resolution, Depth Range, and Sensitivity of Spectral Domain Optical Coherence Tomography at 1.3 µm. J. Korean Phys. Soc. 2009, 55, 2354–2360. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.-W.; Jeong, H.-W.; Kim, B.-M. High-speed spectral domain polarization- sensitive optical coherence tomography using a single camera and an optical switch at 1.3 µm. J. Biomed. Opt. 2010, 15, 10501. [Google Scholar] [CrossRef] [Green Version]
- Fischler, M.; Bolles, R. Random sample consensus: A paradigm for model fitting with applications to image analysis and automated cartography. Commun. ACM 1981, 24, 381–395. [Google Scholar] [CrossRef]
- Rolland, J.P.; Meemon, P.; Murali, S.; Thompson, K.P.; Lee, K.-S. Gabor-based fusion technique for Optical Coherence Microscopy. Opt. Express 2010, 18, 3632–3642. [Google Scholar] [CrossRef] [PubMed]
- Murali, S. Gabor Domain Optical Coherence Microscopy. Ph.D. Thesis, University of Central Florida, Orlando, FL, USA, 2009. [Google Scholar]
- Kang, Y.G.; Jang, H.; Yang, T.D.; Notbohm, J.; Choi, Y.; Park, Y.; Kim, B.-M. Quantification of focal adhesion dynamics of cell movement based on cell-induced collagen matrix deformation using second-harmonic generation microscopy. J. Biomed. Opt. 2018, 23, 065001. [Google Scholar] [CrossRef] [PubMed]
- Jeong, G.S.; Kwon, G.H.; Kang, A.R.; Jung, B.Y.; Park, Y.; Chung, S.; Lee, S.-H. Microfluidic assay of endothelial cell migration in 3D interpenetrating polymer semi-network HA-Collagen hydrogel. Biomed. Microdevices 2011, 13, 717–723. [Google Scholar] [CrossRef]
- Bar-Kochba, E.; Toyjanova, J.; Andrews, E.; Kim, K.-S.; Franck, C. A Fast Iterative Digital Volume Correlation Algorithm for Large Deformations. Exp. Mech. 2014, 55, 261–274. [Google Scholar] [CrossRef]
- Jorge-Peñas, A.; Bové, H.; Sanen, K.; Vaeyens, M.-M.; Steuwe, C.; Roeffaers, M.B.J.; Ameloot, M.; Van Oosterwyck, H. 3D full-field quantification of cell-induced large deformations in fibrillar biomaterials by combining non-rigid image registration with label-free second harmonic generation. Biomaterials 2017, 136, 86–97. [Google Scholar] [CrossRef]
- Landauer, A.K.; Patel, M.; Henann, D.L.; Franck, C. A q-Factor-Based Digital Image Correlation Algorithm (qDIC) for Resolving Finite Deformations with Degenerate Speckle Patterns. Exp. Mech. 2018, 58, 815–830. [Google Scholar] [CrossRef]
- Masters, B.R.; So, P.T. Handbook of Biomedical Nonlinear Optical Microscopy. J. Biomed. Opt. 2009, 14, 19901. [Google Scholar] [CrossRef] [Green Version]
- Tibbitt, M.W.; Anseth, K.S. Hydrogels as extracellular matrix mimics for 3D cell culture. Biotechnol. Bioeng. 2009, 103, 655–663. [Google Scholar] [CrossRef] [Green Version]
- García, A.J.; Vega, M.D.; Boettiger, D.; García, A.J. Modulation of Cell Proliferation and Differentiation through Substrate-dependent Changes in Fibronectin Conformation. Mol. Biol. Cell 1999, 10, 785–798. [Google Scholar] [CrossRef] [Green Version]
- Baker, B.M.; Chen, C.S. Deconstructing the third dimension – how 3D culture microenvironments alter cellular cues. J. Cell Sci. 2012, 125, 3015–3024. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cox, T.R.; Erler, J.T. Remodeling and homeostasis of the extracellular matrix: Implications for fibrotic diseases and cancer. Dis. Model. Mech. 2011, 4, 165–178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mammoto, T.; Mammoto, A.; Ingber, D.E. Mechanobiology and Developmental Control. Annu. Rev. Cell Dev. Biol. 2013, 29, 27–61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Das, R.K.; Zouani, O.F. A review of the effects of the cell environment physicochemical nanoarchitecture on stem cell commitment. Biomaterials 2014, 35, 5278–5293. [Google Scholar] [CrossRef] [PubMed]
- Rosso, F.; Giordano, A.; Barbarisi, M.; Barbarisi, A. From Cell-ECM interactions to tissue engineering. J. Cell. Physiol. 2004, 199, 174–180. [Google Scholar] [CrossRef]
- Huang, S.; Tang, C.; Xu, M.; Qiu, Y.; Lei, Z. BM3D-based total variation algorithm for speckle removal with structure-preserving in OCT images. Appl. Opt. 2019, 58, 6233–6243. [Google Scholar] [CrossRef]
- Chong, B.; Zhu, Y.-K. Speckle reduction in optical coherence tomography images of human finger skin by wavelet modified BM3D filter. Opt. Commun. 2013, 291, 461–469. [Google Scholar] [CrossRef]
- Bobrow, T.; Mahmood, F.; Inserni, M.; Durr, N.J. DeepLSR: A deep learning approach for laser speckle reduction. Biomed. Opt. Express 2019, 10, 2869–2882. [Google Scholar] [CrossRef]
- Li, Y.; Winetraub, Y.; Liba, O.; De La Zerda, A.; Chu, S. Optimization of the Trade-Off Between Speckle Reduction and Axial Resolution in Frequency Compounding. IEEE Trans. Med. Imaging 2018, 38, 107–112. [Google Scholar] [CrossRef]
- Magnain, C.; Wang, H.; Sakadžić, S.; Fischl, B.; Boas, D.A. En face speckle reduction in optical coherence microscopy by frequency compounding. Opt. Lett. 2016, 41, 1925–1928. [Google Scholar] [CrossRef] [Green Version]
- Li, R.; Yin, H.; Hong, J.; Wang, C.; He, B.; Chen, Z.; Li, Q.; Xue, P.; Zhang, X.; Bin, H.; et al. Speckle reducing OCT using optical chopper. Opt. Express 2020, 28, 4021–4031. [Google Scholar] [CrossRef]
- Liba, O.; Lew, M.D.; SoRelle, E.D.; Dutta, R.; Sen, D.; Moshfeghi, D.M.; Chu, S.; De La Zerda, A. Speckle-modulating optical coherence tomography in living mice and humans. Nat. Commun. 2017, 8, 15845. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reu, P. All about Speckles: Aliasing. Exp. Tech. 2014, 38, 1–3. [Google Scholar] [CrossRef]
- Rogowska, J.; Patel, N.; Plummer, S.; E Brezinski, M. Quantitative optical coherence tomographic elastography: Method for assessing arterial mechanical properties. Br. J. Radiol. 2006, 79, 707–711. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Standish, B.; Vuong, B.; Wen, X.-Y.; Yang, V. Digital image correlation–based optical coherence elastography. J. Biomed. Opt. 2013, 18, 121515. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kennedy, B.; McLaughlin, R.A.; Kennedy, K.; Chin, L.; Curatolo, A.; Tien, A.; Latham, B.; Saunders, C.; Sampson, D.D. Optical coherence micro-elastography: Mechanical-contrast imaging of tissue microstructure. Biomed. Opt. Express 2014, 5, 2113–2124. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; Larin, K.V. Optical coherence elastography for tissue characterization: A review. J. Biophotonics 2014, 8, 279–302. [Google Scholar] [CrossRef] [Green Version]
- Meng, F.; Zhang, X.; Wang, J.; Li, C.; Chen, J.; Sun, C. 3D Strain and Elasticity Measurement of Layered Biomaterials by Optical Coherence Elastography based on Digital Volume Correlation and Virtual Fields Method. Appl. Sci. 2019, 9, 1349. [Google Scholar] [CrossRef] [Green Version]



| Type | Error Floor STD 1 (IQR 2) | Displacement Mean (Median) | |||||
|---|---|---|---|---|---|---|---|
| Sampling Rate (µm/px) | 0.8 | 0.53 | 0.4 | 0.8 | 0.53 | 0.4 | GT3 |
| U (µm) | 0.111 | 0.047 | 0.103 | 3.195 | 3.199 | 3.199 | 3.20 |
| (0.147) | (0.071) | (0.160) | (3.198) | (3.200) | (3.199) | ||
| V (µm) | 0.068 | 0.040 | 0.208 | 3.195 | 3.199 | 3.199 | 3.20 |
| (0.094) | (0.060) | (0.377) | (3.198) | (3.199) | (3.199) | ||
| W (µm) | 0.150 | 0.162 | 0.201 | 4.417 | 4.423 | 4.417 | 4.42 |
| (0.220) | (0.209) | (0.313) | (4.422) | (4.424) | (4.419) | ||
| Average (µm) | 0.110 | 0.083 | 0.171 | 3.602 | 3.607 | 3.605 | 3.61 |
| (0.154) | (0.113) | (0.283) | (3.606) | (3.608) | (3.606) | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kang, Y.G.; Jang, H.; Park, Y.; Kim, B.-M. Development of a 3-D Physical Dynamics Monitoring System Using OCM with DVC for Quantification of Sprouting Endothelial Cells Interacting with a Collagen Matrix. Materials 2020, 13, 2693. https://doi.org/10.3390/ma13122693
Kang YG, Jang H, Park Y, Kim B-M. Development of a 3-D Physical Dynamics Monitoring System Using OCM with DVC for Quantification of Sprouting Endothelial Cells Interacting with a Collagen Matrix. Materials. 2020; 13(12):2693. https://doi.org/10.3390/ma13122693
Chicago/Turabian StyleKang, Yong Guk, Hwanseok Jang, Yongdoo Park, and Beop-Min Kim. 2020. "Development of a 3-D Physical Dynamics Monitoring System Using OCM with DVC for Quantification of Sprouting Endothelial Cells Interacting with a Collagen Matrix" Materials 13, no. 12: 2693. https://doi.org/10.3390/ma13122693





