Author Contributions
Conceptualization, S.-S.L.; methodology, Y.-L.H., Y.-H.C., W.-L.W., and Z.-J.P.; investigation, Y.-H.C., H.-P.W., Q.-W.S., T.H.P., C.-K.H., and Z.-J.P.; data curation, S.-S.L. and Z.-J.P.; writing—original draft preparation, S.-S.L. and Z.-J.P.; writing—review and editing, S.-S.L. and Z.-J.P.; supervision, S.-S.L.; visualization, Z.-J.P. All authors have read and agreed to the published version of the manuscript.
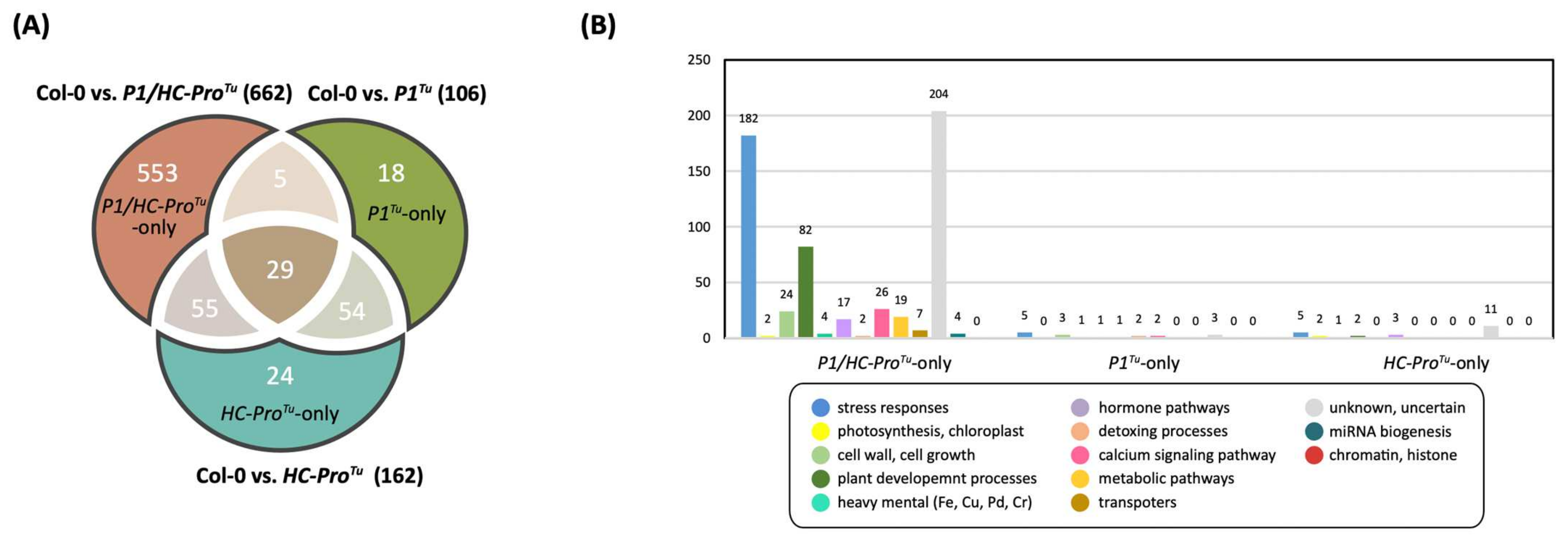
Figure 1.
Network genes among the three comparison datasets obtained from the HTP RNA-seq profiling: Col-0 vs. P1/HC-ProTu, Col-0 vs. P1Tu, and Col-0 vs. HC-ProTu comparative sets. (A) Venn diagram showing the distributions of shared and unique network genes. (B) Functional classification of unique genes in the P1Tu-only, HC-ProTu-only, and P1/HC-ProTu-only sections.
Figure 1.
Network genes among the three comparison datasets obtained from the HTP RNA-seq profiling: Col-0 vs. P1/HC-ProTu, Col-0 vs. P1Tu, and Col-0 vs. HC-ProTu comparative sets. (A) Venn diagram showing the distributions of shared and unique network genes. (B) Functional classification of unique genes in the P1Tu-only, HC-ProTu-only, and P1/HC-ProTu-only sections.
Figure 2.
The P1/HC-ProTu altered ABA-mediated immune responses. (A) P1/HC-ProTu-only genes in the P1/HC-ProTu-only section of the HTP datasets that were functionally annotated to categories and subcategories of genes involved in (i) ABA biosynthesis, (ii) catabolism, (iii) transport, (iv) signaling regulation, (v) posttranscriptional regulation, (vi) calcium signaling, and (vii) ABA responses. (B) Heatmap showing the expression patterns of the genes in the P1/HC-ProTu-only section that are involved in the ABA signaling pathway. ‘+’ and ‘−’ represent genes that positively or negatively regulate the ABA signaling pathway. The symbols in the right panel indicate the functions of genes in various biotic and abiotic stress responses and developmental processes.
Figure 2.
The P1/HC-ProTu altered ABA-mediated immune responses. (A) P1/HC-ProTu-only genes in the P1/HC-ProTu-only section of the HTP datasets that were functionally annotated to categories and subcategories of genes involved in (i) ABA biosynthesis, (ii) catabolism, (iii) transport, (iv) signaling regulation, (v) posttranscriptional regulation, (vi) calcium signaling, and (vii) ABA responses. (B) Heatmap showing the expression patterns of the genes in the P1/HC-ProTu-only section that are involved in the ABA signaling pathway. ‘+’ and ‘−’ represent genes that positively or negatively regulate the ABA signaling pathway. The symbols in the right panel indicate the functions of genes in various biotic and abiotic stress responses and developmental processes.
Figure 3.
Endogenous ABA detection and ABA sensitivity assay. (A) Determination of the endogenous ABA amounts. The mean values ± SD were obtained from three biological repeats. Comparisons between two groups were performed using a Student’s t test. *** p < 0.001. (B) ABA sensitivity assay. The germination phenotypes were classified into four phases: I, rupture of the seed coat; II, radicle protrusion; III, fully-opened cotyledons; and IV, true leaf development.
Figure 3.
Endogenous ABA detection and ABA sensitivity assay. (A) Determination of the endogenous ABA amounts. The mean values ± SD were obtained from three biological repeats. Comparisons between two groups were performed using a Student’s t test. *** p < 0.001. (B) ABA sensitivity assay. The germination phenotypes were classified into four phases: I, rupture of the seed coat; II, radicle protrusion; III, fully-opened cotyledons; and IV, true leaf development.
Figure 4.
Calcium signaling pathway in response to P1/HC-ProTu. (A) Calcium signaling pathway in the P1/HC-ProTu-only section of the HTP profiles in the Col-0 vs. P1/HC-ProTu, Col-0 vs. P1Tu, and Col-0 vs. HC-ProTu comparison sets obtained from the HTP profiles. (B) Heatmap showing the expression patterns of the P1/HC-ProTu-only genes involved in the calcium signaling pathway.
Figure 4.
Calcium signaling pathway in response to P1/HC-ProTu. (A) Calcium signaling pathway in the P1/HC-ProTu-only section of the HTP profiles in the Col-0 vs. P1/HC-ProTu, Col-0 vs. P1Tu, and Col-0 vs. HC-ProTu comparison sets obtained from the HTP profiles. (B) Heatmap showing the expression patterns of the P1/HC-ProTu-only genes involved in the calcium signaling pathway.
Figure 5.
Phenotypes of the P1/HC-ProTu and P1/HC-ProZy plants. Ten-day-old seedlings of (A) Col-0, (B) P1/HC-ProZy, and (C) P1/HC-ProTu transgenic plants. Scale bars = 0.5 cm.
Figure 5.
Phenotypes of the P1/HC-ProTu and P1/HC-ProZy plants. Ten-day-old seedlings of (A) Col-0, (B) P1/HC-ProZy, and (C) P1/HC-ProTu transgenic plants. Scale bars = 0.5 cm.
Figure 6.
The qRT-PCR-based validation of the gene expressions in the P1/HC-Pro-related plants obtained from the HTP profiles. (A–D) DEGs in the ABA signaling pathway. (E–G) DEGs in the Ca2+ signaling pathway. The mean values ± SD were from three biological repeats. Comparisons between two groups were performed with Student’s t test. * p < 0.05, ** p < 0.01, *** p < 0.001.
Figure 6.
The qRT-PCR-based validation of the gene expressions in the P1/HC-Pro-related plants obtained from the HTP profiles. (A–D) DEGs in the ABA signaling pathway. (E–G) DEGs in the Ca2+ signaling pathway. The mean values ± SD were from three biological repeats. Comparisons between two groups were performed with Student’s t test. * p < 0.05, ** p < 0.01, *** p < 0.001.
Figure 7.
Regulatory mechanism controlling stomatal dynamics in response to P1/HC-ProTu. (A) Genes involved in the regulation of stomatal guard cell dynamics and development found in the P1/HC-ProTu-only section. The genes involved in stomatal closure, stomatal opening, starch degradation, and stomatal development are labeled in blue, red, green, and brown, respectively. (B) Heatmap showing the expression patterns of the P1/HC-ProTu-only genes involved in the stomatal closure, opening, and development.
Figure 7.
Regulatory mechanism controlling stomatal dynamics in response to P1/HC-ProTu. (A) Genes involved in the regulation of stomatal guard cell dynamics and development found in the P1/HC-ProTu-only section. The genes involved in stomatal closure, stomatal opening, starch degradation, and stomatal development are labeled in blue, red, green, and brown, respectively. (B) Heatmap showing the expression patterns of the P1/HC-ProTu-only genes involved in the stomatal closure, opening, and development.
Figure 8.
Heatmaps showing the expression patterns of (A) cold response- and (B) senescence-related genes in the P1/HC-ProTu-only section of the HTP datasets.
Figure 8.
Heatmaps showing the expression patterns of (A) cold response- and (B) senescence-related genes in the P1/HC-ProTu-only section of the HTP datasets.
Figure 9.
Comparison of gene expression and networks obtained with the HTP and LTP profiles. (A) PCA plot of gene expression profiles from the HTP and LTP datasets of Col-0, P1Tu, HC-ProTu, and P1/HC-ProTu samples. The top two principal components (X-axis, PC1; Y-axis, PC2) are shown. Each data point represents one biological sample. (B) Venn diagram showing the shared and unique genes in the Col-0 vs. P1/HC-ProTu comparison samples between the HTP and LTP datasets. Network-based comparison of the 75 shared genes in the Col-0 vs. P1/HC-ProTu datasets between (C) the HTP and (D) the LTP datasets. The red and green lines represent positive and negative correlations, respectively. The dark red and green lines indicate conserved connections between the HTP and LTP networks, respectively.
Figure 9.
Comparison of gene expression and networks obtained with the HTP and LTP profiles. (A) PCA plot of gene expression profiles from the HTP and LTP datasets of Col-0, P1Tu, HC-ProTu, and P1/HC-ProTu samples. The top two principal components (X-axis, PC1; Y-axis, PC2) are shown. Each data point represents one biological sample. (B) Venn diagram showing the shared and unique genes in the Col-0 vs. P1/HC-ProTu comparison samples between the HTP and LTP datasets. Network-based comparison of the 75 shared genes in the Col-0 vs. P1/HC-ProTu datasets between (C) the HTP and (D) the LTP datasets. The red and green lines represent positive and negative correlations, respectively. The dark red and green lines indicate conserved connections between the HTP and LTP networks, respectively.
Figure 10.
Heatmaps showing the expression patterns of the 75 overlapped genes in the Col-0 vs. P1/HC-ProTu comparison identified from both the HTP and LTP profiles.
Figure 10.
Heatmaps showing the expression patterns of the 75 overlapped genes in the Col-0 vs. P1/HC-ProTu comparison identified from both the HTP and LTP profiles.
Figure 11.
Network genes among the three comparison sets obtained from the LTP profiles: Col-0 vs. P1/HC-ProTu, Col-0 vs. P1Tu, and Col-0 vs. HC-ProTu comparison sets. (A) Venn diagram showing the distributions of shared and unique network genes. (B) Functional classification of unique genes in the P1Tu-only, HC-ProTu-only, and P1/HC-ProTu-only sections.
Figure 11.
Network genes among the three comparison sets obtained from the LTP profiles: Col-0 vs. P1/HC-ProTu, Col-0 vs. P1Tu, and Col-0 vs. HC-ProTu comparison sets. (A) Venn diagram showing the distributions of shared and unique network genes. (B) Functional classification of unique genes in the P1Tu-only, HC-ProTu-only, and P1/HC-ProTu-only sections.
Table 1.
Identification of the HTP and LTP network genes in the Col-0 vs. P1/HC-ProTu, Col-0 vs. P1Tu, and Col-0 vs. HC-ProTu comparative datasets in the ContigViews system.
Table 1.
Identification of the HTP and LTP network genes in the Col-0 vs. P1/HC-ProTu, Col-0 vs. P1Tu, and Col-0 vs. HC-ProTu comparative datasets in the ContigViews system.
| | HTP | LTP |
|---|
| | Col-0 vs. P1/HC-ProTu | Col-0 vs. P1Tu | Col-0 vs. HC-ProTu | Col-0 vs. P1/HC-ProTu | Col-0 vs. P1Tu | Col-0 vs. HC-ProTu |
|---|
| Passing rate | 80 | 80 | 80 | 80 | 80 | 80 |
| Fold change | 2 | 2 | 2 | 2 | 2 | 2 |
| (+) correlation | 97.5 | 97.5 | 97.5 | 95 | 95 | 95 |
| (−) correlation | 92.5 | 92.5 | 92.5 | 90 | 90 | 90 |
| Sample sets | 10/12 | 10/12 | 10/12 | 10/12 | 10/12 | 10/12 |
| Filtered DEGs | 1601 | 559 | 777 | 700 | 621 | 587 |
| Network genes | 662 | 106 | 162 | 188 | 243 | 193 |
Table 2.
List of genes involved in the ABA signaling pathway that were identified in the P1/HC-ProTu-only section of the HTP profiles.
Table 2.
List of genes involved in the ABA signaling pathway that were identified in the P1/HC-ProTu-only section of the HTP profiles.
| Categories | Subcategories | AGI Locus Code | Gene Name | Annotations |
|---|
| Homeostasis | Biosynthesis | AT3G02570 | PMI1/MEE31 | Mannose-6-phosphate isomerase, type I |
| AT3G51895 | SULTR3;1 | sulfate transporter 3;1 |
| Catabolism | AT5G45340 | CYP707A3 | Cytochrome P450, family 707, subfamily A, polypeptide 3 |
| Transport | AT5G52050 | DTX50 | MATE efflux family protein |
| Regulation | Posttranscriptional regulation | AT5G66070 | ATL27/ATARRE | RING/U-box superfamily protein |
| AT3G52450 | PUB22 | Plant U-box 22 |
| AT2G35930 | PUB23 | Plant U-box 23 |
| AT3G46620 | RDUF1 | Zinc finger (C3HC4-type RING finger) family protein |
| AT5G59550 | RDUF2/BTL09 | Zinc finger (C3HC4-type RING finger) family protein |
| Transcriptional regulation | AT3G61890 | HB12 | Homeobox 12 |
| ABA-responses | Biotic | AT1G69480 | EXS | EXS (ERD1/XPR1/SYG1) family protein |
| AT1G32640 | MYC2 | Basic helix-loop-helix (bHLH) DNA-binding family protein |
| AT2G32030 | NA a | Acyl-CoA N-acyltransferases (NAT) superfamily protein |
| AT2G19810 | OZF1/TZF2 | CCCH-type zinc finger family protein |
| Cold | AT4G25470 | CBF2 | C-repeat/DRE binding factor 2 |
| Cold | AT5G61600 | ERF104 | Ethylene response factor 104 |
| Cold | AT5G51190 | ERF105 | Integrase-type DNA-binding superfamily protein |
| Cold, Development | AT1G74670 | GASA6 | Gibberellin-regulated family protein |
| Development | AT1G56170 | NF-YC2/HAP5B | Nuclear factor Y, subunit C2 |
| Development | AT5G23220 | NIC3 | Nicotinamidase 3 |
| Development | AT5G42050 | NRP | DCD (Development and Cell Death) domain protein, NRP has a positive role in ABA-mediated seed germination. |
| Development, Drought | AT1G13260 | RAV1 | Related to ABI3/VP1 1 |
| Development, Drought | AT5G39760 | HB23 | Homeobox protein 23 |
| Drought | AT2G46510 | AIB/BHLH017 | ABA-inducible BHLH-type transcription factor |
| Drought | AT2G25900 | ATCTH/TZF1 | Zinc finger C-x8-C-x5-C-x3-H type family protein |
| Drought | AT1G74930 | ERF018/ORA47 | Integrase-type DNA-binding superfamily protein |
| Drought | AT5G62470 | MYB96 | MYB domain protein 96 |
| Drought | AT5G67300 | MYBR1/MYB44 | MYB domain protein r1 |
| Drought | AT3G56880 | NA a | VQ motif-containing protein |
| Drought | AT1G54160 | NF-YA5 | Nuclear factor Y, subunit A5 |
| Drought | AT2G39800 | P5CS1 | Delta1-pyrroline-5-carboxylate synthase 1 |
| Drought | AT3G11820 | SYR1/SYP121 | Syntaxin of plants 121 |
| Drought | AT2G46400 | WRKY46 | WRKY DNA-binding protein 46 |
| Drought, Cold | AT5G65300 | SUPA | Unknown protein |
| Drought, Cold | AT1G72240 | SUPA-like | Unknown protein |
| Drought, Senescence | AT1G08920 | ESL1 | ERD (early response to dehydration) six-like 1 |
| Drought, Senescence, Chlorophyll degradation | AT1G69490 | ANAC029/NAP | NAC-like, activated by AP3/PI |
| Multiple stress | AT3G48360 | BT2 | BTB and TAZ domain protein 2 |
| Senescence | AT1G53170 | ERF8 | Ethylene response factor 8 |
| Senescence, Chlorophyll degradation | AT4G24390 | AFB4 | RNI-like superfamily protein |
| Uncertain | AT3G54200 | NHL39 | Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family |
Table 3.
List of genes in the calcium signaling pathway found in the P1/HC-ProTu-only section of the HTP profiles.
Table 3.
List of genes in the calcium signaling pathway found in the P1/HC-ProTu-only section of the HTP profiles.
| Categories | Subcategories | AGI Locus Code | Gene Name | Annotations |
|---|
| Transporter | Uniporter | AT1G27770 | ACA1 | Autoinhibited Ca2+-ATPase 1 |
| AT2G24610 | CNGC14 | Cyclic nucleotide-gated channel 14 |
| AT5G15410 | CNGC2/DND1 | Cyclic nucleotide-regulated ion channel family protein |
| AT5G17850 | CCX2 | Sodium/calcium exchanger family protein |
| AT5G66650 | CMCU | Protein of unknown function (DUF607) |
| Sensor | CaMs/CaMLs | AT1G66400 | CML23 | Calmodulin like 23 |
| AT1G76650 | CML38 | Calmodulin-like 38 |
| AT2G43290 | CML5/MSS3 | Calcium-binding EF-hand family protein |
| AT3G22930 | CML11 | Calmodulin-like 11 |
| AT5G37770 | CML24/TCH2 | EF hand calcium-binding protein family |
| AT5G39670 | CML46 | Calcium-binding EF-hand family protein |
| Other calcium-binding proteins | AT3G01830 | NA a | Calcium-binding EF-hand family protein |
| AT3G16510 | CaLB | Calcium-dependent lipid-binding (CaLB domain) family protein |
| AT3G25600 | NA a | Calcium-binding EF-hand family protein |
| AT4G34150 | CaLB | Calcium-dependent lipid-binding (CaLB domain) family protein |
| Calmodulin-binding proteins | AT2G26190 | IQM4 | Calmodulin-binding family protein |
| AT2G41010 | CAMBP25 | Calmodulin (CAM)-binding protein of 25 kDa |
| AT3G13600 | NA a | Calmodulin-binding family protein |
| AT3G16490 | IQD26 | IQ-domain 26 |
| AT5G26920 | CBP60G | Cam-binding protein 60-like G |
| AT5G35670 | IQD33 | IQ-domain 33 |
| AT5G57010 | NA a | Calmodulin-binding family protein |
| AT5G62570 | CBP60A | Calmodulin binding protein-like |
| CDPKs/CIPK | AT2G30360 | CIPK11/PKS5 | SOS3-interacting protein 4 |
| AT3G57530 | CPK32 | Calcium-dependent protein kinase 32 |
| AT5G66210 | CPK28 | Calcium-dependent protein kinase 28 |
Table 4.
List of genes related to drought responses found in the P1/HC-ProTu-only section of the HTP profiles.
Table 4.
List of genes related to drought responses found in the P1/HC-ProTu-only section of the HTP profiles.
| AGI | Gene Name | Annotations | Categories b | ABA/Ca c |
|---|
| AT2G39800 | P5CS1 | Delta1-pyrroline-5-carboxylate synthase 1 | drought/stomata | ABA |
| AT3G46620 | RDUF1 | Zinc finger (C3HC4-type RING finger) family protein | drought/stomata | ABA |
| AT5G59550 | RDUF2/BTL09 | Zinc finger (C3HC4-type RING finger) family protein | drought/stomata | ABA |
| AT3G11820 | SYR1/SYP121 | Syntaxin of plants 121 | drought/stomata | ABA |
| AT2G46400 | WRKY46 | WRKY DNA-binding protein 46 | drought/stomata | ABA |
| AT1G32640 | MYC2 | Basic helix-loop-helix (bHLH) DNA-binding family protein | drought/stomata | ABA |
| AT3G57230 | AGL16 | AGAMOUS-like 16 | drought/stomata | |
| AT3G23920 | BAM1 | Beta-amylase 1 | drought/stomata | |
| AT1G34245 | EPF2 | Putative membrane lipoprotein | drought/stomata | |
| AT5G10720 | HK5 | Histidine kinase 5 | drought/stomata | |
| AT5G22920 | RZPF34/CHYR1 | CHY-type/CTCHY-type/RING-type Zinc finger protein | drought/stomata | |
| AT3G52400 | SYP122 | Syntaxin of plants 122 | drought/stomata | |
| AT1G74950 | TIFY10B/JAZ2 | TIFY domain/Divergent CCT motif family protein | drought/stomata | |
| AT1G80080 | TMM | Leucine-rich repeat (LRR) family protein | drought/stomata | |
| AT1G52890 | ANAC019/NAC019 | NAC domain containing protein 19 | drought/stomata | |
| AT1G69490 | ANAC029/NAP | NAC-like, activated by AP3/PI | drought | ABA |
| AT2G25900 | ATCTH/TZF1 | Zinc finger C-x8-C-x5-C-x3-H type family protein | drought | ABA |
| AT5G45340 | CYP707A3 | Cytochrome P450, family 707, subfamily A, polypeptide 3 | drought | ABA |
| AT5G52050 | DTX50 | MATE efflux family protein | drought | ABA |
| AT1G74930 | ERF018/ORA47 | Integrase-type DNA-binding superfamily protein | drought | ABA |
| AT1G08920 | ESL1 | ERD (early response to dehydration) six-like 1 | drought | ABA |
| AT5G39760 | HB23 | Homeobox protein 23 | drought | ABA |
| AT5G62470 | MYB96 | MYB domain protein 96 | drought | ABA |
| AT5G67300 | MYBR1/MYB44 | MYB domain protein r1 | drought | ABA |
| AT1G54160 | NF-YA5 | Nuclear factor Y, subunit A5 | drought | ABA |
| AT5G42050 | NRP | DCD (Development and Cell Death) domain protein | drought | ABA |
| AT3G52450 | PUB22 | Plant U-box 22 | drought | ABA |
| AT2G35930 | PUB23 | Plant U-box 23 | drought | ABA |
| AT1G13260 | RAV1 | Related to ABI3/VP1 1 | drought | ABA |
| AT5G65300 | SUPA | Unknown protein | drought | ABA |
| AT2G41010 | CAMBP25 | Calmodulin (CAM)-binding protein of 25 kDa | drought | Ca |
| AT2G26190 | IQM4 | Calmodulin-binding family protein | drought | Ca |
| AT5G17850 | CCX2 | Sodium/calcium exchanger family protein | drought | Ca |
| AT5G66650 | CMCU | Protein of unknown function (DUF607) | drought | Ca |
| AT2G33860 | ARF3/ETT | Transcriptional factor B3 family protein/auxin-responsive factor AUX/IAA-related | drought | |
| AT4G02200 | AtDi19-5 | Drought-responsive family protein | drought | |
| AT3G08760 | ATSIK | Protein kinase superfamily protein | drought | |
| AT5G49450 | BZIP1 | Basic leucine-zipper 1 | drought | |
| AT4G36880 | CP1/RDL1 | Cysteine proteinase1 | drought | |
| AT5G04340 | CZF2/ZAT6 | Zinc finger of Arabidopsis thaliana 6 | drought | |
| AT5G04760 | DIV2 | Duplicated homeodomain-like superfamily protein | drought | |
| AT1G73330 | DR4 | Drought-repressed 4 | drought | |
| AT5G05410 | DREB2A | DRE-binding protein 2A | drought | |
| AT2G45180 | DRN1 | Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein | drought | |
| AT4G17490 | ERF6 | Ethylene responsive element binding factor 6 | drought | |
| AT2G47180 | GOLS1 | Galactinol synthase 1 | drought | |
| AT4G18880 | HSF A4A | Heat shock transcription factor A4A | drought | |
| AT5G12030 | HSP17.6A | Heat shock protein 17.6A | drought | |
| AT4G02410 | LPK1/LECRK-IV.3 | Concanavalin A-like lectin protein kinase family protein | drought | |
| AT3G49580 | LSU1 | Response to low sulfur 1 | drought | |
| AT1G73500 | MKK9 | MAP kinase kinase 9 | drought | |
| AT3G45640 | MPK3 | Mitogen-activated protein kinase 3 | drought | |
| AT5G61290 | NA a | Flavin-binding monooxygenase family protein | drought | |
| AT3G27150 | NA a | Galactose oxidase/kelch repeat superfamily protein | drought | |
| AT5G26260 | NA a | TRAF-like family protein | drought | |
| AT4G33070 | PDC1 | Thiamine pyrophosphate dependent pyruvate decarboxylase family protein | drought | |
| AT3G62260 | PP2C49 | Protein phosphatase 2C family protein | drought | |
| AT5G64905 | PROPEP3 | Elicitor peptide 3 precursor | drought | |
| AT3G18710 | PUB29 | plant U-box 29 | drought | |
| AT3G49810 | PUB30 | ARM repeat superfamily protein | drought | |
| AT1G68840 | RAV2/TEM2 | Related to ABI3/VP1 2 | drought | |
| AT3G08720 | S6K2 | Serine/threonine protein kinase 2 | drought | |
| AT5G62520 | SRO5 | Similar to RCD one 5 | drought | |
| AT3G55980 | SZF1/TZF11 | Salt-inducible zinc finger 1 | drought | |
| AT2G40140 | SZF2 | Zinc finger (CCCH-type) family protein | drought | |
| AT1G27730 | ZAT10 | Salt tolerance zinc finger | drought | |
Table 5.
List of genes related to the cold response found in the P1/HC-ProTu-only section of the HTP profiles.
Table 5.
List of genes related to the cold response found in the P1/HC-ProTu-only section of the HTP profiles.
| AGI | Gene Name | Annotations | Categories a | ABA/Ca b |
|---|
| AT5G61600 | ERF104 | Ethylene response factor 104 | cold | ABA |
| AT5G51190 | ERF105 | Integrase-type DNA-binding superfamily protein | cold | ABA |
| AT1G13260 | RAV1 | Related to ABI3/VP1 1 | cold/drought | ABA |
| AT4G25470 | CBF2 | C-repeat/DRE binding factor 2 | cold | ABA/Ca |
| AT3G50260 | CEJ1/DEAR1 | Cooperatively regulated by ethylene and jasmonate 1 | cold | Ca |
| AT5G59820 | ZAT12 | C2H2-type zinc finger family protein | cold | Ca |
| AT1G27730 | ZAT10 | Salt tolerance zinc finger | cold/drought | Ca |
| AT5G05410 | DREB2A | DRE-binding protein 2A | cold/drought | |
| AT4G17490 | ERF6 | Ethylene responsive element binding factor 6 | cold/drought | |
| AT2G47180 | GOLS1 | Galactinol synthase 1 | cold/drought | |
| AT3G45640 | MPK3 | Mitogen-activated protein kinase 3 | cold/drought | |
| AT3G08720 | S6K2 | Serine/threonine protein kinase 2 | cold/drought | |
| AT1G20823 | ATL80 | RING/U-box superfamily protein | cold | |
| AT1G18740 | B1L | Protein of unknown function (DUF793) | cold | |
| AT5G20230 | BCB | Blue-copper-binding protein | cold | |
| AT3G13310 | DJC66 | Chaperone DnaJ-domain superfamily protein | cold | |
| AT3G49530 | NAC062/NTL6 | NAC domain containing protein 62 | cold | |
| AT1G09070 | SRC2 | Soybean gene regulated by cold-2 | cold | |
| AT1G72940 | TN11 | Toll-Interleukin-Resistance (TIR) domain-containing protein | cold | |
| AT5G57560 | XTH22/TCH4 | Xyloglucan endotransglucosylase/hydrolase family protein | cold | |
Table 6.
List of genes related to senescence, cell death, and chlorophyll degradation found in the P1/HC-ProTu-only section of the HTP datasets.
Table 6.
List of genes related to senescence, cell death, and chlorophyll degradation found in the P1/HC-ProTu-only section of the HTP datasets.
| AGI | Gene Name | Annotations | Categories b |
|---|
| AT5G42050 | NRP | DCD (Development and Cell Death) domain protein | cell death |
| AT3G25250 | AGC2-1/OXI1 | AGC (cAMP-dependent, cGMP-dependent and protein kinase C) kinase family protein | cell death |
| AT5G18270 | ANAC087 | Arabidopsis NAC domain containing protein 87 | cell death |
| AT1G29690 | CAD1/NSL2 | MAC/Perforin domain-containing protein | cell death |
| AT5G58430 | EXO70B1 | Exocyst subunit exo70 family protein B1 | cell death |
| AT1G59910 | FORMIN7 | Actin-binding FH2 (formin homology 2) family protein | cell death |
| AT1G70830 | MLP28 | MLP-like protein 28 | cell death |
| AT4G12720 | NUDT7 | MutT/nudix family protein | cell death |
| AT1G08920 | ESL1 | ERD (early response to dehydration) six-like 1 | senescence |
| AT2G46400 | WRKY46 | WRKY DNA-binding protein 46 | senescence |
| AT5G02760 | APD7 | Protein phosphatase 2C family protein | senescence |
| AT5G08790 | ATAF2/ANAC081 | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | senescence |
| AT1G44830 | ERF014/EPI1 | Integrase-type DNA-binding superfamily protein | senescence |
| AT1G51660 | MKK4 | Mitogen-activated protein kinase kinase 4 | senescence |
| AT1G67470 | NA a | Protein kinase superfamily protein | senescence |
| AT2G28400 | NA a | Protein of unknown function, DUF584 | senescence |
| AT4G21930 | NA a | Protein of unknown function, DUF584 | senescence |
| AT2G44670 | NA a | Protein of unknown function, DUF581 | senescence |
| AT4G04630 | NA a | Protein of unknown function, DUF584 | senescence |
| AT1G55740 | SIP1 | Seed imbibition 1 | senescence |
| AT4G01250 | WRKY22 | WRKY family transcription factor | senescence |
| AT3G21520 | DMP1 | DUF679 domain membrane protein 1 | senescence/cell death |
| AT1G52890 | ANAC019/NAC019 | NAC domain containing protein 19 | senescence/chloroplast degradation |
| AT1G69490 | ANAC029/NAP | NAC-like, activated by AP3/PI | senescence/chloroplast degradation |
| AT4G24390 | AFB4 | RNI-like superfamily protein | senescence/chloroplast degradation |
Table 7.
Statistics of the RNA-seq data and read mapping rates of the Col-0, P1Tu, HC-ProTu, and P1/HC-ProTulibraries obtained with the HTP and LTP profiles.
Table 7.
Statistics of the RNA-seq data and read mapping rates of the Col-0, P1Tu, HC-ProTu, and P1/HC-ProTulibraries obtained with the HTP and LTP profiles.
| Samples a | Read Length
(bp) | Raw Reads | Clear Reads | Mapped Rates
(% of Total) |
|---|
| HTP | Col-0-1 | 125 | 23,692,678 | 23,692,678 | 82.09 |
| Col-0-2 | 125 | 23,536,690 | 23,536,690 | 81.41 |
| Col-0-3 | 125 | 24,361,660 | 24,361,660 | 84.04 |
| P1Tu-1 | 125 | 24,270,634 | 24,270,634 | 79.83 |
| P1Tu-2 | 125 | 23,727,160 | 23,727,160 | 79.97 |
| P1Tu-3 | 125 | 24,663,280 | 24,663,280 | 79.54 |
| HC-ProTu-1 | 125 | 23,502,832 | 23,502,832 | 77.45 |
| HC-ProTu-2 | 125 | 23,074,688 | 23,074,688 | 80.87 |
| HC-ProTu-3 | 125 | 23,817,246 | 23,817,246 | 79.23 |
| P1/HC-ProTu-1 | 125 | 23,104,480 | 23,104,480 | 75.27 |
| P1/HC-ProTu-2 | 125 | 23,766,692 | 23,766,692 | 73.56 |
| P1/HC-ProTu-3 | 125 | 21,935,512 | 21,935,512 | 77.24 |
| LTP | Col-0-1 | 75 | 2,732,672 | 2,611,636 | 78.17 |
| Col-0-2 | 75 | 3,027,192 | 2,895,782 | 78.97 |
| Col-0-3 | 75 | 2,974,324 | 2,844,916 | 79.84 |
| P1Tu-1 | 75 | 3,072,748 | 2,899,822 | 80.16 |
| P1Tu-2 | 75 | 3,100,226 | 2,956,540 | 79.33 |
| P1Tu-3 | 75 | 2,309,898 | 2,151,492 | 80.63 |
| HC-ProTu-1 | 75 | 2,300,776 | 2,196,490 | 77.28 |
| HC-ProTu-2 | 75 | 2,814,492 | 2,664,142 | 78.34 |
| HC-ProTu-3 | 75 | 1,967,520 | 1,888,408 | 77.94 |
| P1/HC-ProTu-1 | 75 | 2,738,024 | 2,609,924 | 76.81 |
| P1/HC-ProTu-2 | 75 | 3,012,322 | 2,866,212 | 77.07 |
| P1/HC-ProTu-3 | 75 | 2,973,652 | 2,817,066 | 77.53 |
Table 8.
List of 75 overlapping genes in the P1/HC-ProTu-only section identified from both the HTP and LTP profiles.
Table 8.
List of 75 overlapping genes in the P1/HC-ProTu-only section identified from both the HTP and LTP profiles.
| AGI | Gene Name | Annotation | Functional Classification b | ABA/Ca c | Stresses d |
|---|
| AT1G48410 | AGO1 | Stabilizer of iron transporter SufD/Polynucleotidyl transferase | ▇ | | |
| AT1G31280 | AGO2 | Argonaute family protein | ▇ | | |
| AT4G12720 | NUDT7 | MutT/nudix family protein | ▇ | | cell death |
| AT3G49530 | NAC062/NTL6 | NAC domain containing protein 62 | ▇ | | cold |
| AT5G57560 | XTH22/TCH4 | Xyloglucan endotransglucosylase/hydrolase family protein | ▇ | | cold |
| AT5G67300 | MYBR1/MYB44 | MYB domain protein r1 | ▇ | ABA | drought |
| AT3G08760 | ATSIK | Protein kinase superfamily protein | ▇ | | drought |
| AT1G73330 | DR4 | Drought-repressed 4 | ▇ | | drought |
| AT4G18880 | HSF A4A | Heat shock transcription factor A4A | ▇ | | drought |
| AT3G49810 | PUB30 | ARM repeat superfamily protein | ▇ | | drought |
| AT3G08720 | S6K2 | Serine/threonine protein kinase 2 | ▇ | | drought, cold |
| AT1G51660 | MKK4 | mitogen-activated protein kinase kinase 4 | ▇ | | senescence |
| AT4G01250 | WRKY22 | WRKY family transcription factor | ▇ | | senescence |
| AT3G48360 | BT2 | BTB and TAZ domain protein 2 | ▇ | ABA | |
| AT1G33560 | ADR1 | Disease resistance protein (CC-NBS-LRR class) family | ▇ | | |
| AT1G08830 | CSD1 | Copper/zinc superoxide dismutase 1 | ▇ | | |
| AT2G28190 | CSD2 | Copper/zinc superoxide dismutase 2 | ▇ | | |
| AT5G41750 | DM1 | Disease resistance protein (TIR-NBS-LRR class) family | ▇ | | |
| AT5G41080 | GDPD2 | PLC-like phosphodiesterases superfamily protein | ▇ | | |
| AT1G25550 | HHO3/NIGT1.1 | MYB-like transcription factor family protein | ▇ | | |
| AT3G10020 | HUP26 | Unknown protein; response to oxidative stress, anaerobic respiration | ▇ | | |
| AT1G17420 | LOX3 | Lipoxygenase 3 | ▇ | | |
| AT1G72520 | LOX4 | PLAT/LH2 domain-containing lipoxygenase family protein | ▇ | | |
| AT2G01180 | LPP1 | Phosphatidic acid phosphatase 1 | ▇ | | |
| AT1G01560 | MPK11 | MAP kinase 11 | ▇ | | |
| AT4G37260 | MYB73 | MYB domain protein 73 | ▇ | | |
| AT1G66090 | TIR-NBS-LRR | Disease resistance protein (TIR-NBS class) | ▇ | | |
| AT5G04720 | PHX21/ADR1-L2 | ADR1-like 2 | ▇ | | |
| AT4G17230 | SCL13 | SCARECROW-like 13 | ▇ | | |
| AT1G19020 | SDA1/HUP35 | Unknown protein | ▇ | | |
| AT2G23810 | TET8 | Tetraspanin8 | ▇ | | |
| AT2G33770 | UBC24/PHO2 | Phosphate 2 | ▇ | | |
| AT1G56250 | VBF/PP2-B14 | Phloem protein 2-B14 | ▇ | | |
| AT1G12520 | CCS | Copper chaperone for SOD1 | ▇ | | |
| AT1G70830 | MLP28 | MLP-like protein 28 | ▇ | | cell death |
| AT5G02760 | APD7 | Protein phosphatase 2C family protein | ▇ | | senescence |
| AT1G74670 | GASA6 | Gibberellin-regulated family protein | ▇ | ABA | |
| AT5G27420 | ATL31/CNI1 | Carbon/nitrogen insensitive 1 | ▇ | | |
| AT2G22840 | GRF1 | Growth-regulating factor 1 | ▇ | | |
| AT3G05690 | NF-YA2/HAP2B | Nuclear factor Y, subunit A2 | ▇ | | |
| AT3G14020 | NF-YA6 | Nuclear factor Y, subunit A6 | ▇ | | |
| AT5G50570 | SPL13A | Squamosa promoter-binding protein-like (SBP domain) transcription factor family protein | ▇ | | |
| AT2G33810 | SPL3 | Squamosa promoter binding protein-like 3 | ▇ | | |
| AT5G60120 | TOE2 | Target of early activation tagged (EAT) 2 | ▇ | | |
| AT1G69530 | EXPA1 | Expansin A1 | ▇ | | |
| AT2G26190 | IQM4 | Calmodulin-binding family protein | ▇ | Ca | drought |
| AT4G34150 | CaLB | Calcium-dependent lipid-binding (CaLB domain) family protein | ▇ | Ca | |
| AT3G16510 | CaLB | Calcium-dependent lipid-binding (CaLB domain) family protein | ▇ | Ca | |
| AT5G26920 | CBP60G | Cam-binding protein 60-like G | ▇ | Ca | |
| AT5G37770 | CML24/TCH2 | EF hand calcium-binding protein family | ▇ | Ca | |
| AT5G66210 | CPK28 | Calcium-dependent protein kinase 28 | ▇ | Ca | |
| AT3G57530 | CPK32 | Calcium-dependent protein kinase 32 | ▇ | Ca | |
| AT1G27770 | ACA1 | Autoinhibited Ca2+-ATPase 1 | ▇ | Ca | |
| AT5G45340 | CYP707A3 | Cytochrome P450, family 707, subfamily A, polypeptide 3 | ▇ | ABA | drought |
| AT5G08790 | ATAF2/ANAC081 | NAC (No Apical Meristem) domain Transcriptional regulator superfamily protein | ▇ | | senescence |
| AT5G66070 | ATL27/ATARRE | RING/U-box superfamily protein | ▇ | ABA | |
| AT3G25780 | AOC3 | Allene oxide cyclase 3 | ▇ | | |
| AT1G28370 | ERF11 | ERF domain protein 11 | ▇ | | |
| AT4G36040 | J11 | Chaperone DnaJ-domain superfamily protein | ▇ | | |
| AT4G23180 | CRK10/RLK4 | Cysteine-rich RLK (RECEPTOR-like protein kinase) 10 | | | |
| AT3G26200 | CYP71B22 | Cytochrome P450, family 71, subfamily B, polypeptide 22 | | | |
| AT1G80610 | NAa | Unknown protein | | | |
| AT1G63750 | TIR-NBS-LRR | Disease resistance protein (TIR-NBS-LRR class) family | | | |
| AT4G32480 | NA a | Protein of unknown function (DUF506) | | | |
| AT3G15450 | NA a | Aluminium induced protein with YGL and LRDR motifs | | | |
| AT4G23870 | NA a | Unknown protein | | | |
| AT5G46780 | NA a | VQ motif-containing protein | | | |
| AT2G39650 | NA a | Protein of unknown function (DUF506) | | | |
| AT1G77270 | NA a | Unknown protein | | | |
| AT2G29300 | NA a | NAD(P)-binding Rossmann-fold superfamily protein | | | |
| AT3G07350 | NAa | Protein of unknown function (DUF506) | | | |
| AT1G52820 | NA a | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | | | |
| AT3G44400 | NA a | Disease resistance protein (TIR-NBS-LRR class) family | | | |
| AT5G44572 | PROSCOOP6 | Unknown protein | | | |
| AT1G16110 | WAKL6 | Wall associated kinase-like 6 | | | |