Artificial Intelligence Models for the Mass Loss of Copper-Based Alloys under Cavitation
Abstract
1. Introduction
- The experimental setup was designed by our team.
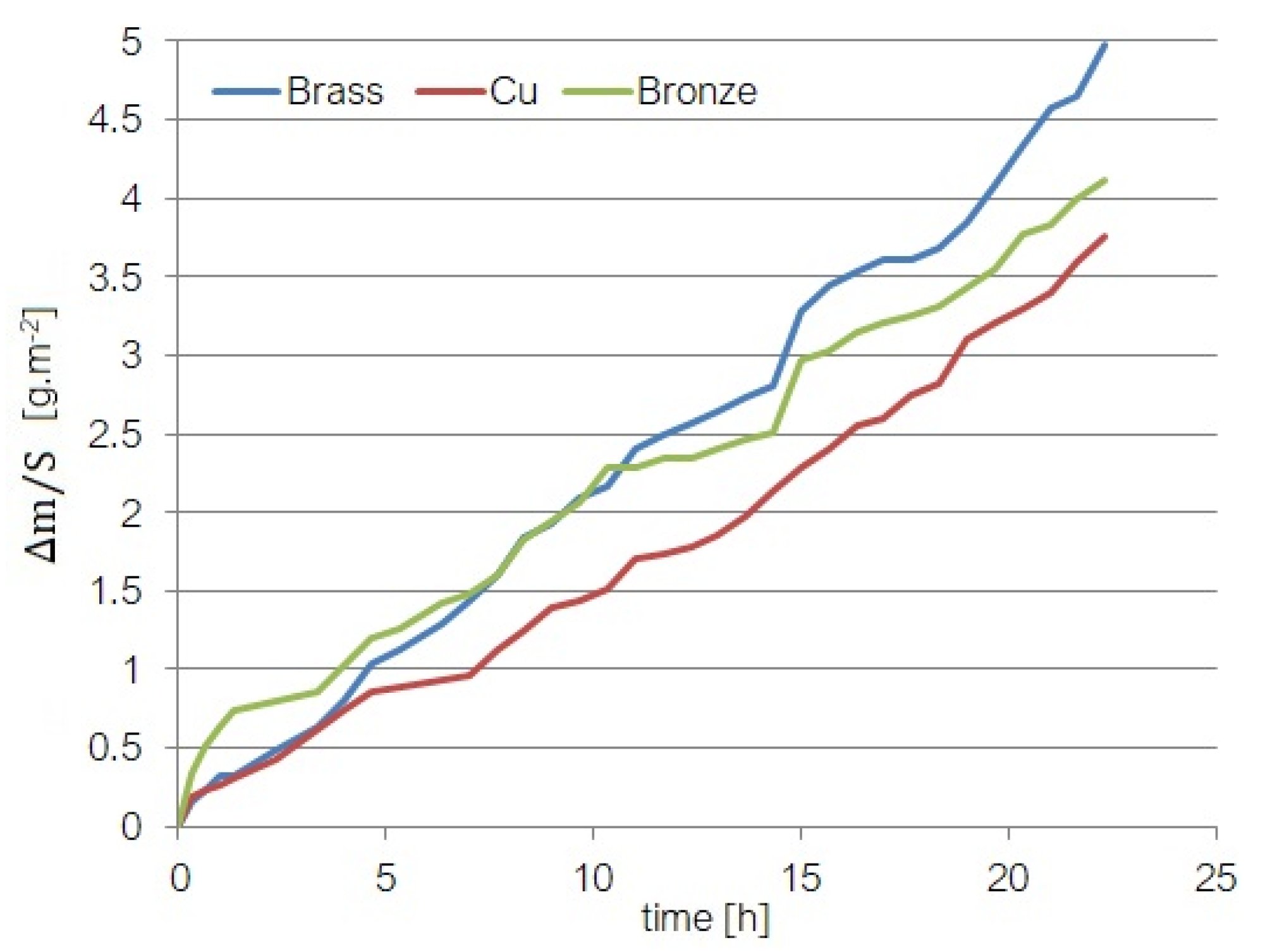
- The experiments on materials mass loss have been performed in the cavitation field produced by ultrasound; this approach is important for knowing the mass-loss behavior of some materials used in naval construction.
- Based on our knowledge, the analysis of the mass loss of copper-based alloys in the ultrasound cavitation field was not extensively performed.
- Modeling the mass loss of such alloys in the cavitation field has not been performed using AI methods.
- Knowing the mass loss is important for predicting the behavior of different components built using such materials; a good model can be used to obtain a forecast that can be utilized for predicting the replacement periods in an integrated reliability study.
2. Materials and Methods
2.1. Experiments
- -
- The tank (1) containing the liquid subject to the ultrasound cavitation;
- -
- The high-frequency ultrasound generator (8), designed to work at 220 V, 18 kHz, and three power levels;
- -
- The piezoceramic transducer (7) that produces cavitation entering into oscillation as a response to the high-frequency signal received from the generator;
- -
- The control panel (command block) (12) from where the ultrasound generator’s working power is selected;
- -
- The cooler (11) that is used to maintain a constant temperature of the liquid;
- -
- The measurement electrodes (13), which are utilized only in the experiments related to capturing the signal induced in the cavitation field;
- -
- The acquisition data unit (14), used only in the experiments related to electrical signals induced by ultrasound cavitation to collect the signals.
- -
- Cu containing small percentages of Fe, Sn, and Zn (0.0395%, 0.0446%, and 0.0747%, respectively);
- -
- A brass with 2.75% Pb, 38.45% Zn besides Cu (57.95 %);
- -
- A bronze containing Zn, Pb, and Sn (4.07%, 4.40%, and 6.4%, respectively) besides Cu.
2.2. Modeling Methodology for the Weight Loss
- -
- Number of genes per chromosome—4.
- -
- Length of the head of a gene—8.
- -
- Number of constant per gene—10.
- -
- The size of the population—50 individuals.
- -
- The maximum number of generations (and without improvement)—2000 (1000).
- -
- The fitness function—MSE, and the hit tolerance—0.01.
- -
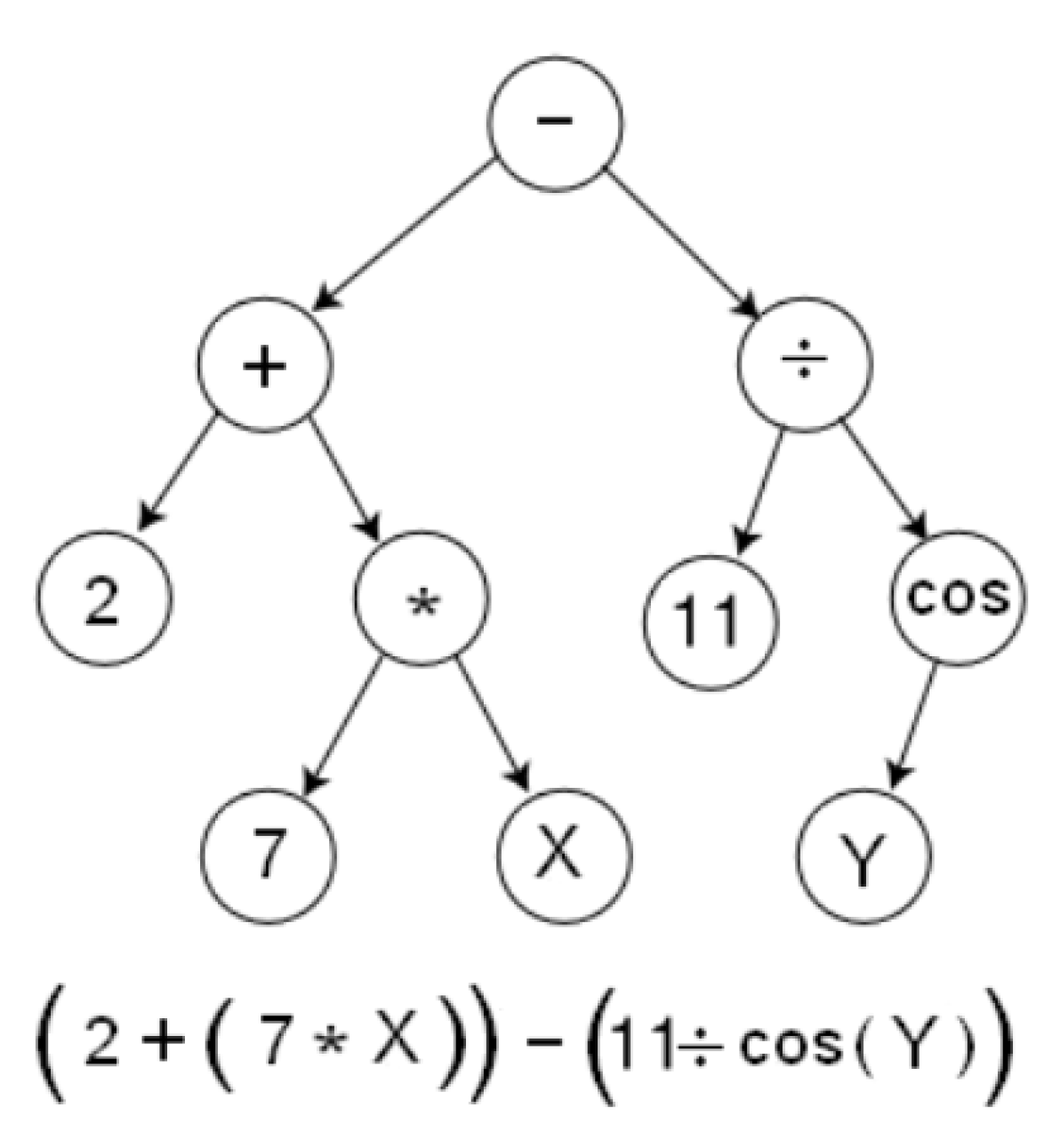
- The functions used to build the final expression—{+, −, *, /, sqrt}.
- -
- The linking function was the addition.
- -
- The algorithm was allowed to do algebraic simplification.
- -
- The mutation (and inversion) rate—0.44 (0.1).
- -
- The transposition rate—0.1.
- -
- The one-point (two-point and gene) recombination rate—0.3 (0.3 and 0.1).
- -
- For the experimental reasons explained above, the regressor in the model was the lag 1 variable, to predict .
- -
- The obtained models are compared with respect to the MSE values obtained on the original data set; in the following, we report only the best model (i.e., the model with the smallest mean standard error) found in all 50 runs of the algorithm.
- -
- Experiments were performed using different ratios between the Training and Test sets, such as 80:20 or 90:10; the best results are presented in this article, which were obtained for the Training set formed by 70% of the data series, and the Test set formed by the rest of the series’ values.
3. Results
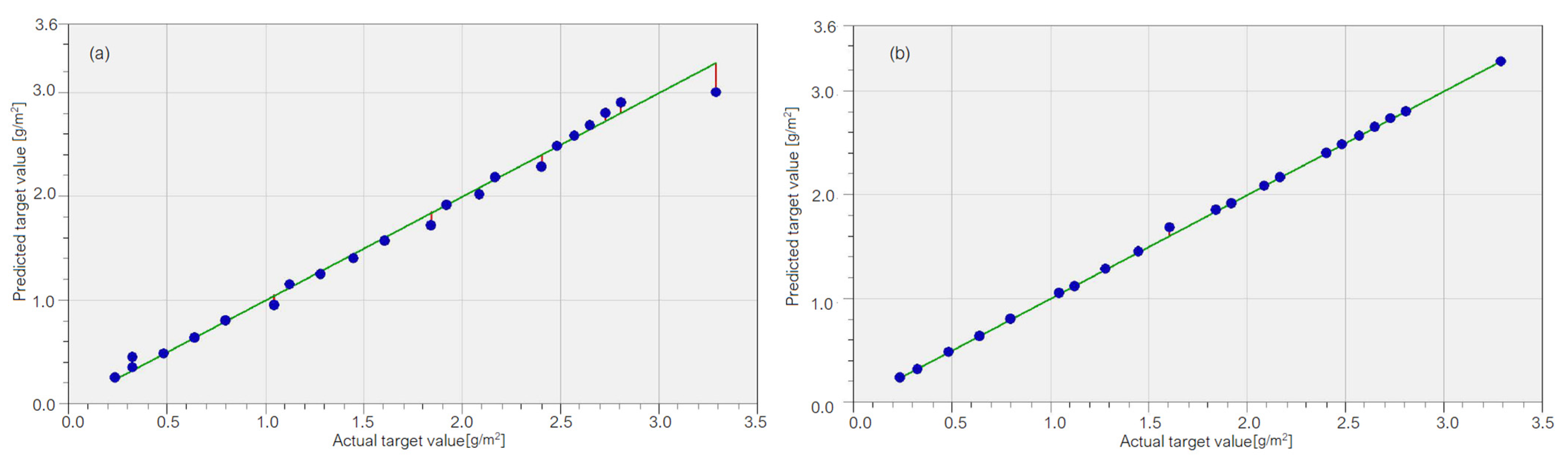
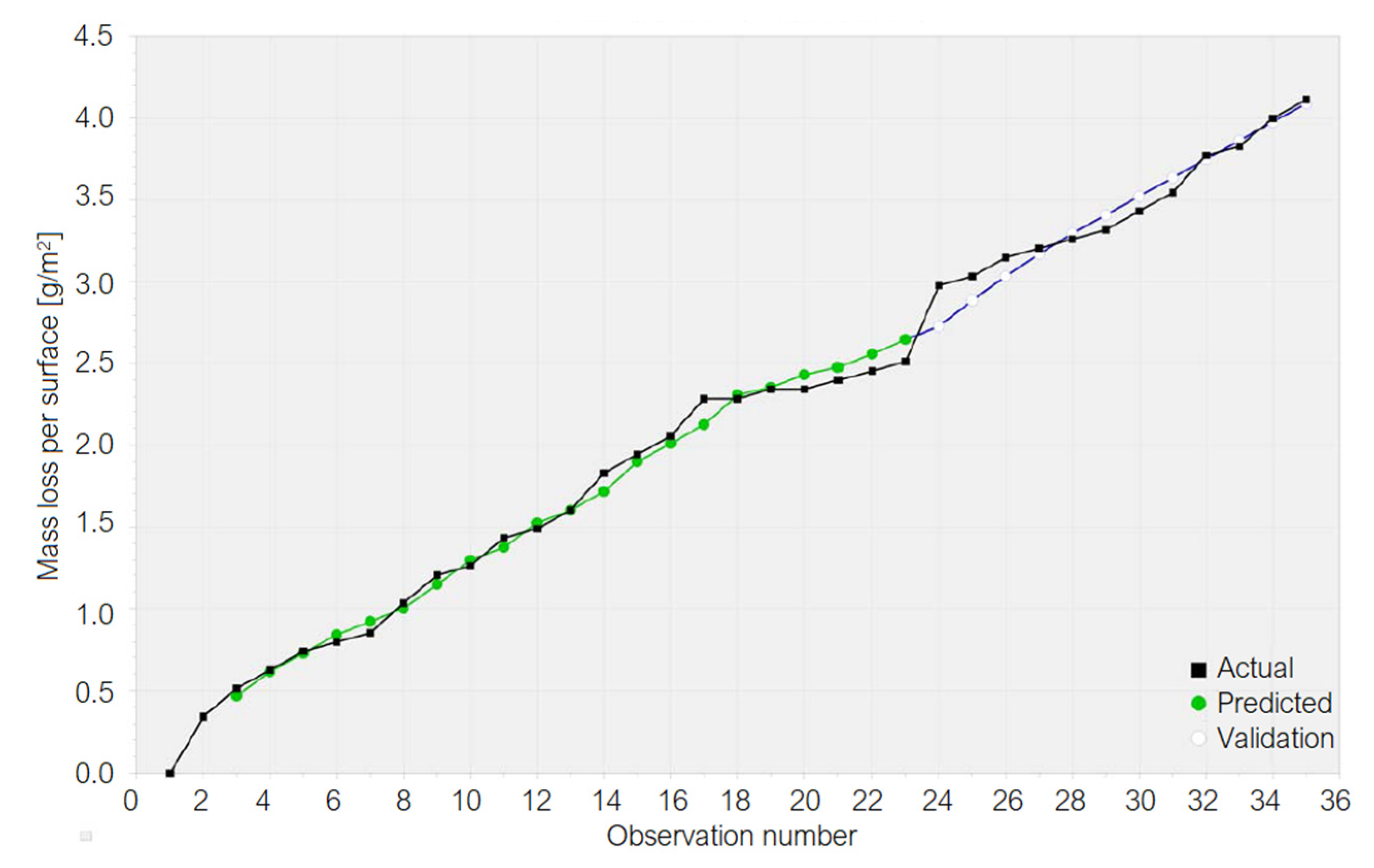
3.1. Results on the Cu Sample Mass Loss
3.2. Results on the Brass Sample Mass Loss
- Point of view of R2, rap, RMSE, and MAE for all algorithms provided the best results on the Training sets;
- Point of view of CV, all but GRNN gave the best results on the Test sets;
- From the MAPE viewpoint, the best results were provided by GRNN on the Test set and by SVR on the Test set.
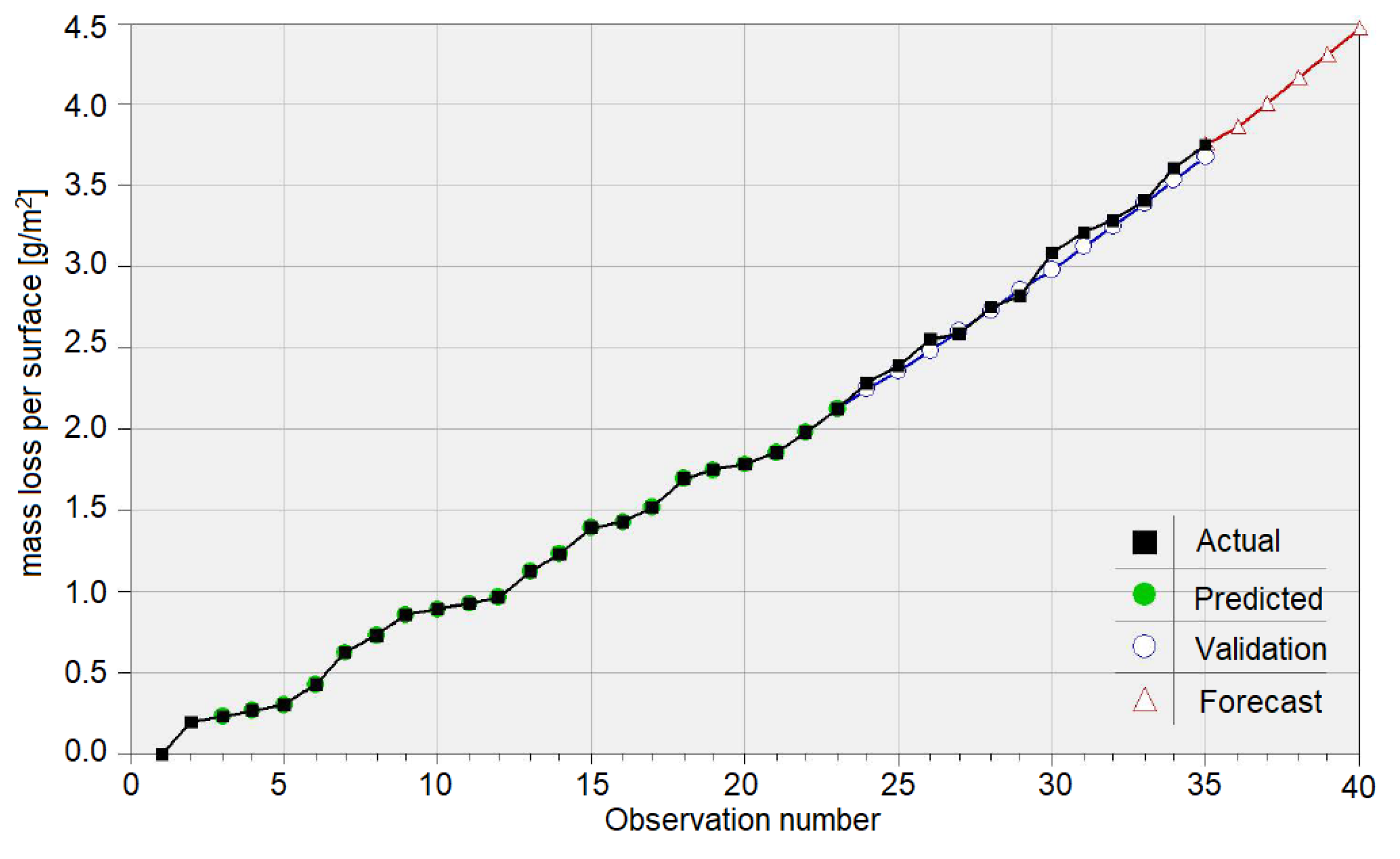
3.3. Results on the Bronze Sample Mass Loss
4. Discussion
- On the Training set: R2 = 99.935%, CV = 0.0151, MAE = 0.0033, RMSE = 0.016, rap = 0.99972, and MAPE = 10−8,
- On the Test set: R2 = 98.369%, CV = 0.0198, MAE=0.0504, RMSE= 0.0590, rap = 0.9964, and MAPE = 1.6719.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Flynn, H.G. Physics of acoustic cavitation in liquids. In Physical Acoustics; Mason, W.P., Ed.; Academic Press: New York, NY, USA, 1964; Volume 1, Part B; pp. 57–172. [Google Scholar]
- Bărbulescu, A. Models of the voltage induced by cavitation in hydrocarbons. Acta Phys. Pol. B 2006, 37, 2919–2931. [Google Scholar]
- Bărbulescu, A.; Dumitriu, C.Ș. Modeling the Voltage Produced by Ultrasound in Seawater by Stochastic and Artificial Intelligence Methods. Sensors 2022, 22, 1089. [Google Scholar] [CrossRef] [PubMed]
- Bărbulescu, A.; Dumitriu, C.S. ARIMA and Wavelet-ARIMA Models for the Signal Produced by Ultrasound in Diesel. In Proceedings of the 25th International Conference on System Theory, Control and Computing (ICSTCC 2021), Iasi, Romania, 20–23 October 2021. [Google Scholar] [CrossRef]
- Bai, L.; Yan, J.; Zeng, Z.; Ma, Y. Cavitation in thin liquid layer: A review. Ultrason. Sonochem. 2020, 66, 105092. [Google Scholar] [CrossRef] [PubMed]
- Young, F.E. Cavitation; Mac Graw-Hill: Maidenhead, UK, 1989. [Google Scholar]
- Rooney, J.A. Ultrasound: Its Chemical, Physical and Biological Effects, Suslick; VCH: New York, NY, USA, 1988. [Google Scholar]
- Dumitriu, C.Ș. On the copper-based materials corrosion. In Physics Studies; Emek, M., Ed.; IKSAD Publishing House: Ankara, Turkey, 2021; pp. 67–100. [Google Scholar]
- Oliphant, R.J. Causes of Copper Corrosion in Plumbing Systems. Foundation for Water Research; Allen House: Marlow, UK, 2003. [Google Scholar]
- Simionov, M. Studies and Research on the Cavitation Destruction of Cylinder Liners from Diesel Engines. Ph.D. Thesis, Dunarea de Jos University of Galati, Galati, Romania, 1997. [Google Scholar]
- Basumatary, J.; Nie, M.; Wood, J.K. The synergistic effects of cavitation erosion-corrosion in ship propeller materials. J. Bio- Tribo-Corros. 2015, 1, 12. [Google Scholar] [CrossRef]
- Basumatary, J.; Wood, R.J.K. Synergistic effects of cavitation erosion and corrosion for nickel aluminium bronze with oxide film in 3.5% NaCl solution. Wear 2017, 376–377, 1286–1297. [Google Scholar] [CrossRef]
- Schüssler, A.; Exner, H.E. The corrosion of nickel-aluminium bronzes in seawater—I. Protective layer formation and the passivation mechanism. Corros. Sci. 1993, 3, 1793–1802. [Google Scholar] [CrossRef]
- Wharton, J.A.; Barik, R.C.; Kear, G.; Wood, R.J.K.; Stokes, K.R.; Walsh, F.C. The corrosion of nickel-aluminium bronze in seawater. Corros. Sci. 2005, 47, 3336–3367. [Google Scholar] [CrossRef]
- Wharton, J.A.; Stokes, K.R. The influence of nickel–aluminium bronze microstructure and crevice solution on the initiation of crevice corrosion. Electrochim. Acta 2008, 53, 2463–2473. [Google Scholar] [CrossRef]
- Bakhshandeh, H.R.; Allahkaram, S.R.; Zabihi, A.H. An investigation on cavitation-corrosion behavior of Ni/β-SiC nanocomposite coatings under ultrasonic field. Ultrason. Sonochem. 2019, 56, 229–239. [Google Scholar] [CrossRef]
- Bărbulescu, A.; Orac, L. Corrosion analysis and models for some composites behavior in saline media. Int. J. Energy Environ. 2008, 1, 35–44. [Google Scholar]
- Peng, S.; Xu, J.; Li, Z.; Jiang, S.; Xie, Z.-H.; Munroe, P. Electrochemical noise analysis of cavitation erosion corrosion resistance of NbC nanocrystalline coating in a 3.5 wt% NaCl solution. Surf. Coat. Technol. 2021, 415, 127133. [Google Scholar] [CrossRef]
- Ivanov, I.V. Corrosion Resistant Materials in Food Industry; Editura Agro-Silvica: Bucharest, Romania, 1959. (In Romanian) [Google Scholar]
- Kumar, S.; Narayanan, T.S.N.S.; Manimaran, A.; Kumar, M.S. Effect of lead content on the dezincification behaviour of leaded brass in neutral and acidified 3.5% NaCl solution. Mater. Chem. Phys. 2007, 10, 134–141. [Google Scholar] [CrossRef]
- Hagen, C.M.H.; Hognestad, A.; Knudsen, O.Ø.; Sørby, K. The effect of surface roughness on corrosion resistance of machined and epoxy coated steel. Prog. Org. Coat. 2019, 130, 17–23. [Google Scholar] [CrossRef]
- Okada, T. Corrosive Liquid Effects on Cavitation Erosion, Reprint UMICh No. 014456-52-1; University of Michigan: Ann Arbor, MI, USA, 1979. [Google Scholar]
- Bărbulescu, A.; Dumitriu, C.Ș. Models of the mass loss of some copper alloys. Chem. Bull. Politehnica Univ. (Timisoara) 2007, 52, 120–123. [Google Scholar]
- Fortes-Patella, R.; Choffat, T.; Reboud, J.L.; Archer, A. Mass loss simulation in cavitation erosion: Fatigue criterion approach. Wear 2013, 300, 205–215. [Google Scholar] [CrossRef]
- Dumitriu, C.S. On the corrosion of two types of bronzes under cavitation. Ann. Dunarea Jos Univ. of Galati Fasc. IX Metall. Mater. Sci. 2021, 4, 12–16. [Google Scholar] [CrossRef]
- Dumitriu, C.S.; Bărbulescu, A. Copper corrosion in ultrasound cavitation field. Ann. Dunarea Jos Univ. of Galati Fasc. IX Metall. Mater. Sci. 2021, 3, 31–35. [Google Scholar] [CrossRef]
- Simian, D.; Stoica, F.; Bărbulescu, A. Automatic Optimized Support Vector Regression for Financial Data Prediction. Neural Comput. Appl. 2020, 32, 2383–2396. [Google Scholar] [CrossRef]
- Uysal, M.; Tanyildizi, H. Estimation of compressive strength of self compacting concrete containing polypropylene fiber and mineral additives exposed to high temperature using artificial neural network. Constr. Build. Mater. 2012, 27, 404–414. [Google Scholar] [CrossRef]
- Bărbulescu, A.; Barbes, L. Modeling the outlet temperature in heat exchangers. Case study. Thermal Sci. 2021, 25, 591–602. [Google Scholar] [CrossRef]
- Javed, M.F.; Amin, M.N.; Shah, M.I.; Khan, K.; Iftikhar, B.; Farooq, F.; Aslam, F.; Alyousef, R.; Alabduljabbar, H. Applications of Gene Expression Programming and Regression Techniques for Estimating Compressive Strength of Bagasse Ash based Concrete. Crystals 2020, 10, 737. [Google Scholar] [CrossRef]
- Farooq, F.; Akbar, A.; Khushnood, R.A.; Muhammad, W.L.B.; Rehman, S.K.U.; Javed, M.F. Experimental investigation of hybrid carbon nanotubes and graphite nanoplatelets on rheology, shrinkage, mechanical, and microstructure of SCCM. Materials 2020, 13, 230. [Google Scholar] [CrossRef] [PubMed]
- Bărbulescu, A.; Șerban, C.; Caramihai, S. Assessing the soil pollution using a genetic algorithm. Rom. J. Phys. 2021, 66, 806. [Google Scholar]
- Vakhshouri, B.; Nejadi, S. Prediction of compressive strength of self-compacting concrete by ANFIS models. Neurocomputing 2018, 280, 13–22. [Google Scholar] [CrossRef]
- Bărbulescu, A.; Dani, A. Statistical analysis and classification of the water parameters of Beas River (India). Rom. Rep. Phys. 2019, 71, 716. [Google Scholar]
- Bustillo, A.; Pimenov, D.Y.; Matuszewski, M.; Mikolajczyk, T. Using artificial intelligence models for the prediction of surface wear based on surface isotropy levels. Robot. Comput.-Integr. Manuf. 2018, 53, 215–227. [Google Scholar] [CrossRef]
- Alsina, E.F.; Chica, M.; Trawiński, K.; Regattieri, A. On the use of machine learning methods to predict component reliability from data-driven industrial case studies. Int. J. Adv. Manuf. Technol. 2018, 94, 2419–2433. [Google Scholar] [CrossRef]
- Aslam, F.; Furqan, F.; Amin, M.N.; Khan, K.; Waheed, A.; Akbar, A.; Javed, M.F.; Alyousef, R.; Alabdulijabbar, H. Applications of Gene Expression Programming for Estimating Compressive Strength of High-Strength Concrete. Adv. Civil Eng. 2020, 2020, 8850535. [Google Scholar] [CrossRef]
- Shen, Z.; Deifalla, A.F.; Kaminski, P.; Dyczko, A. Compressive Strength Evaluation of Ultra-High-Strength Concrete by Machine Learning. Materials 2022, 15, 3523. [Google Scholar] [CrossRef]
- Bărbulescu, A.; Mârza, V.; Dumitriu, C.S. Installation and Method for Measuring and Determining the Effects Produced by Cavitation in Ultrasound Field in Stationary and Circulating Media. Romanian Patent No. RO 123086-B1, 30 April 2010. [Google Scholar]
- Vapnik, V. The Nature of Statistical Learning Theory; Springer: Berlin, Germany, 1995. [Google Scholar]
- Smola, A.J.; Scholkopf, B. A tutorial on support vector regression. Stat. Comput. 2004, 14, 199–222. [Google Scholar] [CrossRef]
- Basak, D.; Pal, S.; Patranabis, D.C. Support vector regression. Neural Inf. Process. Lett. Rev. 2007, 11, 203–224. [Google Scholar]
- Specht, D.F. A General Regression Neural Network. IEEE Trans. Neural Netw. 1991, 2, 568–578. [Google Scholar] [CrossRef] [PubMed]
- Bauer, M.M. 2 General Regression Neural Network (GRNN). Available online: https://minds.wisconsin.edu/bitstream/handle/1793/7779/ch2.pdf?sequence%3D14 (accessed on 26 July 2022).
- Al-Mahasneh, A.J.; Anavatti, S.; Pratama, M.G.M. Applications of General Regression Neural Networks in Dynamic Systems. In Digital Systems; Asadpour, V., Ed.; IntechOpen: London, UK, 2018. [Google Scholar]
- Howlett, R.J.; Jain, L.C. Radial Basis Function Networks 2: New Advances in Design; Physica-Verlag: Heidelberg, Germany, 2001. [Google Scholar]
- Kurban, T.; Beșdok, E. A Comparison of RBF Neural Network Training Algorithms for Inertial Sensor Based Terrain Classification. Sensors 2009, 9, 6312–6329. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zheng, Q.; Shi, Z.; Chen, J. Training radial basis function networks with particle swarms. Lect. Note Comput. Sci. 2004, 3173, 317–322. [Google Scholar]
- Simon, D. Training radial basis neural networks with the extended Kalman filter. Neurocomputing 2002, 48, 455–475. [Google Scholar] [CrossRef]
- Chen, S.; Hong, X.; Harris, C.J. Orthogonal Forward Selection for Constructing the Radial Basis Function Network with Tunable Nodes. In Advances in Intelligent Computing. ICIC 2005. Lecture Notes in Computer Science; Huang, D.S., Zhang, X.P., Huang, G.B., Eds.; Springer: Berlin/Heidelberg, Germany, 2005; Volume 3644, pp. 777–786. [Google Scholar]
- Fernández-Redondo, M.; Hernández-Espinosa, C.; Ortiz-Gómez, M.; Torres-Sospedra, J. Training Radial Basis Functions by Gradient Descent. In Artificial Intelligence and Soft Computing—ICAISC 2004. Lecture Notes in Computer Science; Rutkowski, L., Siekmann, J.H., Tadeusiewicz, R., Zadeh, L.A., Eds.; Springer: Berlin/Heidelberg, Germany, 2004; Volume 3070, pp. 184–189. [Google Scholar]
- Karayiannis, N.B. Reformulated radial basis neural networks trained by gradient descent. IEEE Trans. Neural Netw. 1999, 3, 2230–2235. [Google Scholar] [CrossRef] [PubMed]
- Orr, M.J.L. Introduction to Radial Basis Function Networks. 1966. Available online: https://faculty.cc.gatech.edu/~isbell/tutorials/rbf-intro.pdf (accessed on 6 December 2021).
- Genetic Algorithms for Feature Selection. Available online: https://www.neuraldesigner.com/blog/genetic_algorithms_for_feature_selection (accessed on 26 July 2022).
- Koza, J.R. Genetic Programming: On the Programming of Computers by Means of Natural Selection; MIT Press: Cambridge, MA, USA, 1992. [Google Scholar]
- Cheng, R. Genetic Algorithms and Engineering Design; Wiley: Hoboken, NJ, USA, 2007. [Google Scholar]
- Banzhaf, W.; Nordin, P.; Keller, R.; Francone, F.D. Genetic Programming—An Introduction; On the Automatic Evolution of Computer Programs and Its Applications; Morgan Kaufmann: San Francisco, CA, USA, 1998. [Google Scholar]
- Ferreira, C. Gene Expression Programming: A New Adaptive Algorithm for Solving Problems. Complex Syst. 2001, 13, 85–129. [Google Scholar]
- Zhang, Q.; Zhou, C.; Xiao, W.; Nelson, P.C. Improving Gene Expression Programming Performance by Using Differential Evolution. Available online: https://www.cs.uic.edu/~qzhang/Zhang-GEP.pdf (accessed on 26 July 2022).
- Ferreira, C. Gene Expression Programming: Mathematical Modeling by an Artificial Intelligence; Springer: Berlin/Heidelberg, Germany, 2006. [Google Scholar]
- DTREG. Available online: https://www.dtreg.com/ (accessed on 6 August 2022).


| Set | Training | Test | ||||||
|---|---|---|---|---|---|---|---|---|
| Indicator | SVR | GRNN | RBF | GEP | SVR | GRNN | RBF | GEP |
| R2 (%) | 99.290 | 99.943 | 99.315 | 99.339 | 98.923 | 99.206 | 99.166 | 99.206 |
| CV | 0.0500 | 0.0141 | 0.0491 | 0.0482 | 0.0161 | 0.0137 | 0.0141 | 0.0138 |
| rap | 0.9967 | 0.9997 | 0.9967 | 0.9967 | 0.9963 | 0.9962 | 0.9960 | 0.9962 |
| RMSE | 0.0528 | 0.0149 | 0.0519 | 0.0510 | 0.0479 | 0.0410 | 0.0422 | 0.0412 |
| MAE | 0.0422 | 0.0031 | 0.0453 | 0.0448 | 0.0402 | 0.0345 | 0.0350 | 0.0353 |
| MAPE | 6.5800 | 77 × 10−7 | 7.3097 | 7.0305 | 1.3421 | 1.1612 | 1.1957 | 1.1935 |
| Actual | Computed | Error | % Error |
|---|---|---|---|
| 2.2840 | 2.2734 | 0.0106 | 0.465 |
| 2.4002 | 2.4015 | −0.0013 | 0.056 |
| 2.5550 | 2.5258 | 0.0292 | 1.145 |
| 2.5938 | 2.6500 | −0.0562 | 2.169 |
| 2.7486 | 2.7756 | −0.0270 | 0.982 |
| 2.8260 | 2.9031 | −0.0771 | 2.729 |
| 3.0970 | 3.0329 | 0.0641 | 2.070 |
| 3.2131 | 3.1650 | 0.0481 | 1.500 |
| 3.2906 | 3.2994 | −0.0088 | 0.269 |
| 3.4067 | 3.4363 | −0.0296 | 0.869 |
| 3.6003 | 3.5757 | 0.0246 | 0.683 |
| 3.7551 | 3.7176 | 0.0375 | 0.999 |
| Set | Training | Test | ||||||
|---|---|---|---|---|---|---|---|---|
| Indicator | SVR | GRNN | RBF | GEP | SVR | GRNN | RBF | GEP |
| R2 (%) | 99.231 | 99.969 | 99.619 | 99.555 | 94.501 | 94.197 | 93.574 | 90.628 |
| CV | 0.0563 | 0.0112 | 0.0397 | 0.0429 | 0.0291 | 0.0299 | 0.0315 | 0.0380 |
| rap | 0.9964 | 0.9999 | 0.9981 | 0.9978 | 0.9773 | 0.9724 | 0.9715 | 0.9662 |
| RMSE | 0.0854 | 0.0170 | 0.0602 | 0.0651 | 0.1173 | 0.1205 | 0.1268 | 0.1532 |
| MAE | 0.0580 | 0.0048 | 0.0502 | 0.0546 | 0.1004 | 0.0981 | 0.1022 | 0.1240 |
| MAPE | 6.2502 | 0.2009 | 6.9911 | 6.9481 | 2.463 | 2.4912 | 2.6025 | 3.1965 |
| SVR | GRNN | RBF | GEP |
|---|---|---|---|
| 2.368 | 3.458 | 2.621 | 3.012 |
| 1.676 | 1.757 | 1.119 | 0.745 |
| 0.573 | 0.134 | 0.522 | 1.394 |
| 3.012 | 3.831 | 4.461 | 5.676 |
| 4.474 | 5.537 | 6.106 | 7.506 |
| 3.797 | 4.986 | 5.482 | 6.951 |
| 1.234 | 2.463 | 2.884 | 4.348 |
| 0.986 | 0.244 | 0.600 | 2.040 |
| 2.932 | 1.723 | 1.420 | 0.013 |
| 1.366 | 0.149 | 0.117 | 1.535 |
| 4.676 | 3.520 | 3.296 | 1.942 |
| Set | Training | Test | ||||||
|---|---|---|---|---|---|---|---|---|
| Indicator | SVR | GRNN | RBF | GEP | SVR | GRNN | RBF | GEP |
| R2(%) | 98.629 | 99.066 | 98.730 | 98.849 | 92.271 | 98.686 | 92.029 | 91.312 |
| CV | 0.0594 | 0.0490 | 0.0572 | 0.5746 | 0.0294 | 0.0220 | 0.0299 | 0.0312 |
| rap | 0.9932 | 0.9957 | 0.9938 | 0.9944 | 0.9768 | 0.9834 | 0.9795 | 0.9836 |
| RMSE | 0.0887 | 0.0732 | 0.0854 | 0.0813 | 0.1020 | 0.0762 | 0.1036 | 0.1081 |
| MAE | 0.0676 | 0.0426 | 0.0665 | 0.0613 | 0.0789 | 0.0601 | 0.0794 | 0.0873 |
| MAPE | 5.0594 | 2.4270 | 5.0261 | 4.0533 | 2.4427 | 1.8253 | 2.4556 | 2.6488 |
| Recorded | Computed | Error | Absolute % Error |
|---|---|---|---|
| 2.9724 | 2.8194 | 0.1530 | 5.150 |
| 3.0296 | 2.9639 | 0.0657 | 2.168 |
| 3.1439 | 3.1282 | 0.0157 | 0.501 |
| 3.2011 | 3.1495 | 0.0516 | 1.613 |
| 3.2582 | 3.2993 | −0.0409 | 1.260 |
| 3.3154 | 3.4634 | −0.1470 | 4.463 |
| 3.4297 | 3.4944 | −0.0647 | 1.886 |
| 3.5441 | 3.6393 | −0.0952 | 2.688 |
| 3.7727 | 3.8036 | −0.0309 | 0.818 |
| 3.8298 | 3.8248 | 0.0050 | 0.133 |
| 4.0013 | 3.9745 | 0.0268 | 0.671 |
| 4.1156 | 4.1385 | −0.0229 | 0.556 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dumitriu, C.Ș.; Bărbulescu, A. Artificial Intelligence Models for the Mass Loss of Copper-Based Alloys under Cavitation. Materials 2022, 15, 6695. https://doi.org/10.3390/ma15196695
Dumitriu CȘ, Bărbulescu A. Artificial Intelligence Models for the Mass Loss of Copper-Based Alloys under Cavitation. Materials. 2022; 15(19):6695. https://doi.org/10.3390/ma15196695
Chicago/Turabian StyleDumitriu, Cristian Ștefan, and Alina Bărbulescu. 2022. "Artificial Intelligence Models for the Mass Loss of Copper-Based Alloys under Cavitation" Materials 15, no. 19: 6695. https://doi.org/10.3390/ma15196695
APA StyleDumitriu, C. Ș., & Bărbulescu, A. (2022). Artificial Intelligence Models for the Mass Loss of Copper-Based Alloys under Cavitation. Materials, 15(19), 6695. https://doi.org/10.3390/ma15196695







