1. Introduction
Rice (
Oryza sativa L.) is one of the most important staple foods for billions of people in the world [
1]. Rice is the main food source for more than 50% of the world’s population and a very important commodity for the Indonesian community [
2]. In 2021–2022, rice production, reached 525 million tons, with a consumption of 53.4 kg/year per capita [
3]. Furthermore, rice production in Indonesia reached 31.36 million tons in 2021 [
4]. One of the problems in rice production is iron (Fe) toxicity [
5]. Rice production is also influenced by various environmental factors [
6,
7]. Hence, it is important to develop Fe-tolerant rice, and rice that can adapt to environmental changes, in high yields.
As a staple food, rice is the greatest human energy source. Rice content is generally composed of protein, which mostly consists of glutamate and aspartic acid, carbohydrates, fiber, fat, thiamine (vitamin B1), riboflavin (vitamin B2), and niacin (vitamin B3) [
8,
9]. Thus, rice can be used as a nutritional source for humans.
Recently, rice with high yield potential has been widely developed. High yields are obtained via plant breeding programs. In the process of selecting superior genotypes (high and stable yields), it is obtained via multi-environmental testing. The multi-environment test is a test carried out to evaluate the performance and stability of a variety of plants in various agroecological environments [
7,
10]. Multi-environmental testing is important to carry out in the selection of Fe-tolerant rice with high yield and stability because it can determine genotypes by environmental interactions (GEIs). The existence of GEIs is influenced by biotic and abiotic factors. Biotic factors consist of pests and diseases [
11,
12], while abiotic factors are from the environment [
7]. The existence of GEIs indicates that various genotypes can have different responses to environmental changes. By carrying out multi-environment tests, stable genotypes in various environments that are also adaptive to specific environments can be determined [
13]. However, GEIs make the selection process difficult and inefficient under various environmental conditions [
7,
13,
14]. Several studies have reported that GEIs complicate the selection process in sweet potatoes [
7,
15], durum wheat [
16], soybeans [
17,
18,
19], maize [
20], Stevia [
21], and turmeric [
22]. Therefore, a study is needed to evaluate Fe-resistant rice genotypes under a wide range of environmental conditions to obtain superior genotypes (high and stable yields).
Iron toxicity in rice is a common problem in areas where the soil contains excessive iron. This condition adversely affects plant growth and productivity, leading to significant yield losses [
23]. In order to develop rice genotypes that are resistant to iron toxicity and perform well under diverse environmental conditions, genotypes should be evaluated on various types of land. There are numerous important land types for genotypic evaluation: (1) High iron content soils. Genotypes that exhibit good growth and high yields on high iron content soils can be considered superior candidates for further development. (2) Low iron content soils. Genotypes that show good growth and high yields on low iron content soils can provide solutions for areas experiencing iron deficiency. (3) Different soil pH. Genotypes that can withstand and produce good, as well as low pH (acidic), soils can be considered genotypes that are tolerant to soil pH fluctuations and have the potential to be grown on various types of land [
24,
25].
Several researchers conducted multi-environmental evaluations using various stability measures. Combining stability measurements is considered to be more accurate in selecting genotypes that are stable and have a high yield [
7,
26]. Some of the stability analyses used are parametric estimation models, namely linear regression [
27], Wricke’s equivalence [
28], Shukla’s stability variance [
29], coefficient of variation (
CVi) [
30], GE variance component (
θ(i)) [
31], component of mean-variance (
θi) [
32], and Hanson genotypic stability (
Di) [
33]; non-parametric estimation models, namely the Thennarasu model (
NP(i)) [
34], non-parametric stability (S
(i)) [
35], and Kang ranks (
KR) [
36]; as well as graphical models, i.e., AMMI [
37] and GGE biplot [
38]. Several researchers have also used the sustainability index (SI) to identify or select superior genotypes [
39,
40,
41,
42]. The use of the sustainability index was considered to increase selection accuracy. In addition, the concept of sustainable agriculture was also an important factor in the advancement of agricultural crops [
43]. In plant breeding programs, multi-environment testing is a crucial step to increase productivity and maintain food security [
44]. The use of these various methods was expected to be able to identify and evaluate genotypes in a wide range of environments.
Currently, many researchers have used the combined stability analysis model to select superior genotypes, such as in grass peas [
45], barley [
26], peanuts [
46], maize [
47], sweet potato [
7], and black soybean [
18]. Because the combined stability method was considered more effective and accurate in selecting stable and high-yielding genotypes compared to a single stability estimation [
40]. The advantages of this model are that it is (i) suitable for data with statistical assumptions such as interaction effects and a normal distribution of errors; (ii) simple because estimation is based on genotypic performance and does not require assumptions for homogeneity of variance; and (iii) suitable for visually determining the response of a genotype to the environmental testing [
26,
48]. The objectives of this study were to identify the effects of genotypes, environment, and their interactions (GEIs) on the yields of Fe-tolerant rice; select superior genotypes (stable and high yields) under diverse environmental conditions in Indonesia; and determine the mega-environments (MEs) and representative environments for Fe-tolerant rice development.
4. Discussion
Genotype, environmental, and GEI factors showed a significant effect on Fe-tolerant rice yields. GEIs (50.92%) contributed more, followed by environment (35.78%) and genotype (13.30%). This indicates that the variation in rice yields tested in six different environments was dominantly influenced by external effects, in this case, the environment. The emergence of GEIs in multi-location testing makes plant breeding programs difficult, especially in selecting ideal genotypes [
7]. Several studies have reported that GEIs significantly affect rice yields [
6,
61,
62]. In other studies, GEIs also have a significant effect on crop yields, such as maize [
13,
40,
41], sweet potatoes [
7], butterfly pea [
42], and soybean [
18,
19]. The large contribution of GEIs also indicates a high environmental effect, where the six test environments have different conditions. Environmental differences can be seen in the type of agro-climate and altitude (
Table 2). In addition, the resulting CV also shows a value of 27.28%. This value was included in the high criteria (>20%). The high CV value was caused by various factors, one of which is the environment [
14]. Therefore, highly significant GEIs for rice yields justify using parametric, non-parametric, AMMI, GGE biplot, and sustainability index (SI) stability measures to represent GEIs against tested genotypes and estimate the yield stability of rice genotypes that are evaluated.
In plant breeding programs, the presence of GEIs complicates the plant selection process. This situation indicates that the selection of rice genotypes based on yields must be carried out in certain environments, resulting in inefficient breeding programs [
13]. In addition, the appearance of GEIs indicates that the tested rice genotypes have different potential when planted in different locations [
7], so the development program must be adjusted based on a representative environment. The influence of GEIs also causes sub-optimal genotypic potential under different environmental conditions [
47]. However, the presence of GEIs can also provide an opportunity to select rice genotypes that have high yields in certain environments (adapt-specific). These conditions indicate that the selection of appropriate rice genotypes must be carried out in each environment. This is very interesting because Indonesia has very varied land requirements, so the use of specific (adaptive) genotypes will greatly assist the development of certain areas in terms of rice production. Therefore, plant breeding activities must be directed according to the appropriate environmental conditions so that the plant varieties developed are able to adapt well to various environments.
In the process of selecting high yielding varieties that are stable in a wide range of environments, various methods have been proposed. The use of a single stability measure is still considered difficult for determining stable and high yielding genotypes [
7,
13,
26], so another selection model was needed to obtain the ideal genotype. Several studies have reported the selection of stable genotypes and high yields by applying a combination of stability models, including grass peas [
45], barley [
26], soybeans [
18,
19], turmeric [
22], and maize hybrid [
13,
40]. In this study, parametric and non-parametric stability models, AMMI, GGE biplot, and the sustainability index (SI) were applied to identify stable and high-yielding Fe-tolerant rice genotypes in six different environments. The average rank (AR) values of all stability models (parametric and non-parametric) were used to select high-yielding and stable rice genotypes (with low AR values). This was also expressed by Vaezi et al. (2019) [
26]. Genotypes G4 and G14 had the lowest AR values, indicating a high level of stability and production (above the average overall yield), as shown in
Table 5.
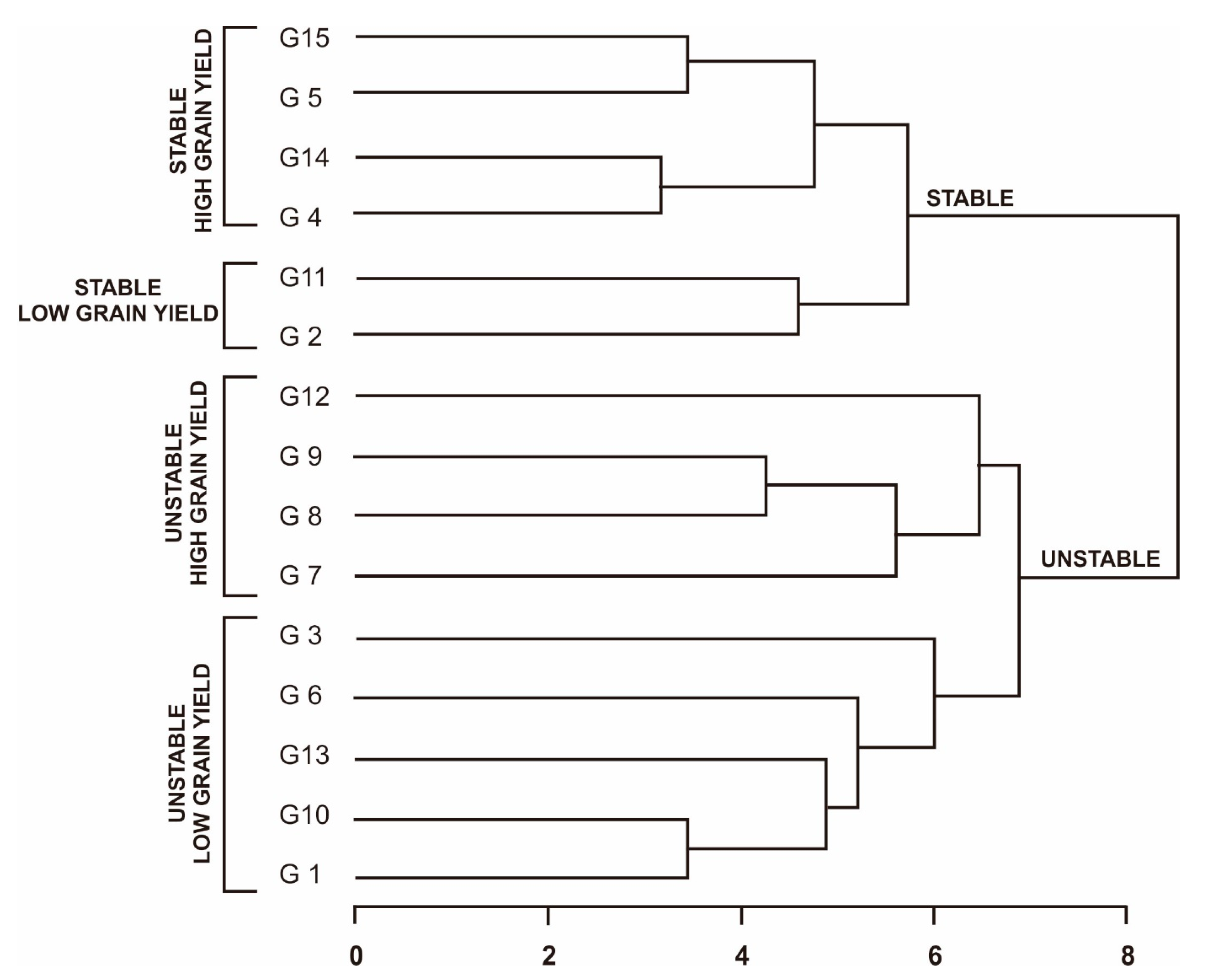
A grouping of Fe-tolerant rice genotypes using a dendrogram based on stability rating was used to confirm these results. The dendrogram was developed based on ranking the average yield of all parametric and non-parametric measurements. The resulting dendrogram divides the Fe-tolerant rice genotypes into two main clusters (
Figure 1): stable and unstable. Each main cluster was divided into two sub-clusters. The stable high-grain-yield sub-cluster consists of genotypes G4, G5, G14, and G15. This group is an ideal group because the genotypes are able to adapt to a wide range of environments and have high yields. This is followed by the stable low-grain-yield sub-cluster, which consists of genotypes G2 and G11. The next sub-cluster is the unstable high grain yield, which contains the genotypes G7, G8, G9, and G12. This group is a group that can be recommended as a genotype with a specific adaptation because the genotypes in this group will have high production in a representative environment [
7]. Finally, the unstable low grain yield group consists of genotypes G1, G3, G6, G10, and G13. This group is the worst and is not recommended for development on a large scale. This was also conveyed by Ruswandi et al. (2022) [
47], where genotypes that were in the stable high-grain-yield group were more desirable, and, conversely, genotypes that were in the unstable low-grain-yield group were not expected and were not used for the development of superior varieties.
The results of Spearman’s rank correlation analysis revealed that the stability measures NP
(4) and θ
i were clearly related to the average yield (Y), indicating that these two were included in the category of dynamic stability measures. Stability measures that are positively correlated with yields can be used to recommend genotypes in a specific environment (adapt) [
63]. Both of these measurements are useful for selecting Fe-tolerant rice genotypes with high average yields in an environment with good growing conditions (representative). In contrast, stability measures were significantly and negatively correlated with crop yields, indicating them to be a static stability term that can be used to select genotypes in unfavorable environments [
26]. W
i2 with σ
2i, s
2d
i, θ
(i), and ASV measurements have a negative and significant correlation with yield (
Table 6), so they are included in the concept of static stability. Stability measurements that have positive and significant correlations indicate that they have a strong strength in selecting stable genotypes [
7]. However, if they have significant and negative correlation coefficients, then they have the opposite power in choosing a stable genotype. For example, in this study, S
(1) has almost the same power as S
(2), S
(3), S
(6), NP
(1), NP
(2), NP
(3), NP
(4),
KR, Wi
2, σ
2i, s
2d
i, θ
(i), ASV, and GSI. In addition, there is also a measure of stability that has the same power in selecting stable genotypes (correlation coefficient = 1.00), namely Wi
2 with σ
2i and θ
(i). In a graphical visualization based on the PCA biplot, the three are also in the same group (
Figure 2), so one can use one of them to select a stable genotype [
13,
19]. The PCA’s visualization of stability measurements divides them into five groups. The first group consists of Di, NP
(2), NP
(3), NP
(4), S
(3), S
(6),
KR, and GSI; the second group consists of S
(1), S
(2), NP
(1), CVi, and
bi; the third group consists of s
2d
i, θ
(i), Wi
2, σ
2i, and ASV measurements; the fourth group consists of yield (Y); and the fifth group consists of θ
i. The first three groups and the fifth group represent the concept of static stability. According to Vaezi et al. 2019 [
26], the concept of stability is divided into two, i.e., the concept of static and dynamic stability. The concept of static stability can be used to select genotypes under unfavorable environmental conditions. Conversely, the concept of dynamic stability can be used to recommend genotypes in favorable environments [
40]. Thus, both parametric and non-parametric measurements provide recommendations for selecting rice in unfavorable environments.
To select stable and high yielding genotypes, graphical visualization is essential. In this study, two graphical measurements were used, namely AMMI and GGE biplots. The AMMI biplot presented in
Figure 3 shows that PC1 contributed 57.1% to diversity and 23.6% to PC2. The large contribution of PC1 to yield diversity implies that the interaction of rice genotypes with the six growing environments in Indonesia was predicted by the first PC from genotype and environment. Several researchers also revealed the same thing, where the PC1 value plays an important role in the diversity of yields in the AMMI biplot [
21,
42,
64]. In the AMMI biplot, genotypes that are close to the biplot axis are stable and have low GEIs [
37]. In this study, five Fe-tolerant rice genotypes were identified that were close to the axis (0.0) and within the radius of the circle. They were G2, G3, G4, G11, and G15. This shows that the five of them have stable yields in the six test environments. Other genotypes identified as adaptive to specific environments, namely G7 and G10 in Lampung; G5, G12, and G13 in Blandean; G14 in Riau; G8 and G9 in West Papua; and G1 and G6 in Bogor. Thus, the identified genotypes can be developed according to their position in relation to their environment. Based on
Figure 3, Lampung has the longest environmental vector. This shows that Lampung is a representative environment in selecting superior genotypes and has a large influence on GEIs. This is in line with the results of the GGE biplot analysis, where Lampung is included in environmental type II, which means it is very representative for testing [
13]. Meanwhile, Banjar Baru and West Papua have the shortest environmental vectors in the AMMI and GGE biplots and were included as a Type I environment, which indicates that both have a small effect on GEIs and do not represent the actual yields.
In the GGE biplot, the ‘representative versus discriminative’ display (
Figure 4) shows that the West Papua, Riau, and Blandean environments are Type I environments, so they should not be used as test environments. Among the six test environments, Lampung is a Type II environment that has selective influence and high representativeness; hence, Lampung is an ideal location to select superior genotypes (
Figure 4) [
7]. The Banjar Baru and Bogor environments are Type III environments and should not be used to select superior genotypes but can be used to select adaptive or region-specific genotypes [
60]. The ‘mean vs. stability’ biplot display shows that five genotypes have yields above the overall average, including G4, G6, G7, G10, and G12 (
Figure 5), while the other ten genotypes had yields below the overall average yield. G4 tends to be stable and has high yields because it is close to the
X-axis and to the left of the
Y-axis [
60]. G7 has the highest average yield (6.05 t·ha
−1), which is indicated by its farthest position from the
Y-axis. On the other hand, G13 is the genotype that has the worst yield in all environments, so this genotype is not recommended for large-scale development.
The ‘which won where’ display shows that the six environments have five sectors with different peak genotypes (
Figure 6). Several researchers revealed that this biplot could indicate the existence of a mega-environment [
7,
40,
48]. There are two sectors that consist of more than one environment so as to form a mega-environment. The first mega-environment is in sector I, which consists of Lampung and Banjar Baru, with G7 as the peak genotype. The second mega-environment is in the second sector, consisting of Bogor and West Papua, with G6 as the peak genotype. These results indicate that there were various environmental groupings during the trials. The first PC explained 54.2% of the total variation due to environmental effects (E) and GEIs during the experiment (
Figure 4,
Figure 5 and
Figure 6). The grouping of environments and mega-environments in various regions in Indonesia with different peak genotypes indicates a genotype-specific adaptation to the mega-environment and the positive utilization of GEIs [
7,
10,
19]. Our findings reveal that several Fe-tolerant rice genotypes have adapted to different environments in Indonesia. This will facilitate the selection process for wider development. In the third sector, G8 is the peak genotype but there is no environment. In sector four, there is the Riau environment with G13 as the peak genotype. Sector five has a Blandean environment with G12 as the peak genotype. Genotypes that are at the top of each sector indicate that these genotypes have high yields in the environment in that sector [
20,
21]. Genotypes that are in a sector containing more than one environment or mega-environment show the ideal genotype [
19,
40,
65]. Conversely, genotypes at the top of the sector but that do not have an environment show low yields [
7]. Therefore, the genotypes in this sector are not recommended for further development.
The evaluation of the yield stability of Fe-tolerant rice based on the sustainability index (SI) divided the genotypes into three groups, namely low, medium, and high (
Table 8). SI estimates for rice yields range from 27.78% (low) to 66.11% (high). Genotypes with high and very high SI categories are expected [
39,
41,
42]. Our results show that the genotypes with the highest SI category were shown by G2, G4, and G6, but only G4 and G6 had yields above the overall average yield (4.18 t·ha
−1), so these two genotypes are included in the selected category. In addition, in another study, Filio et al. (2023) revealed that genotypes with medium SI criteria and that had high average yields were included in the ideal category. Nine rice genotypes were in the medium SI category (40–60%) (
Table 8), including G1, G3, G5, G8, G9, G10, G11, G14, and G15; however, only genotypes G5, G8, and G14 have a yield above the average overall yield (>4.18 t·ha
−1). Genotypes with low SI values indicate unstable yields for these three genotypes, so they are not recommended for further development [
41,
42].
To select superior (stable and high-yielding) rice genotypes, slices were taken from all measurements used. This has also been carried out by several researchers, including grass peas [
45], sweet potatoes [
7,
66], soybeans [
17,
18], and maize [
13,
40,
41]. The information on selected genotypes presented in
Table 9 shows that AMMI, GGE biplot, and the SI each selected stable genotypes by 33.33%. In comparison, the combined stability (parametric and non-parametric) succeeded in selecting 26.67%. However, only one genotype was determined based on all measurements (6.67%), namely G4. Therefore, G4 is highly recommended to be used as a new superior, stable rice variety with a high yield.