Metagenomic Analysis Reveals the Fate of Antibiotic Resistance Genes in a Full-Scale Wastewater Treatment Plant in Egypt
Abstract
:1. Introduction
2. Materials & Methods
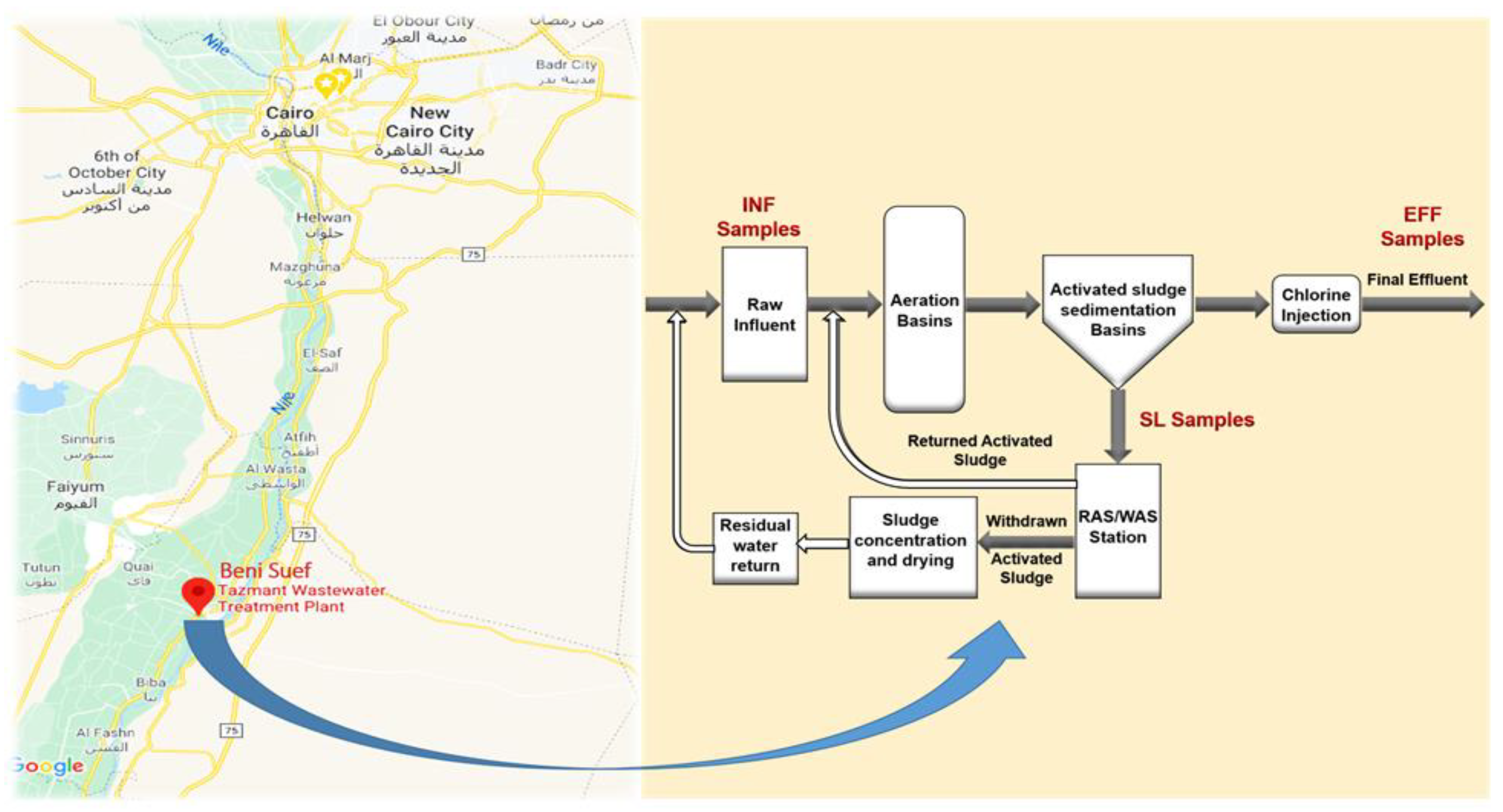
2.1. Sample Collection and Processing
2.2. DNA Extraction
2.3. Metagenomic Sequencing and Bioinformatic Analysis
3. Results
3.1. Metagenomic Sequencing
3.2. Profiling of ARGs
3.3. ARGs Persistence in Influent
4. Removal Efficiency of ARG Types and Subtypes from Influent to Effluent
5. Microbial Composition of Metagenomic Datasets
6. Discussion
7. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Versporten, A.; Bolokhovets, G.; Ghazaryan, L.; Abilova, V.; Pyshnik, G.; Spasojevic, T.; Korinteli, I.; Raka, L.; Kambaralieva, B.; Cizmovic, L. Antibiotic use in eastern Europe: A cross-national database study in coordination with the WHO Regional Office for Europe. Lancet Infect. Dis. 2014, 14, 381–387. [Google Scholar] [CrossRef]
- Pruden, A.; Larsson, D.G.J.; Amézquita, A.; Collignon, P.; Brandt, K.K.; Graham, D.W.; Lazorchak, J.M.; Suzuki, S.; Silley, P.; Snape, J.R. Management options for reducing the release of antibiotics and antibiotic resistance genes to the environment. Environ. Health Perspect. 2013, 121, 878–885. [Google Scholar] [CrossRef] [PubMed]
- Munita, J.M.; Arias, C.A. Mechanisms of antibiotic resistance. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aminov, R.I. Horizontal gene exchange in environmental microbiota. Front. Microbiol. 2011, 2, 158. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qin, K.; Wei, L.; Li, J.; Lai, B.; Zhu, F.; Yu, H.; Zhao, Q.; Wang, K. A review of ARGs in WWTPs: Sources, stressors and elimination. Chin. Chem. Lett. 2020, 31, 2603–2613. [Google Scholar] [CrossRef]
- Crini, G.; Lichtfouse, E. Wastewater Treatment: An Overview. In Adsorption-Oriented Processes Using Conventional and Non-conventional Adsorbents for Wastewater Treatment; Springer: Cham, Switzerland, 2018; pp. 1–21. ISBN 3319921118. [Google Scholar]
- Rizzo, L.; Manaia, C.; Merlin, C.; Schwartz, T.; Dagot, C.; Ploy, M.C.; Michael, I.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: A review. Sci. Total Environ. 2013, 447, 345–360. [Google Scholar] [CrossRef] [Green Version]
- Quach-Cu, J.; Herrera-Lynch, B.; Marciniak, C.; Adams, S.; Simmerman, A.; Reinke, R.A. The effect of primary, secondary, and tertiary wastewater treatment processes on antibiotic resistance gene (ARG) concentrations in solid and dissolved wastewater fractions. Water 2018, 10, 37. [Google Scholar] [CrossRef] [Green Version]
- Michael, I.; Rizzo, L.; McArdell, C.S.; Manaia, C.M.; Merlin, C.; Schwartz, T.; Dagot, C.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for the release of antibiotics in the environment: A review. Water Res. 2013, 47, 957–995. [Google Scholar] [CrossRef] [Green Version]
- Handelsman, J. Metagenomics: Application of genomics to uncultured microorganisms. Microbiol. Mol. Biol. Rev. 2004, 68, 669–685. [Google Scholar] [CrossRef] [Green Version]
- Bengtsson-Palme, J.; Larsson, D.G.J.; Kristiansson, E. Using metagenomics to investigate human and environmental resistomes. J. Antimicrob. Chemother. 2017, 72, 2690–2703. [Google Scholar] [CrossRef]
- Chen, B. Metagenomic profiles of antibiotic resis- tance genes (ARGs) between human impacted estuary and deep ocean sediments. Environ. Sci. Technol. 2013, 47, 12753–12760. [Google Scholar] [CrossRef]
- Li, B.; Yang, Y.; Ma, L.; Ju, F.; Guo, F.; Tiedje, J.M.; Zhang, T. Metagenomic and network analysis reveal wide distribution and co-occurrence of environmental antibiotic resistance genes. ISME J. 2015, 9, 2490–2502. [Google Scholar] [CrossRef] [Green Version]
- Nesme, J.; Cécillon, S.; Delmont, T.O.; Monier, J.-M.; Vogel, T.M.; Simonet, P. Large-scale metagenomic-based study of antibiotic resistance in the environment. Curr. Biol. 2014, 24, 1096–1100. [Google Scholar] [CrossRef] [Green Version]
- Yang, Y.; Li, B.; Zou, S.; Fang, H.H.P.; Zhang, T. Fate of antibiotic resistance genes in sewage treatment plant revealed by metagenomic approach. Water Res. 2014, 62, 97–106. [Google Scholar] [CrossRef]
- Guo, J.; Li, J.; Chen, H.; Bond, P.L.; Yuan, Z. Metagenomic analysis reveals wastewater treatment plants as hotspots of antibiotic resistance genes and mobile genetic elements. Water Res. 2017, 123, 468–478. [Google Scholar] [CrossRef]
- Yoo, K.; Yoo, H.; Lee, J.; Choi, E.J.; Park, J. Exploring the antibiotic resistome in activated sludge and anaerobic digestion sludge in an urban wastewater treatment plant via metagenomic analysis. J. Microbiol. 2020, 58, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Lira, F.; Vaz-Moreira, I.; Tamames, J.; Manaia, C.M.; Martínez, J.L. Metagenomic analysis of an urban resistome before and after wastewater treatment. Sci. Rep. 2020, 10, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Hultman, J.; Tamminen, M.; Pärnänen, K.; Cairns, J.; Karkman, A.; Virta, M. Host range of antibiotic resistance genes in wastewater treatment plant influent and effluent. FEMS Microbiol. Ecol. 2018, 94, fiy038. [Google Scholar] [CrossRef] [PubMed]
- Ben, W.; Wang, J.; Cao, R.; Yang, M.; Zhang, Y.; Qiang, Z. Distribution of antibiotic resistance in the effluents of ten municipal wastewater treatment plants in China and the effect of treatment processes. Chemosphere 2017, 172, 392–398. [Google Scholar] [CrossRef] [PubMed]
- Karkman, A.; Johnson, T.A.; Lyra, C.; Stedtfeld, R.D.; Tamminen, M.; Tiedje, J.M.; Virta, M. High-throughput quantification of antibiotic resistance genes from an urban wastewater treatment plant. FEMS Microbiol. Ecol. 2016, 92, fiw014. [Google Scholar] [CrossRef] [Green Version]
- Paiva, M.C.; Reis, M.P.; Costa, P.S.; Dias, M.F.; Bleicher, L.; Scholte, L.L.S.; Nardi, R.M.D.; Nascimento, A.M.A. Identification of new bacteria harboring qnrS and aac (6′)-Ib/cr and mutations possibly involved in fluoroquinolone resistance in raw sewage and activated sludge samples from a full-scale WWTP. Water Res. 2017, 110, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Bengtsson-Palme, J.; Hammaren, R.; Pal, C.; Östman, M.; Björlenius, B.; Flach, C.-F.; Fick, J.; Kristiansson, E.; Tysklind, M.; Larsson, D.G.J. Elucidating selection processes for antibiotic resistance in sewage treatment plants using metagenomics. Sci. Total Environ. 2016, 572, 697–712. [Google Scholar] [CrossRef] [PubMed]
- Neudorf, K.D.; Huang, Y.N.; Ragush, C.M.; Yost, C.K.; Jamieson, R.C.; Hansen, L.T. Antibiotic resistance genes in municipal wastewater treatment systems and receiving waters in Arctic Canada. Sci. Total Environ. 2017, 598, 1085–1094. [Google Scholar] [CrossRef] [Green Version]
- Christgen, B.; Yang, Y.; Ahammad, S.Z.; Li, B.; Rodriquez, D.C.; Zhang, T.; Graham, D.W. Metagenomics shows that low-energy anaerobic- aerobic treatment reactors reduce antibiotic resistance gene levels from domestic wastewater. Environ. Sci. Technol. 2015, 49, 2577–2584. [Google Scholar] [CrossRef]
- Marti, E.; Jofre, J.; Balcazar, J.L. Prevalence of antibiotic resistance genes and bacterial community composition in a river influenced by a wastewater treatment plant. PLoS ONE 2013, 8, e78906. [Google Scholar] [CrossRef]
- Pazda, M.; Kumirska, J.; Stepnowski, P.; Mulkiewicz, E. Antibiotic resistance genes identified in wastewater treatment plant systems—A review. Sci. Total Environ. 2019, 697, 134023. [Google Scholar] [CrossRef] [PubMed]
- Younes, H.A.; Mahmoud, H.M.; Abdelrahman, M.M.; Nassar, H.F. Seasonal occurrence, removal efficiency and associated ecological risk assessment of three antibiotics in a municipal wastewater treatment plant in Egypt. Environ. Nanotechnol. Monit. Manag. 2019, 12, 100239. [Google Scholar] [CrossRef]
- Elsaied, H.E.; Soliman, T.; Abu-Taleb, H.T.; Goto, H.; Jenke-Kodam, H. Phylogenetic characterization of eukaryotic and prokaryotic gut flora of Nile tilapia, Oreochromis niloticus, along niches of Lake Nasser, Egypt, based on rRNA gene high-throughput sequences. Ecol. Genet. Genom. 2019, 11, 100037. [Google Scholar] [CrossRef]
- Yin, X.; Jiang, X.-T.; Chai, B.; Li, L.; Yang, Y.; Cole, J.R.; Tiedje, J.M.; Zhang, T. ARGs-OAP v2. 0 with an expanded SARG database and Hidden Markov Models for enhancement characterization and quantification of antibiotic resistance genes in environmental metagenomes. Bioinformatics 2018, 34, 2263–2270. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.; Pop, M. ARDB—antibiotic resistance genes database. Nucleic Acids Res. 2009, 37, D443–D447. [Google Scholar] [CrossRef] [Green Version]
- McArthur, A.G.; Waglechner, N.; Nizam, F.; Yan, A.; Azad, M.A.; Baylay, A.J.; Bhullar, K.; Canova, M.J.; De Pascale, G.; Ejim, L. The comprehensive antibiotic resistance database. Antimicrob. Agents Chemother. 2013, 57, 3348–3357. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.; Andersen, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012, 6, 610–618. [Google Scholar] [CrossRef] [PubMed]
- Beghini, F.; McIver, L.J.; Blanco-Míguez, A.; Dubois, L.; Asnicar, F.; Maharjan, S.; Mailyan, A.; Thomas, A.M.; Manghi, P.; Valles-Colomer, M.; et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. bioRxiv 2020. [Google Scholar] [CrossRef]
- Asnicar, F.; Weingart, G.; Tickle, T.L.; Huttenhower, C.; Segata, N. Compact graphical representation of phylogenetic data and metadata with GraPhlAn. PeerJ 2015, 3, e1029. [Google Scholar] [CrossRef] [PubMed]
- Meyer, F.; Paarmann, D.; D’Souza, M.; Olson, R.; Glass, E.M.; Kubal, M.; Paczian, T.; Rodriguez, A.; Stevens, R.; Wilke, A. The metagenomics RAST server–a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinform. 2008, 9, 1–8. [Google Scholar] [CrossRef] [Green Version]
- RStudio Team. Integrated Development Environment for R. RStudio. PBC, Boston, MA. 2020. Available online: http://www.rstudio.com/ (accessed on 22 June 2021).
- Yang, Y.; Li, B.; Ju, F.; Zhang, T. Exploring variation of antibiotic resistance genes in activated sludge over a four-year period through a metagenomic approach. Environ. Sci. Technol. 2013, 47, 10197–10205. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Q.-B.; Guo, M.-T.; Yang, J. Fate of antibiotic resistant bacteria and genes during wastewater chlorination: Implication for antibiotic resistance control. PLoS ONE 2015, 10, e0119403. [Google Scholar] [CrossRef]
- Mao, D.; Yu, S.; Rysz, M.; Luo, Y.; Yang, F.; Li, F.; Hou, J.; Mu, Q.; Alvarez, P.J.J. Prevalence and proliferation of antibiotic resistance genes in two municipal wastewater treatment plants. Water Res. 2015, 85, 458–466. [Google Scholar] [CrossRef]
- Liu, Z.; Klümper, U.; Liu, Y.; Yang, Y.; Wei, Q.; Lin, J.-G.; Gu, J.-D.; Li, M. Metagenomic and metatranscriptomic analyses reveal activity and hosts of antibiotic resistance genes in activated sludge. Environ. Int. 2019, 129, 208–220. [Google Scholar] [CrossRef]
- Khalifa, H.O.; Soliman, A.M.; Ahmed, A.M.; Shimamoto, T.; Nariya, H.; Matsumoto, T.; Shimamoto, T. High prevalence of antimicrobial resistance in Gram-negative bacteria isolated from clinical settings in Egypt: Recalling for judicious use of conventional antimicrobials in developing nations. Microb. Drug Resist. 2019, 25, 371–385. [Google Scholar] [CrossRef]
- Zhang, X.-X.; Zhang, T.; Fang, H.H.P. Antibiotic resistance genes in water environment. Appl. Microbiol. Biotechnol. 2009, 82, 397–414. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhang, X.-X.; Huang, K.; Miao, Y.; Shi, P.; Liu, B.; Long, C.; Li, A. Metagenomic profiling of antibiotic resistance genes and mobile genetic elements in a tannery wastewater treatment plant. PLoS ONE 2013, 8, e76079. [Google Scholar] [CrossRef] [PubMed]
- Caucci, S.; Karkman, A.; Cacace, D.; Rybicki, M.; Timpel, P.; Voolaid, V.; Gurke, R.; Virta, M.; Berendonk, T.U. Seasonality of antibiotic prescriptions for outpatients and resistance genes in sewers and wastewater treatment plant outflow. FEMS Microbiol. Ecol. 2016, 92, fiw060. [Google Scholar] [CrossRef] [Green Version]
- Chen, B.; Hao, L.; Guo, X.; Wang, N.; Ye, B. Prevalence of antibiotic resistance genes of wastewater and surface water in livestock farms of Jiangsu Province, China. Environ. Sci. Pollut. Res. 2015, 22, 13950–13959. [Google Scholar] [CrossRef]
- Harnisz, M.; Kiedrzyńska, E.; Kiedrzyński, M.; Korzeniewska, E.; Czatzkowska, M.; Koniuszewska, I.; Jóźwik, A.; Szklarek, S.; Niestępski, S.; Zalewski, M. The impact of WWTP size and sampling season on the prevalence of antibiotic resistance genes in wastewater and the river system. Sci. Total. Environ. 2020, 741, 140466. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.-C.; Zheng, J.; Wei, Y.-Y.; Chen, T.; Dahlgren, R.A.; Shang, X.; Chen, H. Antibiotic resistance genes in an urban river as impacted by bacterial community and physicochemical parameters. Environ. Sci. Pollut. Res. 2017, 24, 23753–23762. [Google Scholar] [CrossRef]
- Ng, C.; Tan, B.; Jiang, X.-T.; Gu, X.; Chen, H.; Schmitz, B.W.; Haller, L.; Charles, F.R.; Zhang, T.; Gin, K. Metagenomic and resistome analysis of a full-scale municipal wastewater treatment plant in Singapore containing membrane bioreactors. Front. Microbiol. 2019, 10, 172. [Google Scholar] [CrossRef]
- Narciso-da-Rocha, C.; Rocha, J.; Vaz-Moreira, I.; Lira, F.; Tamames, J.; Henriques, I.; Martinez, J.L.; Manaia, C.M. Bacterial lineages putatively associated with the dissemination of antibiotic resistance genes in a full-scale urban wastewater treatment plant. Environ. Int. 2018, 118, 179–188. [Google Scholar] [CrossRef]
- Wagner, M.; Loy, A. Bacterial community composition and function in sewage treatment systems. Curr. Opin. Biotechnol. 2002, 13, 218–227. [Google Scholar] [CrossRef]
- Numberger, D.; Ganzert, L.; Zoccarato, L.; Mühldorfer, K.; Sauer, S.; Grossart, H.-P.; Greenwood, A.D. Characterization of bacterial communities in wastewater with enhanced taxonomic resolution by full-length 16S rRNA sequencing. Sci. Rep. 2019, 9, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Varela, A.R.; Nunes, O.C.; Manaia, C.M. Quinolone resistant Aeromonas spp. as carriers and potential tracers of acquired antibiotic resistance in hospital and municipal wastewater. Sci. Total Environ. 2016, 542, 665–671. [Google Scholar] [CrossRef]
- Piotrowska, M.; Przygodzińska, D.; Matyjewicz, K.; Popowska, M. Occurrence and variety of β-lactamase genes among Aeromonas spp. isolated from urban wastewater treatment plant. Front. Microbiol. 2017, 8, 863. [Google Scholar] [CrossRef] [Green Version]
- Luczkiewicz, A.; Kotlarska, E.; Artichowicz, W.; Tarasewicz, K.; Fudala-Ksiazek, S. Antimicrobial resistance of Pseudomonas spp. isolated from wastewater and wastewater-impacted marine coastal zone. Environ. Sci. Pollut. Res. 2015, 22, 19823–19834. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quintela-Baluja, M.; Abouelnaga, M.; Romalde, J.; Su, J.-Q.; Yu, Y.; Gomez-Lopez, M.; Smets, B.; Zhu, Y.-G.; Graham, D.W. Spatial ecology of a wastewater network defines the antibiotic resistance genes in downstream receiving waters. Water Res. 2019, 162, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Shankar, V.; Gouda, M.; Moncivaiz, J.; Gordon, A.; Reo, N.V.; Hussein, L.; Paliy, O. Differences in gut metabolites and microbial composition and functions between Egyptian and US children are consistent with their diets. Msystems 2017, 2, e00169-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scher, J.U.; Sczesnak, A.; Longman, R.S.; Segata, N.; Ubeda, C.; Bielski, C.; Rostron, T.; Cerundolo, V.; Pamer, E.G.; Abramson, S.B. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. elife 2013, 2, e01202. [Google Scholar] [CrossRef]
- Wen, C.; Zheng, Z.; Shao, T.; Liu, L.; Xie, Z.; Le Chatelier, E.; He, Z.; Zhong, W.; Fan, Y.; Zhang, L. Quantitative metagenomics reveals unique gut microbiome biomarkers in ankylosing spondylitis. Genome Biol. 2017, 18, 1–13. [Google Scholar]
- Pedersen, H.K.; Gudmundsdottir, V.; Nielsen, H.B.; Hyotylainen, T.; Nielsen, T.; Jensen, B.A.H.; Forslund, K.; Hildebrand, F.; Prifti, E.; Falony, G. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 2016, 535, 376–381. [Google Scholar] [CrossRef] [PubMed]
- Allen, H.K.; Donato, J.; Wang, H.H.; Cloud-Hansen, K.A.; Davies, J.; Handelsman, J. Call of the wild: Antibiotic resistance genes in natural environments. Nat. Rev. Microbiol. 2010, 8, 251–259. [Google Scholar] [CrossRef] [PubMed]







| Sample Name | Count of Raw Reads (PE-150 bp) | Count of Reads after QC Filter (PE-150 bp) | Total Read Size (bp) | Q20% | GC% | MG-RAST_ID |
|---|---|---|---|---|---|---|
| INF_W | 137,739,660 | 121,944,908 | 18,291,736,200 | 96.13% | 51.11% | mgm4754064.3 |
| INF_S | 154,503,234 | 119,535,370 | 17,930,305,500 | 97.22% | 57.24% | mgm4757477.3 |
| EFF_W | 112,815,582 | 100,211,020 | 15,031,653,000 | 96.04% | 52.68% | mgm4760706.3 |
| EFF_S | 161,601,122 | 125,803,650 | 18,870,547,500 | 97.15% | 57.27% | mgm4798629.3 |
| SL_W | 167,104,122 | 148,048,024 | 22,207,203,600 | 95.53% | 57.68% | mgm4798679.3 |
| SL_S | 167,341,294 | 126,323,822 | 18,948,573,300 | 97.27% | 60.55% | mgm4798702.3 |
| Totals | 901,105,014 | 741,866,794 | 111,280,019,100 |
| ARG_Type | Average ARG Type Abundance/16s rRNA Gene Copies | Average ARG Type Abundance Percentage | ||||
|---|---|---|---|---|---|---|
| INF | EFF | SL | INF | EFF | SL | |
| aminoglycoside | 0.49249 | 0.27827 | 0.27436 | 14.51% | 17.93% | 23.54% |
| bacitracin | 0.20776 | 0.04949 | 0.12194 | 6.12% | 3.19% | 10.46% |
| beta-lactam | 0.56731 | 0.23430 | 0.15422 | 16.72% | 15.09% | 13.23% |
| bleomycin | 0.00007 | 0.00000 | 0.00000 | 0.00% | 0.00% | 0.00% |
| chloramphenicol | 0.17374 | 0.09708 | 0.04270 | 5.12% | 6.25% | 3.66% |
| fosfomycin | 0.00345 | 0.00133 | 0.00029 | 0.10% | 0.09% | 0.03% |
| fosmidomycin | 0.03518 | 0.00625 | 0.00219 | 1.04% | 0.40% | 0.19% |
| kasugamycin | 0.03480 | 0.00659 | 0.00177 | 1.03% | 0.42% | 0.15% |
| MLS* | 0.17033 | 0.07764 | 0.10525 | 5.02% | 5.00% | 9.03% |
| multidrug | 0.72730 | 0.22423 | 0.12269 | 21.44% | 14.44% | 10.53% |
| polymyxin | 0.02261 | 0.00573 | 0.00336 | 0.67% | 0.37% | 0.29% |
| quinolone | 0.05330 | 0.02437 | 0.01018 | 1.57% | 1.57% | 0.87% |
| rifamycin | 0.01824 | 0.00736 | 0.00370 | 0.54% | 0.47% | 0.32% |
| sulfonamide | 0.18931 | 0.20277 | 0.19647 | 5.58% | 13.06% | 16.86% |
| tetracycline | 0.39135 | 0.22796 | 0.08977 | 11.53% | 14.68% | 7.70% |
| trimethoprim | 0.09795 | 0.05218 | 0.01830 | 2.89% | 3.36% | 1.57% |
| unclassified | 0.20608 | 0.05600 | 0.01498 | 6.07% | 3.61% | 1.29% |
| vancomycin | 0.00174 | 0.00079 | 0.00320 | 0.05% | 0.05% | 0.27% |
| Totals | 3.39303 | 1.55233 | 1.16537 | |||
| ARG Type | Average Abundance/16s-rRNA Gene Copies | Removal Efficiency | |
|---|---|---|---|
| Influent | Effluent | ||
| aminoglycoside | 0.49249 | 0.27827 | 97.25% |
| bacitracin | 0.20776 | 0.04949 | 98.70% |
| beta-lactam | 0.56731 | 0.23430 | 97.93% |
| bleomycin | 0.00007 | 0.00000 | 99.75% |
| chloramphenicol | 0.17374 | 0.09708 | 97.28% |
| fosfomycin | 0.00345 | 0.00133 | 98.05% |
| fosmidomycin | 0.03518 | 0.00625 | 98.97% |
| kasugamycin | 0.03480 | 0.00659 | 98.91% |
| MLS* | 0.17033 | 0.07764 | 97.74% |
| multidrug | 0.72730 | 0.22423 | 98.39% |
| polymyxin | 0.02261 | 0.00573 | 98.63% |
| quinolone | 0.05330 | 0.02437 | 97.73% |
| rifamycin | 0.01824 | 0.00736 | 97.97% |
| sulfonamide | 0.18931 | 0.20277 | 95.02% |
| tetracycline | 0.39135 | 0.22796 | 97.18% |
| trimethoprim | 0.09795 | 0.05218 | 97.40% |
| unclassified | 0.20608 | 0.05600 | 98.55% |
| vancomycin | 0.00174 | 0.00079 | 97.73% |
| Totals | 3.39303 | 1.55233 | 97.73% |
| Phylum | Influent | Effluent | Activated Sludge | ||||||
|---|---|---|---|---|---|---|---|---|---|
| INF_W | INF_S | INF Avg | EFF_W | EFF_S | EFF Avg | SL_W | SL_S | SL Avg | |
| Acidobacteria | 0.01 | 0.10 | 0.05 | 0.00 | 0.12 | 0.06 | 0.06 | 0.96 | 0.51 |
| Actinobacteria | 1.60 | 1.02 | 1.31 | 4.27 | 7.41 | 5.84 | 2.65 | 8.36 | 5.50 |
| Bacteroidetes | 30.98 | 13.66 | 22.32 | 40.81 | 27.55 | 34.18 | 34.10 | 20.63 | 27.37 |
| Chlamydiae | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.29 | 0.00 | 0.14 |
| Chlorobi | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.55 | 0.00 | 0.28 |
| Cyanobacteria | 0.00 | 0.00 | 0.00 | 0.00 | 0.01 | 0.00 | 0.00 | 0.00 | 0.00 |
| Deinococcus_Thermus | 0.00 | 0.00 | 0.00 | 0.02 | 0.13 | 0.07 | 0.00 | 0.00 | 0.00 |
| Euryarchaeota | 0.04 | 0.08 | 0.06 | 0.10 | 0.02 | 0.06 | 0.05 | 0.16 | 0.11 |
| Firmicutes | 19.55 | 8.66 | 14.11 | 11.00 | 11.41 | 11.20 | 2.32 | 5.21 | 3.76 |
| Fusobacteria | 0.09 | 0.00 | 0.05 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
| Lentisphaerae | 0.03 | 0.07 | 0.05 | 0.02 | 0.07 | 0.05 | 0.00 | 0.00 | 0.00 |
| Proteobacteria | 47.47 | 75.40 | 61.43 | 43.45 | 52.28 | 47.86 | 59.98 | 64.61 | 62.29 |
| Spirochaetes | 0.00 | 0.01 | 0.01 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
| Synergistetes | 0.04 | 0.64 | 0.34 | 0.00 | 0.19 | 0.09 | 0.00 | 0.08 | 0.04 |
| Verrucomicrobia | 0.16 | 0.36 | 0.26 | 0.33 | 0.81 | 0.57 | 0.00 | 0.00 | 0.00 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ali, O.S.; Hozayen, W.G.; Almutairi, A.S.; Edris, S.A.; Abulfaraj, A.A.; Ouf, A.A.; Mahmoud, H.M. Metagenomic Analysis Reveals the Fate of Antibiotic Resistance Genes in a Full-Scale Wastewater Treatment Plant in Egypt. Sustainability 2021, 13, 11131. https://doi.org/10.3390/su132011131
Ali OS, Hozayen WG, Almutairi AS, Edris SA, Abulfaraj AA, Ouf AA, Mahmoud HM. Metagenomic Analysis Reveals the Fate of Antibiotic Resistance Genes in a Full-Scale Wastewater Treatment Plant in Egypt. Sustainability. 2021; 13(20):11131. https://doi.org/10.3390/su132011131
Chicago/Turabian StyleAli, Osama S., Walaa G. Hozayen, Abdulwahab S. Almutairi, Sherif A. Edris, Aala A. Abulfaraj, Amged A. Ouf, and Hamada M. Mahmoud. 2021. "Metagenomic Analysis Reveals the Fate of Antibiotic Resistance Genes in a Full-Scale Wastewater Treatment Plant in Egypt" Sustainability 13, no. 20: 11131. https://doi.org/10.3390/su132011131






