Bacterial Community Structure and Predicted Metabolic Function of Landfilled Municipal Solid Waste in China
Abstract
:1. Introduction
2. Materials and Methods
2.1. Samples Collection
2.2. Physical and Chemical Analyses
2.3. DNA Extraction and Sequencing
2.4. High-Throughout Data Processing
2.5. Bacterial Community Function Analysis
3. Results
3.1. Physical and Chemical Properties
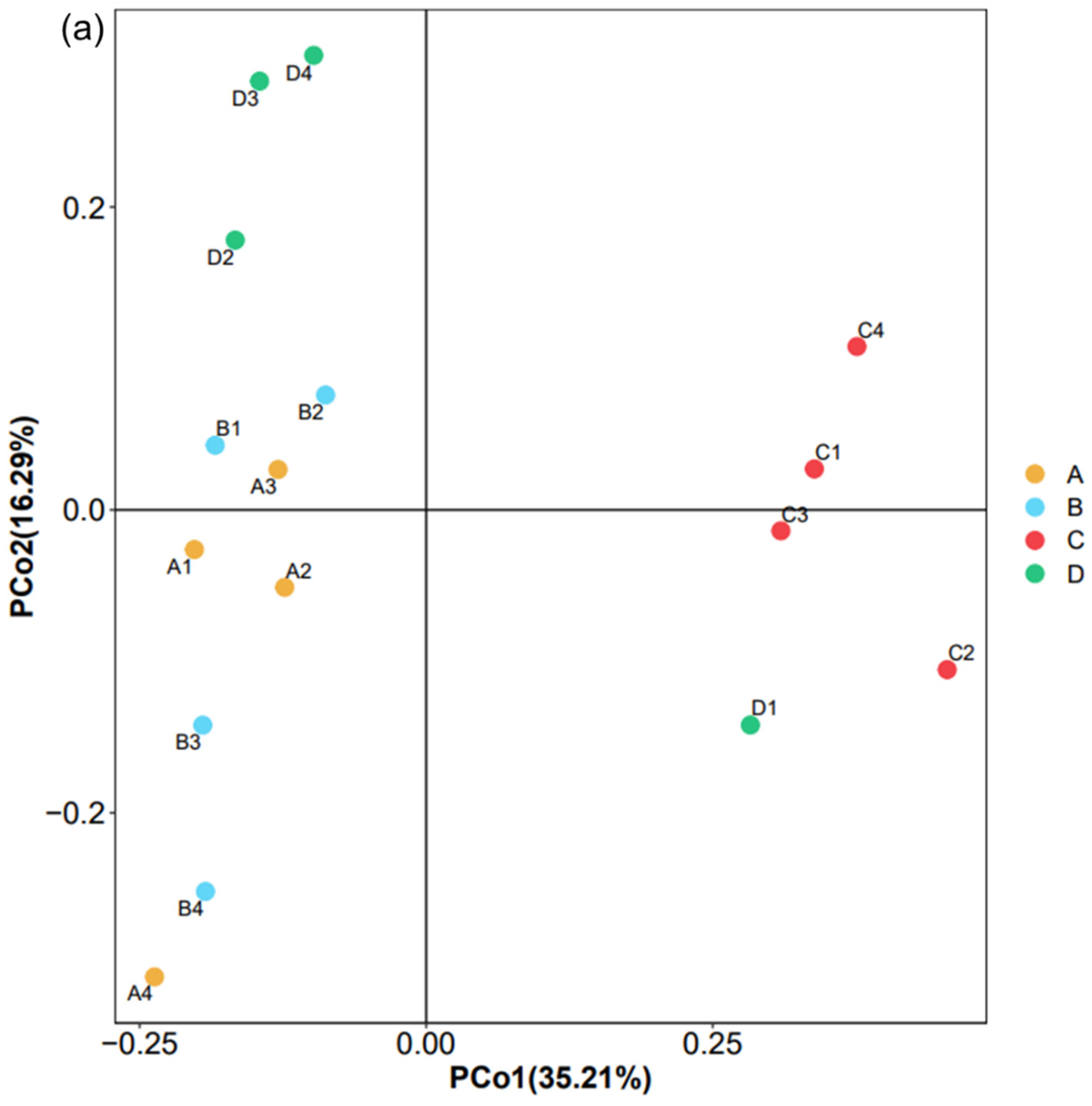
3.2. Sequencing Results and Microbial Diversity
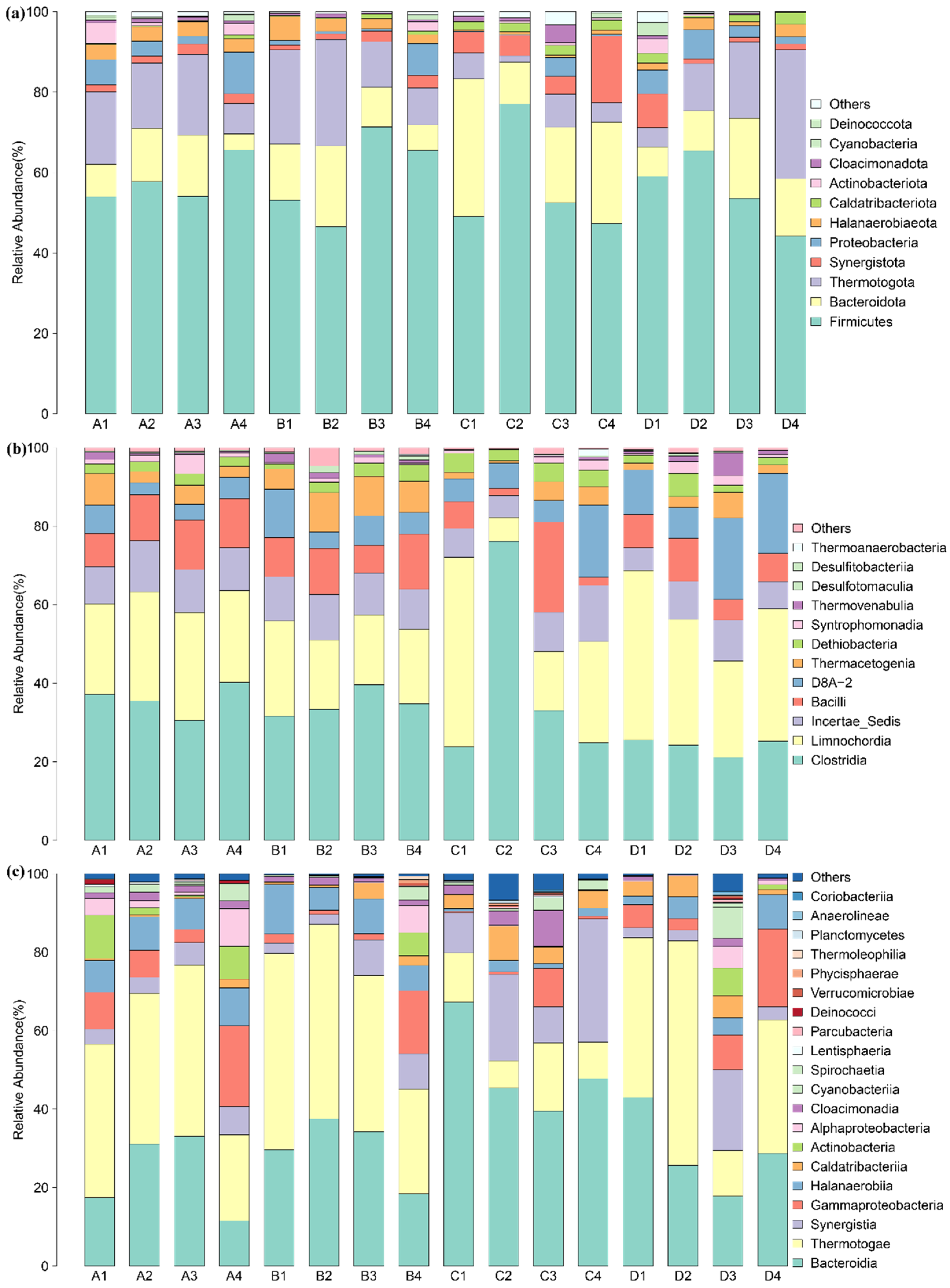
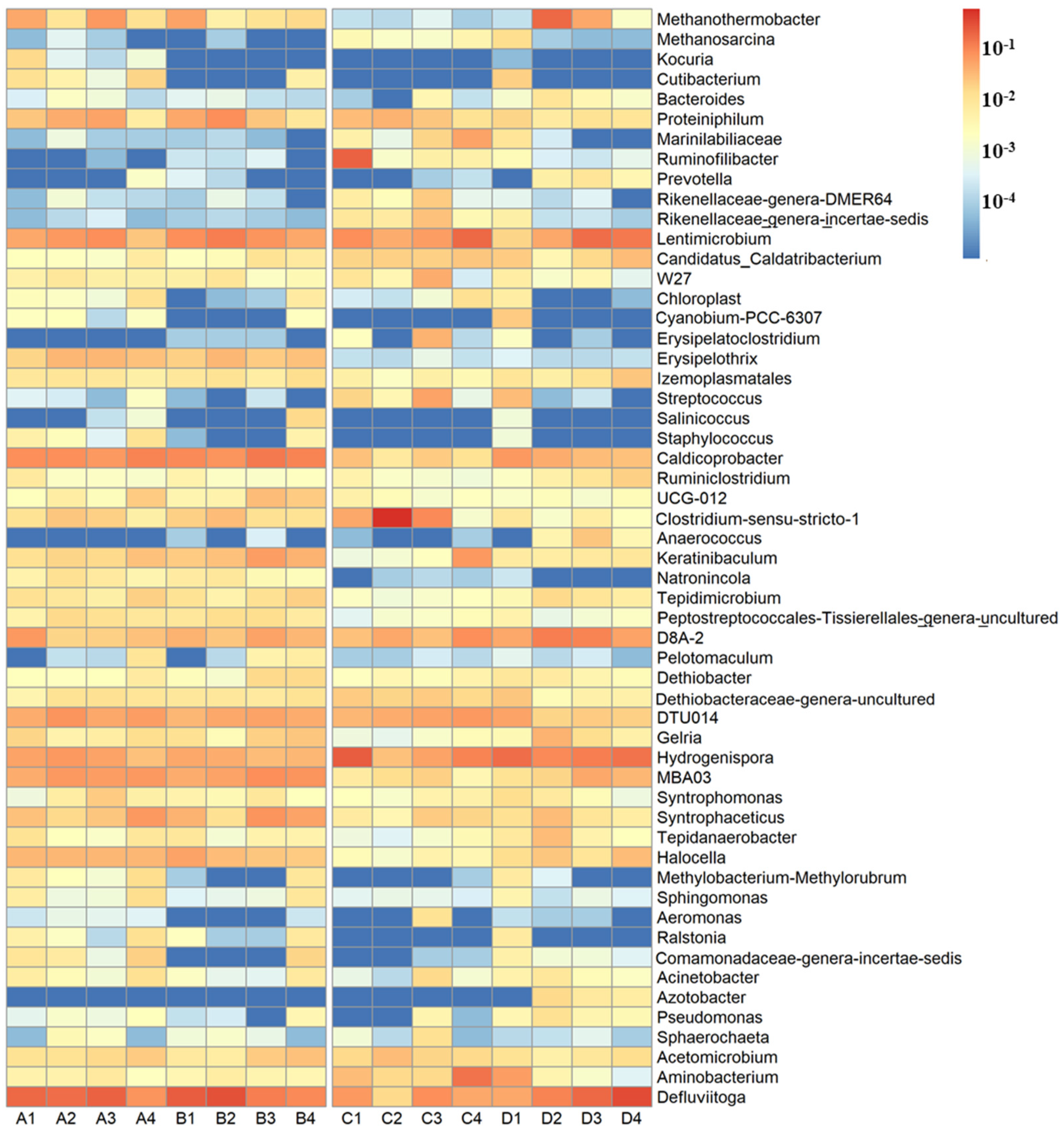
3.3. Bacterial Taxonomic Composition
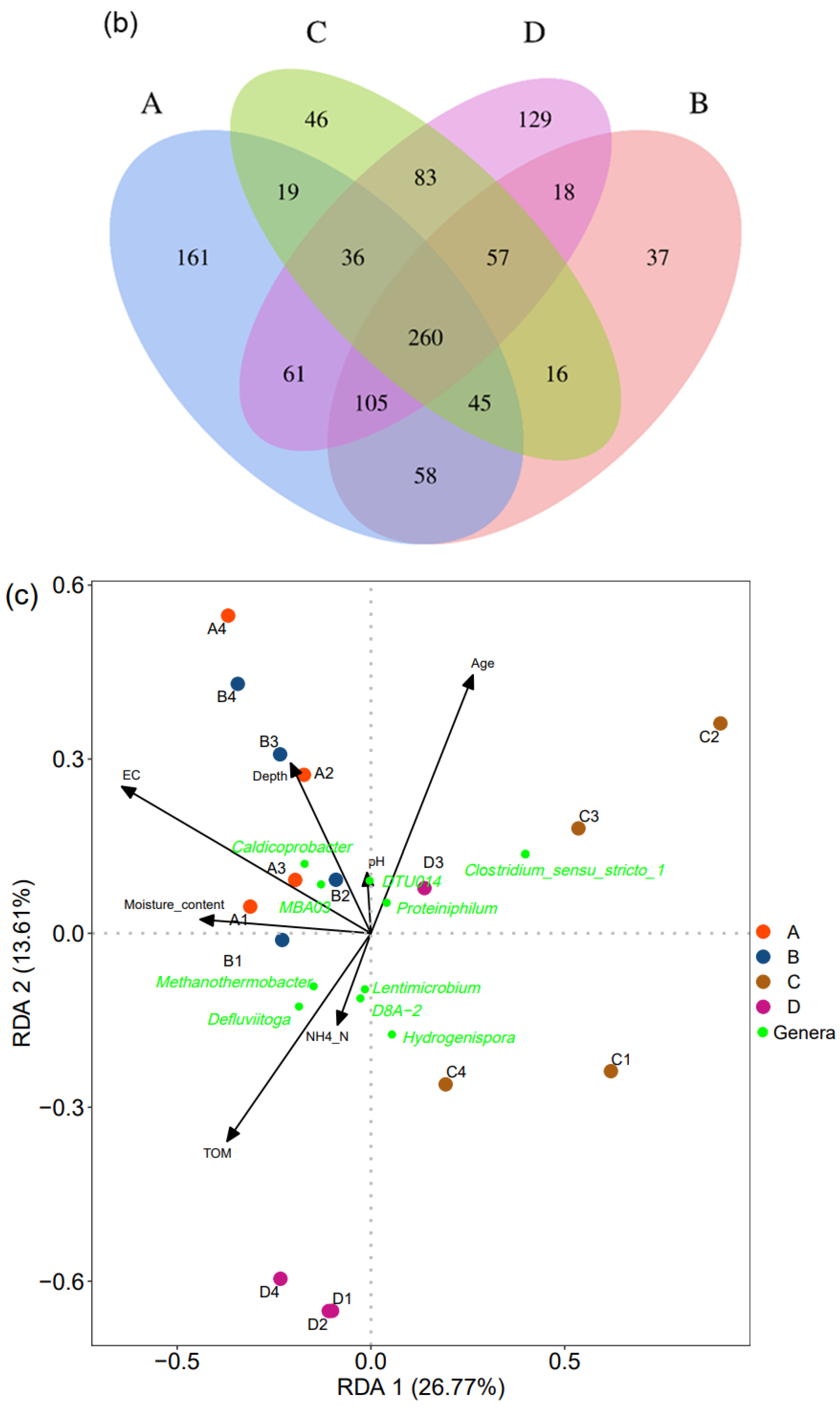
3.4. Impact Factors on Bacterial Community Structure
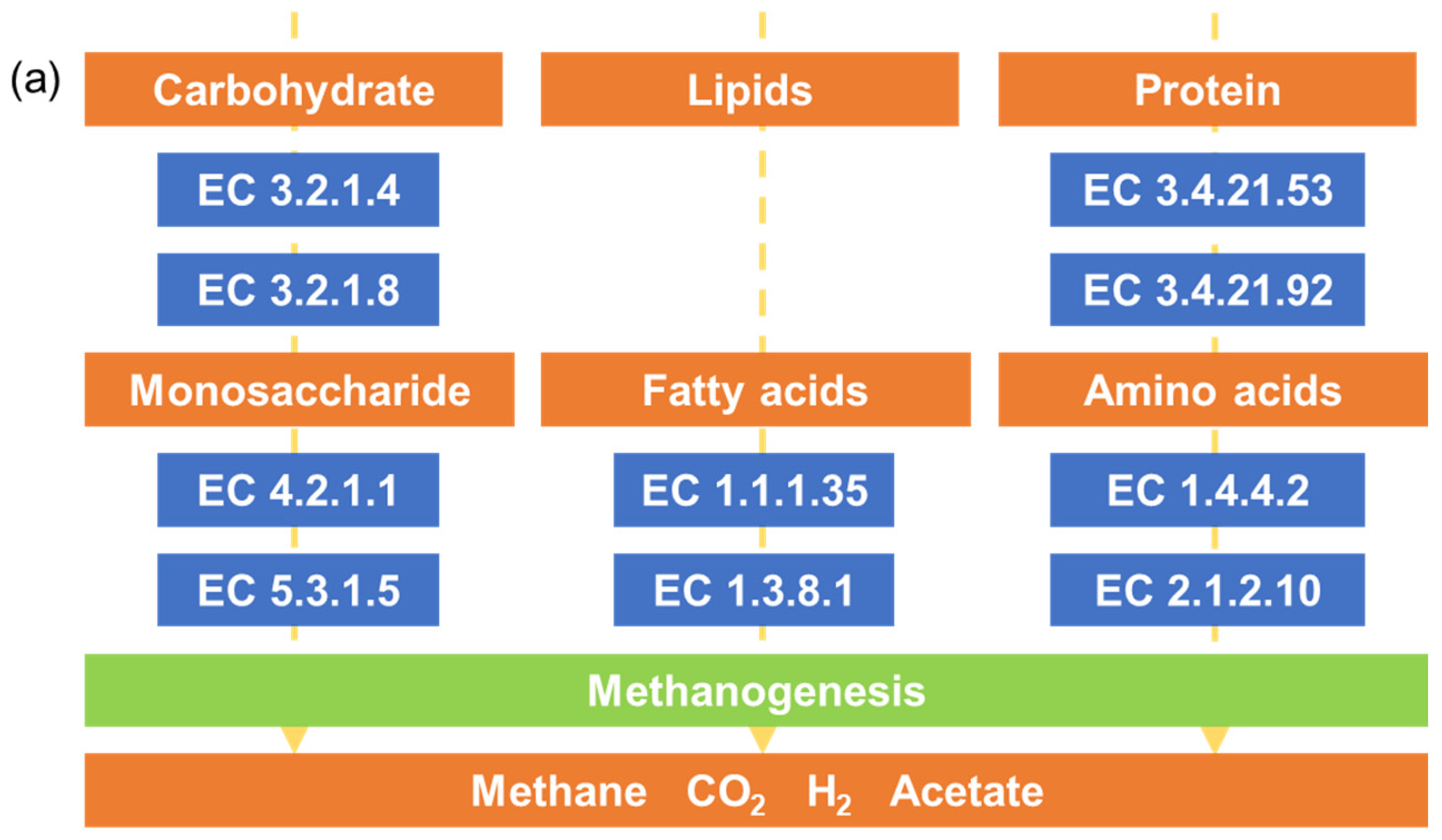
3.5. Functional Analysis of Bacterial Community
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Bank. What a Waste2.0: A Global Snapshot of Solid Waste Management to 2050. Available online: http://hdl.handle.net/10986/30317 (accessed on 24 February 2022).
- Jingxia, Y. Municipal solid waste (MSW)-to-energy in China: Challenges and cost analysis. Energy Sources Part B Econ. Plan. Policy 2017, 13, 116–120. [Google Scholar] [CrossRef]
- Saini, J.K.; Kumar, G.; Singh, S.; Kuhad, R.C. Municipal solid waste biorefinery for sustainable production of bioenergy. In Waste Biorefinery; Elsevier: Amsterdam, The Netherlands, 2021. [Google Scholar]
- National Bureau of Statistics, China. Annual Report; National Bureau of Statistics: Beijing, China, 2020. (In Chinese)
- Chen, Y.; Xu, W.; Ling, D.; Zhan, L.; Gao, W. A degradation–consolidation model for the stabilization behavior of landfilled municipal solid waste. Comput. Geotech. 2020, 118, 103341. [Google Scholar] [CrossRef]
- Barlaz, M.A. Forest products decomposition in municipal solid waste landfills. Waste Manag. 2006, 26, 321–333. [Google Scholar] [CrossRef] [PubMed]
- Ge, S.; Ma, J.; Liu, L.; Yuan, Z. The impact of exogenous aerobic bacteria on sustainable methane production associated with municipal solid waste biodegradation: Revealed by high-throughput sequencing. Sustainability 2020, 12, 1815. [Google Scholar] [CrossRef] [Green Version]
- Reinhart, D.R.; McCreanor, P.T.; Townsend, T. The bioreactor landfill: Its status and future. Waste Manag. Res. J. Sustain. Circ. Econ. 2002, 20, 172–186. [Google Scholar] [CrossRef]
- Valencia, R.; van der Zon, W.; Woelders, H.; Lubberding, H.; Gijzen, H. The effect of hydraulic conditions on waste stabilisation in bioreactor landfill simulators. Bioresour. Technol. 2009, 100, 1754–1761. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Song, L. Succession of bacterial community structure and metabolic function during solid waste decomposition. Bioresour. Technol. 2019, 291, 121865. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Wu, T.; Wang, X.; Qiu, Z.; Lan, J. Study on compressibility and settlement of a landfill with aged municipal solid waste: A case study in taizhou. Sustainability 2021, 13, 4831. [Google Scholar] [CrossRef]
- Gomez, A.M.; Yannarell, A.C.; Sims, G.K.; Cadavid-Restrepo, G.; Herrera, C.X.M. Characterization of bacterial diversity at different depths in the Moravia Hill landfill site at Medellín, Colombia. Soil Biol. Biochem. 2011, 43, 1275–1284. [Google Scholar] [CrossRef]
- Uchida, M.; Hatayoshi, H.; Syuku-Nobe, A.; Shimoyama, T.; Nakayama, T.; Okuwaki, A.; Nishino, T.; Hemmi, H. Polymerase chain reaction-denaturing gradient gel electrophoresis analysis of microbial community structure in landfill leachate. J. Hazard. Mater. 2009, 164, 1503–1508. [Google Scholar] [CrossRef]
- Sawamura, H.; Yamada, M.; Endo, K.; Soda, S.; Ishigaki, T.; Ike, M. Characterization of microorganisms at different landfill depths using carbon-utilization patterns and 16S rRNA gene based T-RFLP. J. Biosci. Bioeng. 2010, 109, 130–137. [Google Scholar] [CrossRef]
- Song, J.; Zhang, W.; Gao, J.; Hu, X.; Zhang, C.; He, Q.; Yang, F.; Wang, H.; Wang, X.; Zhan, X. A pilot-scale study on the treatment of landfill leachate by a composite biological system under low dissolved oxygen conditions: Performance and microbial community. Bioresour. Technol. 2020, 296, 122344. [Google Scholar] [CrossRef]
- Song, L.; Yang, S.; Liu, H.; Xu, J. Geographic and environmental sources of variation in bacterial community composition in a large-scale municipal landfill site in China. Appl. Microbiol. Biotechnol. 2016, 101, 761–769. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Wang, Y.; Zhao, H.; Long, D.T. Composition of bacterial and archaeal communities during landfill refuse decomposition processes. Microbiol. Res. 2015, 181, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Wang, Y.; Tang, W.; Lei, Y. Bacterial community diversity in municipal waste landfill sites. Appl. Microbiol. Biotechnol. 2015, 99, 7745–7756. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Cao, A.; Zhao, G.; Zhou, C.; Xu, R. Microbial community structure and diversity in a municipal solid waste landfill. Waste Manag. 2017, 66, 79–87. [Google Scholar] [CrossRef]
- Xu, S.; Lu, W.; Liu, Y.; Ming, Z.; Liu, Y.; Meng, R.; Wang, H. Structure and diversity of bacterial communities in two large sanitary landfills in China as revealed by high-throughput sequencing (MiSeq). Waste Manag. 2017, 63, 41–48. [Google Scholar] [CrossRef]
- Chien, P.C.T.; Yoo, H.S.; Dykes, G.A.; Lee, S.M. Isolation and characterization of cellulose degrading ability in Paenibacillus isolates from landfill leachate. Malays. J. Microbiol. 2015, 11, 185–194. [Google Scholar]
- Imron, M.F.; Kurniawan, S.B.; Soegianto, A. Characterization of mercury-reducing potential bacteria isolated from Keputih non-active sanitary landfill leachate, Surabaya, Indonesia under different saline conditions. J. Environ. Manag. 2019, 241, 113–122. [Google Scholar] [CrossRef]
- McDonald, J.E.; Lockhart, R.J.; Cox, M.J.; Allison, H.E.; McCarthy, A.J. Detection of novel Fibrobacter populations in landfill sites and determination of their relative abundance via quantitative PCR. Environ. Microbiol. 2008, 10, 1310–1319. [Google Scholar] [CrossRef]
- Pérez-Leblic, M.I.; Turmero, A.; Hernández, M.; Hernández, A.J.; Pastor, J.; Ball, A.S.; Rodríguez, J.; Arias, M.E. Influence of xenobiotic contaminants on landfill soil microbial activity and diversity. J. Environ. Manag. 2012, 95, S285–S290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bareither, C.A.; Wolfe, G.L.; McMahon, K.D.; Benson, C.H. Microbial diversity and dynamics during methane production from municipal solid waste. Waste Manag. 2013, 33, 1982–1992. [Google Scholar] [CrossRef] [PubMed]
- Köchling, T.; Sanz, J.L.; Gavazza, S.; Florencio, L. Analysis of microbial community structure and composition in leachates from a young landfill by 454 pyrosequencing. Appl. Microbiol. Biotechnol. 2015, 99, 5657–5668. [Google Scholar] [CrossRef]
- Mwaikono, K.S.; Maina, S.; Sebastian, A.; Schilling, M.; Kapur, V.; Gwakisa, P. High-throughput sequencing of 16S rRNA Gene Reveals Substantial Bacterial Diversity on the Municipal Dumpsite. BMC Microbiol. 2016, 16, 145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stamps, B.W.; Lyles, C.N.; Suflita, J.M.; Masoner, J.R.; Cozzarelli, I.M.; Kolpin, D.W.; Stevenson, B.S. Municipal Solid Waste Landfills Harbor Distinct Microbiomes. Front. Microbiol. 2016, 7, 534. [Google Scholar] [CrossRef] [Green Version]
- Zainun, M.Y.; Simarani, K. Metagenomics profiling for assessing microbial diversity in both active and closed landfills. Sci. Total Environ. 2018, 616–617, 269–278. [Google Scholar] [CrossRef]
- Yadav, B.; Johri, A.K.; Dua, M. Metagenomic Analysis of the Microbial Diversity in Solid Waste from Okhla Landfill, New Delhi, India. Microbiol. Resour. Announc. 2020, 9, e00921-20. [Google Scholar] [CrossRef]
- Zhao, R.; Liu, J.; Feng, J.; Li, X.; Li, B. Microbial community composition and metabolic functions in landfill leachate from different landfills of China. Sci. Total Environ. 2020, 767, 144861. [Google Scholar] [CrossRef]
- Chen, K.; Deng, X.; Wang, L.; He, R.; Yang, Y.; Jiang, J.; Zhang, C.; Yang, S.; Fan, L.; Wang, L.; et al. Effects of different functional strains on key metabolic pathways of methanogenesis in the domestic waste fermentation reactor. Biomass Bioenergy 2021, 146, 105995. [Google Scholar] [CrossRef]
- Ma, P.C.; Ke, H.; Lan, J.W.; Chen, Y.M.; He, H.J. Field measurement of pore pressures and liquid-gas distribution using drilling and ERT in a high food waste content MSW landfill in Guangzhou, China. Eng. Geol. 2019, 250, 21–33. [Google Scholar] [CrossRef]
- Xu, H.; Zhan, L.-T.; Li, H.; Lan, J.-W.; Chen, Y.-M.; Zhou, H.-Y. Time-and stress-dependent model for predicting moisture retention capacity of high-food-waste-content municipal solid waste: Based on experimental evidence. J. Zhejiang Univ. A 2016, 17, 525–540. [Google Scholar] [CrossRef] [Green Version]
- MOHURD (Ministry of Housing and Urban-Rural Development of the People’s Republic of China). Sampling and Analysis Methods for Domestic Waste; CJ/T 313-2009; Standards Press of China: Beijing, China, 2009. (In Chinese)
- MOHURD (Ministry of Housing and Urban-Rural Development of the People’s Republic of China). General Detecting Methods for the Chemical Characteristic of Domestic Refuse; CJ/T 96-2013; Standards Press of China: Beijing, China, 2013. (In Chinese)
- MEE (Ministry of Ecology and Environment of the People’s Republic of China). Water Quality-Determination of Ammonia Nitrogen Nessler’s Spectrophotometry; HJ 535-2009; Standards Press of China: Beijing, China, 2009. (In Chinese) [Google Scholar]
- Ye, J.; Song, Z.; Wang, L.; Zhu, J. Metagenomic analysis of microbiota structure evolution in phytoremediation of a swine lagoon wastewater. Bioresour. Technol. 2016, 219, 439–444. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Caporaso, J.G. Author Correction: Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Callahan, B.J.; Mcmurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [Green Version]
- Paradis, E.; Schliep, K. ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef]
- Team, R.C. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
- Jari Oksanen, F.; Blanchet, G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. 2019. Available online: https://CRAN.R-project.org/package=vegan (accessed on 24 February 2022).
- Song, L.; Wang, Y.; Tang, W.; Lei, Y. Archaeal community diversity in municipal waste landfill sites. Appl. Microbiol. Biotechnol. 2015, 99, 6125–6137. [Google Scholar] [CrossRef]
- Hill, M.O.; Gauch, H.G. Detrended correspondence analysis: An improved ordination technique. Vegetatio 1980, 42, 47–58. [Google Scholar] [CrossRef]
- Wemheuer, F.; Taylor, J.A.; Daniel, R.; Johnston, E.; Meinicke, P.; Thomas, T.; Wemheuer, B. Tax4Fun2: Prediction of habitat-specific functional profiles and functional redundancy based on 16S rRNA gene sequences. Environ. Microbiome 2020, 15, 11. [Google Scholar] [CrossRef]
- Zhan, L.T.; Xu, H.; Chen, Y.M.; Lü, F.; Lan, J.W.; Shao, L.M.; Lin, W.A.; He, P.J. Biochemical, hydrological and mechanical behaviors of high food waste content MSW landfill: Preliminary findings from a large-scale experiment. Waste Manag. 2017, 63, 27–40. [Google Scholar] [CrossRef]
- Zhao, Y.; Song, L.; Huang, R.; Song, L.; Li, X. Recycling of aged refuse from a closed landfill. Waste Manag. Res. J. Sustain. Circ. Econ. 2007, 25, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Nakasaki, K.; Tran, L.T.H.; Idemoto, Y.; Abe, M.; Rollon, A. Comparison of organic matter degradation and microbial community during thermophilic composting of two different types of anaerobic sludge. Bioresour. Technol. 2009, 100, 676–682. [Google Scholar] [CrossRef]
- Reinhart, D.; Joslyn, R.; Emrich, C.T. Characterization of Florida, U.S. landfills with elevated temperatures. Waste Manag. 2020, 118, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Nesbø, C.L.; Bradnan, D.M.; Adebusuyi, A.; Dlutek, M.; Petrus, A.K.; Foght, J.; Doolittle, W.F.; Noll, K.M. Mesotoga prima gen. nov., sp. nov., the first described mesophilic species of the Thermotogales. Extremophiles 2012, 16, 387–393. [Google Scholar] [CrossRef] [PubMed]
- Ben Hania, W.; Ghodbane, R.; Postec, A.; Hamdi, M.; Ollivier, B.; Fardeau, M.-L. Defluviitoga tunisiensis gen. nov., sp. nov., a thermophilic bacterium isolated from a mesothermic and anaerobic whey digester. Int. J. Syst. Evol. Microbiol. 2012, 62, 1377–1382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, L.; Li, F.; Kuroki, A.; Loh, K.C.; Wang, C.H.; Dai, Y.; Tong, Y.W. Methane yield enhancement of mesophilic and thermophilic anaerobic co-digestion of algal biomass and food waste using algal biochar: Semi-continuous operation and microbial community analysis. Bioresour. Technol. 2020, 302, 122892. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Wang, C.; Sun, F.; Zhu, W.; Wu, W. A comparison of microbial characteristics between the thermophilic and mesophilic anaerobic digesters exposed to elevated food waste loadings. Bioresour. Technol. 2014, 152, 420–428. [Google Scholar] [CrossRef]
- Kim, E.; Lee, J.; Han, G.; Hwang, S. Comprehensive analysis of microbial communities in full-scale mesophilic and thermophilic anaerobic digesters treating food waste-recycling wastewater. Bioresour. Technol. 2018, 259, 442–450. [Google Scholar] [CrossRef]
- Ghollarata, M.; Raiesi, F. The adverse effects of soil salinization on the growth of Trifolium alexandrinum L. and associated microbial and biochemical properties in a soil from Iran. Soil Biol. Biochem. 2007, 39, 1699–1702. [Google Scholar] [CrossRef]
- Sardinha, M.; Müller, T.; Schmeisky, H.; Joergensen, R.G. Microbial performance in soils along a salinity gradient under acidic conditions. Appl. Soil Ecol. 2003, 23, 237–244. [Google Scholar] [CrossRef]
- Townsend, T.G.; Miller, W.L.; Lee, H.-J.; Earle, J.F.K. Acceleration of Landfill Stabilization Using Leachate Recycle. J. Environ. Eng. 1996, 122, 263–268. [Google Scholar] [CrossRef]
- Wu, X.; Tian, Z.; Lv, Z.; Chen, Z.; Liu, Y.; Yong, X.; Zhou, J.; Xie, X.; Jia, H.; Wei, P. Effects of copper salts on performance, antibiotic resistance genes, and microbial community during thermophilic anaerobic digestion of swine manure. Bioresour. Technol. 2020, 300, 122728. [Google Scholar] [CrossRef]
- Jetten, M.S.M. The microbial nitrogen cycle. Environ. Microbiol. 2008, 10, 2903–2909. [Google Scholar] [CrossRef]
- Fierer, N.; Lauber, C.L.; Ramirez, K.S.; Zaneveld, J.; Bradford, M.A.; Knight, R. Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J. 2012, 6, 1007–1017. [Google Scholar] [CrossRef] [Green Version]
- Fierer, N.; Jackson, R.B. The diversity and biogeography of soil bacterial communities. Proc. Natl. Acad. Sci. USA 2006, 103, 626–631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Westlake, K.; Archer, D.; Boone, D. Diversity of cellulolytic bacteria in landfill. J. Appl. Bacteriol. 1995, 79, 73–78. [Google Scholar] [CrossRef]
- Inaba, T.; Su, T.; Aoyagi, T.; Aizawa, H.; Sato, Y.; Suh, C.; Lee, J.H.; Hori, T.; Ogata, A.; Habe, H. Microbial community in an anaerobic membrane bioreactor and its performance in treating organic solid waste under controlled and deteriorated conditions. J. Environ. Manag. 2020, 269, 110786. [Google Scholar] [CrossRef] [PubMed]
- Conklin, A.; Stensel, H.D.; Ferguson, J. Growth Kinetics and Competition between Methanosarcina and Methanosaeta in Mesophilic Anaerobic Digestion. Water Environ. Res. 2006, 78, 486–496. [Google Scholar] [CrossRef] [PubMed]
- Youcai, Z.; Zhugen, C.; Qingwen, S.; Renhua, H. Monitoring and long-term prediction of refuse compositions and settlement in large-scale landfill. Waste Manag. Res. 2001, 19, 160–168. [Google Scholar] [CrossRef] [PubMed]
- Ximenes, F.; Gardner, W.; Cowie, A. The decomposition of wood products in landfills in Sydney, Australia. Waste Manag. 2008, 28, 2344–2354. [Google Scholar] [CrossRef] [PubMed]
- Bothe, H.; Ferguson, S.; Newton, W.E. Biology of the Nitrogen Cycle; Elsevier: Amsterdam, The Netherlands, 2007; pp. 3–20. [Google Scholar]


| Region | Phase | Bacteria Phylum | Dominant Genus | Reference |
|---|---|---|---|---|
| UK | Leachate | Fibrobacters | NA | [23] |
| Western Japan | Solid waste | Firmicutes, Bacteroidota, Proteobacteria | NA | [14] |
| Medellín, Colombia | Solid waste | Bacteroidota, Proteobacteria | Acinetobacter | [12] |
| Madrid, Spain | Solid waste | Proteobacteria, Acidobacteria, Actinobacteria | Sphingosinicella, Microvirga, Amaricoccus, Conexibacter, Marmoricola, Pseudonocardina | [24] |
| USA | Solid waste/leachate | Proteobacteria, Firmicutes, Bacteroidota | Pseudomanas, Arcobacter, Acinetobacter | [25] |
| Brazil | Solid waste | Firmicutes, Bacteroidota, Proteobacteria | Syntrophomonas, Pseudomanas, Clostridium, Petrimonas, Proteiniphilum | [26] |
| Arusha, Tanzania | Solid waste | Firmicutes, Bacteroidota, Proteobacteria, Acidobacteria, Actinobacteria, Spirochaetes | Acinetobacter, Clostridium, Proteinclasticum, Lactobacillus | [27] |
| USA | Leachate | Firmicutes, Proteobacteria | NA | [28] |
| Beijing and Xi’an, China | Solid waste | Firmicutes, Bacteroidota, Proteobacteria | Proteiniphilum, Atopostipes, Petrimonas | [20] |
| Yangzhou, China | Solid waste | Firmicutes, Bacteroidota, Proteobacteria | Halanaerobium, Halanaerobium, Halocella | [19] |
| Malaysia | Solid waste | Firmicutes, Bacteroidota, Proteobacteria, Acidobacteria, Actinobacteria, Spirochaetes | NA | [29] |
| Xi’an, China | Solid waste | Firmicutes, Bacteroidota, Proteobacteria | Olivibacter, Petrimonas | [16] |
| New Delhi, India | Solid waste | Actinobacteria, Proteobacteria, Firmicutes, Bacteroidota, Planctomycetes, Cyanobacteria | NA | [30] |
| 11 cities, China | Leachate | Bacteroidota, Firmicutes, Proteobacteria | Candidatus Cloacamonas, Defluviitoga, Sphaerochaeta, Arcobacter, Syntrophomonas, and Petrimonas | [31] |
| Component | Landfill | Food Waste | Plastic | Paper | Textile | Wood | Glass and Metal | Cinder and Dust | Others |
|---|---|---|---|---|---|---|---|---|---|
| Mass proportion (100%) | Xingfeng | 54.0 | 19.9 | 12.3 | 4.6 | 3.0 | 3.3 | 1.6 | 1.3 |
| Tianziling | 59.4 | 18.0 | 13.5 | 2.0 | 1.1 | 2.3 | 3.7 | 0.0 |
| Sample ID | Site | Depth (m) | Age (a) | Moisture Content (%) | Organic Matter Content (%) | pH | EC (ms/cm) | Ammonia (mg/kg) |
|---|---|---|---|---|---|---|---|---|
| A1 | Xingfeng landfill, Guangzhou | 10 | 2~3 | 49% | 26.73% | 7.7 | 35.1 | 108.15 |
| A2 | 20 | 10~11 | 25% | 23.68% | 6.6 | 32.7 | 165.45 | |
| A3 | 30 | 10~11 | 33% | 22.55% | 8.0 | 28.5 | 180.75 | |
| A4 | 40 | 10~11 | 41% | 19.93% | 8.0 | 27.8 | 166.35 | |
| B1 | 10 | 2~3 | 51% | 28.01% | 7.6 | 33.5 | 49.05 | |
| B2 | 20 | 10~11 | 27% | 32.82% | 6.8 | 33.4 | 150.45 | |
| B3 | 30 | 10~11 | 37% | 28.55% | 8.0 | 28.0 | 166.65 | |
| B4 | 40 | 10~11 | 42% | 26.59% | 8.0 | 29.1 | 183.75 | |
| C1 | Tianziling landfill, Hangzhou | 10 | 7~9 | 33% | 19.88% | 7.8 | 24.5 | 123.75 |
| C2 | 20 | 9~11 | 32% | 22.93% | 7.7 | 18.6 | 187.05 | |
| C3 | 30 | 11~13 | 24% | 21.61% | 7.9 | 24.1 | 30.03 | |
| C4 | 40 | 13~15 | 21% | 15.56% | 7.7 | 24.9 | 299.25 | |
| D1 | 10 | 1~2 | 52% | 32.39% | 7.5 | 30.1 | 169.05 | |
| D2 | 20 | 5~7 | 32% | 32.53% | 7.4 | 24.3 | 226.95 | |
| D3 | 30 | 7~9 | 30% | 35.04% | 7.0 | 28.3 | 28.05 | |
| D4 | 40 | 9~11 | 25% | 29.95% | 7.8 | 19.1 | 116.70 |
| Sample ID | Chao-1 | Shannon | Evenness | Simpson |
|---|---|---|---|---|
| A1 | 451.7 | 4.22 | 0.195 | 0.955 |
| A2 | 497.8 | 4.19 | 0.160 | 0.956 |
| A3 | 603.5 | 3.98 | 0.103 | 0.944 |
| A4 | 362 | 4.37 | 0.264 | 0.975 |
| B1 | 334.9 | 3.72 | 0.126 | 0.930 |
| B2 | 345.9 | 3.47 | 0.094 | 0.905 |
| B3 | 305.8 | 3.91 | 0.164 | 0.959 |
| B4 | 380.2 | 4.39 | 0.247 | 0.973 |
| C1 | 370.4 | 3.54 | 0.094 | 0.921 |
| C2 | 277 | 2.51 | 0.045 | 0.690 |
| C3 | 479.9 | 4.48 | 0.184 | 0.971 |
| C4 | 324.2 | 3.60 | 0.116 | 0.937 |
| D1 | 509.5 | 4.87 | 0.281 | 0.983 |
| D2 | 402 | 4.01 | 0.140 | 0.955 |
| D3 | 349.6 | 3.60 | 0.105 | 0.924 |
| D4 | 312.7 | 3.16 | 0.079 | 0.868 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ke, H.; Li, J.; Zhang, X.; Hu, J. Bacterial Community Structure and Predicted Metabolic Function of Landfilled Municipal Solid Waste in China. Sustainability 2022, 14, 3144. https://doi.org/10.3390/su14063144
Ke H, Li J, Zhang X, Hu J. Bacterial Community Structure and Predicted Metabolic Function of Landfilled Municipal Solid Waste in China. Sustainability. 2022; 14(6):3144. https://doi.org/10.3390/su14063144
Chicago/Turabian StyleKe, Han, Jinghang Li, Xuanqi Zhang, and Jie Hu. 2022. "Bacterial Community Structure and Predicted Metabolic Function of Landfilled Municipal Solid Waste in China" Sustainability 14, no. 6: 3144. https://doi.org/10.3390/su14063144
APA StyleKe, H., Li, J., Zhang, X., & Hu, J. (2022). Bacterial Community Structure and Predicted Metabolic Function of Landfilled Municipal Solid Waste in China. Sustainability, 14(6), 3144. https://doi.org/10.3390/su14063144







